Abstract
The aim of this review was to provide an evidence-based approach to frequently asked questions relating to the risk of transmitting a maternally inherited mitochondrial disorder (MID). We do not address disorders linked with disturbed mitochondrial DNA (mtDNA) maintenance, causing mtDNA depletion or multiple mtDNA deletions, as these are autosomally inherited. The review addresses questions regarding prognosis, recurrence risks and the strategies available to prevent disease transmission. The clinical and genetic complexity of maternally inherited MIDs represent a major challenge for patients, their relatives and health professionals. Since many of the genetic and pathophysiological aspects of MIDs remain unknown, counselling of affected patients and at-risk family members remains difficult. MtDNA mutations are maternally transmitted or, more rarely, they are sporadic, occurring de novo (~25%). Females carrying homoplasmic mtDNA mutations will transmit the mutant species to all of their offspring, who may or may not exhibit a similar phenotype depending on modifying, secondary factors. Females carrying heteroplasmic mtDNA mutations will transmit a variable amount of mutant mtDNA to their offspring, which can result in considerable phenotypic heterogeneity among siblings. The majority of mtDNA rearrangements, such as single large-scale deletions, are sporadic, but there is a small risk of recurrence (~4%) among the offspring of affected women. The range and suitability of reproductive choices for prospective mothers is a complex area of mitochondrial medicine that needs to be managed by experienced healthcare professionals as part of a multidisciplinary team. Genetic counselling is facilitated by the identification of the underlying causative genetic defect. To provide more precise genetic counselling, further research is needed to clarify the secondary factors that account for the variable penetrance and the often marked differential expressivity of pathogenic mtDNA mutations both within and between families.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Genetic counselling for maternally inherited mitochondrial disorders (MIDs) is challenging due to their wide phenotypic and genotypic heterogeneity. |
Reproductive choices are now available for mothers carrying pathogenic mitochondrial DNA mutations. |
1 Introduction
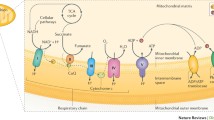
Mitochondrial disorders (MIDs) constitute the most common group of metabolic disorders, with an estimated prevalence of 1:4200 in the population [1, 2] and diagnosed in >1 in 7500 live births [3]. Furthermore, unrecognised oligosymptomatic MIDs have a much higher prevalence than previously thought at about 1 in 400 [4]. MIDs are usually multisystem disorders, either at onset or evolving from single to multiple organ involvement as the disease progresses. The onset of disease can occur any time from the neonatal period to old age. Any combination of tissues or organs can occur, and the severity can be highly variable depending on the underlying mutation, the mutation load and disease duration [5]. MIDs are classically associated with respiratory chain defects that affect the mitochondrial electron transport chain, but it is now well-established that impairment of other mitochondrial pathways namely, β-oxidation, iron metabolism, copper trafficking, mitochondrial translation, cardiolipin remodelling (Barth syndrome), the acyl-glycerol kinase pathway (Sengers syndrome), heme synthesis, the pyruvate dehydrogenase complex (PDC deficiency) or the ketoglutarate dehydrogenase complexes may also cause MIDs [6]. MIDs can be caused by mitochondrial DNA (mtDNA) mutations (primary MIDs) or by mutations in nuclear DNA (nDNA)-located genes (nuclear MIDs). The mitochondrial genome encodes for 37 genes whereas over 1300 nuclear genes encode for critical components of the mitochondrial machinery [7]. MIDs caused by nuclear genetic defects can therefore follow an autosomal dominant (AD), autosomal recessive (AR) or X-linked pattern of inheritance. Most MIDs caused by nDNA-located genes are transmitted via an AR pattern and more rarely via an AD pattern [8]. Pathogenic mtDNA mutations are strictly maternally transmitted, but sporadic mtDNA mutations, in particular single large-scale mtDNA deletions, can also occur. Because of the complex interplay between the mitochondrial and nuclear genomes and the difficulties in predicting the transmission of heteroplasmic mtDNA mutations, genetic counselling for maternally inherited MIDs remains challenging [9]. This literature review addresses clinically relevant questions about disease penetrance, prognosis, recurrence risks and the options for preventing maternal transmission of a pathogenic mtDNA mutation to the next generation.
2 Methods
The data included in this review were identified by a literature search of MEDLINE, Current Contents, EMBASE, Web of Science, Web of Knowledge, LILACS (Literatura Latino Americana em Ciências da Saúde), SCOPUS and Google Scholar using the search terms “maternal inheritance”, “maternally inherited” and “matrilineal”, in combination with “genetic counselling”. Randomised (blinded or open-label) clinical trials, longitudinal studies, case series and case reports were considered. Only articles published in English, French, Spanish or German between 1966 and 2017 were assessed. Appropriate papers were studied and discussed before making a decision on their suitability for inclusion in this review.
3 Results
3.1 Mitochondrial Genome
The human mitochondrial genome is a circular, double-stranded DNA molecule consisting of 16,569 base pairs (bp). It encodes for 13 proteins, which are all essential components of the mitochondrial respiratory chain complexes, two ribosomal RNAs (rRNAs) and 22 transfer RNAs (tRNAs) [10]. Except for the D-loop, which is involved in the initiation of mtDNA replication and transcription, mtDNA is a very compact molecule with overlapping gene regions and there is little non-coding DNA [10]. mtDNA does not contain introns and the genetic code varies from the universal code. A single mitochondrion usually contains five to ten mtDNA copies. Since a single somatic cell can contain up to 1000 mitochondria, <10,000 mtDNA copies are present within a single somatic cell. MtDNA copy number in germ cells differs from that in somatic cells. In mature mammalian oocytes, mtDNA copy number goes up to 100,000–200,000 whereas sperm cells only contain about 100 mtDNA copies [11]. Overall, mtDNA represents ~1% of the total DNA content of a human cell.
3.2 Pathogenic Mitochondrial DNA (mtDNA) Mutations
Primary MIDs can arise from either mtDNA point mutations or genomic rearrangements, namely insertions, duplications, triplications or single large-scale deletions [12]. Primary mtDNA mutations may be present in all mitochondrial genomes of a cell (homoplasmy) or only in some of them (heteroplasmy). In the homoplasmic state, all of the mtDNA molecules in a cell harbour the same mtDNA sequence at a particular nucleotide position, whereas in the heteroplasmic state there is a mixture of two or more distinct types of mitochondrial genome. In the context of a pathogenic mtDNA mutation, the level of the mutant species can have an influence both on penetrance (threshold effect) and the severity of disease expression [12]. Many MIDs only become clinically manifest when the mutant level exceeds a certain threshold, but this figure does vary widely between different mtDNA mutations [3] from as low as 8% [13] to almost homoplasmy. While the frequency of heteroplasmic mtDNA mutations has been estimated to be 1 in 200 in the general population [14], next-generation deep sequencing suggests that it is universal at low levels [15].
