Abstract
Cervical cancer is the fourth most common female cancer worldwide. Long non-coding RNAs (lncRNAs), such as SOX21-AS1, play pivotal roles in the progression and metastasis of cancer. We previously described that SOX21-AS1 was hypomethylated in cervical cancer (CC) and aimed to further explore the relationship between methylation of the SOX21-AS1 promoter and CC using clinical cervical samples. Pyrosequencing was performed to detect the methylation status of the SOX21-AS1 promoter in 33 cervical specimens. Additionally, expression levels of related genes in 43 clinical cervical specimens were measured using quantitative real-time PCR (qRT-PCR). The SOX21-AS1 promoter was significantly hypomethylated in CC (P < 0.01). SOX21-AS1 hypomethylation was also significantly associated with an advanced Federation of Gynecology and Obstetrics (FIGO) stage (P < 0.01). The expression levels of SOX21-AS1 and SOX21 were noted to be higher in cancer vs. normal cervix (all P < 0.001). Moreover, the expression of SOX21-AS1 was positively correlated with SOX21 in all samples (r = 0.891, P < 0.001). Methylation statue of the SOX21-AS1 promoter region was negatively correlated with the expression levels of SOX21-AS1 and SOX21 (SOX21-AS1, r = − 0.628; SOX21, r = − 0.648; both P < 0.001). The methylation status of SOX21-AS1 displayed promising diagnostic potential for CC, exhibiting good sensitivity (100.0%) and specificity (69.2%), with an area under the curve of 0.846. In addition, bioinformatic analyses identified a potential link between SOX21-AS1 and the Wnt signaling pathway. Furthermore, methylation status of SOX21-AS1 was negatively correlated with β-catenin/c-myc/cyclin D1 mRNA levels (rs = − 0.529, − 0.462 ,and − 0.383, respectively, P < 0.05). Our findings illuminated that lncRNA SOX21-AS1 showed hypomethylation in cervical cancer and SOX21-AS1 could serve as a novel biomarker for CC diagnosis or a potential therapeutic target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 2018, there were approximately 570,000 new cases and 311,000 new deaths worldwide owing to cervical cancer (CC), making cervical cancer the fourth most common female cancer [1]. In developing regions such as Middle, Southern, and Eastern Africa, the number of cases was disproportionately high (> 85%) [2]. Persistent human papillomavirus (HPV) infection is the leading cause of CC, although there are many other contributory factors, such as epigenetic changes [3]. Basic screening tests for HPV detection and liquid-based cytology (LBC) diagnoses have somewhat decreased the mortality associated with CC [4,5,6]. However, these tests have their limitations and are plagued by poor specificity [7]. Thus, it is a necessity to identify novel and reliable biomarkers for CC.
Long non-coding RNAs (lncRNAs) are composed of more than 200 nucleotides and do not encode proteins [8]. Although not translated, lncRNAs have emerged as indispensable regulators within various biological processes [9]. In particular, lncRNAs are crucial participants in chromatin dynamics, gene expression, and cancer [10]. They can serve as decoys, activators, or guides with the ability to bind DNA, RNA, or protein [8, 11]. As lncRNAs are involved in transcriptional, post-transcriptional, and epigenetic regulation, there have been an increased number of studies implicating lncRNAs within the progression of cancer [12]. Long non-coding RNA CCAT1 promotes the proliferation and invasion of colon cancer [13]. Long non-coding RNA H19 promotes lung cancer proliferation and metastasis by inhibiting miR-200a function [14]. Finally, lncRNA MEG3 can induce cell cycle arrest and apoptosis to reduce the proliferation of cervical carcinoma cells [15]. Highlighting that lncRNAs can be used as an interesting diagnostic and therapeutic target within cancer.
DNA methylation plays a critical role in transcriptional regulation and is linked to various diseases, especially cancer [16]. In normal cells, DNA methylation is involved in regulating gene expression [17], and aberrant methylation can result in gene silencing or gene activation directly [18]. In previous studies, we utilized a genome-wide methylation chip and described that hypomethylation of the SOX21-AS1 promoter was associated with the pathogenesis of CC [13]. SOX21-AS1 is the non-coding ribonucleic acid of 3230 bp on human chromosome 13q32.1:94712716-94715945, which is the antisense of SOX21 and shares a head-to-head promoter with SOX21 [19]. We hypothesized that abnormal methylation of the SOX21-AS1 promoter may alter the expression of the associated genes and could impact the progress of CC. Thus, more clinical samples were required to clearly identify the relationship between SOX21-AS1 methylation and CC.
