Abstract
In this study, we focus on the phenylalanine ammonia lyase (PAL) as one of the possible defense factors of black rice (a black colour mutation of Oryza sativa, containing a high amount of anthocyanin and other antioxidants) against Xanthomonas oryzae pv. oryzae (Xoo), causing bacterial leaf blight of rice. Two black rice varieties namely Pari Ireng and Melik, and three white rice varieties (IRBB21, Java14, and IR64) as control were used in this study. The analysis of the resistance trait was performed by determining disease intensity (DI) and area under the disease progress curve (AUDPC). The determination of PAL activity was performed using HPLC by analyzing trans-cinnamic acid as a product of PAL enzyme by in vitro bioassay. Protein profile analysis using SDS-PAGE was also performed to compare the profile of the infected and non-infected plants. The results showed that Java 14 (positive control) has the highest AUDPC (107.59) followed by Pari Ireng (128.33), Melik (199.63), and IRBB21 (250.19). Meanwhile, IR64 has the lowest AUDPC (304.63) which was consistent as a negative control. There was a significant increase in PAL enzyme activity in Pari Ireng, Melik and Java 14 but not in IR64 and IRBB21 following Xoo infection. Those results indicated that PAL contributes to the resistance mechanism against Xoo. The distinctive protein profile (ranged from 48 to 75 kDa) was also observed between the infected and non-infected cultivar. Based on our results, Pari Ireng was classified as a resistant cultivar against Xoo.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial leaf blight (BLB) disease is one of the most important biotic stresses on rice in Indonesia and in other rice-producing countries in Asia (Mahmood et al. 2006). BLB is caused by Xanthomonas oryzae pv. oryzae (Xoo). Bacterial leaf blight disease was noted as a limiting factor in increasing rice production (Pascuzzi and McCouch 2007) and reducing rice yields by up to 30–40% as well as reducing the quality of rice (Kadir 2009).
In rice plants, there are many enzymes that play a role in increasing the resistance towards Xoo and one of them is phenylalanine ammonia lyase (PAL) (Song et al. 2016; Yasmin et al. 2016). PAL is an enzyme induced in the phenyl-propanoid pathway in the biosynthesis of various natural products of phenyl-propanoid such as lignin, pigment, flavonoid, and phytoalexin. PAL activity is influenced by biotic and abiotic responses including pathogenic attacks, injured tissue, UV radiation, heavy metal exposure, low temperature, and low levels of nitrogen, phosphate, or ions (Dixon and Paiva 1995). PAL induction and activity is a stress response that can be used as regulatory control. The control may occur through several mechanisms of product inhibition, transcriptional and translational regulation, post-translational inactivation and proteolysis, subcellular enzymes/compartments, and metabolite feedback regulation (Dixon 2001). Therefore, by monitoring the PAL activity, it can be used to predict and support the defense response of a certain mechanism in black rice.
Therefore, in this study, we aim to understand the mechanism of the defense by monitoring the upstream enzyme (PAL) activity following Xoo infection in black rice as one of the model pathways of plant defense.
Materials and methods
Rice plant material
The rice cultivars used in this study included two black rice cultivars (Pari Ireng and Melik), IRBB21 as a moderate resistance control which contain Xa21 gene, IR64 as susceptible control, and Java 14 as resistant control (Susanto and Sudir 2012). Pari Ireng and Melik seeds were obtained from Balai Pengkajian dan Teknologi Pertanian (BPTP) Yogyakarta. The seeds of the other varieties were obtained from Balai Besar Penelitian Tanaman Padi Sukamandi, Subang, West Java. After sowing in the soil (soil to compost fertilizer ratio 3:1) the rice plants were maintained under greenhouse conditions (30–32 °C, 70–85% RH, 12 h dark and light period in tropical conditions) until reach the heading period.
Inoculation of Xoo
The inoculation of Xoo was performed by using leaf clipping method (Kauffman et al. 1993). To avoid overheating and prevent the bacteria from dying, the inoculation took place in the evening. The scissor was soaked into the bacterial suspension (108cells ml−1), two centimeters from the leaf tip. The mock treatment was used by soaking the scissor into sterile water in order to monitor the response of PAL due to injury effect instead of Xoo infection. Each treatment was replicated ten times. Three days after inoculation, the leaves then were harvested and used for further downstream analysis.
