Abstract
Purpose of Review
Acceleration of biological processes of aging is hypothesized to drive excess morbidity and mortality in socially disadvantaged populations. DNA methylation measures of biological aging provide tools for testing this hypothesis.
Recent Findings
Next-generation DNA methylation measures of biological aging developed to predict mortality risk and physiological decline are more predictive of morbidity and mortality than the original epigenetic clocks developed to predict chronological age. These new measures show consistent evidence of more advanced and faster biological aging in people exposed to socioeconomic disadvantage and may be able to record the emergence of socially determined health inequalities as early as childhood. Next-generation DNA methylation measures of biological aging also indicate race/ethnic disparities in biological aging. More research is needed on these measures in samples of non-Western and non-White populations.
Summary
New DNA methylation measures of biological aging open opportunities for refining inference about the causes of social disparities in health and devising policies to eliminate them. Further refining measures of biological aging by including more diversity in samples used for measurement development is a critical priority for the field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Individuals who are socioeconomically disadvantaged or marginalized based on their racial/ethnic identity tend to develop aging-related diseases at younger ages and suffer earlier mortality as compared to individuals who are wealthier and White [1, 2]. Macro-structural social determinants of health, including racism, classism, sexism, and their intersections, drive these disparities [3, 4••]. One mechanism hypothesized to link social determinants of health with a shorter healthy lifespan is an acceleration of biological processes of aging [5, 6].
Biological aging is the gradual and progressive decline in system integrity that occurs with advancing age [7]. This age-dependent decline in system integrity is thought to arise from an accumulation of molecular changes, known as hallmarks, that undermine the functioning of molecular networks and organ systems, driving vulnerability to disease and death [8]. Now, the emerging field of geroscience aims to prevent and treat disease through intervention on these hallmarks. The core hypothesis of geroscience is that slowing or reversing the molecular hallmarks of aging can slow or reverse the decline in system integrity, preventing or delaying disease and disability [8, 9].
The geroscience field has been pioneered by researchers studying model organisms under laboratory conditions and is focused on developing clinical treatments for diseases of aging [10,11,12,13]. However, there are important connections between the basic biology of aging and social determinants of health in humans. Many of the molecular changes that form the basis of aging, including cell senescence, inflammation, mitochondrial dysfunction, and epigenetic alternations, are also affected by environmental exposures ranging from chemical toxicants to social stressors that are concentrated in socially disadvantaged populations [14,15,16,17,18]. Geroscience, therefore, promises new opportunities for understanding the causes of social gradients in health and can help devise new strategies for building aging health equity.
Realizing the promise of geroscience to improve human health requires an integration of aging biology with behavioral and social sciences [19]. However, studies to investigate biological aging as a mediator of social determinants of health have faced the barrier that the molecular changes that form the biological basis of aging are difficult to observe in epidemiologic studies. There is no gold standard measure of aging and, historically, a little consensus around valid aging biomarkers [20, 21]. Now, this is beginning to change. A new family of measurements based on analysis of DNA methylation shows promise and opens new opportunities for the integration of research into aging biology and social determinants of health.
In this article, we review progress in applications of DNA methylation–based measures of biological aging to study how socially determined inequalities drive disparities in healthy aging. Early studies applying these new DNA methylation measures suggest opportunities for refining inference about the causes of social disparities in health and devising programs and policies to eliminate them. Specifically, because these new measures can reveal differences in the progress and pace of aging decades before chronic diseases become established, they can help isolate when in the life course and through what specific exposures health inequalities in aging become established. In addition, these new measures can inform the evaluation of interventions to address health inequalities by providing a readout on the short- and medium-term effects of programs and policies in “pre-symptomatic” individuals who have not yet begun to manifest aging-related disease and disability.
This promise comes within the context of an important limitation: DNA methylation measurements of biological aging have often been developed in convenience samples not designed to represent particular populations. Even when samples are socioeconomically representative, they represent populations that are overwhelmingly White. This underrepresentation of non-White individuals parallels data gaps noted in human genetics research [22, 23]. Despite the reliance on mostly White samples for measurement development, DNA methylation measures of biological aging tend to show similar magnitudes of association with risk for disease, disability, and mortality across different race/ethnic groups within the USA [24,25,26,27]. Where there are differences, associations tend to be somewhat stronger in White as compared with non-White samples. Research in more non-White samples and in diverse cohorts is needed to better establish the validity of DNA methylation measures of biological aging in populations of diverse genetic ancestries and race/ethnic, social identities. Further development of aging measures in more diverse samples is a priority.
The remainder of this review is organized as follows: the “Biological Aging and Social Determinants of Health” section provides a conceptual overview of how social determinants of health affect biological aging. The “Quantification of Biological Aging for Social Determinants of Health Research” section introduces approaches to the measurement of biological aging in social determinants of health research, with a focus on recently introduced methods based on analysis of DNA methylation. The “DNA Methylation Clocks and Social Determinants of Health” section reviews recent work testing how social determinants of health are associated with DNA methylation measures of aging. A key finding from these studies is that the new generation of DNA methylation measures of biological aging derived from analysis of mortality risk and physiological decline, which are more predictive of morbidity and mortality, are also more strongly associated with social determinants of health as compared with the original epigenetic clocks developed from analysis of chronological age. The “Challenges and Recommendations” section reviews limitations of existing DNA methylation measures of biological aging and makes recommendations to overcome them with the goal of maximizing the utility of measures of biological aging in promoting aging health equity.
Biological Aging and Social Determinants of Health
How Do Social Determinants of Health Affect Biological Aging?
Healthspan and lifespan disparities at the intersection of socioeconomic status and socially constructed dimensions of race/ethnicity, as well as other identity characteristics, are profound [4••, 28]. Exposure to environmental toxicants, opportunities for restorative leisure and exercise, physical and psychological safety, social support, and access to nutritious food and healthcare, among other factors, differ across these social positions [29, 30]. In turn, these differences in health-damaging exposures and health-promoting resources drive biological changes that contribute to more or less healthy aging [18, 31,32,33,34,35]. Connections among social identities, mechanisms of inequality, processes of biological aging, and disparities in healthy aging are illustrated in Fig. 1.
The social environment is associated with multiple environments that affect health across the lifespan. Epidemiological research has documented healthspan and lifespan disparities across dimensions of social identities (e.g., social class and racial/ethnic identity; blue circle). Mechanisms of social inequality (e.g., income inequality and policing; red circle) lead to disparate access to health-enhancing resources (e.g., nutrition, leisure; yellow circle) and disparate exposure to health risks (e.g., toxicants, stress) between social identities. These cause social disparities in aging-related disease, disability, and mortality (green circle), which may reinforce dimensions of social inequality in the next generation
When in Development Do Social Determinants of Health Affect Biological Aging?
Social determinants of health clearly affect aging in later life. People living in or near poverty and those with marginalized racial/ethnic identities experience more rapid functional decline, earlier accumulation of disease and disability, and earlier mortality [36,37,38,39,40]. The processes driving these later-life disparities begin much earlier in the life course; a range of adverse early-life conditions, including socioeconomic disadvantage, maltreatment by caregivers, and unsafe or unstable living conditions, are associated with a shorter lifespan, earlier onset of aging-related disease, and more rapid decline in physiological integrity from young adulthood to midlife [41,42,43,44,45]. A possible mechanism linking early-life social determinants with unhealthy aging is an acceleration of biological aging.
The biological process of aging, which is characterized by a breakdown in resilience mechanisms, damage accumulation, and loss of system integrity, is distinct from programmed development, which assembles reproductively viable life [46]. However, accumulation of molecular damage commences at the very earliest stages of development, suggesting the possibility that aging is ongoing almost from conception [47, 48]. Observations of biological aging at the early stages of development are few. However, epigenetic marks associated with aging are removed from genomes during embryogenesis and begin to accumulate thereafter, suggesting that aging may indeed begin at the earliest stages of life [49]. And there is substantial evidence for the effects of the prenatal environment on outcomes in aging [50]. However, molecular analysis of aging in early-life humans remains limited. Blood analysis of telomere length, a biomarker of cellular aging [51, 52], indicates more advanced/faster aging in human infants exposed to perinatal adversity and in children exposed to early-life adversity [53,54,55,56,57]. But the science of telomere length as a true biomarker of aging remains unsettled [58,59,60]. Even if the early accumulation of molecular damage represents a disruption to development rather than the onset of aging, such damage may still condition the rate of aging later in life as a consequence of reduced resilience capacity [61]. Therefore, social determinants of health may affect the biological processes of aging from very early in development.