Nuclear genes that cause MIDs can result in mtDNA maintenance defects in the form of either multiple mtDNA deletions or mtDNA depletion. A rapidly expanding list of nuclear genes causing multiple mtDNA deletions have now been identified, including POLG1, C10orf2 (PEO1), TK2, RRM2B, RNASEH1 and OPA1 [16]. Depletion is present when the amount of mtDNA is decreased by >70% of normal [17]. Mitochondrial depletion syndromes have been reported in patients carrying mutations in nuclear genes such as POLG1, TK2, SUCLA2, MPV17, FBXL4, RRM2B, TYMP, PEO1, SUCLG1, DGUOK, or OPA1 [18]. These nuclear genes are outside the scope of this review because they cause disease with an AD or AR mode of inheritance.
3.2.1 MtDNA Point Mutations
MtDNA point mutations are maternally inherited, but de novo mutations can occur (~25%) [18], which are usually in the heteroplasmic state [19,19,20,21,22]. They can occur in the 22 tRNAs, the two rRNAs, one of the 13 protein-encoding genes, or in non-coding regions. The identification of a mtDNA mutation in only the proband (de novo mutation) can be due to: (1) other family members in the maternal lineage harbouring the mutation, but below the detection limit for the laboratory assay used; (2) the proband is not biologically related to other family members; or (3) the mutation has truly arisen de novo [21], with [22] or without germline mosaicism.
3.2.2 MtDNA Rearrangements
3.2.2.1 Single, large-scale mtDNA Deletions
Compared with mtDNA point mutations, single large-scale mtDNA deletions usually represent de novo mutational events occurring either in an oocyte or during embryogenesis [5]. Therefore, only the proband is usually affected in the family, but there is nevertheless a low risk of transmission by the mother of an affected child. The recurrence risk for a single, large-scale mtDNA deletion has been estimated at 1 in 117 (4.1%) [20]. Single, large-scale mtDNA deletions have a wide phenotypic spectrum ranging from isolated chronic progressive external ophthalmoplegia to the more severe Kearns-Sayre Syndrome (KSS) or Pearson Syndrome (PS) [5]. Since not all large-scale mtDNA rearrangements can be detected reliably in blood samples, muscle is the preferred tissue for long-range PCR analysis [5].
3.2.2.2 Insertions, Duplications and Triplications
Pathogenic mtDNA insertions, duplications or triplications have been occasionally reported as causing MIDs, either as large-scale [23] or small-scale [24] rearrangements. In a particular case with maternally inherited diabetes and deafness (MIDD) and complex-I deficiency in fibroblasts, mtDNA sequencing revealed a heteroplasmic insertion of one or two cytosine residues in the coding region of the MT-ND6 gene (m.14535_14536insC or CC), leading to a premature stop codon [25]. A 9-bp triplication in the MT-CYTB gene was detected in an African patient with isolated mitochondrial myopathy [24]. In a patient with dilated cardiomyopathy, the causative mutation was a 15 bp duplication in the tRNA Pro gene [26].
3.2.3 Confirmation of a Mutation as Pathogenic
Because the mutation rate of mtDNA is rapid, neutral polymorphisms are common. Distinguishing between pathogenic mtDNA mutations and neutral mtDNA variants can generally be achieved using several approaches:
-
1.
In silico computational prediction, applying programs such as CADD, CONDEL, FATHMM, MutationAssessor, PANTHER, PolyPhen-2, PROVEAN or SIFT. Computational algorithms predict the pathogenicity of a mutation based on the phylogenetic similarity among homologues (orthologues and paralogues), 3-dimensional structures, and epidemiological, clinical and genetic characteristics of the variants.
-
2.
In vitro biochemical methods, which include confirmation of decreased mitochondrial enzyme activity in muscle, liver or fibroblasts or in an in vitro transmitochondrial cybrid system. The latter is an artificial eukaryotic cell line produced by introducing mitochondria from one organism into a cell of the same or different species, from which the mtDNA has been removed. A disadvantage of in vitro studies is that they cannot represent the physiological condition and cannot test as yet unrecognised activities or functions [27, 28].
-
3.
By correlating the mutation load with the histochemical phenotype in skeletal muscle.
-
4.
In vivo studies of knock-in transgenic animals [27].
Additionally, a mutation can be classified as pathogenic if the mutation is heteroplasmic, if the mutation is not present in healthy controls, if the type of mutation and the mutated gene are consistent with biochemical and histopathological findings, if the mutation is located in a structurally and functionally important region, or if the amino acid involved is evolutionarily conserved [29, 30].
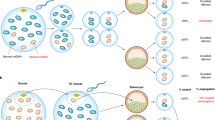
3.3 The Mitochondrial Bottleneck
An important phenomenon during early oogenesis is the marked reduction in mtDNA copy number within the maturing cell from 200,000 copies to 200–2000 copies [31], which is referred to as the bottleneck effect [3, 5, 32]. Although the effective size of the germline mtDNA bottleneck has no precise figure, it has been estimated as between 10 and ~30-35 [31, 33]. If a mother carries a heteroplasmic mtDNA mutation, the mitochondrial bottleneck can result in significant shifts in the level of the mutant species in her population of oocytes. The mechanisms driving this bottleneck effect is under intense investigation and there is still ongoing debate whether it is an entirely random event and/or determined by the nature of the underlying mtDNA mutation [34]. Homoplasmic mtDNA mutations are not affected by the bottleneck effect.
3.3.1 Random Bottleneck Effect
The bottleneck effect is frequently defined as a rapid intergenerational switching of mtDNA types that has been largely finished by the time oocytes are mature [35]. This seems to occur during the expansion in mtDNA copy number that occurs between early and late oogenesis [33, 36,36,37,38]. The major component of the bottleneck occurs between the primordial germ cell and the primary oocyte stage [35]. The mitochondrial bottleneck effect may explain why an asymptomatic mother or a mother with a mild MID can have offspring with a highly variable phenotype ranging from normal to severe life-threatening phenotypes [5]. The bottleneck effect may also explain the intrafamilial phenotypic heterogeneity within a generation and between generations. However, the mean amount of mutant mtDNA per cell in the offspring and in their oocytes (for female children) is likely to be close to that of the mother [35, 39, 40].