Disease intensity and area under disease curve progress (AUDPC) calculation
The observed leaf damage (see below for formula) was converted into disease intensity every seven days for one month. The disease intensity figures were then used to determine the AUDPC value. The damage scale was calculated using the formula according to Suparyono (2003):
The damage scale obtained was then categorized into five categories/scores and the disease intensity was calculated based on the formula according to Tjubarat et al. (1999) cit. in Djatmiko et al. (2011) and the AUDPC calculation was performed according to Djatmiko et al. (2011). To determine the significant differences of each treatment, the Duncan’s multiple range test (DMRT) at 95% confidence level was also used.
Category/score of Xoo damage scale:
Disease intensity formula:
AUDPC formula:
Phenylalanine ammonia lyase (PAL) extraction
One hundred milligram of symptomatic leaf sample was ground with liquid nitrogen by using a mortar until fine powder. The sample was then put into 1.5 ml tube, and the enzyme was extracted by adding 1 ml of 0.2 M borate buffer pH 8.8 containing proteinase inhibitor cocktail (Cat. No. 11836153001, Roche, Swiss) then centrifuged at 12000 g for 15 min at 4 °C. The supernatant was transferred into a new tube and stored at 4 °C until used. The aliquot of the crude extract was used to determine the protein concentration by using Bradford microplate assay method (Bradford 1976). Bovine serum albumin (BSA) was used as a protein standard to make the standard curve. The standard as well as the sample at the microplate, replicated three times, was measured at wavelength 595 nm using visible spectrophotometry (GENESYS™ 10 UV Scanning, Thermo Scientific).
Determination of PAL activity
The in vitro bioassay to measure PAL activity was performed by analyzing trans-cinnamic acid as a product of PAL enzyme, using high-performance liquid chromatography (HPLC). A total of 100 μl of the crude extract was incubated at 40 °C in the waterbath for 5 min, then added 50 μl of L-phenylalanine 30 mM and the incubation was continued for 1 h followed by addition of 5 ml of HCl 5 N addition to the reaction. The extract then was stored in a dark room with a temperature of 4 °C until the analysis by HPLC performed (HPLC conditions column = YMC-Triat C18 Reserved - Phase (250 mm × 4.6 mm, 5 m), Mobile phase = Glacial acetic acid 1%: Methanol (70: 30), Detector = UV-Vis 278 nm). The concentration of trans-cinnamic acid then used to determine the PAL activity in mM/g according to Darmanti (2015) based on the following formula:
To determine the significant differences of each treatment, the DMRT test at 95% confidence level was also used.
SDS-page
The Laemmli (1970) method for separating proteins by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) was used to determine the protein profile by using 12% (w/v) of separating gel and 4% (w/v) of stacking gel. The protein concentration of 15 μg was used in each sample and loaded onto the gels. The sample was then electrophoresed for 3 h at gradually increased voltage including 50 V for 20 min, 80 V for 1 h, and 100 V for the final step by using standard electrophoresis chamber (Atto, Japan). In the gel, the prestained wide range molecular weight (GoldBio, USA) was included to determine the molecular weight (MW) of the protein bands. The silver staining method accrording to Rabilloud (1992) was used to visualize the protein band in the gel. The molecular weight of the protein bands subsequently determined by extrapolation method based on the protein marker bands (Khunsook et al. 2003).
Results and discussion
Disease intensity and AUDPC
Disease intensity was calculated by monitoring the disease progress within four weeks. The determined disease intensities were used to determine the percentage of the area of plant tissue that is attacked by pathogens from the total area observed (Agrios 2005). Results are presented in Table 1, where it is clear that each cultivar experiences an increase in disease intensity values every week. Based on Gnanamanickam et al. (1999) the resistance of a rice variety to bacterial leaf blight is determined by several factors, one of which is the incubation period of the disease. Therefore, the disease intensity observation during four weeks was necessary to carried out to evaluate the black rice cultivars resistance following Xoo infection.