Quantification of Biological Aging for Social Determinants of Health Research
Measurement Approaches Across Biological Levels of Analysis
There is no gold standard measure of aging [21, 62]. Several approaches have been proposed at different levels of biological organization [63]. A conceptual overview of the progression of aging across levels of analysis and measures associated with different levels is presented in Fig. 2.
Levels of analysis of biological aging. Biological aging is the gradual and progressive decline in system integrity that occurs with advancing age. The figure illustrates the progression of biological aging across levels of analysis, from an accumulation of molecular changes to declines in organ system integrity, functional decline, disease, disability, and mortality. Measures of biological aging can be implemented at different levels of analysis: Molecular changes are commonly measured as omics clocks,1 telomere attributes,2 and mitochondrial DNA copy number.3 Decline in system integrity is commonly quantified as allostatic load measures4 and in blood chemistry clocks.5 Functional decline is typically measured with various frailty indices.6 DNA methylation measures of biological aging are implemented at the molecular level (i.e., omics clocks). Some DNA methylation measures, including the PhenoAge and GrimAge and the DunedinPoAm pace of aging, also incorporate information from the level of organ system integrity
The level of biological organization most proximate to disease, disability, and death is commonly measured using indices of organism-level functional capacities. These include tests of balance, walking speed, strength, and cognitive performance, as well as summary scores counting deficits in functional domains and biological systems known as frailty indices [64, 65]. Measurements based on deficits in functional capacities and frailty are most sensitive to changes occurring at the end of life when aging processes are advanced.
Beneath this organism-level functional capacity is the process of decline in system integrity, commonly measured using indices of organ- and organ-system-level functions. Broadly, there are three types of these indices. One type consists of counts of deficits in physiological parameters, including blood chemistry analytes and organ function tests, such as allostatic load indices [66, 67]. A second type consists of blood chemistry clocks and related algorithms that combine continuous information from multiple blood chemistry analytes and organ function tests to estimate the state of system integrity in an organism [68,69,70,71,72]. A third type uses longitudinal data on blood chemistry analytes and organ function tests to model the rate of decline in system integrity, such as Pace of Aging measures [73,74,75]. Measurements based on organ- and organ-system-level functions are sensitive to aging-related changes from young adulthood when trajectories of aging-related decline in system integrity begin to take shape. These types of measurements all show clear evidence of socioeconomic and racial/ethnic disparities [45, 71, 76,77,78].
Within the geroscience model, accumulating molecular changes underpin declines in system integrity. These molecular changes are abundant, and most are challenging to measure in humans. For example, telomere attrition and mitochondrial dysfunction are among the hallmarks of aging and are theorized as mediators of early-life adversity effects on aging [56, 79]. But telomere- and mitochondria-related measurements easily quantified in the blood are imperfect biomarkers of aging hallmarks [58, 59, 80, 81]. Expression of p16INK4a is linked with cellular senescence, but may be most informative about aging when measured in specific lymphocyte subpopulations [82, 83]. The development of mechanistic biomarkers of aging hallmarks that can be assayed in studies of humans remains a work in progress [84, 85].
At present, the most promising molecular-level biomarkers of aging for clinical and epidemiologic studies have emerged from “omics”-based approaches that capture biological changes downstream of mechanistic hallmarks of aging. Recent developments in proteomic and metabolomic analyses suggest promise [86,87,88]. Currently, the best-established omics-based biomarkers of aging processes are based on analysis of genome-wide patterns of DNA methylation, in particular a family of measurements broadly known as “clocks” [89••].
DNA Methylation Clock Measures of Biological Aging
DNA methylation marks are chemical tags on the DNA sequence that contribute to the regulation of gene expression (90). DNA methylation states are dynamic across the life course. As humans age, they experience a global loss of DNA methylation [91]. However, at specific sites on the genome, methylation marks are both gained and lost with age [92]. These site-specific changes with aging are so regular across individuals it is possible, through machine learning analysis, to develop algorithms that can predict a person’s chronological age from DNA methylation analysis to a precision of within a few years [93,94,95]. These algorithms, introduced in the early 2010s, became known as “clocks.” Clocks have received substantial attention in aging research because of the hypothesis that clock ages that are older than a person’s chronological age indicate an advanced state of biological aging, whereas clock ages younger than a person’s chronological age indicate delayed biological aging [89••].
DNA methylation clocks have so far progressed through two generations of development, with further generations now emerging. The first generation of clocks were developed by comparing older individuals to younger ones. For these clocks, the goal of machine learning analysis was to predict how many years a person had lived up to the time their DNA were collected, i.e., their chronological age. These “first-generation” clocks are modestly predictive of mortality [96], but less consistent in predictions of other aging-related phenotypes, including disease, disability, and physiological and functional decline [97,98,99,100]. Moreover, the most precise clocks, those for which chronological age predictions were closest to the truth, have tended to be less predictive of health and mortality [101].
The second generation of clocks were developed by comparing individuals based on survival [25, 71, 102]. For these second-generation clocks, the goal of machine learning analysis was to predict how many years a person would continue to live following the collection of their DNA, i.e., remaining lifespan. The most prominent of these, the PhenoAge and GrimAge clocks, include an intermediate step in which physiological features of aging are modeled from DNA methylation. In the case of the PhenoAge clock, mortality risk was first modeled from physiological markers and chronological age. This first-stage algorithm was then applied to a new sample in which it was modeled from DNA methylation to derive the final DNA methylation clock. In the case of the GrimAge clock, a set of physiological indicators were modeled from DNA methylation and then these DNA methylation predictions along with age, sex, and a DNA methylation prediction of smoking history were applied to model mortality. The resulting PhenoAge and GrimAge clocks are substantially more predictive of morbidity and mortality as compared to the first-generation clocks [103, 104].
A third generation of clocks is now emerging, including measures based on analysis of longitudinal within-person change [105••]. These clocks, referred to as Pace of Aging measures, are derived from the analysis of trajectories of physiological decline. The first stage of analysis models within-person change in a panel of physiological indicators. The second stage composites each participants’ rates of change across the panel of indictors to form a single index of their personal rate of physiological decline. Finally, this composite Pace of Aging is modeled from DNA methylation measured at the end of the follow-up interval to derive the final algorithm. Whereas the first- and second-generation clocks aim to predict how old a person is biological, Pace of Aging measures aim to predict how fast a person is aging. First- and second-generation clocks take on values interpretable as ages. The Pace of Aging measures take on values interpretable as rates. The Pace of Aging measures have not yet received the same level of research attention as the earlier clocks. But the available evidence suggests that they are comparably predictive of health risks to the other clocks, although they are not as predictive of mortality as GrimAge [27, 103, 105••].
Several other DNA methylation biomarkers have been developed, including those that measure mortality risk [102] and mitotic age [106, 107]. In some datasets, these biomarkers outperform the second-generation clocks in the prediction of morbidity and mortality [108]. But, to date, these measures have been less widely used in social determinants of health research.
The first two generations of DNA methylation clocks and the Pace of Aging measures are described in Table 1.
DNA Methylation Clocks and Social Determinants of Health
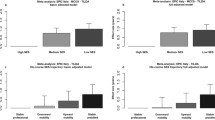
Research testing associations between social determinants of health and DNA methylation clocks is still in its early stages. There are not yet enough studies with consistent methods to undertake meaningful meta-analyses. However, some patterns are emerging. Specifically, while the first-generation DNA methylation clocks show weak and inconsistent associations with social determinants of health, later generations of measures show stronger and more consistent associations. Figure 3 graphs effect sizes for five DNA methylation measures of aging from analysis of socioeconomic inequality and racial/ethnic identity. Details of the studies are reported in Supplementary Table S1. The socioeconomic inequality measures are varied, ranging from educational attainment to socioeconomic disadvantage indices to neighborhood conditions. But the pattern of results is consistent. Overall, the GrimAge clock and DunedinPoAm Pace of Aging show the strongest associations with social determinants of health. Differences in these measures of biological aging between high and low socioeconomic status groups and between White and marginalized racial/ethnic groups are consistent with the hypothesis that social disadvantage contributes to an acceleration of biological aging.