In the current study, the methylation status of the SOX21-AS1 promoter and expression levels of SOX21-AS1 and SOX21 in clinical cervical samples were evaluated through pyrosequencing and qRT-PCR, respectively. Besides, we investigated the association between the methylation status of SOX21-AS1 and CC to evaluate the diagnostic potential of this relationship. Here, the methylation status of SOX21-AS1 promoter was significantly lower in CC, suggesting that SOX21-AS1 hypomethylation may be a novel biomarker for the diagnosis of CC.
Materials and Methods
Clinical Samples
All the specimens were randomly selected from a prospective collection of samples stored in our biological sample banks between 2016 and 2018 at the Third Affiliated Hospital of Zhengzhou University. In this study, the patients included in the cancer group displayed cytological and pathological indications of squamous cervical carcinoma (SCC) and HPV infection. Conversely, the patients included in the normal group displayed cytological features of mild inflammation without HPV infection. All patients were sexually active, not pregnant, without total hysterectomy, and no history of miscarriage, medical treatment, or surgery. Finally, a total of 43 clinical samples, including 23 normal cervical specimens (median age 48 years, range 26–65 years) and 20 cervical cancer specimens (median age 50 years, range 28–68 years), were randomly selected for this research. The clinical features of the CC samples are listed in Table 1. Ethical consent was obtained from the Ethics Committee of the Third Affiliated Hospital, Zhengzhou University.
DNA Extraction and Bisulfite Modification
DNA was extracted from the clinical samples using a QIAamp DNA Blood Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. The concentration and purity of DNA was detected on a NanoDrop ND-2000 spectrophotometer (Thermo Scientific, Wilmington, USA). The final concentration of DNA samples was ≥ 30 ng/μl, and the OD260/OD280 values were between 1.7 and 1.9. There were 10 specimens that did not meet the necessary standard, and only 33 specimens were investigated in follow-up experiments. Bisulfite conversion was performed using the EpiTect Bisulfite Kit (Qiagen, Germany) according to the manufacturer’s instructions, and the bisulfite-treated DNA was purified on columns.
DNA Amplification and Pyrosequencing
Pyrosequencing was performed to detect the methylation levels. The target sequence focused on chromosome (chr) 13: 94714102-94715945 and the CpG site (nucleotide positions 4202) located in the 5′UTR of the promoter region of SOX21-AS1 (Fig. 1a). Amplification and sequencing primers used in the study were designed using Primer Premier 5.0 software (Premier, Canada; Table 2). To perform the polymerase chain reactions (PCRs), 10 μl 5 × PCR reaction buffer, 1 μl of a dNTP mix (10 mM/each), 1 μl of forward primer (50 pM/μl), 1 μl of reverse primer (50 pM/μl), 2 μl of DNA template, 0.2 μl of Taq polymerase (5 U/μl), and 34.8 μl of H2O (total reaction volume of 50 μl) were added to PCR tubes. The PCR cycle conditions were as follows: preheating at 95 °C for 3 min, followed by 40 cycles of denaturation at 94 °C for 30 s, annealing at 50 °C for 30 s, extension at 72 °C for 1 min, and 1 cycle of sequencing at 72 °C for 7 min. The size and specificity of the PCR product was verified by agarose gel electrophoresis. A PyroMark Q96 ID (Qiagen, Germany) was used to perform pyrosequencing, and Pyro Q-CG software was used to perform the assay setup, run the sequence, and analyze results.