Based on the results of the DMRT test, each cultivar showed a significant increase in disease intensity values in the first week up to the fourth week. As expected, Java 14 cultivars (positive control) had the lowest disease intensity value compared to other cultivars, and IR64 (negative control) had the highest disease intensity value. Pari Ireng cultivar was more resistant than Melik and IRBB21 cultivars.
Based on the disease intensity value, the progress of the disease in the form of AUDPC is represented in Fig.1. The higher the value of AUDPC, the more the variety is susceptible to the disease. Our results show that AUDPC was in line with the disease intensity. Pari Ireng cultivar was more resistant than Melik. Overall, Java 14 (positive control) has the highest AUDPC (107.59) followed by Pari Ireng (128.33), Melik (199.63), and IRBB21 (250.19). Meanwhile, IR64 has the lowest AUDPC (304.63) which was consistent as a negative control.
PAL enzyme activity of black rice following Xoo infection
In plants, phenyl-propanoid pathways produce important essential metabolites such as flavonoids, isoflavonoids, lignins, anthocyanins, antimicrobial compounds, and phenolic esters which act as a defense response to microbial infections and abiotic stress (Ritter and Schulz 2004). PAL is an enzyme in the phenyl-propanoid pathway, which converts L-phenylalanine to trans-cinnamic acid through the non-oxidative removal of ammonia. Trans-cinnamic acid is used as a precursor for lignification and flavonoids biosynthesis, one of which is anthocyanin (Balakrishnan and Subramaniam 2009). In addition, PAL enzymes are also important in the formation of phenolic components that play a role in the recovery of wounds that occur in plants. Increased PAL activity can be expressed as a defense reaction in plants as protection against pathogen infection (Chet 1993; Seki et al. 1999; Cui et al. 2000; Logemann et al. 2000).
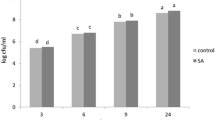
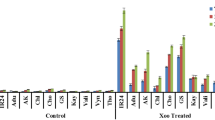
To monitor the PAL enzyme activity following Xoo infection, in vitro bioassay of PAL enzyme was performed by analyzing trans-cinnamic acid concentration with HPLC. The trans-cinnamic acid chromatogram is represented in Fig. 2. The results of the analysis showed that PAL enzyme activity in control treatment of all cultivars was the lowest. However, mock and infection treatment results in increasing activity, which was highest in the inoculated plants (Table 2) indicating that PAL enzyme activity was induced by wounding. The comparison between treatments showed an increase trend of PAL enzyme activity in every cultivar. This result confirmed that Xoo infection affects the expression of PAL as one of the defense systems in black rice plant.
Interestingly, PAL enzyme activity was consistent with the AUDPC value in all cultivars tested. Java14 cultivar which is known as the resistant cultivar, has the highest PAL enzyme activity (90.41 mM/g) whereas IR64 cultivar which was the susceptible cultivar has the lowest PAL enzyme activity (4.80 mM/g). In addition, Pari Ireng which has a higher content of anthocyanin as compared to Melik also has a higher PAL enzyme activity compared to Melik. The activity of PAL enzyme also increased significantly in Pari Ireng (72%) as well as Melik (66%) cultivars following Xoo infection. The dramatic change of PAL activity happened in Java 14 in control treatment (11.56 mM/g) to mock (48.43 mM/g) and infection treatment (90.41 mM/g) indicating that PAL enzyme plays important role in the defense system of the black rice following Xoo infection. This kind of mechanism did not occur in Pari Ireng. Although Pari Ireng has the highest PAL enzyme activity in control treatment (16.00 mM/g), the change is not as high as Java14 following Xoo infection. The high level of PAL enzyme activity in Pari Ireng is probably due to the basal metabolism of this cultivar. Pari Ireng has the highest anthocyanin content among all cultivars tested, indicating that a precursor of PAL enzyme may be involved in the biosynthesis of PAL. In other words, the defense mechanism of black rice (in PAL pathway) may be slightly different with Java14 as compared to the other cultivar studied, but the PAL pathway may still contribute to the defense. This is supported by low PAL activity in IR64 and IRBB21. In addition, the regulation of PAL activity in plants is increasingly complex due to many PAL encoding genes, some of which are only expressed in specific tissues or only under certain environmental conditions (Emiliani et al. 2009). Therefore, further study elucidating PAL encoding genes in different black rice cultivars especially in Pari Ireng cultivar is interesting and necessary.