Standardized effect sizes of associations of socioeconomic status (SES), education, and racial/ethnic identity with DNA methylation measures of biological aging. Effect sizes are reported in the metric of Cohen’s d. Effect sizes reported in the different studies were harmonized to this metric as follows: For studies that reported comparisons between groups, coefficients denominating group differences were divided by the standard deviation of the aging measure. In studies not reporting standard deviations [122, 152], we used the standard deviations reported by the US Health and Retirement Study [151]. For studies reporting associations between continuous measures of SES and aging, we first converted coefficients to the metric of Pearson’s r and then to Cohen’s d. Conversions to Pearson’s r were made by dividing the coefficient by the standard deviation of the aging measure and multiplying by the standard deviation of the SES measure. Details of the samples and measurements of the studies included in the figure are reported in Supplementary Table S1
Studies are also beginning to examine the question of how early in the life course socioeconomic patterning of DNA methylation measures of aging may emerge. A recent review of studies examining how socioeconomic disadvantage related to first-generation DNA methylation clocks in children found an inconsistent pattern of results [109]. An analysis of saliva DNA methylation in children that we published with the Texas Twin Project identified associations between both socioeconomic and White vs. Latinx identity differences in the DunedinPoAm Pace of Aging, but not the other clocks [110]. More studies of this question are needed to establish confidence in results. Studies including blood sample data will be especially valuable. In addition, new DNA methylation measures of biological aging have been developed in samples of children, including methods designed for tissues more readily available in pediatric samples [111,112,113]. Studies are needed to establish how these measures relate to family-level socioeconomic disadvantage.
In sum, based on the limited evidence available so far, the DNA methylation clocks that are more predictive of morbidity and mortality (i.e., second-generation clocks, third-generation Pace of Aging measures) are also more strongly associated with social determinants of health. It is not yet clear when in the life course these associations become established. However, social gradients in biological aging may already be evident during childhood. As DNA methylation data are better integrated into longitudinal studies, research can begin to test life-course models of how socioeconomic status in childhood and adulthood shape biological aging [27, 114, 115].
Challenges and Recommendations
Ancestry and Genetic Confounding
The emerging evidence linking social disadvantage to accelerated biological aging as measured by DNA methylation clocks must be interpreted within the context of several limitations. A first limitation has to do with the potential confounding of DNA methylation measurements of aging by genetic ancestry. There is a substantial bias in DNA-based research to study people solely of recent European ancestries [22, 23]. Genetic variation is an important determinant of DNA methylation states across the genome [116]. Genetic ancestry differences, therefore, have the potential to generate artifacts in DNA methylation datasets [117]. Genetic variants that affect DNA methylation and that have a low frequency or are absent in European-ancestry populations but are more common in other populations, therefore, have the potential to generate bias or noise in DNA methylation clock measures of aging.
While genetic ancestry is not the same thing as socially constructed racial/ethnic identity, people solely of recent European ancestries are likely to identify as White [118]. The samples used to develop the DNA methylation measures of aging that are most predictive of health and mortality and most sensitive to social disadvantage (PhenoAge and GrimAge clocks and DunedinPoAm) are mostly or entirely White [24, 25, 105••]. Establishing that these widely used clocks represent comparably valid measurements of aging across race/ethnic groups is a priority, as is the development of DNA methylation measures of aging from more diverse samples. Progress is now being made to establish validity across ancestry populations, including in the multi-ethnic samples in the US Health and Retirement Study [27], and work within non-White samples such as the American Indian participants of the Strong Heart Study [119], African-Americans in the Strong African American Healthy Adults Project [120], the Chinese National Twin Registry [121], and others.
So far, there is limited evidence for genetic ancestry-related confounding in DNA methylation clock research. Effect sizes for clock associations with healthy aging phenotypes are similar between groups of Black and White Americans, although effect sizes tend to be slightly larger for White Americans [24, 25, 27, 122]. The inclusion of genetic principal components as covariates in the analysis may provide some correction for ancestry-related artifacts in DNA methylation data [123]. Going forward, studies that use DNA methylation clocks to test differences in biological aging between socially constructed racial/ethnic identity groups that differ in genetic ancestry should be cautious in their interpretation of data. Designs that incorporate measures of healthy aging endpoints to establish parallel criterion validity of clocks between ancestry groups can build confidence in inferences that group differences in DNA methylation measures indicate differences in biological aging.
Longitudinal Analysis and Measurement Reliability
A second limitation is that nearly all research to date relating social determinants of health to DNA methylation measures of aging relies on cross-sectional data, at least for measurements of the aging outcomes. (This limitation also applies to nearly all studies relating DNA methylation measures of aging to health and mortality.) Research on human development establishes that differences in development and aging between individuals observed from a single time point of data can be a poor representation of the changes that occur within individuals over time [124,125,126]. It remains unclear whether or how DNA methylation measures of aging may be modified [127]. Social determinants of health research into biological aging are premised on the idea that interventions to modify social circumstances can slow the pace of aging and contribute to the elimination of health disparities. A critical next step is for longitudinal studies with repeated measures of DNA methylation to establish if changes in social determinants of health are associated with changes in DNA methylation measures of aging.
Two key challenges facing longitudinal repeated-measures studies of DNA methylation clocks are assay batch effects and technical reliability concerns. Batch effects refer to the variation in measurements arising from features of the measurement process that are shared among groups of samples measured together and different between groups of samples measured separately, such as samples grouped on assay plates or which DNA extractions or bisulphite conversions were performed at different times. The issue of batch effects in DNA methylation has been well described, and a number of corrections have been proposed [128]. However, these corrections are not perfect and have the potential to induce biases of their own [129,130,131]. Therefore, repeated measures analysis based on DNA methylation datasets in which time point is fully confounded by assay batch must be interpreted with caution.
Even when repeated measures are generated from the same assay batch (when repeated DNA samples from an individual are extracted and bisulphite-converted together and assayed on the same plate), low test–retest reliability of DNA methylation measurements can present challenges. DNA methylation arrays generate highly reliable genome-wide measurements of total DNA methylation [132]. However, at the level of individual CpG sites, the dinucleotide locations on the genome at which DNA methylation levels are assayed, reliabilities are strikingly poor [133,134,135]. DNA methylation clocks and Pace of Aging measures are algorithms that combine information on the methylation states of dozens to hundreds of CpG sites. Clock CpGs tend to have somewhat higher reliabilities than the average [135], and the clocks themselves are substantially more reliable than the individual CpGs from which they are composed [136]. Nevertheless, test–retest reliability as measured by the intraclass correlation coefficient (ICC) for most clocks is well below 0.9 [136]. This suggests that at least 20% of the variation in most clock measurements is error or noise. In an analysis of change across two time points, measurement error is additive. A consequence is that the statistical signal arising from an effect of social determinants of health on biological aging will be significantly diluted. The GrimAge clock and DunedinPACE measure both have ICCs well above 0.9 and so may be less subject to this limitation [137]. New methods may substantially increase the reliabilities of other clocks [136].
Finally, DNA methylation measures that are well established to predict morbidity and mortality and correlate with social determinants of health were developed for blood tissue. Methylation varies substantially by tissue type [138]. However, it may be infeasible to collect blood samples within many large cohort studies. Blood collections typically require medical personnel that would further increase the already high cost of DNA methylation sampling. In contrast, saliva samples are more amenable to large-scale studies, including pediatric participants. While some measures have shown high correspondence between blood and saliva samples [139], more research is needed to establish associations of DNA methylation measures of aging taken in non-blood tissues with healthy aging endpoints, such as morbidity and mortality, before correlations between these measurements and social determinants of health can be interpreted with confidence.
Future Directions
Within the bounds of these challenges, there is clear evidence that socially disadvantaged individuals show more advanced and faster biological aging as compared to more socially advantaged individuals of the same chronological age. The next steps in research to establish the effects of social determinants on biological aging involve refinements to study designs to strengthen internal and external validity.
Using Natural Experiment and Randomized Trial Designs to Improve Internal Validity
Research relating social determinants of health to DNA methylation measures of aging consists mostly of correlational study designs with limited ability to establish causality of associations. So-called “natural experiments,” in which an event or policy change alters socioeconomic circumstances for a segment of the population, provide one path to strengthening causal inference [140]. Changes over time and/or differences across borders in schooling reforms mandating additional years of education, minimum wage thresholds, or other anti-poverty policies are settings in which to investigate causal effects of social determinants on biological aging. Randomized controlled trials of anti-poverty interventions and other programs to address social determinants represent another promising research direction [141, 142].