The methylation status of a CpG dinucleotide in SOX21-AS1 was assessed by pyrosequencing. a The detection site of SOX21-AS1, which is located in the promoter regions. Cg11007120 is the number of the CpG site. b Methylation status of this site in CC. c Methylation status of this site in normal group. CC, cervical cancer
Quantitative Real-Time PCR
Primer design: Primers to amplify the SOX21-AS1 and SOX21 gene sequences (accession numbers NR_046514.1 and NM_007084.4, respectively) were designed using PyroMark Assay Design 2.0 software. GAPDH was used as an internal reference gene for calibration (Tables 3). RNA extraction: Total RNA was extracted using Trizol (Toyobo, Japan) in 43 clinical specimens, and the RNA concentrations were quantified on a NanoDrop ND-2000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Reverse transcription: 1 μg RNA was reverse transcribed into cDNA using the ReverTra Ace qPCR RT Kit (Toyobo, Japan). Quantitative real-time PCR: qRT-PCR was conducted using the fluorescence quantitative PCR kit (Toyobo, Japan) on a StepOnePlus™ Real-Time PCR System (ABI, Foster City, CA, USA) according to the manufacturer’s instructions. PCR was performed in a 20 μl reaction mixture containing 10 μl 1 × PCR Master Mix, 0.8 μl forward primer (0.4 μM/μl), 0.8 μl of reverse primer (0.4 μM/μl), 2 μl of DNA template, and 6.4 μl of H2O. The PCR reaction was conducted using the following conditions: preheating at 95 °C for 1 min, followed by 40 cycles of 95 °C for 15 s, 60 °C for 15 s, and 72 °C for 45 s. The specificity of the amplified products was verified via a melting curve.
Bioinformatic Prediction
Multi-experimental matrix (MEM: http://biit.cs.ut.ee/mem/) is a web resource which provides common available gene expression datasets for a large number of different tissues, cells, and diseases [20]. MEM was used to predict genes co-expressed of SOX21-AS1 by selecting suitable species and microarray platform types. Enrichment analysis was performed using the Database for Annotation, Visualization, and Integrated Discovery (DAVID, https://david.ncifcrf.gov/home.jsp) v6.8 to predict potential biological pathways associated with SOX21-AS1.
Statistical Analysis
SPSS 23.0 software (SPSS, Inc., Chicago, IL, USA) was used for statistical analysis. t test and one-way ANOVA were used for comparing the methylation status of SOX21-AS1 between the different groups. The Mann-Whitney test was used to compare the expression levels of SOX21-AS1 and SOX21 between groups. The Spearman rank correlation was used to evaluate the changes of SOX21-AS1/SOX21 expression and the association between SOX21-AS1 methylation and corresponding gene expression. P values of < 0.05 were considered statistically significant.
Results
Hypomethylation Status of SOX21-AS1 in CC
The promoter methylation status of the CpG site (4202) in SOX21-AS1 was quantified in 33 cervical LBC samples (Fig. 1b–c). We found that methylation of the SOX21-AS1 promoter was significantly lower in CC samples compared with the normal samples (t = 3.459, P < 0.01; Fig. 2a). All cervical cancer samples were classified into a hypermethylation group (n = 4) and a hypomethylation group (n = 9) according to the median methylation level of SOX21-AS1. Here we observed that the methylation level of SOX21-AS1 in FIGO stage III samples was significantly lower than that of stage I or stage II samples (P < 0.01, Fig. 2b). However, no significant differences were observed between methylation and other clinical features (P > 0.05).
Overexpression of SOX21-AS1 and SOX21 in CC
In this study, qRT-PCR was performed using total RNA from all 43 samples to detect the expression levels of SOX21-AS1 and SOX21. The expression levels of SOX21-AS1 and SOX21 was significantly increased in CC compared with the normal cervix (SOX21-AS1: Z = − 4.602, P < 0.001; SOX21; Z = − 4.992, P < 0.001, Fig. 3a). Moreover, SOX21-AS1 expression was directly correlated with SOX21 expression in the 43 LBC specimens (r = 0.891, P < 0.001, Fig. 3b).
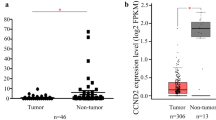
Expression levels of SOX21-AS1 and SOX21 in 43 LBC specimens. a lncRNA SOX21-AS1 and mRNA SOX21 were expressed in cervical cancer and normal samples. b Correlation coefficients of SOX21-AS1 and SOX21 expressions in the LBC specimens. c Correlation coefficients of SOX21-AS1&SOX21 expressions and methylation levels, respectively. LBC, liquid-based cytology. ***P < 0.001
The Relationship Between Methylation and Expression of SOX21-AS1 in CC
To further analyze the relationship between promoter methylation and gene expression of SOX21-AS1, the qRT-PCR results were compared with the methylation status of the SOX21-AS1 promoter. The expression level of SOX21-AS1 in the cancer group was significantly higher than that of normal group, and this increase in SOX21-AS1 expression was correlated with a decrease in its methylation status (r = − 0.628, P < 0.001; Fig. 3c). Similarly, the expression of SOX21 also increased with the decrease in SOX21-AS1 methylation (r = − 0.648, P < 0.001; Fig. 3c).
