Abstract
Background and Objectives
To investigate a potential association between single-nucleotide polymorphisms (SNPs) and haplotypes at the TNFA-LTA locus and the development of oral cancer in an Indian population.
Materials and Methods
In this study, 150 oral precancer/cancer samples (50 precancer and 100 cancer), along with an equal number of control samples, were genotyped. Six SNPs at the TNF-LTA locus (i.e., −238G/A, −308G/A, −857C/T, −863C/A, −1031T/C, and +252A/G) were analyzed by use of a polymerase chain reaction–restriction fragment length polymorphism method, the assay was validated by sequencing 10 % of samples.
Results
The allelic frequencies of TNFA and LTA SNPs were found to be significantly associated with the risk of oral cancer and precancerous lesions in comparison with controls (P < 0.0003). Further haplotypic analysis showed that two haplotypes (ATCTGG and ACACGG) served as risk haplotypes for oral cancer. These haplotypes were also found to be significantly and positively associated with lifestyle habits (tobacco chewing P = 0.04, odds ratio [OR] 3.4) and socioeconomic status (P = 0.01, OR 3.4). We noticed an increased percentage of risk haplotypes correlating with the aggressiveness of oral cancer. The percentages of risk haplotypes were found to be threefold higher in precancer and fourfold higher in advanced stages of oral cancer in comparison with controls.
Conclusion
Five SNPs at the TNF-LTA locus (i.e., −308G>A, −857C>T, −863C>A, −1031T>C, and +252A>G) were found to be associated with the development of oral cancer. Two haplotypes (ATCTGG and ACACGG) emerged as major risk haplotypes for oral carcinoma progression and were also found to be associated with lifestyle factors and clinical aggressiveness. These findings make the TNF-LTA locus a suitable candidate for a future biomarker, which may be used either for early detection or for helping to improve treatment efficacy and effectiveness.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
SNPs at the TNFA-LTA locus are associated with susceptibility to oral cancer progression in an Indian population. |
Two haplotypes, ATCTGG and ACACGG, show a risk for oral cancer in India. |
The TNF-LTA locus may serve as a biomarker for tobacco-associated oral cancer predisposition. |
1 Introduction
Oral cancer is one of the most prevelent diseases worldwide. According to the Global Burden of Cancer 2013, lip and oral cancer ranked 11th globally, but in developing countries such as India, it ranked in second position [1]. Most of the cases (80 %) are diagnosed only in the final stage of cancer, leading to a low patient survival rate despite various advanced therapeutic efforts. Responses to similar existing treatments vary from patient to patient even in the same stage of cancer; therefore, new strategies are needed to improve survival rates.
In the Indian population, the high incidence of oral cancer is attributed to a number of etiological factors. The etiology of oral malignancy is very complex and multifactorial, as its pathogenesis and progression include many environmental factors (smoking, tobacco chewing, and alcohol drinking) and genetic factors (oncogenes, immune response and suppressor genes) [2]. Nicotine exposure has been established (as either tobacco chewing or tobacco smoking) as a risk factor for oral squamous cell carcinoma (OSCC) in several previous studies [3, 4]. Despite the risk of nicotine exposure, some patients do not develop oral cancer; therefore, there must be other factors that also influence the susceptibility of tobacco-exposed individuals to malignancy, and these may include a combination of total tobacco exposure and genetic susceptibility. This suggests that genes involved in immunomodulation are worth studying.
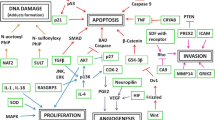
Cytokines, a group of immune proteins, are involved in inflammation, immunity, and protection against infections. Cytokines that have previously been implicated in tumor and inflammation events include tumor necrosis factor (TNF)-α and TNF-β, and lymphotoxin-α (LTA), which are encoded by the TNFA and LTA genes, respectively. These immunomodulating genes are located within the major histocompatibility complex (MHC) centromeric to HLA-B and telomeric to class III genes. Expression of TNF-α is mostly synchronized at the transcriptional level and cell cycle regulation [5]. Polymorphisms in TNF have been intensively studied as a potential determinant of disease susceptibility in a number of diseases where TNF levels are considered to be important.
Several single-nucleotide polymorphisms (SNPs) have been identified primarily in the promoter region of TNF. Most of the polymorphisms studied were focused on TNFA −308G>A (rs1800629) and −238G>A (rs361525). Besides the above, the TNFA −857C>T (rs1799724), −863C>A (rs1800630), and −1031T>C (rs1799964) loci have emerged as important candidates accounting for the increased risk of cancer development [6–8]. A polymorphism in LTA 252A>G (rs909253) has also been associated with oral carcinoma, but the functional assays described in the literature remain conflicting [9, 10], and the outcomes of those studies were inconsistent.
To date, there has been little documented evidence about these SNPs in the context of the Indian population. Therefore, we investigated the frequency of these SNPs in oral cancer patients of Indian origin. This study also suggested a role of the reported SNPs in disease progression in the context of lifestyle factors.
