Abstract
Drastic fluctuation in temperature during crop growth challenges its yield to a greater extent, thus diminishing the agricultural productivity globally. At the cellular level, protein homeostasis is altered or disturbed directly or indirectly by the high temperature. Improper protein folding due to heat stress creates ER stress in the endoplasmic reticulum (ER). In order to balance the protein folding demand and its capacity and to maintain ER homeostasis, cells activate unfolded protein response (UPR), as a homeostatic mechanism. The transcription factors, bZip60 and bZip28/17 have been reported as the key modules of UPR for alleviating ER stress. Despite UPR mechanisms has been extensively investigated, the intervention of heat stress with ER homeostasis remains unclear. This review summarizes the key findings on this potential nexus. Additionally, the link between ER associated degradation and plant growth regulators during heat stress were also addressed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Unpredictable climate anomalies accompanied with severe constraints on arable lands and available water enact serious threats to agriculture crops. The increased temperature prompts heat stress on plants and it seeks immediate attention for sustainable crop production due to its adverse impact on growth and development. Indeed, 1 °C raise in global mean temperature have drastically declined the yields of major crops (Zhao et al. 2017). Sensing elements are of utmost importance to understand the mechanisms of heat stress tolerance despite extensive reports on downstream genes and its regulation under heat stress were known. Plasma membrane (PM), a lipid-bilayer with intrinsic and transmembrane proteins, could be the site for the initiation of sensing external thermal fluctuations of the cell (Niu and Xiang 2018; Ruelland and Zachowski 2010). Heat stress affects the plasma membrane integrity through increased fluidity. The PM associated calcium channels sense this fluctuation and initiate the heat-shock responses (Demidchik et al. 2018; Ward et al. 2009) which are further amplified and orchestrated by the secondary transducers viz., histone sensors in the nucleus (Kumar and Wigge 2010) and unfolded protein response sensors in the endoplasmic reticulum (Mittler et al. 2012; Saidi et al. 2009; Srivastava et al. 2014). Unfolded protein response (UPR) is one of the crucial mechanisms like autophagy, ubiquitin proteasome degradation systems in plants. Extensive studies have focused on the details of UPR pathways, its structure and ER stress responses (Liu and Howell 2016; Nawkar et al. 2018; Liu and Li 2019; Park and Park 2019). However, the influence of heat stress on UPR has still remained unclear. In this review, we have summarized the recent advances of the heat stress influences on ER homeostasis with the special emphasis on the molecular players associated with these responses in crop plants.
ER homeostasis
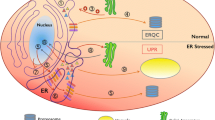
Endoplasmic Reticulum (ER) is one of the essential organelles of plant cells and is involved in key functions like protein synthesis, folding and exporting, phospholipid biosynthesis, and calcium storage (Anelli and Sitia 2008). ER is the necessary site for the protein to attain its functional conformation through post-translational modifications such as glycosylation, phosphorylation, disulfide bond formation, and other modifications. It is highly sensitive to both external and internal stimuli and causes the production and accumulation of misfolded proteins (Howell 2013; Walter and Ron 2011). This aggregation of misfolded proteins in the ER generates “ER stress”. Molecular players such as chaperones and co- chaperones in the ER membrane, alleviate this stress through unfolded protein response by means of upregulated lipid biosynthesis, aided protein folding, repressed translation, and expanded ER space (Pastor-Cantizano et al. 2020; Braakman and Hebert 2013). In general, the cellular imbalance due to the secretion and landing of polypeptides in the ER membrane increases the level of protein misfolding (Nawkar et al. 2018). Apart from these factors, biotic and abiotic stresses, disruption of calcium homeostasis, inhibition of glycosylation, and depletion of glucose have triggered the level of protein misfolding (Patil and Walter 2001; Ron and Walter 2007; Angelos et al. 2017; Verchot 2016). During ER stress, the UPR sensors cause pro-death processes and when they are not powerful enough to attenuate, the programmed cell death is induced (Liu et al. 2011a; Schwarz and Blower 2016). The detailed overview of UPR mechanisms for ER homeostasis has been presented in Fig. 1.
Protein folding and the ER Quality control system under normal and heat stress conditions. (Steps 1–8). Under normal conditions, proteins entering the ER are acted upon by the chaperones and foldases present in the ER lumen. The entered proteins are properly folded and undergone quality control check within the lumen before being transported to their final destinations. The proteins which are not getting through the quality check, will be sent back to the calnexin/calreticulin/EBS2 to restart their folding mechanisms. In case of persistence of misfolding in the consequent quality checks, the misfolded proteins are ubiquitinated and targeted for ER associated degradation. Thereby, the ER maintains a balance between protein folding demand and protein folding capacity. (Steps 9–14). On the advent of stress, the homeostasis in the ER gets affected as a result of increased protein folding demand and lowered protein folding capacity. Thus, an imbalance prevails in ER which in turn would activate the UPR pathways (IRE1-bZip60 and bZip28/ bZip17). (Steps 15–22). The activated UPR pathways self-regulate its key players and upregulates chaperones and foldases of the ER folding machinery to enhance the folding capacity and inhibits further protein load into ER. If it is still unable to establish homeostasis, the autophagy and ubiquitination pathways get activated
Unfolded protein response (UPR) arms in plants
A pair of UPR arms is functional in plants for undertaking ER stress perception and signal transduction which has been explained in detail in Fig. 2. The first arm, IRE1 (Inositol-requiring enzyme 1) mediates unconventional splicing of bZip60 in the cytosol, and the second arm, bZip17/bZip28 regulates intramembrane sequential proteolysis (Deng et al. 2011; Liu and Howell 2016). The processing and activation mechanisms are unique to each bZip TFs, which involves either proteolytic cleavage or alternate splicing (Lu et al. 2012; Sita et al. 2017; Takahashi et al. 2012).
UPR arms and its signaling under heat stress. UPR pathways in plant are mainly regulated by IRE1-bZip60 and bZip28/bZip17 modules located in the ER lumen which are also activated in response to heat stress. a BiP binds to IRE1 and maintains it in a dormant state under normal conditions. When heat stress induced the protein misfolding, the BiP unbinds from the IRE1 and attaches to the misfolded protein. On the release of BiP, the IRE1 dimerizes and in turn splices bZip60 mRNA. The spliced mRNA enters the nucleus and upregulates the expression of UPR downstream genes such as BiP and PDI. b bZip28 and BAG7 are kept inactive under normal conditions by the binding of BiPs. On sensing of heat stress, the BiPs get released from bZip28 and BAG7. This process is followed by the SUMOylation and translocation of BAG7 to the nucleus and bZip28 transported to the Golgi body via COPII vesicles. At the Golgi body, it is spliced by S2P and translocated to nucleus, thereby upregulating the downstream genes (BiP3). The activation of bZip17 follows the pathway similar to bZip28 by which it is proteolytically spliced by S1P/S2P and translocated to the nucleus. c a possible bZip28 compensatory pathway exists under heat stress. In the absence of bZip28, H2O2 accumulates in the cytosol and further, HSFA2 activates the ascorbate peroxidase pathway in response to heat stress
IRE1, an endoribonuclease and serine/threonine kinase, is the most conserved ER sensor among the eukaryotes. IRE1 is negatively regulated by the BiP, an ER-resident chaperone. During ER stress, the BiP unbinds from membrane-associated IRE1. The released IRE1 dimerizes and executes unconventional splicing of the membrane-associated transcription factor (MTF), bZip60 mRNA through a process called ‘regulated IRE1-dependent splicing’ (RIDS). The spliced bZip60 then shuttles to the nucleus and activates UPR responsive genes (Mori et al. 1996; Sun et al. 2013). The IRE gene family in Arabidopsis has three paralogues AtIRE1a, AtIRE1b, and AtIRE1c, while in rice single IRE member has been reported (Koizumi et al. 2001; Lu et al. 2012; Okushima et al. 2002; Pu et al. 2019). Stress-mediated unconventional splicing by IRE1 resulted frameshifting by the loss of 23 ribonucleotides, leads to the elimination of trans membrane domain of bZip60 (Deng et al. 2011; Nagashima et al. 2011). bZip60 homologues identified in Arabidopsis and rice are AtbZip60 and OsbZip74 respectively (Hayashi et al. 2012; Lu et al. 2012) Srivastava et al. (2018). reported two maize IRE1 homologues, that splice ZmbZip60 under ER stress. Apart from activating UPR genes, bZip60 shuttled transcellularly for transmitting UPR signal to distal tissues in Arabidopsis (Lai et al. 2018).