Developing Population-Representative Samples to Improve External Validity
Participation in biomedical research in general and DNA-based research, in particular, tends to be lower for persons with less education and members of a certain race/ethnic identity groups [143,144,145]. Underrepresentation of such groups is especially pronounced in large biobank datasets, and although there are many strategies to address this challenge, none are perfect and some may induce their own biases [146,147,148,149,150]. Oversampling of underrepresented populations and the application of survey probability weights can help generate more population-representative estimates [151]. However, a concern is that low socioeconomic status and non-White individuals who choose to participate in DNA research may differ from those who do not in ways that may be consequential for aging, resulting in selection bias. There has not yet been systematic consideration of these types of selection bias issues in relation to measures of biological aging. As the field matures out of its early days, closer attention is needed to which individuals and groups may be missing or underrepresented in existing samples. This need is being recognized and addressed with initiatives such as National Institute on Aging and National Institute on Minority Health and Health Disparities priority funding for social epigenomics research (https://www.nimhd.nih.gov/programs/extramural/investigator-initiated-research/socioepigenomics-grants.html). Data generated under this initiative and other datasets from underrepresented groups and populations should receive the careful attention to evaluate the extent to which findings accumulated in samples of mostly White and higher socioeconomic status individuals replicate in different and more diverse samples.
Conclusion
Novel measures quantified in DNA methylation are being used to capture processes of biological aging in ways that may inform why and how social inequality is associated with aging-related disparities in health. These new tools have the potential to help evaluate how exposures contribute to risk in people who are still “pre-symptomatic” and ultimately may provide surrogate end points for testing the effects of social programs on healthy aging decades before effects on aging-related chronic disease or mortality would be apparent.
References
Papers of particular interest, published recently, have been highlighted as: •• Of major importance
Pérez-Stable EJ, Collins FS. Science visioning in minority health and health disparities. Am J Public Health. 2019 [cited 2021 Feb 17];109(S1):S5–S5. Available from: https://doi.org/10.2105/AJPH.2019.304962
Snyder-Mackler N, Burger JR, Gaydosh L, Belsky DW, Noppert GA, Campos FA, et al. Social determinants of health and survival in humans and other animals. Science. 2020 [cited 2020 Jun 3];368(6493). Available from: https://science.sciencemag.org/content/368/6493/eaax9553
Creanga AA. Maternal mortality in the United States. Clin Obstet Gynecol. 2018;61(2):1.
••Krieger N. Measures of racism, sexism, heterosexism, and gender binarism for health equity research: from structural injustice to embodied harm—an ecosocial analysis. Annu Rev Public Health. 2020;41(1):37–62. This review provides a thorough overview of dimensions and mechanisms of social dimensions of health.
Geronimus AT. The weathering hypothesis and the health of African-American women and infants: evidence and speculations. Ethn Dis. 1992;2(3):207–21.
Vineis P, Kelly-Irving M, Rappaport S, Stringhini S. The biological embedding of social differences in ageing trajectories. J Epidemiol Community Health. 2016;70(2):111–3.
Kirkwood TBL. Understanding the Odd Science of Aging. Cell. 2005;120(4):437–47.
López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The Hallmarks of Aging. Cell. 2013;153(6):1194–217.
Kennedy BK, Berger SL, Brunet A, Campisi J, Cuervo AM, Epel ES, et al. Geroscience: linking aging to chronic disease. Cell. 2014;159(4):709–13.
Barzilai N, Cuervo AM, Austad S. Aging as a biological target for prevention and therapy. JAMA. 2018;320(13):1321–2.
Campisi J, Kapahi P, Lithgow GJ, Melov S, Newman JC, Verdin E. From discoveries in ageing research to therapeutics for healthy ageing. Nature. 2019 [cited 2019 Jul 16];571(7764):183. Available from: https://www.nature.com/articles/s41586-019-1365-2
Justice JN, Nambiar AM, Tchkonia T, LeBrasseur NK, Pascual R, Hashmi SK, et al. Senolytics in idiopathic pulmonary fibrosis: results from a first-in-human, open-label, pilot study. EBioMedicine. 2019 [cited 2020 Feb 13];40:554–63. Available from: http://www.sciencedirect.com/science/article/pii/S2352396418306297
Kaeberlein M, Galvan V. Rapamycin and Alzheimer’s disease: time for a clinical trial? Science Translational Medicine. 2019 [cited 2019 May 9];11(476):eaar4289. Available from: https://stm-sciencemag-org.ezproxy.cul.columbia.edu/content/11/476/eaar4289
Bastain T, Breton C, Farzan S, Habre R, Johnston J, Tabor DC, et al. Physical environment, and minority health and health disparities research. In: The Science of Health Disparities Research. Wiley; 2021 [cited 2021 May 23]. p. 95–108. Available from: https://doi.org/10.1002/9781119374855.ch6
Crimmins EM. Social hallmarks of aging: suggestions for geroscience research. Ageing Res Rev. 2020 [cited 2021 Feb 15];63:101136. Available from: https://www.sciencedirect.com/science/article/pii/S1568163720302713
Emeny RT, Carpenter DO, Lawrence DA. Health disparities: intracellular consequences of social determinants of health. Toxicol Appl Pharmacol. 2021 [cited 2021 May 23];416:115444. Available from: https://www.sciencedirect.com/science/article/pii/S0041008X2100051X
Epel ES. The geroscience agenda: Toxic stress, hormetic stress, and the rate of aging. Ageing Res Rev. 2020[cited 2021 Feb 15];63:101167. Available from: https://www.sciencedirect.com/science/article/pii/S1568163720303020
Peters A, Nawrot TS, Baccarelli AA. Hallmarks of environmental insults. Cell. 2021 [cited 2021 May 13];184(6):1455–68. Available from: https://www.sciencedirect.com/science/article/pii/S0092867421000866
Moffitt TE. Behavioral and social research to accelerate the geroscience translation agenda. Ageing Res Rev. 2020 [cited 2021 Jan 11];63:101146. Available from: http://www.sciencedirect.com/science/article/pii/S1568163720302816
Cohen AA, Kennedy BK, Anglas U, Bronikowski AM, Deelen J, Dufour F, et al. Lack of consensus on an aging biology paradigm? A global survey reveals an agreement to disagree, and the need for an interdisciplinary framework. Mech Ageing Dev. 2020 [cited 2020 Aug 13];111316. Available from: http://www.sciencedirect.com/science/article/pii/S0047637420301123
Ferrucci L, Gonzalez‐Freire M, Fabbri E, Simonsick E, Tanaka T, Moore Z, et al. Measuring biological aging in humans: a quest. Aging Cell. 2020 [cited 2020 Aug 7];19(2):e13080. Available from: https://doi.org/10.1111/acel.13080
Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet. 2019;51(4):584–91.
Mills MC, Rahal C. A scientometric review of genome-wide association studies. Commun Biol. 2019 [cited 2020 Feb 12];2(1):1–11. Available from: https://www.nature.com/articles/s42003-018-0261-x
Levine ME, Lu AT, Quach A, Chen BH, Assimes TL, Bandinelli S, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY). 2018 [cited 2018 May 23];10(4):573–91. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5940111/
Lu AT, Quach A, Wilson JG, Reiner AP, Aviv A, Raj K, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY). 2019;11(2):303–27.
Schmitz LL, Zhao W, Ratliff SM, Goodwin J, Miao J, Lu Q, et al. The socioeconomic gradient in epigenetic aging clocks: evidence from the multi-ethnic study of atherosclerosis and the health and retirement study. medRxiv. 2021[cited 2021 Mar 26];2021.03.01.21252660. Available from: https://doi.org/10.1101/2021.03.01.21252660v1
Graf GH, Crowe CL, Kothari M, Kwon D, Manly JJ, Turney IC, Valeri L, Belsky DW. Testing Black-White disparities in biological aging in older adults in the United States: analysis of DNA-methylation and blood-chemistry methods. American J Epidemiol 2021. https://doi.org/10.1093/aje/kwab281
Cogburn CD. Culture, race, and health: implications for racial inequities and population health. Milbank Q. 2019;97(3):736–61.
Goosby BJ, Cheadle JE, Mitchell C. Stress-related biosocial mechanisms of discrimination and African American health inequities. Ann Rev Sociol. 2018;44(1):319–40.
Williams DR, Priest N, Anderson NB. Understanding associations among race, socioeconomic status, and health: patterns and prospects. Health Psychol. 2016;35(4):407–11.