3.3.2 Conditional Bottleneck Effect
There is preliminary evidence that the shift in heteroplasmy levels during oogenesis may not only be due to a random drift process, but could also be influenced by the specific nature of a particular mtDNA variant, implying that the mitochondrial bottleneck could have selective properties [14, 39]. In one study of 577 mother–child pairs transmitting the m.11778G > A, m.3460G > A, m.8344A > G, m.8993T > G/C and m.3243A > G mtDNA mutations, there was no evidence for selection during transmission, but in another study there appeared to be selection against novel heteroplasmies, which are likely to be deleterious [41]. Different mutations segregated at different rates in human pedigrees [14]. Patients harbouring the m.8993T > G/C mutation showed more rapid segregation of heteroplasmy than the m.11778G > A, m.8344A > G or m.3243A > G mutations [14]. This trend is consistent with a tighter mitochondrial bottleneck in m.8993T > G/C pedigrees, suggesting that there are different bottlenecks depending on the underlying mtDNA mutation [14]. It is therefore conceivable that non-synonymous, single-base pair substitutions in the background mtDNA haplotype could also result in geographic, ethnic or inter-familial differences of segregation patterns [12]. To make the story more complex, genotypic shifts probably occur post-zygotic [42, 43] as has been shown in mice [42]. These intriguing findings need to be explored further to better understand the different inheritance patterns observed in human pedigrees transmitting pathogenic mtDNA mutations [14].
3.4 Mitochondrial Heteroplasmy
Heteroplasmy levels can vary considerably, not only within a generation or between generations but also between tissues and organs of the same individual. Conceptually, higher levels of the mutant mtDNA species should result in a more severe phenotype, but there is not always a straightforward correlation between mutant load and disease severity [44]. For example, a 48-year-old woman with diabetes mellitus, hearing loss and retinal dystrophy carried the m.3243A > G mutation at heteroplasmy rates of 8% in blood, 6% in hair follicles and 16% in buccal mucosa cells, whereas her asymptomatic 28-year-old son carried the mutation at higher levels of 23, 15 and 16%, respectively [30]. An important conclusion to note from this is that the amount of heteroplasmy in blood or unaffected tissues does not always reflect the mutant load in affected post-mitotic tissues [30]. The mutant load is almost always lower in blood than in muscle and for the m.3243A > G mutation there is good evidence that it falls with time [45]. The load of mutant mtDNA appears to be relatively uniform at birth [46], but often falls in blood [44] and increases in post-mitotic tissues such as brain and muscle [9, 47]. The mechanisms underlying these changes are not well-understood but likely depend both on differences between wild-type and mutant mtDNA in the rate of mtDNA synthesis [48] and mtDNA turnover. Decrease of the m.3243A > G mutant load in blood could be also due to death of highly mutated stem cells in the hematopoietic tissue [49]. This makes estimating mutant loads difficult, as levels of mutant mtDNA in blood, which is accessible for sampling, may be a poor reflection of the mutant load in tissues causing symptoms. In other patients with MIDs, however, the level of heteroplasmy correlates more closely, but not invariably, with the phenotype, such as in carriers of the m.8993T > G mutation [30]. Depending on the mutant load, a carrier of this mutation will develop a severe neurodegenerative Leigh-like syndrome (>90%), the neuropathy, ataxia, and retinitis pigmentosa (NARP) phenotype characterised by neurogenic weakness, ataxia and retinitis pigmentosa (60–75%), or remain asymptomatic (<60%) [30]. The mutant load for these groupings are correspondingly higher for the milder m.8993T > C mutation [50].
As a result of the mitochondrial bottleneck, the degree of heteroplasmy varies considerably in the oocyte population of a female carrier harbouring a heteroplasmic mtDNA mutation [51]. Prenatal testing, in which mtDNA is sampled through amniocentesis or chorionic villus sampling (CVS) therefore carries the risk that it may not accurately reflect the mutation load in the child’s tissues, especially those most at risk of disease [52]. Nevertheless, it definitely reduces the risk of severe disease and some couples find this acceptable [45]. Phenotypic heterogeneity can be additionally explained by the involvement of disease-modifying nuclear genes [30].
3.5 Variable Penetrance
Some mtDNA mutations exist mostly in the homoplasmic state although, despite this, they can exhibit variable penetrance. A classical example is Leber hereditary optic neuropathy (LHON), which is the most common primary MID in the population with an estimated minimum prevalence of about 1 in 30,000 [53]. LHON is characterised by bilateral subacute visual loss with a peak age of onset in the second or third decades of life. About 90% of cases are due to one of three point mtDNA mutations (m.3460G > A in MT-ND1, m.11778G > A in MT-ND4 or m.14484T > C in MT-ND6), which all affect critical complex-I subunits, and the majority of these mutations are detected at homoplasmic levels. The paradox is that only ~50% of male carriers and ~10% of female carriers will lose vision in their lifetime due to the influence of secondary genetic, hormonal or environmental modifiers [54,54,, 55]. Although the majority of those who will lose vision will do so before the age of 50 years and the risk is significantly lower for female carriers, it is not currently possible to accurately predict who is going to lose vision and when. MtDNA copy number appears to influence penetrance, given the observation that symptomatic individuals have less mtDNA per cell than do asymptomatic carriers of the same mutation [55]. This could be due to different rates of mitochondrial biogenesis [55] and/or turnover [56].
3.6 Tissue Specificity
MIDs frequently present with non-specific multisystem manifestations and, as a result, their diagnosis and proper management is often delayed [5]. To compound the diagnostic challenges, patients can present at any age, with almost any affected body system and with a wide range of clinical severity [5]. The phenotype may also differ between paediatric and adult cases. In paediatric cases, encephalopathy, muscle hypotonia, lactic acidosis, epilepsy, cardiomyopathy and psychomotor delay are the most frequently reported features [57]. In adult-onset cases, the predominant phenotypic features include sensorineural deafness, diabetes, myopathy (ptosis, ophthalmoplegia, limb weakness, respiratory insufficiency), peripheral neuropathy, visual failure (cataract, optic neuropathy, pigmentary retinopathy), cardiac, gastrointestinal, renal, other endocrine or central nervous system involvement (ataxia, movement disorders, seizures, cognitive impairment). The heterogeneous tissue specificity and variable expressivity that can be associated with the same pathogenic mtDNA mutation contribute to the challenging nature of genetic counselling for this group of disorders with regards to prevention, recurrence risk and prognosis.
3.7 Genetic Counselling
Genetic counselling is the process of providing individuals and families with information on the nature, inheritance and implications of genetic disorders to help them make informed medical and personal decisions [58]. A detailed family history and genetic testing to clarify the genetic status of both clinically affected and unaffected family members are essential in guiding the discussion. Up to 80% of the childhood-onset MIDs are due to mutations in nDNA genes whereas 75% of adult-onset MIDs are due to mtDNA mutations [59]. Genetic counselling will therefore differ in these two categories of patients and other considerations will take different levels of importance, such as educational needs, occupational rehabilitation or reproductive options.
3.7.1 Prerequisites for Genetic Counselling
From a management and prevention standpoint, appropriate genetic investigations in a patient with suspected MID is essential [5]. A confirmed molecular diagnosis will help to better understand the natural history of the disease, the recurrence risk to other family members and the option of familial contact tracing [5]. With regard to genetic counselling, it is additionally important to know if a given mtDNA mutation is homoplasmic or heteroplasmic [60]. A homoplasmic mtDNA mutation will be transmitted to all offspring, but disease penetrance can vary as in LHON [61]. For a heteroplasmic mtDNA mutation, the level can vary markedly in the same sibship as a result of the mitochondrial bottleneck, which can influence the pattern and severity of disease expression.