The second arm of plant UPR signalling is mainly carried out by the membrane associated transcription factors such as bZip17 and bZip28. On the onset of stress, BiP dissociates from the C-terminal domain of the bZip28 and then pursues regulated intramembrane proteolysis (RIP). This process released bZip28 from ER membrane thus resulting in its activation. The activated transmembrane protein translocated to Golgi apparatus, where it undergoes cleavage by site-1 protease (S1P) and site-2 protease (S2P) (Iwata et al. 2017). The cleaved protein enters nucleus and upregulates UPR downstream genes (Niu and Xiang 2018). OsbZip16 (OsbZip39) and OsbZip60 in rice have been identified as homologues to AtbZip28 and AtbZip17 respectively (Takahashi et al. 2012) Their significance attributed to their evolutionary conservation across the plant species (Seo 2014).
Other than the UPR mechanisms, plant IRE1 is involved in the selective degradation of mRNA, through the pathway called as regulated IRE1-dependent decay (RIDD) (Mishiba et al. 2013). Bao et al. (2018) reported that AtIRE1B regulates the transcriptomes by degrading the mRNA of various secretory proteins like β-glucosidase related protein BGLU21, pathogenesis related protein PR-14 and MD2-related lipid recognition domain protein ROSY1/ML under ER induced autophagy. Generally, UPR in the proximity of both the cytoplasm and the ER was mounted by heat shock proteins and bZip transcription factors respectively (Mittler et al. 2012; Saidi et al. 2009). Various reports have emphasized the key features of UPR mediating ER homeostasis under control and heat stress which are altogether furnished in Table.1.
Key modules linking UPR with heat stress
Despite general plant mechanisms known for heat stress response such as flower development, circadian clock regulation, immunity responses (Liu et al. 2015a, b), UPR mechanisms specific to heat stress remain unclear. However, few reports have demonstrated the link between thermal sensing and UPR through phytochrome null mutants, which function as thermosensor as well as the negative regulator of UPR (Jung et al. 2016; Nawkar et al. 2017). Importantly, heat stress has been hypothesized as the major cause of ER stress among other abiotic stressors (Afrin et al. 2020). Besides, recent reports have shed lights on heat stress mediated ER homeostasis pathways viz., IRE1 mediated bZip60 pathway and S2-protease spliced bZip28 pathway (Kataoka et al. 2017; Li et al. 2017a, b, c). The activated bZip60 and bZip28 play key roles in UPR mediated gene regulatory network under heat stress (Che et al. 2010; Kataoka et al. 2017; Li et al. 2017a, b, c). They are stabilized by ER localized chaperones such as HSP70, ERDJ3 (Howell 2014), CNX/CRT (Nawkar et al. 2018), foldases such as PDI (Kamauchi et al. 2005) and ERO1(Onda et al. 2009). Up-regulation of these chaperones and foldases might be one of the early responses to alleviate the misfolded protein accumulation under heat stress. Mainly, there are two key modules of UPR associated with heat stress responses and have been described below.
IRE1-bZip60 module
IRE1, a dual protein kinase, is mainly involved in the splicing of bZip60 for the activation of stress-responsive genes. Indeed, bZip60 is considered as a biomarker for UPR (Li Yangjie et al. 2012). Under normal conditions, BiP is associated with the lumen facing the N terminal domain of IRE1 and maintains in a dormant state (Deng et al. 2011; Howell 2014). On the accumulation of misfolded proteins, the BiP dissociates from IRE1 and transiently binds to the misfolded proteins. This transient binding signals the downstream transcriptional activity. Concurrently, overexpression of BiP reduced the ER stress induced by DTT and heat (Yang et al. 2016). Likewise, CaBiP1 silenced pepper plants were susceptible to ER and heat stress while its overexpression conferred heat tolerance (Wang et al. 2017). Also, thermosensitive male sterile 1 (TMS1) protein is one of the interaction partners of BiP under heat stress (Ma et al. 2015). TMS1 belongs to DnaJ protein group, which function as co-chaperones of BiP. DnaJ domain of TMS1 interacted with BiP1 and BiP3 and stimulated its ATPase activity, thus imparting thermotolerance in plants (Nishikawa et al. 2005; Yamamoto et al. 2008).
A non-canonical interaction was observed between BiP and HOP3 (HSP70-HSP90 organizing protein) via the IRE1 dependent pathway in presence of ER stress inducers (Fernández-Bautista et al. 2017). Previous works on HOP3 induction under heat stress revealed its interaction with BiP under heat mediated ER stress. Thus, the accumulation of misfolded proteins could activate HOP3-BiP interaction, followed by the dissociation of BiP from IRE1. This process further leads to the autophosphorylation of the N-terminal domain of IRE1(Fernández-Bautista et al. 2018; Yángüez et al. 2013) that activates IRE1 dimerization, for recognizing the consensus sequence of the kissing loop present in its target bZip mRNA (Geng et al. 2018; Shamu and Walter 1996).
On splicing, bZip60s are activated and relocated to the nucleus to induce the transcription of BiP genes through ER stress-responsive cis-element (ERSE) and/or through plant-unfolded protein response element (P-UPRE) (Iwata et al. 2009; Iwata and Koizumi 2005). Since the splicing is attributed to IRE1, and in vitro ribonuclease activity is presented in IRE1a and IRE1b, the attempts has been done to determine which of the two forms is primarily responsible for splicing of bZip60 under heat stress (Humbert et al. 2012). Deng et al. (2011), noticed that loss-of-function mutant of IRE1b or point mutation in the bZip60 mRNA splicing site hampered BiP3 induction in Arabidopsis under heat/ Tunicamycin (Tm) stress. Also, the variants of IRE1b mutants developed by the RNAi approach have shown an impairment in bZip60 splicing with higher accumulation of IRE1b transcripts under heat stress treatment. Thus, the modification of IRE1/bZip60 activity was found to subvert heat stress responses through UPR (Humbert et al. 2012).
Likewise, heat-induced bZip60 mRNA processing implied that AtIRE1b is necessary for splicing of bZip60 under heat stress (Moreno et al. 2012). In contradiction, Nagashima et al. (2011) reported that ire1a ire1b double mutants abstained from splicing of bZip60 mRNA, whereas its spliced form was present in ire1a and ire1b single mutants. Thus, IRE1a and IRE1b are functionally redundant in performing the unconventional splicing of bZip60 mRNA under heat/Tm-induced UPR pathway. ZmbZip60, an ortholog of AtbZip60, was found in maize (Li Yanjie et al. 2012) whose splicing was followed immediately after heat/Tm/DTT treatment resulting in the excision of 20 bp intron. Despite ZmbZip60 splicing was strongly observed in the seedlings, same level was also noted in the tassels. Interestingly, ire1a ire1b double mutant was found fertile under normal conditions but sterile at high temperature. This conditional male sterility perhaps due to the structural defects tapetum, which thereby failed to deposit the pollen coat under high temperature conditions (Deng et al. 2016).
While most of the works on IRE1-bZip60 splicing have been reported in dicots Hayashi et al. (2012), found IRE1-bZip pathway in the monocots, where OsIRE1 regulates the expression of OsbZip50/OsbZip74, thus demonstrating the association between OsbZip50 and ER stress sensors. Also, OsbZip50/OsbZip74 spliced by OsIRE1 after treating with ER stress inducers like tunicamycin (Tm) and DTT; environmental stressors like heat stress and salicylic acid (Lu et al. 2012). However, IRE1 recognition sites in monocots and dicots differ through shorter interloop bridge sequences (Li Yanjie et al. 2012). A dual-luciferase system reported that OsbZip74s induced the promoter activity of a NAC factor (OsNTL3) under ER stress. Additionally, the relative expression of OsNTL3 reduced in bZip74 mutant under heat stress (Liu et al. 2020). Subsequently, bZip factor identified from a full-length wheat cDNA library was reported to confer multiple abiotic stress tolerance. However, its role in ER stress response is not elucidated (Zhang et al. 2015). Later, it has been demonstrated the evolutionary conservation between the TabZip60 splicing pathway and signal transductions in wheat. Invariably, the overexpression lines of TabZip60s exhibited a high level of stress tolerance than the wild type. Through ChIP-qPCR it was found that conventional ER or heat stress related transcripts (AtBI-1, BiP3, and PDI genes) were directly upregulated by TabZip60s (Geng et al. 2018). Deng et al. (2013a, b) demonstrated that an active RNase domain of IRE1 is required for stress tolerance through complementation of multiple mutants of UPR signal transducers, with IRE1b with mutations in its kinase and/or RNase domains.