SOX21-AS1 Methylation as a Potential Diagnostic Biomarker of CC
Receptor operating characteristic curve (ROC) analysis was performed to assess the diagnostic potential of SOX21-AS1. We observed that the methylation status of SOX21-AS1 had high potential to reliably diagnose CC (area under the curve of 0.846; Table 4). When the Youden index was at its maximum value, SOX21-AS1 methylation showed 100.0% sensitivity and 69.2% specificity in the ROC analysis (Fig. 4a).
ROC curves analysis and bioinformatic analysis for SOX21-AS1 in CC. a ROC curve analysis for SOX21-AS1 methylation status as a diagnostic tool for CC; each point on the ROC curve represents a sensitivity/specificity pair. b Co-expression genes and related pathways of SOX21-AS1 and hsa04310: Wnt signaling pathway. CC, cervical cancer; ROC, receiver operating characteristic
SOX21-AS1-Related Pathways in CC
The MEM was used in order to search for genes co-expressed with SOX21-AS1. We chose a collection named the Affymetrix GeneChip Human Genome U133 Plus 2.0 with 1794 datasets as the platform, which included the largest number of genes co-expressed with SOX21-AS1. DAVID was used to perform gene enrichment analysis and identify possible pathways associated with SOX21-AS1 expression. Co-expression results showed that SOX21-AS1 was associated with FZD3/FRAT2/VANGL2, suggesting the potential involvement of the Wnt signaling pathway (Fig. 4b).
Analysis of the Relationship Between SOX21-AS1 Methylation and Wnt Signaling Pathway
In this study, qRT-PCR was performed to detect the expression of β-catenin/c-myc/cyclinD1 mRNA in 43 cases of CC, and the results were compared with SOX21-AS1 methylation status. The β-catenin/c-myc/cyclin D1 expression levels were significantly higher than that in normal cervix (all P < 0.01; Fig. 5a). Besides, the β-catenin/c-myc/cyclin D1 expression levels increased with the decrease of SOX21-AS1 methylation (rs = − 0.529, − 0.462 and − 0.383, respectively, P < 0.05; Fig. 5b).
Expression levels of β-catenin/c-myc/cyclin D1 mRNA in 43 samples and the relationship between SOX21-AS1 methylation and Wnt signaling pathway. a Expression levels of β-catenin/c-myc/cyclin D1 mRNA in normal group and cancer group. b Correlation coefficients of β-catenin/c-myc/cyclin D1 expression levels and methylation status of SOX21-AS1, respectively. **P < 0.01
Discussion
The biological responses of hosts infected with different types of HPV are very different; even the same type of HPV infection can cause different clinical outcomes. It was therefore suggested that there are other important factors involved in the malignant process associated with CC. For instance, Hoppe-Seyle et al. inferred that lncRNAs may regulate the carcinogenic processes associated with HPV infection [21]. Besides, epigenetic modifications, especially DNA methylation, are an essential trigger in cancer progression [17, 22]. LncRNA SOX21-AS1 promoter region contains a number of CpG dinucleotides in the conserved palindromic sequence, which is a potential target for DNA methylation in the host cell. Our previous study reported that the SOX21-AS1 promoter region has a hypomethylated site in cervical cancer through the comprehensive application of high-throughput chip technology [23]. Thus, we used clinical samples to detect the promoter methylation status of SOX21-AS1. In the current study, the methylation status of a CpG site in the SOX21-AS1 promoter region was successfully detected through pyrosequencing. Our analysis showed that SOX21-AS1 undergoes hypomethylated in CC, which is in line with our previous studies. In addition, clinical assays indicated that SOX21-AS1 hypomethylation was significantly associated with advanced clinical progression, including FIGO stage, indicating that dysregulation of SOX21-AS1 may influence the clinical outcome of CC. ROC curve analysis indicated that SOX21-AS1 methylation had good specificity and sensitivity in the diagnosis of CC. Taken together, our study provided important evidence for SOX21-AS1 hypomethylation in CC and could be used as a potential biomarker.