Protein profile of black rice following Xoo infection
Plant attacked by viruses, bacteria, pathogens, parasites, and herbivorous insects will have an impact on plant imbalances (Pimentel 2001) and respond to these stresses by producing several defense compounds and protein such as pathogenesis-related (PR) proteins (Edreva 2005). To this end the protein profile of black rice to find out the full description of the regulated protein as a defense response caused by Xoo bacterial infection was performed. Protein analysis in this study was done by separating proteins using electrophoresis method SDS-PAGE 1 dimension. Separation of proteins with electrophoresis aims to observe if there is a difference in protein profile produced by infection of Xoo bacteria.
Based on the protein profile (Fig. 3), there are no big differences between all cultivars as well as in each treatment. The protein bands ranged from 8 kDa to 91 kDa and showed some similarity. However, a distinctive protein band can be observed in Pari Ireng and Java14 (ranging from 48 to 75 kDa) in infection treatment, indicating that protein may play a role in defense mechanism (red box in Fig. 3). Interestingly, at 75 kDa there is a thick band which may correspond as PAL protein which has molecular weight of 75 kDa (UniProt). This assumption needs further confirmation by protein sequencing. Nevertheless, further study is needed to indentify more protein possibly involved in the defense mechanism of black rice following Xoo infection.
Conclusions
PAL contributes to the resistance mechanism of black rice by increasing its activity in the cells following Xoo infection. The activity of PAL enzyme increased significantly in Pari Ireng (72%) as well as Melik (66%) cultivars following Xoo infection. However, the activity of PAL enzyme in Pari Ireng cultivar was higher than in Melik. The value of AUDPC, PAL activity, and protein profile analysis confirmed that black cultivar Pari Ireng was more resistant than Melik cultivar following Xoo infection.
Change history
10 December 2019
The original article was published with a spelling error to contributor name Rofiatun Solekha.
References
Agrios GN (2005). Plant pathology 5 edition. Elsevier: Academic Press-London, UK
Balakrishnan V, Subramaniam S (2009) Early investigation on cloning of a phenylalanine ammonia-lyase (PAL) promoter gene fragment from Dendrobium crumenatum. Nat Appl Sci 3:314–323
Bradford M (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein dye binding. Anal Biochem 72:248–254
Chet I (1993) Biotechnology in plant disease control. WileyLiss, New York
Cui Y, Bell AA, Joost O, Magill C (2000) Expression of potential defense response genes in cotton. Physiol Mol Plant Pathol 56:25–31
Darmanti S. (2015). The tolerance of soybeans [Glycine max (l.) merr. cv. grobogan] against interference of Cyperus rotundus L.in drought-stressed conditions. Ph.D. Thesis. Universitas Gadjah Mada. Indonesia
Dixon RA (2001) Natural products and plant disease resistance. Nature. 411:843–847
Dixon RA, Paiva NL (1995) Stress induced phenilpropanoid metabolism. Plant Cell 7:1085–1097
Djatmiko HA, Budi P, Nur P (2011) Determination of genetic pathotype and diversity of Xanthomonas oryzae pv. oryzae on rice plants in the residency area of Banyumas. Jurnal Hama Penyakit Tanaman Tropika 11(1):35–46
Edreva A (2005) Pathogenesis-related proteins:research progress in the last 15 years. Gen App Plant Physiol 31:20
Emiliani G, Fondi M, Fani R, Gribaldo S (2009) A horizontal gene transfer at the origin of phenylpropanoid metabolism: a key adaptation of plants to land. Biol Direct 4(7):1–12
Gnanamanickam SS, Priyadarisini VB, Narayanan NN, Vasudevan P, Kavitha S (1999) An overview of bacterial blight disease of rice and strategies for its management. Curr Sci 77:1435–1443
Kadir TS (2009) Ward off BLB by rotating varieties. Warta Penelitian dan Pengembangan Pertanian 31(5):1–3