Interestingly, it was found that heat stress mediated UPR downstream genes expressed at different developmental stages of the plant. For instance, the expression of ER stress mediated genes under heat stress was well documented in the vegetative stage (Deng et al. 2011; Moreno et al. 2012). UPR activation in the vegetative tissues leads to the constitutive production of spliced forms of bZip60 in anthers and expression of UPR genes in flowers under control conditions (Fragkostefanakis et al. 2016; Iwata et al. 2008, 2010; Quilichini et al. 2014). Similarly, upregulated bZip60 promoter activity in the reproductive tissues and in the developing seeds as associated with the UPR activity in the secretory cells (Deng et al. 2016; Iwata et al. 2008; Lu et al. 2012). Besides being heat susceptible, ire1a ire1b bZip28 triple mutant is involved in male gametophyte development, as evident from its defective tapetum in Arabidopsis (Deng et al. 2013b). Zhang et al. (2017a, b) reported the enrichment of UPR related genes in the reproductive tissues under heat stress through transcriptome analysis of bZip28 and bZip60 mutants which also showed reduced silique length and fertility phenotypes. Besides, the role of UPR in pollen development and function has also been demonstrated under heat stress (Raja et al. 2019).
bZip28/bZip17 module
The importance of bZip28/bZip17 module in UPR was interpreted through heat- sensitive phenotype of bZip28 knockout mutant (Gao et al. 2008), which suggests that this sustained bZip28 activity may be required for thermotolerance in Arabidopsis. The dissociation of BiPs from bZip28 under heat stress could be interpreted in many ways viz., the high affinity of BiP for unfolded protein over bZip28, changing redox conditions, and the interaction of DNAJ protein with the BiP (Srivastava et al. 2014). Recently, Herath et al. (2020) reported three BiPs viz., StBiP1, StBiP2, and StBiP3 identified by genome—wide association in Solanum tuberosum, whose overexpression enriched the binding of ER stress element (ERSE-I) in AtbZip28 promoter under high temperature. The nuclear-localized bZip28 proteins bind to ERSE and activate genes involved in thermotolerance (Liu and Howell 2010; Yamamoto et al. 2004).
AtBAG7, an ER localized Arabidopsis homologue of mammalian BAG (B cell lymphoma associated athanogen) coordinated the ER-nucleus signalling in collaboration with AtBiP2 and AtbZip28 during heat stress. AtBAG7 maintains AtbZip28 in the dormant stage in the ER membrane (Williams et al. 2010) and AtBAG7 knockouts mutants were hypersensitive to heat stress. The selective up-regulation of the AtBiP3 in AtBAG7 knockouts mutant retained AtbZip28 on the ER membrane. Thus, the released bZip28 enhances the upregulation of AtBiP3. Together these results demonstrated the role of BAG7 as a cytoprotective co- chaperone during UPR. During the onset of stress, the dissociation of BiP from the AtBAG7 and AtbZip28 triggers a cascade of events starting from the proteolytic release of BAG7 and bZip28 from the ER membrane leads to the SUMOylation of BAG7 (Williams et al. 2010). The SUMOylated BAG7 translocated to the nucleus and interacted with WRKY29, for regulating the expression of stress-responsive genes (Yurong Li et al., 2017a, b, c; Williams et al. 2010). Simultaneously, the AtbZip28 undergone S2P-mediated proteolysis in Golgi and then translocated to the nucleus where it binds to ERSE, and induces downstream stress responsive genes like BiP3 (Liu and Howell 2010; Yamamoto et al. 2004).
During the heat stress response, bZip28 has several direct targets (both canonical and noncanonical UPR targets) in different tissues. For instance, 133 putative direct targets of bZip28 in Arabidopsis revealed through ChIP-seq experiments with Myc-bZip28-expressing plants under heat stress (Zhang et al. 2017a, b). Also, the reduced expression of heat-inducible genes such as BiP2 and small heat shock proteins like HSP26.5 in the bZip28 null mutant revealed the importance of bZip28 in heat stress tolerance (Gao et al. 2008). Also, heat- responsive molecular chaperone BiP3 acted as a direct target of bZip28 (Williams et al. 2010; Zhang et al. 2017a, b). Interestingly, it was reported that a compensatory heat response pathway operating through HSFA2 could compensate for bZip28 deficiency by activating the ascorbate peroxidase pathway (Kataoka et al. 2017).
In addition to bZip28, the role of bZip17 in both ER homeostasis and heat stress had been demonstrated in thermotolerant finger millet through a transgenic line overexpressing EcbZip17. Also, EcbZip17E, EcbZip17S, and EcbZip17W overexpression plants exhibited high seedling survival rate, chlorophyll retention, and seedling fresh weight compared to the wild type under heat stress (Ramakrishna et al. 2018). Heat sensitivity was alleviated in an s2p proteolysis mutant when it was complemented with activated forms of either bZip17 or bZip28. Thus, the partial functional redundancy of bZip17 and bZip28 under heat stress was observed (Che et al. 2010). This functional redundancy provided flexibility in perceiving as well as responding to stress. UPR connection with phytochrome is another interesting area and needs to be explored further. The temperature perception of phytochrome, a red-light receptor in Arabidopsis was reported by Jung et al. (2016) and Nawkar et al. (2017). It was found that HY5, a bZip TF as well as a downstream component of phytochrome signaling exhibited elongated hypocotyl due to the modulation of UPR. This evidence has opened an avenue for HY5 mediated crosstalk between heat stress and UPR (Park and Park 2019).
Other key players mediating ER homeostasis under heat stress
NAC factors interaction with UPR arms
In general, NAM/ATAF1/2/CUC2 (NAC) transcription factors are involved in the developmental processes, phytohormonal interactions, and biotic and abiotic stress responses of plants (He et al. 2005; Nuruzzaman et al. 2013; Xie et al. 2000; Zhong et al. 2006). Intriguingly, extensive studies substantiated the role of NAC in mediating UPR and heat stress responses (Deng et al. 2016; Kataoka et al. 2017; Yu and Kanehara 2020). These evidences demonstrated that NAC factors act as secondary players in the heat mediated UPR signalling network and are regulated at both transcriptional and post-transcriptional level (Shao et al. 2015).
A characteristic NAC protein has a conserved N terminal domain of 150 amino acids, known as the NAC domain, and a varying length of amino acids in the C terminal domain (Ooka et al., 2003). NAC domain is divided into five subdomains (A-E) (Ooka et al. 2003) and is responsible for DNA binding, nucleus relocation, and dimerization (Olsen et al. 2005). The diversified C terminal domain functions as a transcriptional regulatory domain, which imparts individual function to the NAC proteins (Jensen et al. 2010; Olsen et al. 2005). The C terminal domain of few NAC proteins has transmembrane motifs, which aids NAC TFs bind to the PM or ER membrane (Puranik et al. 2012). Those transmembrane possessing NACs are named as NTLs (NAC with TRANSMEMBRANE MOTIF1-LIKE) (Ernst et al. 2004; Seo and Park 2010).