Febbraio MA. Health benefits of exercise — more than meets the eye! Nat Rev Endocrinol. 2017 Feb [cited 2021 May 25];13(2):72–4. Available from: https://www.nature.com/articles/nrendo.2016.218
Hodes RJ, Sierra F, Austad SN, Epel E, Neigh GN, Erlandson KM, et al. Disease drivers of aging. Ann N Y Acad Sci. 2016 [cited 2019 Feb 7];1386(1):45–68. Available from: https://doi.org/10.1111/nyas.13299
Picard M, McEwen BS. Psychological stress and mitochondria: a conceptual framework. Psychosom Med. 2018 [cited 2021 May 25];80(2):126–40. Available from: https://journals.lww.com/psychosomaticmedicine/Fulltext/2018/02000/Psychological_Stress_and_Mitochondria__A.2.aspx
Santos AL, Sinha S. Obesity and aging: molecular mechanisms and therapeutic approaches. Ageing Res Rev. 2021 [cited 2021 May 25];67:101268. Available from: https://www.sciencedirect.com/science/article/pii/S1568163721000155
Shalev I, Entringer S, Wadhwa PD, Wolkowitz OM, Puterman E, Lin J, et al. Stress and telomere biology: a lifespan perspective. Psychoneuroendocrinology. 2013 [cited 2017 Jan 29];38(9):1835–42. Available from: https://www.sciencedirect.com/science/article/pii/S0306453013001054
Chetty R, Stepner M, Abraham S, Lin S, Scuderi B, Turner N, et al. The association between income and life expectancy in the United States, 2001–2014. JAMA. 2016 [cited 2017 Oct 10];315(16):1750–66. Available from: https://jamanetwork.com/journals/jama/fullarticle/2513561
Dwyer-Lindgren L, Bertozzi-Villa A, Stubbs RW, Morozoff C, Mackenbach JP, Lenthe FJ van, et al. Inequalities in life expectancy among US counties, 1980 to 2014: Temporal Trends and Key Drivers. JAMA Intern Med. 2017 [cited 2017 May 31]; Available from: http://jamanetwork.com/journals/jamainternalmedicine/fullarticle/2626194
Mackenbach JP, Stirbu I, Roskam A-JR, Schaap MM, Menvielle G, Leinsalu M, et al. Socioeconomic inequalities in health in 22 European countries. N Engl J Med. 2008;358(23):2468–81.
Minkler M, Fuller-Thomson E, Guralnik JM. Gradient of disability across the socioeconomic spectrum in the United States. N Engl J Med. 2006 [cited 2016 Dec 26];355(7):695–703. Available from: https://doi.org/10.1056/NEJMsa044316
d’Orsi E, Xavier AJ, Steptoe A, de Oliveira C, Ramos LR, Orrell M, et al. Socioeconomic and lifestyle factors related to instrumental activity of daily living dynamics: results from the English Longitudinal Study of Ageing. J Am Geriatr Soc. 2014;62(9):1630–9.
Galobardes B, Lynch JW, Smith GD. Is the association between childhood socioeconomic circumstances and cause-specific mortality established? Update of a systematic review. J Epidemiol Commun Health. 2008 [cited 2021 Jul 12];62(5):387–90. Available from: https://jech.bmj.com/content/62/5/387
Galobardes B, Smith GD, Lynch JW. Systematic review of the influence of childhood socioeconomic circumstances on risk for cardiovascular disease in adulthood. Ann Epidemiol. 2006;16:91–104.
Lidfeldt J, Li TY, Hu FB, Manson JE, Kawachi I. A prospective study of childhood and adult socioeconomic status and incidence of type 2 diabetes in women. Am J Epidemiol. 2007 [cited 2021 Jul 12];165(8):882–9. Available from: https://doi.org/10.1093/aje/kwk078
Tom SE, Phadke M, Hubbard RA, Crane PK, Stern Y, Larson EB. Association of demographic and early-life socioeconomic factors by birth cohort with dementia incidence among US adults born between 1893 and 1949. JAMA Netw Open. 2020 [cited 2021 Jul 12];3(7):e2011094. Available from: https://doi.org/10.1001/jamanetworkopen.2020.11094
Belsky DW, Caspi A, Cohen HJ, Kraus WE, Ramrakha S, Poulton R, et al. Impact of early personal-history characteristics on the Pace of Aging: implications for clinical trials of therapies to slow aging and extend healthspan. Aging Cell. 2017;16(4):644–51.
Feltes BC, Poloni J de F, Bonatto D. Development and Aging: Two opposite but complementary phenomena. aging and health – a systems biology perspective. 2015 [cited 2021 May 27];40:74–84. Available from: https://www.karger.com/Article/FullText/364932
Gladyshev VN. The Ground Zero of Organismal Life and Aging. Trends Mol Med. 2021;27(1):11–9. https://doi.org/10.1016/j.molmed.2020.08.012.
Kinzina ED, Podolskiy DI, Dmitriev SE, Gladyshev VN. Patterns of aging biomarkers, mortality, and damaging mutations illuminate the beginning of aging and causes of early-life mortality. Cell Rep. 2019 [cited 2020 Nov 10];29(13):4276–4284.e3. Available from: http://www.sciencedirect.com/science/article/pii/S221112471931589X
Kerepesi C, Zhang B, Lee S-G, Trapp A, Gladyshev VN. Epigenetic clocks reveal a rejuvenation event during embryogenesis followed by aging. Sci Adv. 2021 [cited 2021 Aug 9];7(26):eabg6082. Available from: https://advances.sciencemag.org/content/7/26/eabg6082
Preston JD, Reynolds LJ, Pearson KJ. Developmental origins of health span and life span: a mini-review. Gerontology. 2018;64(3):237–45.
Blackburn EH, Epel ES, Lin J. Human telomere biology: a contributory and interactive factor in aging, disease risks, and protection. Science. 2015;350(6265):1193–8.
Chakravarti D, LaBella KA, DePinho RA. Telomeres: history, health, and hallmarks of aging. Cell. 2021 [cited 2021 May 23];184(2):306–22. Available from: https://www.sciencedirect.com/science/article/pii/S0092867420317505
Humphreys KL, Esteves K, Zeanah CH, Fox NA, Nelson CA, Drury SS. Accelerated telomere shortening: tracking the lasting impact of early institutional care at the cellular level. Psychiatry Res. 2016;246:95–100.
Middeldorp CM. Childhood stress and psychopathology: it’s not too early to look at biological aging. J Am Acad Child Adolesc Psychiatry. 2019;59(1):38–9.
Mitchell C, McLanahan S, Schneper L, Garfinkel I, Brooks-Gunn J, Notterman D. Father Loss and Child Telomere Length. Pediatrics. 2017 [cited 2021 May 27];140(2). Available from: https://pediatrics.aappublications.org/content/140/2/e20163245
Ridout KK, Khan M, Ridout SJ. Adverse childhood experiences run deep: toxic early life stress, telomeres, and mitochondrial DNA copy number, the biological markers of cumulative stress. BioEssays. 2018 [cited 2021 Jul 12];40(9):1800077. Available from: https://doi.org/10.1002/bies.201800077
Ridout KK, Levandowski M, Ridout SJ, Gantz L, Goonan K, Palermo D, et al. Early life adversity and telomere length: a meta-analysis. Mol Psychiatry. 2018 [cited 2020 Mar 23];23(4):858–71. Available from: https://www.nature.com/articles/mp201726
Nettle D, Gadalla SM, Lai T-S, Susser E, Bateson M, Aviv A. Measurement of Telomere Length for Longitudinal Analysis: Implications of Assay Precision. American J Epidemiol. 2021;190(7):1406–13. https://doi.org/10.1093/aje/kwab025.
Hastings WJ, Shalev I, Belsky DW. Translating measures of biological aging to test effectiveness of geroprotective interventions: what can we learn from research on telomeres? Front Genet. 2017 [cited 2017 Nov 28];8. Available from: https://doi.org/10.3389/fgene.2017.00164/full
Sanders JL, Newman AB. Telomere length in epidemiology: a biomarker of aging, age-related disease, both, or neither? Epidemiol Rev. 2013 [cited 2020 May 15];35(1):112–31. Available from: https://academic.oup.com/epirev/article/35/1/112/552544
Gavrilov LA, Gavrilova NS. The reliability-engineering approach to the problem of biological aging. Ann N Y Acad Sci. 2004;1019(1):509–12.
Jylhävä J, Pedersen NL, Hägg S. Biological age predictors. EBioMedicine. 2017 [cited 2018 Jan 12];21:29–36. Available from: http://www.ebiomedicine.com/article/S2352-3964(17)30142-1/abstract
Ferrucci Luigi, Levine Morgan E., Kuo Pei-Lun, Simonsick Eleanor M. Time and the metrics of aging. Circ Res. 2018 [cited 2020 Jul 6];123(7):740–4. Available from: https://doi.org/10.1161/CIRCRESAHA.118.312816
Fried LP, Cohen AA, Xue Q-L, Walston J, Bandeen-Roche K, Varadhan R. The physical frailty syndrome as a transition from homeostatic symphony to cacophony. Nat Aging. 2021 [cited 2021 Jul 11];1(1):36–46. Available from: https://www.nature.com/articles/s43587-020-00017-z
Rockwood K, Mitnitski A. Frailty in relation to the accumulation of deficits. J Gerontol A Biol Sci Med Sci. 2007 [cited 2021 Jul 11];62(7):722–7. Available from: https://doi.org/10.1093/gerona/62.7.722
Rosso AL, Sanders JL, Arnold AM, Boudreau RM, Hirsch CH, Carlson MC, et al. Multisystem physiologic impairments and changes in gait speed of older adults. J Gerontol A Biol Sci Med Sci. 2015;70(3):319–24.