3.7.2 Objectives of Genetic Counselling
Genetic counselling aims to provide accurate information about the prognosis (prediction) of affected individuals, the recurrence risk for other family members and reproductive choices (prevention) [10]. Genetic counselling for patients carrying mtDNA mutations is determined by the specific mutation involved, whether the mutation is heteroplasmic or homoplasmic, by disease penetrance and by the associated phenotypic spectrum [5]. Genetic counsellors are trained health professionals that work in close collaboration with physicians and clinical scientists as part of a multidisciplinary team. Such a concerted approach is needed to develop the best management approach based on the individual needs and expectations of patients and their families, with the aim of helping them to build effective coping strategies [62].
3.7.3 Prediction of Phenotype and Progression
It is very difficult to predict the clinical progression associated with most pathogenic mtDNA mutations. Caution should also be exercised when mutation levels have been quantified in non-affected tissues, such as blood, as the mutant load might be significantly higher in the tissues most at risk of disease. Even for homoplasmic mtDNA mutations, there can be differential vulnerability to mitochondrial dysfunction in different tissues or organs [30]. LHON, for example is mostly caused by homoplasmic mtDNA mutations, but this MID is characterised by both marked incomplete penetrance and a male bias [63]. The m.14484T > C mutation carries the best visual prognosis, but it is not possible to predict which patients will experience spontaneous visual recovery, nor the timing or the extent [64].
3.7.4 Recurrence Risk
3.7.4.1 Parents of a Proband
The father of a proband carrying an mtDNA mutation will not be a carrier due to the strict maternal inheritance of mtDNA [58]. The mother of a proband will invariably carry the pathogenic mtDNA mutation, except in rarer cases of a de novo mutation, and she may or may not be clinically affected (Table 1) [58].
3.7.4.2 Offspring of a Proband
MtDNA Point Mutations A male patient carrying an mtDNA point mutation can be reassured that his children are not at risk of inheriting his genetic defect [63]. A female carrier of a homoplasmic mtDNA mutation will transmit her pathogenic mtDNA variant to all of her children. If the mutation is heteroplasmic, the level that is transmitted to each child can fluctuate considerably due to the bottleneck effect [63] but reflecting the mother’s load of mutant mtDNA. For instance, ~30% of children born to mothers harbouring 5–25% of m.3243A>G in urinary epithelial cells with have <5% and in many undetectable m.3243A>G [65]. For mothers harbouring >25% m.3243A>G, mutant will be detectable in almost all offspring [65], though the published figures probably over-estimate the risks due to ascertainment bias. Since the level of the mutation in amniocytes or chorionic villi may differ from the mutation load in other cell populations, prenatal testing for females carrying a heteroplasmic mtDNA mutation poses significant challenges [63]. In a study of 33 children and 72 adults carrying an mtDNA point mutation, a de novo frequency of 24.6% was reported [19]. Prenatal diagnosis (PND) for the mothers of four patients with de novo mtDNA mutations revealed the absence of the mutant mtDNA species in all 11 oocytes/embryos tested [19]. Based on a literature survey, 137 patients harbouring de novo mtDNA mutations have been previously reported and PND was performed for the mothers of nine probands [19]. Recurrence occurred in only one of these nine mothers in two of her subsequent pregnancies, presumably due to germline mosaicism [19]. The overall recurrence risk of de novo mtDNA point mutations to the same parents therefore seems to be relatively low [19].
MtDNA Rearrangements For genetic counselling of a patient carrying a single mtDNA deletion, it is essential to assess if the deletion is de novo or, more rarely, maternally inherited [30]. In the rare cases with maternally inherited major rearrangements [67], i.e. tandem duplications and/or deletion dimers are almost always present in addition to the single deletions in skeletal muscle [67]. The most parsimonious explanation is that the first mutant molecule was a tandem duplication (likely occurring during replication) and that related molecules were generated by homologous recombination [68]. If a single mtDNA deletion is de novo, having arisen sporadically during embryonic development, the probability of transmitting it to the next generation is extremely low, except in cases of mosaicism of the germline oocytes [12]. A clinically unaffected mother is highly unlikely to have more than one affected child. However, a clinically affected mother has about a 1 in 24 risk of having a clinically affected child, which can be associated with the presence of a partially duplicated mtDNA in the mother [69].
3.7.4.3 Siblings of a Proband
The risk to siblings of an affected proband depends on the genetic and clinical status of the mother. If the mother carries an mtDNA point mutation, all siblings of a proband are at risk of inheriting the pathogenic variant, although the level can vary depending on whether the mutation is heteroplasmic or homoplasmic [58]. If the proband harbours a single mtDNA deletion, the recurrence risk to a sibling is 4% if the mother is clinically affected [69].
3.8 Preventing Maternal Transmission of mtDNA Mutations
3.8.1 Oocyte Donation
Oocyte donation is a low-risk strategy to eliminate the transmission of a pathogenic mtDNA mutation from mother to child. The donated oocyte is fertilised with the partner’s sperm and the resulting embryo is placed in the uterus of the carrier mother who then becomes the biological, but not the genetic mother. Potential issues relating to genetic versus biological parentage should be put in perspective. As >10% of the UK population do not identify their genetic fathers correctly [71], a genetic link does not appear to be a prerequisite for good intrafamilial relationships. Overall, oocyte donation is not a common reproductive choice for women of childbearing age who are known to harbour a pathogenic mtDNA mutation.
3.8.2 Prenatal Diagnosis
Prenatal testing and prenatal counselling after conception is only possible if the causative pathogenic mutation is known [3]. A mother who carries a homoplasmic mtDNA mutation should not be offered CVS or an amniocentesis to evaluate the risk of disease for the fetus, which will also be homoplasmic. This is because it will not help assess the risk of recurrence (see Sect. 3.5 for discussion of unknown factors affecting penetrance). Interpretation of PND testing in mothers who carry a heteroplasmic mtDNA mutation is complex since the mutational load in fetal tissues tested (amniocytes or chorionic vili) may differ from the eventual level in the tissues that are most at risk of a particular mtDNA mutation [58]. Nervertheless, most studies show a positive correlation between mutant loads in fetal tissue and amniocytes [42, 72].
In the case of heteroplasmic mutations, interpretation of PND depends on factors such as the heteroplasmy rate, threshold level, phenotypic expression of the mutation in maternal relatives and the strength of the genotype/phenotype correlation, which should be considered and carefully discussed with the mother [3]. Counselling is particularly difficult if fetuses harbour mtDNA mutations with intermediate heteroplasmy and unclear genotype/phenotype correlation [3]. The most common mtDNA mutation studied for PND is the mutation m.8993T > G [30]. In a study of 62 prenatal samples (17 mtDNA and 45 nDNA), it was observed that the analysis of heteroplasmy levels in other family members can be helpful in interpreting the prenatal mtDNA test result, particularly in cases of rare mtDNA mutations or when intermediate heteroplasmy rates (30–70%) have been detected [3]. After PND and termination of pregnancy, at least 11 cases of MID were prevented, three of which were related to mtDNA mutations (m.3243A > G, m.14453G > A, m.13513G > A) [3].