Epigenetic plays an important role in biological processes of cancer [24, 25]. The hypomethylation of SOX21-AS1 promoter region can upregulate expression of related gene and participate in the occurrence and development of CC. Thus, this study further compared the differential expression patterns of SOX21-AS1 and SOX21. Our data showed that the expression levels of SOX21-AS1 in CC were significantly higher than that in the control group, consistent with that of Zhang et al., who suggested that SOX21-AS1 expression was significantly upregulated in both CC tissues and cell lines [26]. Importantly, we first reported that the expression of SOX21 was significantly upregulated in CC. Combined with SOX21-AS1 methylation, our results suggested that the upregulation of SOX21-AS1 and SOX21 expression in CC was influenced by SOX21-AS1 hypomethylation. Besides, we also found that the expression level of SOX21-AS1 was positively correlated with SOX21, suggesting that they may have a shared biological function. Our previous study found that SOX21 was involved in epidermal development and nucleic acid binding of transcription factors through GSEA analysis [23]. To sum up, these finding revealed that SOX21-AS1 may enhance the transcriptional activity of some factors to stimulate cell proliferation. Of note, the differences in gene expression were much larger in comparison with the differences in methylation, indicating that there were other factors involved in the dysregulation of SOX21-AS1 in CC, which needed further study.
Previous studies focused on the expression level of SOX21-AS1 in tumors; however, there were few studies on SOX21-AS1 methylation. Yang et al. reported that SOX21-AS1 methylation is more common in oral cancer, which promotes the growth and invasion of oral cancer cells [19]. In contrast to this study, our study showed that hypomethylation of SOX21-AS1 occurred in CC. There are two main reasons for these inconsistencies. First, DNA methylation was detected using a variety of techniques, and some assays could not quantify methylation adequately at individual CpG sites. Second, the source of the samples used for these experiments is different. Yang’s research was carried out on oral tissues, whereas our research was carried out on cervical exfoliated cells. However, our data suggested that SOX21-AS1 appears to play different roles in different cancers, implying that SOX21-AS1 has obvious tumor heterogeneity and important diagnostic potential.
In this study, we predicted that SOX21-AS1 may influence the progress of CC through Wnt signaling pathway based on bioinformatic analysis. Wang et al. also reported that silencing of SOX21-AS1 inhibits apoptosis and reduces oxidative stress damage by activating the Wnt signaling pathway [27]. The quantitative RT-PCR results confirmed that the expression level of β-catenin/c-myc/cyclinD1 in CC was significantly increased. In addition, our data showed that the β-catenin/c-myc/cyclin D1 expression levels increased with the decrease of SOX21-AS1 methylation. Thus, the results of our research further support the hypothesis that SOX21-AS1 hypomethylation may promote the progress of CC via the Wnt signaling pathway.
However, the small number of clinical samples used in this study was a limitation, and future studies will expand on the clinical sample size and follow-up with these patients. In addition, we would investigate the methylation status of SOX21-AS1 in a cell model to further understand the molecular mechanisms underlying the progression of CC.
Conclusions
In summary, our work provided new evidence regarding the methylation status of the SOX21-AS1 promoter region in CC. A decrease in the promoter methylation status of SOX21-AS1 was associated with an increase in cervical lesion severity and increased expression of SOX21-AS1 and SOX21. This study further confirmed that epigenetic alterations to SOX21-AS1 are related to the pathophysiology of CC. Therefore, hypomethylation of SOX21-AS1 may be regarded as a promising biomarker for CC diagnosis and therapy.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359–86.
Bosch FX, Lorincz A, Muñoz N, Meijer CJ, Shah KV. The causal relation between human papillomavirus and cervical cancer. J Clin Pathol. 2002;55(4):244–65.
Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol. 1999;189(1):12–9.
Feng Q, Balasubramanian A, Hawes SE, Toure P, Sow PS, Dem A, et al. Detection of hypermethylated genes in women with and without cervical neoplasia. J Natl Cancer Inst. 2005;97(4):273–82.
Bosch FX, Manos MM, Muñoz N, Sherman M, Jansen AM, Peto J, et al. Prevalence of human papillomavirus in cervical cancer: a worldwide perspective. International biological study on cervical cancer (IBSCC) study group. J Natl Cancer Inst. 1995;87(11):796–802.
Kontostathi G, Zoidakis J, Anagnou NP, Pappa KI, Vlahou A, Makridakis M. Proteomics approaches in cervical cancer: focus on the discovery of biomarkers for diagnosis and drug treatment monitoring. Expert Rev Proteomics. 2016;13(8):731–45.