Seeman TE, McEwen BS, Rowe JW, Singer BH. Allostatic load as a marker of cumulative biological risk: MacArthur studies of successful aging. Proc Natl Acad Sci U S A. 2001;98:4770–5.
Cohen AA, Milot E, Yong J, Seplaki CL, Fülöp T, Bandeen-Roche K, et al. A novel statistical approach shows evidence for multi-system physiological dysregulation during aging. Mech Ageing Dev. 2013;134(3–4):110–7.
Gaydosh L, Belsky DW, Glei DA, Goldman N. Testing proposed quantifications of biological aging in Taiwanese older adults. J Gerontol A Biol Sci Med Sci. [cited 2020 Mar 29];glz223. Available from: https://doi.org/10.1093/gerona/glz223/5578440
Levine ME. Modeling the rate of senescence: can estimated biological age predict mortality more accurately than chronological age? J Gerontol A Biol Sci Med Sci. 2013 [cited 2013 Oct 15];68(6):667–74. Available from: http://biomedgerontology.oxfordjournals.org/content/68/6/667
Liu Z, Kuo P-L, Horvath S, Crimmins E, Ferrucci L, Levine M. A new aging measure captures morbidity and mortality risk across diverse subpopulations from NHANES IV: A cohort study. PLoS Med. 2018 [cited 2019 Jan 7];15(12):e1002718. Available from: https://doi.org/10.1371/journal.pmed.1002718
Mamoshina P, Kochetov K, Putin E, Cortese F, Aliper A, Lee W-S, et al. Population specific biomarkers of human aging: a big data study using South Korean, Canadian, and Eastern European patient populations. J Gerontol A Biol Sci Med Sci. 2018 [cited 2021 Jul 12];73(11):1482–90. Available from: https://doi.org/10.1093/gerona/gly005
Belsky DW, Caspi A, Houts R, Cohen HJ, Corcoran DL, Danese A, et al. Quantification of biological aging in young adults. Proc Natl Acad Sci U S A. 2015;112(30):E4104-4110.
Glei DA, Goldman N, Rodríguez G, Weinstein M. Beyond self-reports: changes in biomarkers as predictors of mortality. Popul Dev Rev. 2014;40(2):331–60.
Sanders JL, Ding V, Arnold AM, Kaplan RC, Cappola AR, Kizer JR, et al. Do changes in circulating biomarkers track with each other and with functional changes in older adults? J Gerontol A Biol Sci Med Sci. 2014 [cited 2017 Mar 13];69A(2):174–81. Available from: https://academic.oup.com/biomedgerontology/article/69A/2/174/515152/Do-Changes-in-Circulating-Biomarkers-Track-With
Hastings WJ, Shalev I, Belsky DW. Comparability of biological aging measures in the National Health and Nutrition Examination Study, 1999–2002. Psychoneuroendocrinology. 2019 [cited 2019 May 19];106:171–8. Available from: http://www.sciencedirect.com/science/article/pii/S0306453018308084
Levine ME, Crimmins EM. Evidence of accelerated aging among African Americans and its implications for mortality. Soc Sci Med. 2014;118:27–32.
Dowd JB, Simanek AM, Aiello AE. Socio-economic status, cortisol and allostatic load: a review of the literature. Int J Epidemiol. 2009;38(5):1297–309.
Shalev I. Early life stress and telomere length: investigating the connection and possible mechanisms: a critical survey of the evidence base, research methodology and basic biology. BioEssays . 2012;34(11):943–52. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22991129
Dolcini J, Wu H, Nwanaji-Enwerem JC, Kiomourtozlogu M-A, Cayir A, Sanchez-Guerra M, et al. Mitochondria and aging in older individuals: an analysis of DNA methylation age metrics, leukocyte telomere length, and mitochondrial DNA copy number in the VA normative aging study. Aging (Albany NY). 2020;12(3):2070–83.
Castellani CA, Longchamps RJ, Sun J, Guallar E, Arking DE. Thinking outside the nucleus: Mitochondrial DNA copy number in health and disease. Mitochondrion. 2020 [cited 2021 Aug 9];53:214–23. Available from: https://www.sciencedirect.com/science/article/pii/S1567724920300659
Kim WY, Sharpless NE. The regulation of INK4/ARF in cancer and aging. Cell. 2006 [cited 2021 Jul 12];127(2):265–75. Available from: https://www.sciencedirect.com/science/article/pii/S0092867406012840
Liu Y, Sanoff HK, Cho H, Burd CE, Torrice C, Ibrahim JG, et al. Expression of p16INK4a in peripheral blood T-cells is a biomarker of human aging. Aging Cell. 2009 [cited 2021 Jul 12];8(4):439–48. Available from: https://doi.org/10.1111/j.1474-9726.2009.00489.x
Justice JN, Ferrucci L, Newman AB, Aroda VR, Bahnson JL, Divers J, et al. A framework for selection of blood-based biomarkers for geroscience-guided clinical trials: report from the TAME Biomarkers Workgroup. GeroScience. 2018 [cited 2019 May 19];40(5):419–36. Available from: https://doi.org/10.1007/s11357-018-0042-y
Crimmins EM, Thyagarajan B, Kim JK, Weir D, Faul J. Quest for a summary measure of biological age: the health and retirement study. GeroScience. 2021 [cited 2021 Aug 9];43(1):395–408. Available from: https://doi.org/10.1007/s11357-021-00325-1
Jansen R, Han LK, Verhoeven JE, Aberg KA, van den Oord EC, Milaneschi Y, et al. An integrative study of five biological clocks in somatic and mental health. eLife. 2021 [cited 2021 Feb 17];10:e59479. Available from: https://doi.org/10.7554/eLife.59479
Lehallier B, Gate D, Schaum N, Nanasi T, Lee SE, Yousef H, et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat Med. 2019 [cited 2021 Jan 27];25(12):1843–50. Available from: https://www.nature.com/articles/s41591-019-0673-2
Robinson O, Hyam MC, Karaman I, Pinto RC, Ala-Korpela M, Handakas E, et al. Determinants of accelerated metabolomic and epigenetic aging in a UK cohort. Aging Cell. 2020 [cited 2021 Jan 27];19(6):e13149. Available from: https://doi.org/10.1111/acel.13149
••Horvath S, Raj K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat Rev Genet. 2018 [cited 2018 Jul 17];19(6):371–84. Available from: https://www.nature.com/articles/s41576-018-0004-3. This review provides a thorough overview of DNA methylation and aging.
Jaenisch R, Bird A, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet. 2003 [cited 2021 Jul 13];33(3):245–54. Available from: https://www.nature.com/articles/ng1089z
Wilson VL, Jones PA. DNA methylation decreases in aging but not in immortal cells. Science. 1983 [cited 2017 Nov 12];220(4601):1055–7. Available from: http://science.sciencemag.org/content/220/4601/1055
Fraga MF, Esteller M. Epigenetics and aging: the targets and the marks. Trends Genet. 2007 [cited 2021 Jul 13];23(8):413–8. Available from: https://www.sciencedirect.com/science/article/pii/S0168952507001862
Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013;49(2):359–67.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013 [cited 2014 Jun 2];14(10):R115. Available from: http://genomebiology.com/2013/14/10/R115/abstract
Weidner CI, Lin Q, Koch CM, Eisele L, Beier F, Ziegler P, et al. Aging of blood can be tracked by DNA methylation changes at just three CpG sites. Genome Biol. 2014 [cited 2014 Feb 11];15(2):R24. Available from: http://genomebiology.com/2014/15/2/R24/abstract
Chen BH, Marioni RE, Colicino E, Peters MJ, Ward-Caviness CK, Tsai P-C, et al. DNA methylation-based measures of biological age: meta-analysis predicting time to death. Aging (Albany NY). 2016;8(9):1844–65.
Bell CG, Lowe R, Adams PD, Baccarelli AA, Beck S, Bell JT, et al. DNA methylation aging clocks: challenges and recommendations. Genome Biol. 2019[cited 2019 Dec 2];20(1):249. Available from:https://doi.org/10.1186/s13059-019-1824-y
Belsky DW, Moffitt TE, Cohen AA, Corcoran DL, Levine ME, Prinz JA, et al. Eleven telomere, epigenetic clock, and biomarker-composite quantifications of biological aging: do they measure the same thing? Am J Epidemiol. 2018;187(6):1220–30.