Being a secondary transducer, NAC TFs bind to the consensus sequence of its downstream genes under abiotic stress in plants (Hong et al. 2016; Yuan et al. 2019). Transcriptional upregulation of NAC factors by various biotic and abiotic stimuli have been documented in plants (Nuruzzaman et al. 2013; Seo and Park 2010). Indeed, locating a cis-regulatory motif on the NAC promoter responding to both heat and ER stress is imperative in elucidating the possible role of NAC in integrating ER and heat stress signalling. Recently, Liu et al. (2020) reported functional characterization of OsNTL3, which encodes an NAC TF with a predicted C-terminal domain. The constitutively expressed OsNTL3 was upregulated under heat stress and ER stress. RNA-Seq analysis revealed that the expression of OsNTL3 was dependent on 771genes responsive to heat stress and UPR. The direct binding of OsNTL3 to the OsbZip74 promoter under heat stress was revealed by ChIP sequencing. Further, through transient expression assays, OsbZip74 activates the promoter of OsNTL3 which suggests a positive feedback loop occurring between OsNTL3 and OsbZip74 in coordinating ER and heat stress signalling in rice (Liu et al. 2020). This study also emphasized the need for an attenuator in the post-stress period, so that the normal growth recovery remains unaffected. Although NAC TFs play a key role in responding to ER and heat stress, further characterization of UPR responsive NACs under heat stress would help in understanding how NACs relay the heat stress signals for ER homeostasis. Thus, the identification of putative NACs could be the potential targets of crop improvement for heat stress tolerance.
Phytohormone interplaying with UPR arms
UPR integrates multiple cues for normal plant growth and development. Given its association with phytohormones like Brassinosteroid, Gibberellin, ABA, and other plant growth regulators, the multi-faceted nature of UPR signalling has been revealed. Brassinosteroid is one of the key hormones associated with UPR signalling which sheds the light on the multi-layered regulation of ER homeostasis by UPR. For instance, high temperature leads to the inhibition of protein glycosylation of brassinosteroid signaling factors (Che et al. 2010). The S2P-mediated RIP activates bZip17 and bZip28, which directly affects the activation of BES1, a key TF involved in BR signalling in s2p mutant under heat stress (Che et al. 2010). This mechanism was further refined by EcbZip17 that activates BR signalling for normal growth responses and ER signalling for stress responses (Ramakrishna et al. 2018). Barba-Espín et al. (2014) investigated barley aleurone layers that secrete proteins in response to GA3 treatment in vitro. When the GA3 treated aleurone is subjected to heat and ER stress, the overlapping effects of both stresses at the intracellular and secretory proteomics level were observed. Salicylic acid- induced expression of ER proteins such as BiP2 and BiP3 and ER-stress related genes through the typical UPR mechanisms of proteolytic processing of bZip28 and IRE1 mediated splicing of bZip60 mRNA in Arabidopsis (Nagashima et al. 2014). CPR5, one of the negative regulators of salicylic acid, also negatively regulated bZip28 and bZip60 through interactions with them (Meng et al. 2017). ABA was predicted to be a counterpart of UPR through the evidence that overexpression of bZip17 in maize was associated with ABA-mediated ER response through the binding of bZip17 with cis-elements of ABA-responsive genes (Yang et al. 2016). Apart from phytohormones, UPR was also influenced by plant growth retardants. For instance, heat stress treatment combined with the paclobutrazol application in maize inbred lines leads to the splicing of bZip60 and sensitized the plant to UPR responses (Neill et al. 2019).
ER associated degradation (ERAD) under heat stress
When the UPR pathways are unsuccessful to achieve proper folding and maintain homeostasis within a stipulated time frame, the misfolded proteins were ubiquitinated for degradation through proteasomes in the cytoplasm through ERAD pathway (ER associated degradation) (Bernales et al. 2006; Strasser 2018). Most of the knowledge on key constituents of ERAD was revealed through the studies on yeast and mammals. The ERAD pathway consists of four steps, viz., recognition, ubiquitination, dislocation, and degradation (Deng et al. 2013a). These steps are driven by various intermediators like Hrd3 and Yos9, which belong to E3 ubiquitin ligase and ER-resident lectins respectively. The ubiquitination process was carried out with the help of UBC (ubiquitin conjugating enzymes) and Cue1, an ER membrane protein that employs the UBC7 to the Hrd1 and Doa10. The process of dislocating the ubiquitinated proteins from the ER to the cytoplasm was performed by CDC48 and its associated proteins such as NP14 and UFD1. The final degradation process was mediated by CDC48, UFD2, RAD23 and RPN1 (Deng et al. 2013a). Overexpression of a few of these intermediators like UBC1, UBC6 and UBC7 imparted tolerance to heat stress and H2O2 in yeast (Liu et al. 2011b).
While the ERAD process is conserved across the eukaryotes, the information on the molecular players of ERAD in plants is limited. However, yeast orthologues of HRD1, HRD 3, YOS9, UBC6, CDC48, and DOA10 have been reported in Arabidopsis (Cui et al. 2012; Hüttner et al. 2012; Li et al. 2017a, b, c; Liu Liang et al. 2011b; Ruddock and Molinari 2006). Additionally, plants possess unique ERAD components like EMS-mutagenized brassinosteroid-insensitive 1 suppressor 7 (EBS7), ERAD-mediating ring finger (EMR), protein associated with Hrd1-1 (PAWH1) & PAWH2 in Arabidopsis and E3 ligase in Medicago falcata salt & tunicamycin-induced ring finger protein (MfSTMIR) (Lin et al. 2019; Liu Yidan et al. 2015; Park et al. 2018; Zhang et al. 2019).
Li et al. (2017a, b, c) demonstrated UPR and ERAD are evolutionarily conserved quality- control mechanisms; playing an important role under heat stress in Arabidopsis. Further the ERAD genes CER9 and HRD1A/1B have been demonstrated to play the negative roles in heat stress responses, as the triple mutant cer9-2 hrd1a hrd1b exhibited a high survival rate than the corresponding single and double mutants. The mutants were observed to induce the upregulation of downstream UPR genes such as bZip60 and BiP1 under both normal and heat stress conditions in Arabidopsis, thus suggesting the possibility of crosstalk between ERAD and heat stress.
As the intensity of ER stress increases, ubiquitin independent pathway mediated by autophagy gets activated and misfolded proteins are transported to vacuole for degradation Liu et al. (2012). observed the portions of ER inside autophagic bodies in response to TM or DTT treatment, thus demonstrating the prevalence of autophagy in plants during ER stress. In Arabidopsis, the unfolded proteins accumulated under heat stress lead to ER stress followed by the expression of ATG genes (Yang et al. 2016; Zhou et al. 2013). Also, this ER-mediated autophagy partly depends on the IRE1b signalling pathway to mitigate unfolded protein accumulation (Yang et al. 2016). The cargo proteins generated under stress conditions were transported to autophagosomes by autophagy adaptors. Arabidopsis NBR1 is one such autophagy cargo adaptor that interacts with ATG8 under heat stress. The atg5, atg7, and nbr1 mutants were sensitive to heat stress which was reflected in their poor recovery rates (Zhou et al. 2013). Besides, overexpression of AtBiP1 reduced the effects of autophagy following heat stress (Yang et al. 2016).
Conclusion
UPR is a master regulator for maintaining ER homeostasis in plants under both biotic and abiotic stresses. Indeed, it integrates and acts as a signalling hub to different stress combinations that are encountered by crop plants. Despite being uncovered some of the key regulators in the UPR signalling, it is noteworthy to explore further on the novel UPR modules particularly under the current climate scenario with due importance to heat stress resilience in crops. Aside from considering the significance of UPR and ER stress in proteostasis, understanding their importance in heat stress responses is indeed imperative. Considering the facts like ER homeostasis is vital for plant growth and heat stress directly sensitizes the plants for UPR pathways, the succeeding research should address the cross-talk between ER stress and heat stress along with its multi-layered signaling in crop plants under the climate change. This could be one of the promising areas of future research to generate better breeding material for heat stress tolerance and to improve the resilience of crops for enhancing their productivity.
References
Afrin, T., Diwan, D., Sahawneh, K., & Pajerowska-Mukhtar, K. (2020). Multilevel regulation of endoplasmic reticulum stress responses in plants: Where old roads and new paths meet. Journal of Experimental Botany, 71(5), 1659–1667. https://doi.org/10.1093/jxb/erz487.
Anelli, T., & Sitia, R. (2008). Protein quality control in the early secretory pathway. The EMBO journal, 27(2), 315–327. https://doi.org/10.1038/sj.emboj.7601974.
Angelos, E., Ruberti, C., Kim, S.-J., & Brandizzi, F. (2017). Maintaining the factory: the roles of the unfolded protein response in cellular homeostasis in plants. The Plant journal. https://doi.org/10.1111/tpj.13449.