Yang G, Lu X, Yuan L. LncRNA: a link between RNA and cancer. Biochim Biophys Acta. 2014;1839(11):1097–109.
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77(15):3965–81.
Bhan A, Mandal SS. LncRNA HOTAIR: A master regulator of chromatin dynamics and cancer. Biochim Biophys Acta. 2015;1856(1):151–64.
Tsai MC, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71(1):3–7.
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472(7341):120–4.
He X, Tan X, Wang X, Jin H, Liu L, Ma L, et al. C-Myc-activated long noncoding RNA CCAT1 promotes colon cancer cell proliferation and invasion. Tumour Biol. 2014;35(12):12181–8.
Hadji F, Boulanger MC, Guay SP, Gaudreault N, Amellah S, Mkannez G, et al. Altered DNA methylation of long noncoding RNA H19 in calcific aortic valve disease promotes mineralization by silencing NOTCH1. Circulation. 2016;134(23):1848–62.
Qin R, Chen Z, Ding Y, Hao J, Hu J, Guo F. Long non-coding RNA MEG3 inhibits the proliferation of cervical carcinoma cells through the induction of cell cycle arrest and apoptosis. Neoplasma. 2013;60(5):486–92.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358(11):1148–59. https://doi.org/10.1056/NEJMra072067.
Kulis M, Esteller M. DNA methylation and cancer. Adv Genet. 2010;70:27–56.
Leonard SM, Wei W, Collins SI, Pereira M, Diyaf A, Constandinou-Williams C, et al. Oncogenic human papillomavirus imposes an instructive pattern of DNA methylation changes which parallel the natural history of cervical HPV infection in young women. Carcinogenesis. 2012;33(7):1286–93.
Yang CM, Wang TH, Chen HC, Li SC, Lee MC, Liou HH, et al. Aberrant DNA hypermethylation-silenced SOX21-AS1 gene expression and its clinical importance in oral cancer. Clin Epigenetics. 2016;8:129.
Adler P, Kolde R, Kull M, Tkachenko A, Peterson H, Reimand J, et al. Mining for coexpression across hundreds of datasets using novel rank aggregation and visualization methods. Genome Biol. 2009;10(12):R139.
Hoppe-Seyler F, Hoppe-Seyler K. Emerging topics in human tumor virology. Int J Cancer. 2011;129(6):1289–99.
Morgan AE, Davies TJ, Mc Auley MT. The role of DNA methylation in ageing and cancer. Proc Nutr Soc. 2018;77(4):412–22.
Wang R, Li Y, Du P, Zhang X, Li X, Cheng G. Hypomethylation of the lncRNA SOX21-AS1 has clinical prognostic value in cervical cancer. Life Sci. 2019;233:116708.
Kanwal R, Gupta K, Gupta S. Cancer epigenetics: an introduction. Methods Mol Biol. 2015;1238:3–25.
Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. 2012;150(1):12–27.
Zhang X, Zhao X, Li Y, Zhou Y, Zhang Z. Long noncoding RNA SOX21-AS1 promotes cervical cancer progression by competitively sponging miR-7/VDAC1. J Cell Physiol. 2019;234(10):17494–504.
Zhang L, Fang Y, Cheng X, Lian YJ, Xu HL. Silencing of long noncoding RNA SOX21-AS1 relieves neuronal oxidative stress injury in mice with Alzheimer’s disease by upregulating FZD3/5 via the Wnt signaling pathway. Mol Neurobiol. 2019;56(5):3522–37.
Acknowledgments
We would like to thank Linlin Zhang, Yaming Liu, and Haiyang Yu for the prominent technical assistance and valuable suggestions.
Funding
This study was supported by the Science and Technology Department of Henan, China (Grant Number: 22170005) and Science and Technology Department of Henan, China (Grant Number: LHGJ20190392).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
All the research has been approved by the ethics committee of The Third Affiliated Hospital of Zhengzhou University.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peipei Du and Yanfang Zhi are joint first authors.
Rights and permissions
About this article
Cite this article
Du, P., Zhi, Y., Wang, R. et al. Aberrant Methylation of the SOX21-AS1 Promoter Region Promotes Gene Expression and Its Clinical Value in Cervical Cancer. Reprod. Sci. 28, 532–540 (2021). https://doi.org/10.1007/s43032-020-00335-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43032-020-00335-y