Breitling LP, Saum K-U, Perna L, Schöttker B, Holleczek B, Brenner H. Frailty is associated with the epigenetic clock but not with telomere length in a German cohort. Clin Epigenetics. 2016;8:21.
Murabito JM, Zhao Q, Larson MG, Rong J, Lin H, Benjamin EJ, et al. Measures of biologic age in a community sample predict mortality and age-related disease: the Framingham offspring study. J Gerontol A Biol Sci Med Sci. 2017 [cited 2018 Jan 8];(glx144). Available from: https://doi.org/10.1093/gerona/glx144/4034776
Zhang Q, Vallerga CL, Walker RM, Lin T, Henders AK, Montgomery GW, et al. Improved precision of epigenetic clock estimates across tissues and its implication for biological ageing. Genome Med. 2019 [cited 2019 Aug 28];11(1):54. Available from: https://doi.org/10.1186/s13073-019-0667-1
Zhang Y, Wilson R, Heiss J, Breitling LP, Saum K-U, Schöttker B, et al. DNA methylation signatures in peripheral blood strongly predict all-cause mortality. Nat Commun. 2017 [cited 2017 Jul 28];8:ncomms14617. Available from: https://www.nature.com/articles/ncomms14617
Hillary RF, Stevenson AJ, McCartney DL, Campbell A, Walker RM, Howard DM, et al. Epigenetic measures of ageing predict the prevalence and incidence of leading causes of death and disease burden. Clin Epigenet. 2020 [cited 2021 Mar 26];12(1):115. Available from:https://doi.org/10.1186/s13148-020-00905-6
McCrory C, Fiorito G, Hernandez B, Polidoro S, O’Halloran AM, Hever A, et al. GrimAge outperforms other epigenetic clocks in the prediction of age-related clinical phenotypes and all-cause mortality. Le Couteur D, editor. J Gerontol A Biol Sci Med Sci. 2021;76(5):741–9.
••Belsky D, Caspi A, Arseneault L, Baccarelli A, Corcoran D, Gao X, et al. Quantification of the pace of biological aging in humans through a blood test, the DunedinPoAm DNA methylation algorithm. eLife. 2020;9:e54870. https://doi.org/10.7554/eLife.54870. https://elifesciences.org/articles/73420. This study provides a detailed description of DNA methylation and pace of aging.
Yang Z, Wong A, Kuh D, Paul DS, Rakyan VK, Leslie RD, et al. Correlation of an epigenetic mitotic clock with cancer risk. Genome Biol. 2016;17(1):205.
Youn A, Wang S. The MiAge Calculator: a DNA methylation-based mitotic age calculator of human tissue types. Epigenetics. 2018;13(2):192–206.
Gao X, Colicino E, Shen J, Just AC, Nwanaji-Enwerem JC, Wang C, Coull B, Lin X, Pantel V, Zheng Y, Hou L, Schwartz J, Baccarelli AA. Comparative validation of an epigenetic mortality risk score with three aging biomarkers for predicting mortality risks among older adult males. Int J Epidemiol. 2019;48(6):1958–71. https://doi.org/10.1093/ije/dyz082.
Colich NL, Rosen ML, Williams ES, McLaughlin KA. Biological aging in childhood and adolescence following experiences of threat and deprivation: a systematic review and meta-analysis. Psychol Bull. 2020 [cited 2020 Aug 5]; Available from: http://ezproxy.lib.utexas.edu/login?url=http://search.ebscohost.com/login.aspx?direct=true&db=pdh&AN=2020-56119-001&site=ehost-live
Raffington L, Belsky DW, Kothari M, Malanchini M, Tucker-Drob EM, Harden KP. Socioeconomic disadvantage and the pace of biological aging in children. Pediatrics 2021;147(6):e2020024406. https://doi.org/10.1542/peds.2020-024406
McEwen LM, O’Donnell KJ, McGill MG, Edgar RD, Jones MJ, MacIsaac JL, et al. The PedBE clock accurately estimates DNA methylation age in pediatric buccal cells. Proc Natl Acad Sci. 2019;14:201820843.
Knight AK, Craig JM, Theda C, Bækvad-Hansen M, Bybjerg-Grauholm J, Hansen CS, et al. An epigenetic clock for gestational age at birth based on blood methylation data. Genome Biol. 2016;17(1):206.
Li C, Gao W, Gao Y, Yu C, Lv J, Lv R, et al. Age prediction of children and adolescents aged 6–17 years: an epigenome-wide analysis of DNA methylation. Aging. 2018;10(5):1015–26.
Joyce BT, Gao T, Koss K, Zheng Y, Cardenas A, Heiss J, Just A, Zhang K, van Horn L, Allen NB, Greenland P, Cohen S, Gordon-Larsen P, Mitchell C, McLanahan S, Schneper L, Notterman D, Rifas-Shiman SL, Oken E, Hivert M-F, Wright R, Baccarelli A, Lloyd-Jones D, Hou L. Impact of paternal education on epigenetic ageing in adolescence and mid-adulthood: a multi-cohort study in the USA and Mexico. Int J Epidemiol 2021. https://doi.org/10.1093/ije/dyab196
Dunn EC, Soare TW, Zhu Y, Simpkin AJ, Suderman MJ, Klengel T, et al. Sensitive periods for the effect of childhood adversity on DNA methylation: results from a prospective, longitudinal study. Biol Psychiatry. 2019;85(10):838–49.
Dongen J van, Nivard MG, Willemsen G, Hottenga J-J, Helmer Q, Dolan CV, et al. Genetic and environmental influences interact with age and sex in shaping the human methylome. Nat Commun. 2016 [cited 2017 Apr 18];7:11115. Available from: http://www.nature.com/ncomms/2016/160330/ncomms11115/full/ncomms11115.html
Heijmans BT, Mill J. Commentary: The seven plagues of epigenetic epidemiology. Int J Epidemiol. 2012 [cited 2014 Feb 4];41(1):74–8. Available from: http://ije.oxfordjournals.org/content/41/1/74
Raffington L, Mallard T, Harden KP. Polygenic scores in developmental psychology: Invite Genetics In, Leave Biodeterminism Behind. Annu Rev Dev Psychol. 2020;2(1):389–411.
Navas-Acien A, Domingo-Relloso A, Subedi P, Riffo-Campos AL, Xia R, Gomez L, et al. Blood DNA methylation and incident coronary heart disease: evidence from the strong heart study. JAMA Cardiol. 2021;6(11):1237.
Ehrlich KB, Yu T, Sadiq A, Brody GH. Neighborhood poverty, allostatic load, and changes in cellular aging in African American young adults: the moderating role of attachment. Attach Hum Dev. 2021;7:1–14.
Peng H, Gao W, Cao W, Lv J, Yu C, Wu T, et al. Combined healthy lifestyle score and risk of epigenetic aging: a discordant monozygotic twin study. Aging. 2021;13(10):14039–52.
Liu Z, Chen BH, Assimes TL, Ferrucci L, Horvath S, Levine ME. The role of epigenetic aging in education and racial/ethnic mortality disparities among older U.S. Women. Psychoneuroendocrinology. 2019;104:18–24.
Birney E, Smith GD, Greally JM. Epigenome-wide association studies and the interpretation of disease -Omics. PLoS Genet. 2016 [cited 2017 Feb 3];12(6):e1006105. Available from: https://doi.org/10.1371/journal.pgen.1006105
Lindenberger U, von Oertzen T, Ghisletta P, Hertzog C. Cross-sectional age variance extraction: what’s change got to do with it? Psychol Aging. 2011;26(1):34–47.
Nyberg L, Salami A, Andersson M, Eriksson J, Kalpouzos G, Kauppi K, et al. Longitudinal evidence for diminished frontal cortex function in aging. PNAS. 2010;107(52):22682–6.
Schaie KW. Age changes and age differences. Gerontologist. 1967;7(2):128–32.