Bao, Y., Pu, Y., Yu, X., Gregory, B. D., Srivastava, R., Howell, S. H., & Bassham, D. C. (2018). IRE1B degrades RNAs encoding proteins that interfere with the induction of autophagy by ER stress in Arabidopsis thaliana. Autophagy, 14(9), 1562–1573. https://doi.org/10.1080/15548627.2018.1462426.
Barba-Espín, G., Dedvisitsakul, P., Hägglund, P., Svensson, B., & Finnie, C. (2014). Gibberellic acid-induced aleurone layers responding to heat shock or tunicamycin provide insight into the N-glycoproteome, protein secretion, and endoplasmic reticulum stress. Plant Physiology, 164(2), 951–965. https://doi.org/10.1104/pp.113.233163.
Bernales, S., Papa, F. R., & Walter, P. (2006). Intracellular Signaling by the Unfolded Protein Response. Annual Review of Cell and Developmental Biology, 22(1), 487–508. https://doi.org/10.1146/annurev.cellbio.21.122303.120200.
Braakman, I., & Hebert, D. N. (2013). Protein folding in the endoplasmic reticulum. Cold Spring Harbor perspectives in biology, 5(5), a013201. https://doi.org/10.1101/cshperspect.a013201.
Che, P., Bussell, J. D., Zhou, W., Estavillo, G. M., Pogson, B. J., & Smith, S. M. (2010). Signaling from the endoplasmic reticulum activates brassinosteroid signaling and promotes acclimation to stress in Arabidopsis. Science Signaling. https://doi.org/10.1126/scisignal.2001140.
Cui, F., Liu, L., Zhao, Q., Zhang, Z., Li, Q., Lin, B., et al. (2012). Ubiquitin Conjugase UBC32 Is an ERAD Component That Functions in Brassinosteroid-Mediated Salt Stress Tolerance. The Plant Cell, 24(1), 233–244. https://doi.org/10.1105/tpc.111.093062.
Demidchik, V., Shabala, S., Isayenkov, S., Cuin, T. A., & Pottosin, I. (2018). Calcium transport across plant membranes: mechanisms and functions. New Phytologist, 220(1), 49–69. https://doi.org/10.1111/nph.15266.
Deng, Y., Humbert, S., Liu, J. X., Srivastava, R., Rothstein, S. J., & Howell, S. H. (2011). Heat induces the splicing by IRE1 of a mRNA encoding a transcription factor involved in the unfolded protein response in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 108(17), 7247–7252. https://doi.org/10.1073/pnas.1102117108.
Deng, Y., Srivastava, R., & Howell, S. H. (2013a). Endoplasmic reticulum (ER) stress response and its physiological roles in plants. International Journal of Molecular Sciences, 14(4), 8188–8212. https://doi.org/10.3390/ijms14048188.
Deng, Y., Srivastava, R., & Howell, S. H. (2013b). Protein kinase and ribonuclease domains of IRE1 confer stress tolerance, vegetative growth, and reproductive development in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 110(48), 19633–19638. https://doi.org/10.1073/pnas.1314749110.
Deng, Y., Srivastava, R., Quilichini, T. D., Dong, H., Bao, Y., Horner, H. T., & Howell, S. H. (2016). IRE1, a component of the unfolded protein response signaling pathway, protects pollen development in Arabidopsis from heat stress. The Plant Journal, 88(2), 193–204. https://doi.org/10.1111/tpj.13239.
Ernst, H. A., Olsen, A. N., Skriver, K., Larsen, S., & Lo Leggio, L. (2004). Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Reports, 5(3), 297–303. https://doi.org/10.1038/sj.embor.7400093.
Fernández-Bautista, N., Fernández-Calvino, L., Muñoz, A., & Castellano, M. M. (2017). HOP3, a member of the HOP family in Arabidopsis, interacts with BiP and plays a major role in the ER stress response. Plant Cell and Environment, 40(8), 1341–1355. https://doi.org/10.1111/pce.12927.
Fernández-Bautista, N., Fernández-Calvino, L., Muñoz, A., Toribio, R., Mock, H. P., & Castellano, M. M. (2018). HOP family plays a major role in long-term acquired thermotolerance in Arabidopsis. Plant, Cell & Environment, 41(8), 1852–1869. https://doi.org/10.1111/pce.13326.
Fragkostefanakis, S., Mesihovic, A., Hu, Y., & Schleiff, E. (2016). Unfolded protein response in pollen development and heat stress tolerance. Plant Reproduction, 29(1–2), 81–91. https://doi.org/10.1007/s00497-016-0276-8.
Gao, H., Brandizzi, F., Benning, C., & Larkin, R. M. (2008). A membrane-tethered transcription factor defines a branch of the heat stress response in Arabidopsis thaliana. Proceedings of the National Academy of Sciences of the United States of America, 105(42), 16397–16403. https://doi.org/10.1073/pnas.0808463105.
Geng, X., Zang, X., Li, H., Liu, Z., Zhao, A., Liu, J., et al. (2018). Unconventional splicing of wheat TabZip60 confers heat tolerance in transgenic Arabidopsis. Plant Science, 274(May), 252–260. https://doi.org/10.1016/j.plantsci.2018.05.029.
Hayashi, S., Wakasa, Y., Takahashi, H., Kawakatsu, T., & Takaiwa, F. (2012). Signal transduction by IRE1-mediated splicing of bZip50 and other stress sensors in the endoplasmic reticulum stress response of rice. Plant Journal, 69(6), 946–956. https://doi.org/10.1111/j.1365-313X.2011.04844.x.
He, X. J., Mu, R. L., Cao, W. H., Zhang, Z. G., Zhang, J. S., & Chen, S. Y. (2005). AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant Journal, 44(6), 903–916. https://doi.org/10.1111/j.1365-313X.2005.02575.x.
Herath, V., Gayral, M., Adhikari, N., Miller, R., & Verchot, J. (2020). Genome-wide identification and characterization of Solanum tuberosum BiP genes reveals the role of the promoter architecture in BiP gene diversity. bioRxiv. https://doi.org/10.1101/2020.05.16.098244.
Hong, Y., Zhang, H., Huang, L., Li, D., & Song, F. (2016). Overexpression of a stress- responsive NAC transcription factor gene ONAC022 improves drought and salt tolerance in rice. Frontiers in Plant Science. https://doi.org/10.3389/fpls.2016.00004.
Howell, S. H. (2013). Endoplasmic Reticulum Stress Responses in Plants. Annual Review of Plant Biology, 64(1), 477–499. https://doi.org/10.1146/annurev-arplant-050312-120053.
Howell, S. H. (2014). Molecular biology. Molecular Biology. https://doi.org/10.1007/978-1-.
Humbert, S., Zhong, S., Deng, Y., Howell, S. H., & Rothstein, S. J. (2012). Alteration of the bZip60/IRE1 pathway affects plant response to ER stress in Arabidopsis Thaliana. PLoS ONE, 7(6), 1–8. https://doi.org/10.1371/journal.pone.0039023.
Hüttner, S., Veit, C., Schoberer, J., Grass, J., & Strasser, R. (2012). Unraveling the function of Arabidopsis thaliana OS9 in the endoplasmic reticulum-associated degradation of glycoproteins. Plant Molecular Biology, 79(1–2), 21–33. https://doi.org/10.1007/s11103-012-9891-4.
Iwata, Y., Ashida, M., Hasegawa, C., Tabara, K., Mishiba, K.-I., & Koizumi, N. (2017). Activation of the Arabidopsis membrane-bound transcription factor bZip28 is mediated by site-2 protease, but not site-1 protease. The Plant journal, 91(3), 408–415. https://doi.org/10.1111/tpj.13572.
Iwata, Y., Fedoroff, N. V., & Koizumi, N. (2008). Arabidopsis bZip60 is a proteolysis- activated transcription factor involved in the endoplasmic reticulum stress response. The Plant Cell, 20(11), 3107–3121. https://doi.org/10.1105/tpc.108.061002.
Iwata, Y., & Koizumi, N. (2005). An Arabidopsis transcription factor, AtbZip60, regulates the endoplasmic reticulum stress response in a manner unique to plants. Proceedings of the National Academy of Sciences of the United States of America, 102(14), 5280–5285. https://doi.org/10.1073/pnas.0408941102.