3.8.3 Preimplantation Genetic Diagnosis
Although preimplantation genetic diagnosis (PGD) remains controversial, it is a potential option that could be applied to lower the recurrence risk. This approach has raised a number of concerns [73], in particular that the mutant load cannot be accurately predicted in all tissues of the resulting embryo [5]. This concern is because of a single case where the child’s mutant load exceeded that of the blastocyst biopsy [74]. However, in this instance, the embryo was not biopsied until the blastocyst stage. Previous studies have shown that the variable tissue segregation of mtDNA variants that occurs in later stages of embryonic development is not yet present in the preimplantation cleavage-stage embryo, in which heteroplasmic mtDNA tends to be uniformly distributed in all blastomeres [32, 39]. Thus, accurate assessment of the embryo’s mutant load may be crucially dependent upon a biopsy procedure being done <4 days after fertilisation, just before the blastocyst forms [75]. Hence, if the zygote is sampled at an earlier stage of development, the mutant load can be more accurately assessed. Nevertheless, in women where only a small number of embryos are generated it can be ethically challenging to decide which embryos should be implanted to maximise the chance of having an unaffected child. PGD is not an option for mothers who carry homoplasmic mtDNA mutations.
3.8.4 Prevention of Germline Transmission
Mitochondrial replacement therapy (MRT), which involves a modification of conventional in vitro fertilization (IVF) techniques, has been proposed as a possible reproductive strategy [76]. The two techniques of mitochondrial donation being optimised are pronuclear transfer and metaphase-II spindle transfer. In pronuclear transfer, the mother’s oocyte is first fertilised with the father’s sperm, followed by the removal of the parental pronuclei. These are then transferred into an enucleated mitochondrial donor zygote at the same stage, harbouring only wild-type mtDNA [77, 78]. In metaphase-II spindle transfer, the mother’s metaphase-II spindle, which contains the maternal nDNA, is transferred into an enucleated mitochondrial donor oocyte and then fertilised by intracytoplasmatic sperm injection [79, 80]. Both IVF techniques have shown promising results with minimal carryover of mutant mtDNA, but a number of safety concerns have been raised, magnified by the fact that the germline is being altered [81, 82]. While the technique is likely to reduce the load of mutant mtDNA, segregation may be unpredictable, resulting in reversion towards the pathogenic mtDNA type [78, 83]. The cause of this segregation is not clear, though one group identified a polymorphism that may drive a replicative advantage [83]. Two strategies have been suggested for countering this phenomenon. Firstly, segregation may be reduced by matching the mtDNA haplotype of the donor and recipient [84, 85]. Secondly, it might be possible to recruit the cell’s own quality control mechanisms to target the pathogenic mutant mtDNAs [86]. There remains the possibility of significant metabolic problems arising from a mismatch [87, 88].
There are also important legal and ethical issues that need to be carefully addressed before considering the wider clinical implementation of mitochondrial donation for prospective mothers harbouring homoplasmic mtDNA mutations [63, 89]. Concerns have been raised as reversal to the mutant cell line may occur in some MRT children, possibly due to polymorphisms in the CSB-II region of the D-loop [83]. Certain mtDNA haplotypes may confer embryonic stem cells with faster growth and proliferative advantages [83]. Despite these potential risks, the UK Parliament voted in favour of changing the law in February 2015 to allow mitochondrial donation to be offered as a reproductive option if the technique can be shown to be effective with the benefits outweighing the potential risks [90]. In the first reported case of mitochondrial donation, which was carried out in Mexico in 2016, the child was reported to be healthy at 7 months of age with a mutational load of 2.36–9.23% in his tested tissues. The boy’s mother is heteroplasmic for the m.8993T>G mtDNA mutation in MT-ATP6, which in addition to NARP can cause Leigh syndrome when present at high mutational levels. Whilst this is encouraging, the effectiveness of reducing the mutant load remains unclear as the mutant load in the oocyte was not reported [91]. Furthermore, long term follow up is needed as mtDNA segregation could occur later in the child's life.
3.9 The Patient and Family Perspective
The diagnostic process is often perceived as complex by patients and their families, and it is frequently accompanied by strong emotions that are partly influenced by the patient’s support system and the approach of the healthcare professionals overseeing their care [5]. Due to the uncertainty about the diagnosis, psychological distress may arise, including feelings of blame and guilt, which can precipitate maladaptive coping mechanisms [5]. Based on a study of 14 parents searching for a diagnosis for their children, their perception of the diagnostic process was accompanied by both emotional and sociological responses [92]. Their emotional experience meant coming to terms with the health status of their affected child, whereas their sociological experience reflected their interactions as parents with members of the healthcare system and their access to support systems [92]. Frustration was a major reaction as they encountered obstacles when searching for their child’s diagnosis [92]. Only a few studies have explored the psychological consequences to patients and their relatives following the diagnosis of a MID and further work is needed regarding how to best support these individuals in coping with these emotionally charged and difficult personal life events [5].
4 Conclusions
Genetic counselling for MIDs remains an imperfect science due to the genetic complexity of this group of disorders and our incomplete understanding of the complex factors that drive tissue specificity and disease expressivity. A multidisciplinary and holistic approach is crucial to better understand how patients and their relatives deal with the anxiety and fears brought by an unexpected genetic diagnosis. Genetic counsellors have an important role to play in effectively communicating the information provided by a genetic test, and in making sure that it is properly processed to empower patients and their relatives in making the best decisions based on their own personal circumstances. A number of reproductive choices are now available for prospective mothers carrying pathogenic mtDNA mutations with the advent of mitochondrial donation and other potential germline strategies emerging. More than ever, the proper framework needs to be put in place to allow for careful regulation and to encourage an informed societal debate about the risks and benefits of these experimental therapies.
Change history
04 July 2017
An erratum to this article has been published.
References
Parikh S, Goldstein A, Keening MK, Scaglia F, Enns GM, Saneto R, et al. Diagnosis and management of mitochondrial disease: a consensus statement from the Mitochondrial Medicine Society. Genet Med. 2015;17:689–701.
Gorman G, Schaefer A, Ng Y, Gomez N, Blakely EL, Alston CL, et al. Prevalence of nuclear and mitochondrial DNA mutations related to adult mitochondrial disease. Ann Neurol. 2015;77:753–9.
Nesbitt V, Alston CL, Blakely EL, Fratter C, Feeney CL, Poulton J, et al. A national perspective on prenatal testing for mitochondrial disease. Eur J Hum Genet. 2014;22:1255–9.