Justice JN, Kritchevsky SB. Putting epigenetic biomarkers to the test for clinical trials. eLife. 2020 [cited 2020 Jul 21];9:e58592. Available from:https://doi.org/10.7554/eLife.58592
Price EM, Robinson WP. Adjusting for batch effects in DNA methylation microarray data, a lesson learned. Front Genet. 2018 [cited 2021 Aug 9];0. Available from: https://doi.org/10.3389/fgene.2018.00083/full
Canty AJ, Paterson AD. Evidence of batch effects masking treatment effect in GAW20 methylation data. BMC Proc. 2018 [cited 2021 Jul 13];12(9):32. Available from:https://doi.org/10.1186/s12919-018-0129-6
Zindler T, Frieling H, Neyazi A, Bleich S, Friedel E. Simulating ComBat: how batch correction can lead to the systematic introduction of false positive results in DNA methylation microarray studies. BMC Bioinformatics. 2020 [cited 2021 Jul 13];21(1):271. Available from: https://doi.org/10.1186/s12859-020-03559-6
Nygaard V, Rødland EA, Hovig E. Methods that remove batch effects while retaining group differences may lead to exaggerated confidence in downstream analyses. Biostatistics. 2016;17(1):29–39.
Pidsley R, Zotenko E, Peters TJ, Lawrence MG, Risbridger GP, Molloy P, et al. Critical evaluation of the Illumina MethylationEPIC BeadChip microarray for whole-genome DNA methylation profiling. Genome Biol. 2016 [cited 2018 Jul 17];17(1):208. Available from: https://doi.org/10.1186/s13059-016-1066-1
Forest M, O’Donnell KJ, Voisin G, Gaudreau H, MacIsaac JL, McEwen LM, et al. Agreement in DNA methylation levels from the Illumina 450K array across batches, tissues, and time. Epigenetics. 2018;13(1):19–32. Available from: https://doi.org/10.1080/15592294.2017.1411443
Logue MW, Smith AK, Wolf EJ, Maniates H, Stone A, Schichman SA, et al. The correlation of methylation levels measured using Illumina 450K and EPIC BeadChips in blood samples. Epigenomics. 2017;9(11):1363–71.
Sugden K, Hannon EJ, Arseneault L, Belsky DW, Corcoran DL, Fisher HL, et al. Patterns of reliability: assessing the reproducibility and integrity of DNA methylation measurement. Patterns. 2020 [cited 2020 Jul 15];1(2):100014. Available from: http://www.sciencedirect.com/science/article/pii/S2666389920300143
Higgins-Chen AT, Thrush KL, Wang Y, Kuo P-L, Wang M, Minteer CJ, et al. A computational solution for bolstering reliability of epigenetic clocks: implications for clinical trials and longitudinal tracking. bioRxiv. 2021 [cited 2021 May 13];2021.04.16.440205. Available from: https://doi.org/10.1101/2021.04.16.440205v1
Belsky D, Caspi A, Corcoran D, Sugden K, Poulton R, Arseneault L, et al. DunedinPACE: A DNA methylation biomarker of the Pace of Aging. Epidemiology; 2021 [cited 2021 Dec 7]. Available from: https://doi.org/10.1101/2021.08.30.21262858
Bakulski KM, Halladay A, Hu VW, Mill J, Fallin MD. Epigenetic research in neuropsychiatric disorders: the “Tissue Issue.” Curr Behav Neurosci Rep. 2016;3(3):264–74.
Raffington L, Belsky DW, Malanchini M, Tucker-Drob EM, Harden KP. Analysis of socioeconomic disadvantage and pace of aging measured in saliva DNA methylation of children and adolescents. bioRxiv. 2020 [cited 2020 Oct 19];2020.06.04.134502. Available from: https://doi.org/10.1101/2020.06.04.134502v1
Glymour MM. Natural experiments and instrumental variable analyses in social epidemiology. In: Methods in social epidemiology. Hoboken: Jossey-Bass/Wiley; 2006. p. 429–60.
Muennig P, McEwen B, Belsky DW, Noble KG, Riccio J, Manly J. Determining the optimal outcome measures for studying the social determinants of health. Int J Environ Res Public Health. 2020 [cited 2020 Apr 27];17(9):3028. Available from: https://www.mdpi.com/1660-4601/17/9/3028
Courtin E, Aloisi K, Miller C, Allen HL, Katz LF, Muennig P. The health effects of expanding the earned income tax credit: results from New York City. Health Aff. 2020 [cited 2021 Mar 6];39(7):1149–56. Available from: https://doi.org/10.1377/hlthaff.2019.01556
Garza MA, Quinn SC, Li Y, Assini-Meytin L, Casper ET, Fryer CS, et al. The influence of race and ethnicity on becoming a human subject: factors associated with participation in research. Contemp Clin Trials Commun. 2017;7:57–63.
Fisher ER, Pratt R, Esch R, Kocher M, Wilson K, Lee W, et al. The role of race and ethnicity in views toward and participation in genetic studies and precision medicine research in the United States: a systematic review of qualitative and quantitative studies. Mol Genet Genom Med. 2020 [cited 2021 Aug 9];8(2):e1099. Available from: https://doi.org/10.1002/mgg3.1099
Middleton A, Milne R, Thorogood A, Kleiderman E, Niemiec E, Prainsack B, et al. Attitudes of publics who are unwilling to donate DNA data for research. Eur J Med Genet. 2019 [cited 2021 Aug 9];62(5):316–23. Available from: https://www.sciencedirect.com/science/article/pii/S1769721218307316
Huang JY. Representativeness is not representative: addressing major inferential threats in the UK Biobank and other big data repositories. Epidemiology. 2021 [cited 2021 Aug 9];32(2):189–93. Available from: https://journals.lww.com/epidem/fulltext/2021/03000/representativeness_is_not_representative_.5.aspx?casa_token=BEq5TtqUwhsAAAAA:7y4lfTdyEEjhlOEAYjPGMwJVhDKCOex6kJv9m2qDXvUshivej10do4wLAiQSUqtv-JXgQoPRBwd_W6W2J7z0zZw
Keyes KM, Westreich D. UK Biobank, big data, and the consequences of non-representativeness. Lancet. 2019 [cited 2020 Feb 12];393(10178):1297. Available from: http://www.sciencedirect.com/science/article/pii/S0140673618330678
Stamatakis E, Owen KB, Shepherd L, Drayton B, Hamer M, Bauman AE. Is cohort representativeness passé? Poststratified associations of lifestyle risk factors with mortality in the UK Biobank. Epidemiology. 2021 [cited 2021 Aug 9];32(2):179–88. Available from: https://journals.lww.com/epidem/Fulltext/2021/03000/Is_Cohort_Representativeness_Pass___Poststratified.4.aspx
Haworth S, Mitchell R, Corbin L, Wade KH, Dudding T, Budu-Aggrey A, et al. Apparent latent structure within the UK Biobank sample has implications for epidemiological analysis. Nat Commun. 2019 [cited 2019 Mar 25];10(1):333. Available from: https://www.nature.com/articles/s41467-018-08219-1
Fry A, Littlejohns TJ, Sudlow C, Doherty N, Adamska L, Sprosen T, et al. Comparison of sociodemographic and health-related characteristics of UK Biobank participants with those of the general population. Am J Epidemiol. 2017;186(9):1026–34.
Eileen M Crimmins, PhD, Bharat Thyagarajan, MD, PhD, Morgan E Levine, PhD, David R Weir, PhD, Jessica Faul, PhD, MPH, Associations of Age, Sex, Race/Ethnicity, and Education With 13 Epigenetic Clocks in a Nationally Representative U.S. Sample: The Health and Retirement Study. J Gerontol A Biol Sci Med Sci 2021;76(6):1117–1123. https://doi.org/10.1093/gerona/glab016
Fiorito G, McCrory C, Robinson O, Carmeli C, Rosales CO, Zhang Y, et al. Socioeconomic position, lifestyle habits and biomarkers of epigenetic aging: a multi-cohort analysis. Aging. 2019;11(7):2045–70.
Nelson PG, Promislow DEL, Masel J. Biomarkers for aging identified in cross-sectional studies tend to be non-causative. J Gerontol A Biol Sci Med Sci. 2020 [cited 2020 May 20];75(3):466–72. Available from: https://academic.oup.com/biomedgerontology/article/75/3/466/5540066
Funding
LR is supported by the German Research Foundation (DFG). DWB is supported by Russell Sage Foundation BioSS grant 1810–08987, National Institute on Aging grants R01AG066887 and R01AG061378, a pilot grant from the Columbia Population Research Center, the Canadian Institute for Advanced Research Child Brain Development Network, and the Jacobs Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
This article does not contain any studies with human or animal subjects performed by any of the authors.
Conflict of Interest
Laurel Raffington declares no conflict of interest. Daniel W. Belsky is listed as an inventor on a Duke University and University of Otago invention that was licensed to a commercial entity.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Environment and Aging
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Raffington, L., Belsky, D.W. Integrating DNA Methylation Measures of Biological Aging into Social Determinants of Health Research. Curr Envir Health Rpt 9, 196–210 (2022). https://doi.org/10.1007/s40572-022-00338-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40572-022-00338-8