Iwata, Y., Sakiyama, M., Lee, M.-H., & Koizumi, N. (2010). Transcriptomic response of Arabidopsis thaliana to tunicamycin-induced endoplasmic reticulum stress. Plant Biotechnology, 27(2), 161–171. https://doi.org/10.5511/plantbiotechnology.27.161.
Iwata, Y., Yoneda, M., Yanagawa, Y., & Koizumi, N. (2009). Characteristics of the nuclear form of the Arabidopsis transcription factor AtbZip60 during the endoplasmic reticulum stress response. Bioscience, Biotechnology and Biochemistry, 73(4), 865–869. https://doi.org/10.1271/bbb.80779.
Jensen, M. K., Kjaersgaard, T., Nielsen, M. M., Galberg, P., Petersen, K., O’Shea, C., & Skriver, K. (2010). The Arabidopsis thaliana NAC transcription factor family: structure–function relationships and determinants of ANAC019 stress signalling. Biochemical Journal, 426(2), 183–196. https://doi.org/10.1042/BJ20091234.
Jung, J.-H., Domijan, M., Klose, C., Biswas, S., Ezer, D., Gao, M., et al. (2016). Phytochromes function as thermosensors in Arabidopsis. Science, 354(6314), 886–889. https://doi.org/10.1126/science.aaf6005.
Kamauchi, S., Nakatani, H., Nakano, C., & Urade, R. (2005). Gene expression in response to endoplasmic reticulum stress in Arabidopsis thaliana. The FEBS Journal, 272(13), 3461–3476. https://doi.org/10.1111/j.1742-4658.2005.04770.x.
Kataoka, R., Takahashi, M., & Suzuki, N. (2017). Coordination between bZip28 and HSFA2 in the regulation of heat response signals in Arabidopsis. Plant Signaling and Behavior, 12(11), 1–6. https://doi.org/10.1080/15592324.2017.1376159.
Koizumi, N., Martinez, I. M., Kimata, Y., Kohno, K., Sano, H., & Chrispeels, M. J. (2001). Molecular characterization of two Arabidopsis Ire1 homologs, endoplasmic reticulum-located transmembrane protein kinases. Plant physiology, 127(3), 949–962.
Kumar, S. V., & Wigge, P. A. (2010). H2A.Z-Containing Nucleosomes Mediate the Thermosensory Response in Arabidopsis. Cell, 140(1), 136–147. https://doi.org/10.1016/j.cell.2009.11.006.
Lai, Y.-S., Stefano, G., Zemelis-Durfee, S., Ruberti, C., Gibbons, L., & Brandizzi, F. (2018). Systemic signaling contributes to the unfolded protein response of the plant endoplasmic reticulum. Nature Communications, 9(1), 3918. https://doi.org/10.1038/s41467-018-.
Li, L.-M., Lü, S.-Y., & Li, R.-J. (2017). The Arabidopsis endoplasmic reticulum associated degradation pathways are involved in the regulation of heat stress response. Biochemical and Biophysical Research Communications, 487(2), 362–367. https://doi.org/10.1016/j.bbrc.2017.04.066.
Li, L. M., Lü, S. Y., & Li, R. J. (2017). The Arabidopsis endoplasmic reticulum associated degradation pathways are involved in the regulation of heat stress response. Biochemical and Biophysical Research Communications, 487(2), 362–367. https://doi.org/10.1016/j.bbrc.2017.04.066.
Li, Y., Humbert, S., & Howell, S. H. (2012). ZmbZip60 mRNA is spliced in maize in response to ER stress. BMC Research Notes. https://doi.org/10.1186/1756-0500-5-144.
Li, Y., Williams, B., & Dickman, M. (2017). Arabidopsis B-cell lymphoma2 (Bcl-2)- associated athanogene 7 (BAG7)-mediated heat tolerance requires translocation, sumoylation and binding to WRKY29. New Phytologist, 214(2), 695–705. https://doi.org/10.1111/nph.14388.
Lin, L., Zhang, C., Chen, Y., Wang, Y., Wang, D., Liu, X., et al. (2019). PAWH1 and PAWH2 are plant-specific components of an Arabidopsis endoplasmic reticulum-associated degradation complex. Nature Communications, 10(1), 3492. https://doi.org/10.1038/s41467-019-11480-7.
Liu, J., Feng, L., Li, J., & He, Z. (2015). Geneticandepigeneticcontrolofplantheatresponses. Frontiers in Plant Science, 6, 1–21. https://doi.org/10.3389/fpls.2015.00267.
Liu, J. X., & Howell, S. H. (2010). Endoplasmic reticulum protein quality control and its relationship to environmental stress responses in plants. The Plant Cell, 22(9), 2930–2942. https://doi.org/10.1105/tpc.110.078154.
Liu, J. X., & Howell, S. H. (2016). Managing the protein folding demands in the endoplasmic reticulum of plants. The New phytologist, 211(2), 418–428. https://doi.org/10.1111/nph.13915.
Liu, L., Cui, F., Li, Q., Yin, B., Zhang, H., Lin, B., et al. (2011). The endoplasmic reticulum- associated degradation is necessary for plant salt tolerance. Cell research, 21(6), 957–969. https://doi.org/10.1038/cr.2010.181.
Liu, L., & Li, J. (2019). Communications between the endoplasmic reticulum and other organelles during abiotic stress response in plants. Frontiers in Plant Science. https://doi.org/10.3389/fpls.2019.00749.
Liu, X. H., Lyu, Y. S., Yang, W., Yang, Z. T., Lu, S. J., & Liu, J. X. (2020). A membrane- associated NAC transcription factor OsNTL3 is involved in thermotolerance in rice. Plant Biotechnology Journal, 18(5), 1317–1329. https://doi.org/10.1111/pbi.13297.
Liu, Y., Burgos, J. S., Deng, Y., Srivastava, R., Howell, S. H., & Bassham, D. C. (2012). Degradation of the endoplasmic reticulum by autophagy during endoplasmic reticulum stress in Arabidopsisc w. The Plant Cell, 24(11), 4635–4651. https://doi.org/10.1105/tpc.112.101535.
Liu, Y., Zhang, C., Wang, D., Su, W., Liu, L., Wang, M., & Li, J. (2015). EBS7 is a plant- specific component of a highly conserved endoplasmic reticulum-associated degradation system in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 112(39), 12205–12210.
Lu, S. J., Yang, Z. T., Sun, L., Sun, L., Song, Z. T., & Liu, J. X. (2012). Conservation of IRE1- regulated bZip74 mRNA unconventional splicing in rice (Oryza sativa L.) involved in ER stress responses. Molecular Plant, 5(2), 504–514. https://doi.org/10.1093/mp/ssr115.
Ma, Z. X., Leng, Y. J., Chen, G. X., Zhou, P. M., Ye, D., & Chen, L. Q. (2015). The thermosensitive male sterile 1 interacts with the BiPs via DnaJ domain and stimulates their ATPase enzyme activities in arabidopsis. PLoS ONE, 10(7), 1–13. https://doi.org/10.1371/journal.pone.0132500.
Meng, Z., Ruberti, C., Gong, Z., & Brandizzi, F. (2017). CPR5 modulates salicylic acid and the unfolded protein response to manage tradeoffs between plant growth and stress responses. Plant Journal, 89(3), 486–501. https://doi.org/10.1111/tpj.13397.
Mishiba, K. I., Nagashima, Y., Suzukia, E., Hayashi, N., Ogata, Y., Shimada, Y., & Koizumi, N. (2013). Defects in IRE1 enhance cell death and fail to degrade mRNAs encoding secretory pathway proteins in the Arabidopsis unfolded protein response. Proceedings of the National Academy of Sciences of the United States of America, 110(14), 5713–5718. https://doi.org/10.1073/pnas.1219047110.
Mittler, R., Finka, A., & Goloubinoff, P. (2012). How do plants feel the heat? Trends in Biochemical Sciences, 37(3), 118–125. https://doi.org/10.1016/j.tibs.2011.11.007.
Moreno, A. A., Mukhtar, M. S., Blanco, F., Boatwright, J. L., Moreno, I., Jordan, M. R., et al. (2012). IRE1/bZip60-mediated unfolded protein response plays distinct roles in plant immunity and abiotic stress responses. PLoS ONE, 7(2), E31944. https://doi.org/10.1371/journal.pone.0031944.