Kauffman HE, Reddy APK, Heish SPY, Meera SD (1993) An improved technique for evaluation of resistance to rice varieties to Xanthomonas oryzae. Plant Dis Rep 65:663–666
Khunsook S, Bary SB, Susan RMG, Jack AA (2003) Purification and characterization of plasma membrane-associated human sperm α-L-fucosidase. Biol Reprod 709:716
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 227:680–685
Logemann E, Tavernaro A, Shulz W, Somssish IE, Hahlbrock K (2000) Uv light selectively co induces supply pathway from primary metabolism and flavonoid secondary product formation in parsley. Proc Natl Acad Sci U S A 97:1903–1907
Mahmood T, Jan A, Kakishima M, Komatsu S (2006) Proteomic analysis of bacterial-blight defense responsive proteins in rice leaf blades. Proteomics. 6:6053–6065
Pascuzzi IAS, McCouch SR (2007) Recessive resistance genes and the Oryza sativa-Xanthomonas oryzae pv. oryzae pathosystem. Mol Plant Mic Intr 20(7):731–739
Pimentel D. (2001). Biological Invasions: Economic and Environmental Costs of Alien Plant, Animal and Microbe Species. College of Agriculture and Life Sciences. Cornell University, Ithaca, NY 14850–0901, USA, 384
Rabilloud TA (1992) Comparison between low background silver diammine and silver nitrate protein stains. Electrophoresis. 13(7):429–439
Ritter H, Schulz GE (2004) Sructural basis for the entrance into the phenylpropanoid metabolism catalyzed by phenylalanine ammonia-lyase. Plant Cell 16:3426–3436
Seki H, Nagasugi Y, Shiraishi T, Yamada T (1999) Changes in vivo dna-protein interactions in pea phenylalanine ammonia-lyase and chalcone synthase gene promoter induced by fungal signal molecules. Plant Cell Physiol 40:88–95
Song A, Xue G, Cui P, Fan F, Liu H, Yin C, Sun W, Liang Y (2016) The role of silicon in enhancing resistance to bacterial blight of hydroponic- and soil-cultured rice. Sci Rep 6:24640
Susanto U, Sudir (2012) Ketahanan genotipe padi terhadap Xanthomonas oryzae pv. oryzae patotype iii, iv, and viii. Jurnal Penyakit dan Perlindungan Tanaman Pertanian 31(2):108–116
Tjubarat TTS, Kadir E, Sumadi (1999) Skrining Varietas Terhadap Hawar Daun Bakteri. Prosiding Kongres Nasional XV dan Seminar Ilmiah FPI, Purwokerto
Yasmin S, Zaka A, Imran A, Zahid MA, Yousaf S, Rasul G, Arif M, Mirza MS (2016) Plant growth promotion and suppression of bacterial leaf blight in rice by inoculated bacteria. PLoS One 11(8):e0160688. https://doi.org/10.1371/journal.pone
Acknowledgements
We thank Dr. Kristamtini from the Institute for Agricultural Technology (BPTP) Yogyakarta for providing the black rice seeds used in this study. We express our gratitude to Dr. Sudir from the Indonesian Center for Rice Research Sukamandi, Subang, West Java for providing the Xoo isolates well as the white rice seeds used in this study.
Funding
This research was funded by the Ministry of Research, Technology, and Higher Education The Republic of Indonesia by Universities Leading Research Project 2016 to YAP (contract number 863/UN1-P.III/LT/DIT-LIT/2016) and Program Penguatan Kelembagaan Pusat Studi Menuju Keunggulam Akademik UGM Tahun Anggaran 2018 (contract number 4104/UNI/DITLIT/DIT-LIT/LT/2018).
Author information
Authors and Affiliations
Contributions
RS, FAS performed the field and laboratory works and analyzed the data, All authors wrote the manuscript. YAP, TJ, TRN conceived and designed the study. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Research involving human participants and/or animals
Not Applicable.
Informed consent
Not Applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Solekha, R., Susanto, F.A., Joko, T. et al. Phenylalanine ammonia lyase (PAL) contributes to the resistance of black rice against Xanthomonas oryzae pv. oryzae. J Plant Pathol 102, 359–365 (2020). https://doi.org/10.1007/s42161-019-00426-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-019-00426-z