Manwaring N, Jones MM, Wang JJ, Rochtchina E, Howard C, Mitchell P, Sue CM. Population prevalence of the MELAS A3243G mutation. Mitochondrion 2007;7:230–3.
Vento JM, Pappa B. Genetic counseling in mitochondrial disease. Neurotherapeutics. 2013;10:243–50.
Huang X, Bedoyan JK, Demirbas D, Harris DJ, Miron A, Edelheit S, et al. Succinyl-CoA synthetase (SUCLA2) deficiency in two siblings with impaired activity of other mitochondrial oxidative enzymes in skeletal muscle without mitochondrial DNA depletion. Mol Genet Metab. 2017;120(3):213–22.
Calvo SE, Clauser KR, Mootha VK. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 2016;44:D1251–7.
Tranchant C, Anheim M. Movement disorders in mitochondrial diseases. Rev Neurol (Paris). 2016;172:524–9.
Poulton J, O’Rahilly S, Morten KJ, Clark A. Mitochondrial DNA, diabetes and pancreatic pathology in Kearns–Sayre syndrome. Diabetologia. 1995;38:868–71.
Thorburn DR, Dahl HH. Mitochondrial disorders: genetics, counseling, prenatal diagnosis and reproductive options. Am J Med Genet. 2001;106:102–14.
Jansen RP. Germline passage of mitochondria: quantitative considerations and possible embryological sequelae. Hum Reprod. 2000;15(suppl 2):112–28.
Alston CL, Rocha MC, Lax NZ, Turnbull DM, Taylor RW. The genetics and pathology of mitochondrial disease. J Pathol. 2017;241:236–50.
Sacconi S, Salviati L, Nishigaki Y, Walker WF, Hernandez-Rosa E, Trevisson E, et al. A functionally dominant mitochondrial DNA mutation. Hum Mol Genet. 2008;17:1814–20.
Wilson IJ, Carling PJ, Alston CL, Floros VI, Pyle A, Hudson G, et al. Mitochondrial DNA sequence characteristics modulate the size of the genetic bottleneck. Hum Mol Genet. 2016;25(5):1031–41.
Payne BA, Wilson IJ, Yu-Wai-Man P, Coxhead J, Deehan D, Horvath R, Taylor RW, Samuels DC, Santibanez-Koref M, Chinnery PF. Universal heteroplasmy of human mitochondrial DNA. Hum Mol Genet 2013;22:384-90.
Reyes A, Melchionda L, Nasca A, Carrara F, Lamantea E, Zanolini A, et al. RNASEH1 mutations impair mtDNA replication and cause adult-onset mitochondrial encephalomyopathy. Am J Hum Genet. 2015;97:186–93.
Navarro-Sastre A, Tort F, Garcia-Villoria J, Pons MR, Nascimento A, Colomer J, et al. Mitochondrial DNA depletion syndrome: new descriptions and the use of citrate synthase as a helpful tool to better characterise the patients. Mol Genet Metab. 2012;107:409–15.
Finsterer J, Ahting U. Mitochondrial depletion syndromes in children and adults. Can J Neurol Sci. 2013;40:635–44.
Sallevelt SC, de Die-Smulders CE, Hendrickx AT, Hellebrekers DM, de Coo IF, Alston CL, et al. De novo mtDNA point mutations are common and have a low recurrence risk. J Med Genet. 2017;54(2):73–83. doi:10.1136/jmedgenet-2016-103876.
Chinnery PF. Mitochondrial disorders overview. 2000 Jun 8 [updated 2014 Aug 14]. In: Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, et al., editors. GeneReviews® [internet]. Seattle: University of Washington, Seattle; 1993–2016. http://www.ncbi.nlm.nih.gov/books/NBK1224/. Accessed Mar 2017
de Laat P, Janssen MC, Alston CL, Taylor RW, Rodenburg RJ, Smeitink JA. Three families with ‘de novo’ m.3243A > G mutation. BBA Clin. 2016;6:19–24.
Marchington D, Malik S, Banerjee A, Turner K, Samuels D, Macaulay V, et al. Information for genetic management of mtDNA disease: sampling pathogenic mtDNA mutants in the human germline and in placenta. J Med Genet. 2010;47:257–61.
Ballinger S, Shoffner J, Gebhart S, Koontz D, Wallace D. Mitochondrial diabetes revisited. Nat Genet. 1994;7:458–9.
Mkaouar-Rebai E, Felhi R, Tabebi M, Alila-Fersi O, Chamkha I, Maalej M, et al. Mitochondrial DNA triplication and punctual mutations in patients with mitochondrial neuromuscular disorders. Biochem Biophys Res Commun. 2016;473:578–85.
Bannwarth S, Abbassi M, Valéro R, Fragaki K, Dubois N, Vialettes B, et al. A novel unstable mutation in mitochondrial DNA responsible for maternally inherited diabetes and deafness. Diabetes Care. 2011;34:2591–3.
Cardena MM, Mansur AJ, Pereira Ada C, Fridman C. A new duplication in the mitochondrially encoded tRNA proline gene in a patient with dilated cardiomyopathy. Mitochondrial DNA. 2013;24:46–9.
Kasahara T, Ishiwata M, Kakiuchi C, Fuke S, Iwata N, Ozaki N, et al. Enrichment of deleterious variants of mitochondrial DNA polymerase gene (POLG1) in bipolar disorder. Psychiatry Clin Neurosci. 2016;. doi:10.1111/pcn.12496.
Ruzzenente B, Rötig A, Metodiev MD. Mouse models for mitochondrial diseases. Hum Mol Genet. 2016;25:R115–22.
Uehara N, Mori M, Tokuzawa Y, Mizuno Y, Tamaru S, Kohda M, et al. New MT-ND6 and NDUFA1 mutations in mitochondrial respiratory chain disorders. Ann Clin Transl Neurol. 2014;1:361–9.
Wong LJ. Diagnostic challenges of mitochondrial DNA disorders. Mitochondrion. 2007;7:45–52.
Rebolledo-Jaramillo B, Su MS, Stoler N, McElhoe JA, Dickins B, Blankenberg D, et al. Maternal age effect and severe germ-line bottleneck in the inheritance of human mitochondrial DNA. Proc Natl Acad Sci USA. 2014;111:15474–9.
Poulton J, Marchington DR. Segregation of mitochondrial DNA (mtDNA) in human oocytes and in animal models of mtDNA disease: clinical implications. Reproduction. 2002;123:751–5.
Johnston IG, Burgstaller JP, Havlicek V, Kolbe T, Rülicke T, Brem G, et al. Stochastic modelling, Bayesian inference, and new in vivo measurements elucidate the debated mtDNA bottleneck mechanism. Elife. 2015;2(4):e07464. doi:10.7554/eLife.07464.
Marlow FL. Mitochondrial matters: mitochondrial bottlenecks, self-assembling structures, and entrapment in the female germline. Stem Cell Res. 2017;. doi:10.1016/j.scr.2017.03.004.