Mori, K., Kawahara, T., Yoshida, H., Yanagi, H., & Yura, T. (1996). Signalling from endoplasmic reticulum to nucleus: transcription factor with a basic-leucine zipper motif is required for the unfolded protein-response pathway. Genes to Cells, 1(9), 803–817.
Nagashima, Y., Iwata, Y., Ashida, M., Mishiba, K., & Koizumi, N. (2014). Exogenous salicylic acid activates two signaling arms of the unfolded protein response in Arabidopsis. Plant and Cell Physiology, 55(10), 1772–1778. https://doi.org/10.1093/pcp/pcu108.
Nagashima, Y., Mishiba, K. I., Suzuki, E., Shimada, Y., Iwata, Y., & Koizumi, N. (2011). Arabidopsis IRE1 catalyses unconventional splicing of bZip60 mRNA to produce the active transcription factor. Scientific Reports. https://doi.org/10.1038/srep00029.
Nawkar, G. M., Kang, C. H., Maibam, P., Park, J. H., Jung, Y. J., Chae, H. B., et al. (2017). HY5, a positive regulator of light signaling, negatively controls the unfolded protein response in Arabidopsis. Proceedings of the National Academy of Sciences. https://doi.org/10.1073/pnas.1609844114.
Nawkar, G. M., Lee, E. S., Shelake, R. M., Park, J. H., Ryu, S. W., Kang, C. H., & Lee, S. Y. (2018). Activation of the transducers of unfolded protein response in plants. Frontiers in Plant Science, 9, 1–10. https://doi.org/10.3389/fpls.2018.00214.
Neill, E. M., Byrd, M. C. R., Billman, T., Brandizzi, F., & Stapleton, A. E. (2019). Plant growth regulators interact with elevated temperature to alter heat stress signaling via the Unfolded Protein Response in maize. Scientific Reports, 9(1), 1–10. https://doi.org/10.1038/s41598-019-46839-9.
Nishikawa, S., Brodsky, J. L., & Nakatsukasa, K. (2005). roles of molecular chaperones in endoplasmic reticulum (ER) quality control and er-associated degradation (ERAD). The Journal of Biochemistry, 137(5), 551–555. https://doi.org/10.1093/jb/mvi068.
Niu, Y., & Xiang, Y. (2018). An overview of biomembrane functions in plant responses to high-temperature stress. Frontiers in Plant Science, 9(July), 1–18. https://doi.org/10.3389/fpls.2018.00915.
Nuruzzaman, M., Sharoni, A. M., & Kikuchi, S. (2013). Roles ofNAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Frontiers in Microbiology, 4, 1–16. https://doi.org/10.3389/fmicb.2013.00248.
Okushima, Y., Koizumi, N., Yamaguchi, Y., Kimata, Y., Kohno, K., & Sano, H. (2002). Isolation and characterization of a putative transducer of endoplasmic reticulum stress in Oryza sativa. Plant and Cell Physiology, 43(5), 532–539. https://doi.org/10.1093/pcp/pcf063.
Olsen, A. N., Ernst, H. A., Leggio, L. L., & Skriver, K. (2005). NAC transcription factors: structurally distinct, functionally diverse. Trends in Plant Science, 10(2), 79–87. https://doi.org/10.1016/j.tplants.2004.12.010.
Onda, Y., Kumamaru, T., & Kawagoe, Y. (2009). ER membrane-localized oxidoreductase Ero1 is required for disulfide bond formation in the rice endosperm. Proceedings of the National Academy of Sciences of the United States of America, 106(33), 14156–14161. https://doi.org/10.1073/pnas.0904429106.
Ooka, H., Satoh, K., Doi, K., Nagata, T., Otomo, Y., Murakami, K., et al. (2003). Comprehensive Analysis of NAC Family Genes in Oryza sativa and Arabidopsis thaliana. DNA Research, 10(6), 239–247. https://doi.org/10.1093/dnares/10.6.239.
Park, C. J., & Park, J. M. (2019). Endoplasmic reticulum plays a critical role in integrating signals generated by both biotic and abiotic stress in plants. Frontiers in Plant Science. https://doi.org/10.3389/fpls.2019.00399.
Park, J. H., Kang, C. H., Nawkar, G. M., Lee, E. S., Paeng, S. K., Chae, H. B., et al. (2018). EMR, a cytosolic-abundant ring finger E3 ligase, mediates ER-associated protein degradation in Arabidopsis. New Phytologist, 220(1), 163–177. https://doi.org/10.1111/nph.15279.
Pastor-Cantizano, N., Ko, D. K., Angelos, E., Pu, Y., & Brandizzi, F. (2020). Functional Diversification of ER Stress Responses in Arabidopsis. Trends in Biochemical Sciences, 45(2), 123–136. https://doi.org/10.1016/j.tibs.2019.10.008.
Patil, C., & Walter, P. (2001). Intracellular signaling from the endoplasmic reticulum to the nucleus: the unfolded protein response in yeast and mammals. Current Opinion in Cell Biology, 13(3), 349–355. https://doi.org/10.1016/s0955-0674(00)00219-2.
Pu, Y., Ruberti, C., Angelos, E. R., & Brandizzi, F. (2019). AtIRE1C, an unconventional isoform of the UPR master regulator AtIRE1, is functionally associated with AtIRE1B in Arabidopsis gametogenesis. Plant Direct. https://doi.org/10.1002/pld3.187.
Puranik, S., Sahu, P. P., Srivastava, P. S., & Prasad, M. (2012). NAC proteins: regulation and role in stress tolerance. Trends in Plant Science, 17(6), 369–381. https://doi.org/10.1016/j.tplants.2012.02.004.
Quilichini, T. D., Douglas, C. J., & Samuels, A. L. (2014). New views of tapetum ultrastructure and pollen exine development in Arabidopsis thaliana. Annals of Botany, 114(6), 1189–1201. https://doi.org/10.1093/aob/mcu042.
Raja, M. M., Vijayalakshmi, G., Naik, M. L., Basha, P. O., Sergeant, K., Hausman, J. F., & Khan, P. S. S. V. (2019). Pollen development and function under heat stress: from effects to responses. Acta Physiologiae Plantarum, 41(4), 1–20. https://doi.org/10.1007/s11738-019-2835-8.
Ramakrishna, C., Singh, S., Raghavendrarao, S., Padaria, J. C., Mohanty, S., Sharma, T. R., & Solanke, A. U. (2018). The membrane tethered transcription factor EcbZip17 from finger millet promotes plant growth and enhances tolerance to abiotic stresses. Scientific Reports, 8(1), 1–14. https://doi.org/10.1038/s41598-018-19766-4.
Ron, D., & Walter, P. (2007). Signal integration in the endoplasmic reticulum unfolded protein response. Nature reviews. Molecular cell biology, 8(7), 519–529. https://doi.org/10.1038/nrm2199.
Ruddock, L. W., & Molinari, M. (2006). N-glycan processing in ER quality control. Journal of Cell Science, 119(21), 4373–4380. https://doi.org/10.1242/jcs.03225.
Ruelland, E., & Zachowski, A. (2010). How plants sense temperature. Environmental and Experimental Botany, 69(3), 225–232. https://doi.org/10.1016/j.envexpbot.2010.05.011.
Saidi, Y., Finka, A., Muriset, M., Bromberg, Z., Weiss, Y. G., Maathuis, F. J. M., & Goloubinoff, P. (2009). The heat shock response in moss plants is regulated by specific calcium-permeable channels in the plasma membrane. The Plant Cell, 21(9), 2829–2843. https://doi.org/10.1105/tpc.108.065318.
Schwarz, D. S., & Blower, M. D. (2016). The endoplasmic reticulum: structure, function and response to cellular signaling. Cellular and Molecular Life Sciences, 73(1), 79–94. https://doi.org/10.1007/s00018-015-2052-6.
Seo, P. J. (2014). Recent advances in plant membrane-bound transcription factor research: Emphasis on intracellular movement. Journal of Integrative Plant Biology, 56(4), 334–342. https://doi.org/10.1111/jipb.12139.
Seo, P. J., & Park, C. (2010). A membrane bound transcription factor as an integrator of biotic and abiotic stress signals. Plant Signaling and Behaviour., 5(5), 481–483.
Shamu, C. E., & Walter, P. (1996). Oligomerization and phosphorylation of the Ire1p kinase during intracellular signaling from the endoplasmic reticulum to the nucleus. The EMBO Journal, 15(12), 3028–3039.
Shao, H., Wang, H., & Tang, X. (2015). NAC transcription factors in plant multiple abiotic stress responses: progress and prospects. Frontiers in Plant Science, 6, 902. https://doi.org/10.3389/fpls.2015.00902.
Sita, K., Sehgal, A., Hanumantharao, B., Nair, R. M., Vara Prasad, P. V., Kumar, S., et al. (2017). Food legumes and rising temperatures: Effects, adaptive functional mechanisms specific to reproductive growth stage and strategies to improve heat tolerance. Frontiers in Plant Science, 8, 1–30. https://doi.org/10.3389/fpls.2017.01658.
Srivastava, R., Deng, Y., & Howell, S. H. (2014). Stress sensing in plants by an ER stress sensor/transducer, bZip28. Frontiers in Plant Science, 5, 1–6. https://doi.org/10.3389/fpls.2014.00059.
Srivastava, R., Li, Z., Russo, G., Tang, J., Bi, R., Muppirala, U., et al. (2018). Response to persistent er stress in plants: A multiphasic process that transitions cells from prosurvival activities to cell death. The Plant Cell, 30(6), 1220–1242. https://doi.org/10.1105/tpc.18.00153.
Strasser, R. (2018). Protein quality control in the endoplasmic reticulum of plants. Annual Review of Plant Biology, 69, 147–172. https://doi.org/10.1146/annurev-arplant-042817-040331.
Sun, L., Yang, Z. T., Song, Z. T., Wang, M. J., Sun, L., Lu, S. J., & Liu, J. X. (2013). The plant-specific transcription factor gene NAC103 is induced by bZip60 through a new cis- regulatory element to modulate the unfolded protein response in Arabidopsis. Plant Journal, 76(2), 274–286. https://doi.org/10.1111/tpj.12287.
Takahashi, H., Kawakatsu, T., Wakasa, Y., Hayashi, S., & Takaiwa, F. (2012). A rice transmembrane bZip transcription factor, OsbZip39, regulates the endoplasmic reticulum stress response. Plant and Cell Physiology, 53(1), 144–153. https://doi.org/10.1093/pcp/pcr157.
Verchot, J. (2016). How does the stressed out ER find relief during virus infection? Current Opinion in Virology, 17, 74–79. https://doi.org/10.1016/j.coviro.2016.01.018.
Walter, P., & Ron, D. (2011). The unfolded protein response: from stress pathway to homeostatic regulation. Science, 334(6059), 1081–1086. https://doi.org/10.1126/science.1209038.
Wang, H., Niu, H., Zhai, Y., & Lu, M. (2017). Characterization of BiP genes from pepper (Capsicum annuum L.) and the role of CaBiP1 in response to endoplasmic reticulum and multiple abiotic stresses. Frontiers in Plant Science, 8, 1–15. https://doi.org/10.3389/fpls.2017.01122.
Ward, J. M., Mäser, P., & Schroeder, J. I. (2009). Plant ion channels: gene families, physiology, and functional genomics analyses. Annual Review of Physiology, 71, 59–82. https://doi.org/10.1146/annurev.physiol.010908.163204.
Williams, B., Kabbage, M., Britt, R., & Dickman, M. B. (2010). AtBAG7 an Bcl-2-associated athanogene, resides in the endoplasmic reticulum and is involved in the unfolded protein response. Proceedings of the National Academy of Sciences, 107(13), 6088–6093. https://doi.org/10.1073/pnas.0912670107.
Xie, Q., Frugis, G., Colgan, D., & Chua, N. H. (2000). Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes and Development, 14(23), 3024–3036. https://doi.org/10.1101/gad.852200.
Yamamoto, K., Yoshida, H., Kokame, K., Kaufman, R. J., & Mori, K. (2004). Differential contributions of ATF6 and XBP1 to the activation of endoplasmic reticulum stress- responsive cis-acting elements ERSE UPRE and ERSE-II. Journal of Biochemistry, 136(3), 343–350. https://doi.org/10.1093/jb/mvh122.
Yamamoto, M., Maruyama, D., Endo, T., & Nishikawa, S. (2008). Arabidopsis thaliana has a set of J proteins in the endoplasmic reticulum that are conserved from yeast to animals and plants. Plant and Cell Physiology, 49(10), 1547–1562. https://doi.org/10.1093/pcp/pcn119.
Yang, X., Srivastava, R., Howell, S. H., & Bassham, D. C. (2016). Activation of autophagy by unfolded proteins during endoplasmic reticulum stress. Plant Journal, 85(1), 83–95. https://doi.org/10.1111/tpj.13091.
Yángüez, E., Castro-Sanz, A. B., Fernández-Bautista, N., Oliveros, J. C., & Castellano, M. M. (2013). Analysis of genome-wide changes in the translatome of Arabidopsis seedlings subjected to heat Stress. PLoS ONE, 8(8), e71425. https://doi.org/10.1371/journal.pone.0071425.
Yu, C. Y., & Kanehara, K. (2020). The unfolded protein response modulates a phosphoinositide-binding protein through the IRE1-bZip60 pathway1[open]. Plant Physiology, 183(5), 221–235. https://doi.org/10.1104/pp.19.01488.
Yuan, X., Wang, H., Cai, J., Bi, Y., Li, D., & Song, F. (2019). Rice NAC transcription factor ONAC066 functions as a positive regulator of drought and oxidative stress response. BMC Plant Biology, 19(1), 1–19. https://doi.org/10.1186/s12870-019-1883-y.
Zeng, Y., Chung, K. P., Li, B., Lai, C. M., Lam, S. K., Wang, X., et al. (2015). Unique COPII component AtSar1a/AtSec23a pair is required for the distinct function of protein ER export in Arabidopsis. Proceedings of the National Academy of Sciences, 112(46), 14360–14365. https://doi.org/10.1073/pnas.1519333112.
Zhang, L., Zhang, L., Xia, C., Zhao, G., Liu, J., Jia, J., & Kong, X. (2015). AnovelwheatbZip transcription factor, TabZip60, confers multiple abiotic stress tolerances in transgenic Arabidopsis. Physiologia Plantarum, 153(4), 538–554. https://doi.org/10.1111/ppl.12261.
Zhang, R., Chen, H., Duan, M., Zhu, F., Wen, J., Dong, J., & Wang, T. (2019). Medicago falcata MfSTMIR, an E3 ligase of endoplasmic reticulum-associated degradation, is involvedinsaltstressresponse. The Plant Journal, 98(4), 680–696. https://doi.org/10.1111/tpj.14265.
Zhang, S.-S., Yang, H., Ding, L., Song, Z.-T., Ma, H., Chang, F., & Liu, J.-X. (2017). Tissue-specific transcriptomics reveals an important role of the unfolded protein response in maintaining fertility upon heat stress in Arabidopsis. The Plant cell, 29(5), 1007–1023. https://doi.org/10.1105/tpc.16.00916.
Zhao, C., Liu, B., Piao, S., Wang, X., Lobell, D. B., Huang, Y., et al. (2017). Temperature increase reduces global yields of major crops in four independent estimates. Proceedings of the National Academy of Sciences, 114(35), 9326–9331. https://doi.org/10.1073/pnas.1701762114.
Zhong, R., Demura, T., & Ye, Z.-H. (2006). SND1, a NAC domain transcription factor, is a key regulator of secondary wall synthesis in fibers of arabidopsis. The Plant Cell, 18(11), 3158–3170. https://doi.org/10.1105/tpc.106.047399.
Zhou, J., Wang, J., Cheng, Y., Chi, Y. J., Fan, B., Yu, J. Q., & Chen, Z. (2013). NBR1-mediated selective autophagy targets insoluble ubiquitinated protein aggregates in plant stress responses. PLoS Genetics, 9(1), e1003196. https://doi.org/10.1371/journal.pgen.1003196.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Ramalingaswami Fellow-DBT.
Rights and permissions
About this article
Cite this article
Malini, M.K., Lekshmy, V.S., Pal, M. et al. Unfolded protein response (UPR) mediated under heat stress in plants. Plant Physiol. Rep. 25, 569–582 (2020). https://doi.org/10.1007/s40502-020-00548-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-020-00548-y