Marchington DR, Macaulay V, Hartshorne GM, Barlow D, Poulton J. Evidence from human oocytes for a genetic bottleneck in an mtDNA disease. Am J Hum Genet. 1998;63:769–75.
Cao L, Shitara H, Sugimoto M, Hayashi J, Abe K, Yonekawa H. New evidence confirms that the mitochondrial bottleneck is generated without reduction of mitochondrial DNA content in early primordial germ cells of mice. PLoS Genet. 2009;5:e1000756.
Cree LM, Samuels DC, de Sousa Lopes SC, Rajasimha HK, Wonnapinij P, Mann JR, et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat Genet. 2008;40:249–54.
Wai T, Teoli D, Shoubridge EA. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat Genet. 2008;40:1484–8.
Poulton J, Marchington DR. Progress in genetic counselling and prenatal diagnosis of maternally inherited mtDNA diseases. Neuromuscul Disord. 2000;10:484–7.
Brown DT, Samuels DC, Michael EM, Turnbull DM, Chinnery PF. Random genetic drift determines the level of mutant mtDNA in human primary oocytes. Am J Hum Genet. 2001;68:533–6.
Li M, Rothwell R, Vermaat M, Wachsmuth M, Schröder R, Laros JF, et al. Transmission of human mtDNA heteroplasmy in the Genome of the Netherlands families: support for a variable-size bottleneck. Genome Res. 2016;26:417–26.
Monnot S, Gigarel N, Samuels DC, Burlet P, Hesters L, Frydman N, et al. Segregation of mtDNA throughout human embryofetal development: m.3243A > G as a model system. Hum Mutat. 2011;32:116–25.
Freyer C, Cree LM, Mourier A, Stewart JB, Koolmeister C, Milenkovic D, et al. Variation in germline mtDNA heteroplasmy is determined prenatally but modified during subsequent transmission. Nat Genet. 2012;44:1282–5.
Barbiroli B, Montagna P, Cortelli P, Iotti S, Lodi R, Barboni P, et al. Defective brain and muscle energy metabolism shown by in vivo 31P magnetic resonance spectroscopy in nonaffected carriers of 11778 mtDNA mutation. Neurology. 1995;45:1364–9.
Rahman S, Poulton J, Marchington D, Suomalainen A. Decrease of 3243 A–> G mtDNA mutation from blood in MELAS syndrome: a longitudinal study. Am J Hum Genet. 2001;68:238–40.
Matthews PM, Hopkin J, Brown RM, Stephenson JB, Hilton-Jones D, Brown GK. Comparison of the relative levels of the 3243 (A–> G) mtDNA mutation in heteroplasmic adult and fetal tissues. J Med Genet. 1994;31:41–4.
Weber K, Wilson JN, Taylor L, Brierley E, Johnson MA, Turnbull DM, et al. A new mtDNA mutation showing accumulation with time and restriction to skeletal muscle. Am J Hum Genet. 1997;60:373–80.
Emmerson CF, Brown GK, Poulton J. Synthesis of mitochondrial DNA in permeabilised human cultured cells. Nucleic Acids Res. 2001;29:E1.
Rajasimha HK, Chinnery PF, Samuels DC. Selection against pathogenic mtDNA mutations in a stem cell population leads to the loss of the 3243A–> G mutation in blood. Am J Hum Genet. 2008;82:333–43.
White SL, Collins VR, Wolfe R, Cleary MA, Shanske S, DiMauro S, Dahl HH, Thorburn DR. Genetic counseling and prenatal diagnosis for the mitochondrial DNA mutations at nucleotide 8993. Am J Hum Genet 1999;65:474–82.
Blok RB, Gook DA, Thorburn DR, Dahl HH. Skewed segregation of the mtDNA nt 8993 (TRG) mutation in human oocytes. Am J Hum Genet. 1997;60:1495–501.
Marchington DR, Scott-Brown M, Barlow DH, Poulton J. Mosaicism for mitochondrial DNA polymorphic variants in placenta has implications for the feasibility of prenatal diagnosis in mtDNA diseases. Eur J Hum Genet. 2006;14:816–23.
Yu-Wai-Man P, Griffiths PG, Brown DT, Howell N, Turnbull DM, Chinnery PF. The epidemiology of Leber hereditary optic neuropathy in the North East of England. Am J Hum Genet. 2003;72:333–9.
Yu-Wai-Man P, Howell N, Mackey DA, Nørby S, Rosenberg T, Turnbull DM, et al. Mitochondrial DNA haplogroup distribution within Leber hereditary optic neuropathy pedigrees. J Med Genet. 2004;41:e41.
Giordano C, Iommarini L, Giordano L, Maresca A, Pisano A, Valentino ML, et al. Efficient mitochondrial biogenesis drives incomplete penetrance in Leber’s hereditary optic neuropathy. Brain. 2014;137:335–53.
Dombi E, Diot A, Morten K, Carver J, Lodge T, Fratter C, et al. The m.13051G > A mitochondrial DNA mutation results in variable neurology and activated mitophagy. Neurology. 2016;86:1921–3.
Debray FG, Lambert M, Mitchell GA. Disorders of mitochondrial function. Curr Opin Pediatr. 2008;20:471–82.
DiMauro S, Hirano M. MERRF [internet]. 2003 Jun 3 [updated 2015 Jan 29]. In: Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, et al., editors. GeneReviews® [internet]. Seattle: University of Washington, Seattle; 1993–2016. http://www.ncbi.nlm.nih.gov/books/NBK1520/. Accessed Mar 2017.
Schaefer AM, McFarland R, Blakely EL, He L, Whittaker RG, Taylor RW, et al. Prevalence of mitochondrial DNA disease in adults. Ann Neurol. 2008;63:35–9.
Carrasco Salas P, Palma Milla C, López Montiel J, Benito C, Franco Freire S, López Siles J. Leber hereditary optic neuropathy: usefulness of next generation sequencing to study mitochondrial mutations on apparent homoplasmy. Med Clin (Barc). 2016;146:163–6.
Gorman GS, Chinnery PF, DiMauro S, Hirano M, Koga Y, McFarland R, et al. Mitochondrial diseases. Nat Rev Dis Primers. 2016;20(2):16080. doi:10.1038/nrdp.2016.80.
Lipinski SE, Lipinski MJ, Biesecker LG, Biesecker BB. Uncertainty and perceived personal control among parents of children with rare chromosome conditions: the role of genetic counseling. Am J Med Genet C Semin Med Genet. 2006;142C:232–40.
Yu-Wai-Man P, Votruba M, Moore AT, Chinnery PF. Treatment strategies for inherited optic neuropathies: past, present and future. Eye (Lond). 2014;28:521–37.
Yu-Wai-Man P, Votruba M, Burté F, La Morgia C, Barboni P, Carelli V. A neurodegenerative perspective on mitochondrial optic neuropathies. Acta Neuropathol. 2016;132:789–806.
de Laat P, Koene S, Heuvel LP, Rodenburg RJ, Janssen MC, Smeitink JA. Inheritance of the m.3243A > G mutation. JIMD Rep. 2013;8:47–50.
Dunbar D, Moonie P, Swingler R, Davidson D, Roberts R, Holt I. Maternally transmitted partial direct tandem duplication of mitochondrial DNA associated with diabetes mellitus. Hum Mol Genet. 1993;2:1619–24.
Poulton J, Deadman ME, Bindoff L, Morten K, Land J, Brown G. Families of mtDNA re-arrangements can be detected in patients with mtDNA deletions: duplications may be a transient intermediate form. Hum Mol Genet. 1993;2:23–30.
Poulton J, Holt I. Mitochondrial DNA: does more lead to less? Nat Genet. 1994;8:313–5.
Chinnery PF, DiMauro S, Shanske S, Schon EA, Zeviani M, Mariotti C, et al. Risk of developing a mitochondrial DNA deletion disorder. Lancet. 2004;364:592–6.
Yu-Wai-Man P, Chinnery PF. Leber hereditary optic neuropathy. 2000 Oct 26 [updated 2016 Jun 23]. In: Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, et al., editors. GeneReviews® [internet]. Seattle: University of Washington, Seattle; 1993–2017. http://www.ncbi.nlm.nih.gov/books/NBK1174/. Accessed Mar 2017
Macintyre S, Sooman A. Non-paternity and prenatal genetic screening. Lancet. 1991;338:869–71.
Steffann J, Gigarel N, Corcos J, Bonnière M, Encha-Razavi F, Sinico M, et al. Stability of the m.8993T- > G mtDNA mutation load during human embryofetal development has implications for the feasibility of prenatal diagnosis in NARP syndrome. J Med Genet. 2007;44:664–9.
Bredenoord AL, Dondorp W, Pennings G, De Die-Smulders CE, De Wert G. PGD to reduce reproductive risk: the case of mitochondrial DNA disorders. Hum Reprod. 2008;23:2392–401.
Mitalipov S, Amato P, Parry S, Falk MJ. Limitations of preimplantation genetic diagnosis for mitochondrial DNA diseases. Cell Rep. 2014;7:935–7.
Steffann J, Gigarel N, Samuels DC, Monnot S, Borghese R, Hesters L, et al. Data from artificial models of mitochondrial DNA disorders are not always applicable to humans. Cell Rep. 2014;7:933–4.
Poulton J, Oakeshott P. Nuclear transfer to prevent maternal transmission of mitochondrial DNA disease. BMJ. 2012;345:e6651.
Craven L, Tuppen HA, Greggains GD, Harbottle SJ, Murphy JL, Cree LM, et al. Pronuclear transfer in human embryos to prevent transmission of mitochondrial DNA disease. Nature. 2010;465:82–5.
Hyslop LA, Blakeley P, Craven L, Richardson J, Fogarty NM, Fragouli E, et al. Towards clinical application of pronuclear transfer to prevent mitochondrial DNA disease. Nature. 2016;534:383–6.
Tachibana M, Sparman M, Sritanaudomchai H, Ma H, Clepper L, Woodward J, et al. Mitochondrial gene replacement in primate offspring and embryonic stem cells. Nature. 2009;461:367–72.
Tachibana M, Amato P, Sparman M, Woodward J, Sanchis DM, Ma H, et al. Towards germline gene therapy of inherited mitochondrial diseases. Nature. 2013;493:627–31.
Chinnery PF, Craven L, Mitalipov S, Stewart JB, Herbert M, Turnbull DM. The challenges of mitochondrial replacement. PLoS Genet. 2014;10(4):e1004315. doi:10.1371/journal.pgen.1004315.
Morrow EH, Reinhardt K, Wolff JN, Dowling DK. Risks inherent to mitochondrial replacement. EMBO Rep. 2015;16:541–4.
Kang E, Wu J, Gutierrez NM, Koski A, Tippner-Hedges R, Agaronyan K, et al. Mitochondrial replacement in human oocytes carrying pathogenic mitochondrial DNA mutations. Nature. 2016;540:270–5.
Burgstaller JP, Johnston IG, Poulton J. Mitochondrial DNA disease and developmental implications for reproductive strategies. Mol Hum Reprod. 2015;21:11–22.
Burgstaller J, Johnston I, Jones N, Albrechtová J, Kolbe T, Vogl C, et al. MtDNA segregation in heteroplasmic tissues is common in vivo and modulated by haplotype differences and developmental stage. Cell Rep. 2014;7:2031–41.
Diot A, Dombi E, Lodge T, Liao C, Morten K, Carver J, et al. Modulating mitochondrial quality in disease transmission: towards enabling mitochondrial DNA disease carriers to have healthy children. Biochem Soc Trans. 2016;44:1091–100.
Latorre-Pellicer A, Moreno-Loshuertos R, Lechuga-Vieco AV, Sanchez-Cabo F, Torroja C, Acin-Perez R, et al. Mitochondrial and nuclear DNA matching shapes metabolism and healthy ageing. Nature. 2016;535:561–5.
Chinnery PF, Zeviani M. Mitochondrial matchmaking. N Engl J Med. 2016;375:1894–6.
Klopstock T, Klopstock B, Prokisch H. Mitochondrial replacement approaches: challenges for clinical implementation. Genome Med. 2016;25(8):126.
Yu-Wai-Man P. Harnessing the power of genetic engineering for patients with mitochondrial eye diseases. J Neuroophthalmol. 2017;37:56–64.
Zhang J, Liu HJ, Luo S, Chavez-Badiola A, Liu Z, Yang M, et al. First live birth using human oocytes reconstituted by spindle nuclear transfer for mitochondrial DNA mutation causing Leigh syndrome. Fertil Steril. 2016;106(suppl 3):e375–6.
Lewis C, Skirton H, Jones R. Living without a diagnosis: the parental experience. Genet Test Mol Biomarkers. 2010;14:807–15.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
JP, JF and PYWM declare no conflicts of interest.
Funding
PYWM is supported by a Clinician Scientist Fellowship Award (G1002570) from the Medical Research Council (MRC) (UK). PYWM also receives funding from Fight for Sight (UK), the UK National Institute of Health Research (NIHR) as part of the Rare Diseases Translational Research Collaboration, and the NIHR Biomedical Research Centre based at Moorfields Eye Hospital NHS Foundation Trust and UCL Institute of Ophthalmology. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health.
Additional information
An erratum to this article is available at https://doi.org/10.1007/s40291-017-0286-8.
Rights and permissions
About this article
Cite this article
Poulton, J., Finsterer, J. & Yu-Wai-Man, P. Genetic Counselling for Maternally Inherited Mitochondrial Disorders. Mol Diagn Ther 21, 419–429 (2017). https://doi.org/10.1007/s40291-017-0279-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-017-0279-7