Therefore, the present study was designed to evaluate the association between SNPs and haplotypes in the TNFA-LTA locus (rs361525, rs1800629, rs1799724, rs1800630, rs1799964, and rs909253) and susceptibility to oral carcinoma. In addition, we also investigated the combined effect of these SNPs through haplotype analysis and we correlated the impact of demographic characteristics on oral cancer. To the best of our knowledge, this is the first such study carried out in India, as well as globally.
2 Material and Methods
2.1 Sample Collection
In the present study, samples from a total of 300 consecutive subjects, consisting of 150 histologically confirmed oral cancer tissue biopsy samples (50 precancer and 100 cancer cases) and 150 scraped cell samples from healthy control subjects (persons undergoing routine physical checkups or with some other oral-related problem such as muscular swelling or bleeding, etc.) of similar age and ethnicity, were included. Categorization and grading of the precancer (variation in normal cells associated with an increased risk of cancer) [11] and cancerous lesions were done according to World Health Organization (WHO) criteria. Written consent was obtained from all subjects, and the study was carried out in accordance with the principles of the Declaration of Helsinki. In the written consents, we also documented information about the lifestyle habits of both cases and controls, who included alcohol drinkers (average 100–150 mL daily at least 3–4 times weekly), tobacco chewers (4–6 packs per day) and smokers (three or more cigarettes per day). These criteria were decided upon after consultation with doctors and study of guidelines from the Centers for Disease Control and Prevention and the WHO, and previous reports [12, 13]. The subjects’ socioeconomic status was also recorded according to Kuppuswamy’s socioeconomic status [14]. The study was approved by the ethics committee of the institute.
2.2 DNA Extraction
Genomic DNA was extracted from freshly collected tissue samples from oral cancer/precancer patients and oral scrape cells from healthy control subjects by a standard method using proteinase K followed by phenol/chloroform/isopropanol treatment [15].
2.3 SNP Genotyping of TNFA and LTA Genes by PCR–RFLP
Five polymorphic sites in the promoter region of the TNF gene (−308, −238, −857, −863, and −1031) and one in LTA (+252) were genotyped with PCR according to the method mentioned in a previous study from our laboratory [7]. The six amplicons (−308, −238, −857, −863, and −1031) were digested by NcoI, BglI, TaiI, TaiI, BbsI, and NcoI restriction enzymes, respectively. The digested products were run on 10 % native polyacrylamide gels. The PCR primers and restriction enzyme list for RFLP analysis is presented in Table 1 in the Electronic Supplementary Material.
2.4 DNA Sequencing
We randomly selected 10 % of the samples for sequencing, to validate the PCR-RFLP assay. Sequencing reactions were carried out according to the conventional dideoxy chain termination method, using an ABI Prism® 310 automated DNA sequencer (Applied Biosystems, USA).
2.5 Statistical Analysis
Data analysis was performed using the computer software Quanto 1.1 (for the power of the study) and GraphPad Instat version 3.3 (for statistical significance in the study). Chi-squared tests/Fisher’s exact tests (for smaller numbers in subgroup analysis) were used to compare the distributions of TNFA and LTA polymorphisms between cancer patients, precancer patients, and controls. The effects of lifestyle habits on the risk of oral cancer were analyzed in the same way. Haplotype patterns and multiple testing corrections were made by Plink software. Confirmation of Hardy–Weinberg equilibrium, linkage disequilibrium statistics, and association of haplotypes with lifestyle habits were analyzed by Haploview software (http://www.broad.mit.edu/mpg/haploview) [16].
3 Results
3.1 Population Characteristics
Demographic data on the studied population are shown in Table 1. The mean ages of the oral cancer patients, precancer patients, and controls were 49.56 ± 14.2, 45.52 ± 13.8, and 41.14 ± 10.37 years, respectively. The cancer and precancer groups contained twice as many men as women (P < 0.05). The percentages of smokers, tobacco chewers, and alcohol drinkers were 50, 64, and 12 %, respectively, in the precancer group; 37, 70, and 8 %, respectively, in the cancer group; and 56, 80, and 40 %, respectively, in controls. Statistical significance was observed more for cancerous patients than for precancer patients in all three habits.
On further analysis, we noticed high percentages and significant data for the risks of oral cancer and precancer among low-socioeconomic-group subjects (67 % [P < 0.0001] for oral cancer; 66 % [P = 0.0032] for precancer) in comparison with high-/medium-socioeconomic-group subjects, which may have been due to low-quality food and lifestyle habits adopted by those subjects. In terms of histological grades, 15 % of samples had poorly differentiated squamous cell carcinoma (PDSCC), 27 % had moderately differentiated squamous cell carcinoma (MDSCC), and 58 % had well differentiated squamous cell carcinoma (WDSCC) (Table 1).
3.2 Genotypic Analysis of SNPs in the TNFA-LTA Locus
The distribution of TNFA promoter SNPs (−238G/A, −308G/A, −857C/T, −863C/A, and −1031T/C) and the TNFB/LTA SNP (+252A/G) genotype for oral cancer cases, precancer cases, and controls is depicted in Table 2. Genotype frequencies of all polymorphisms were further checked and were found to be in Hardy–Weinberg equilibrium in both cases and controls (P < 0.05).
3.2.1 TNFA −238G/A Polymorphism (rs361525) and Risk of Oral Cancer
Higher frequencies of the TNFA −238 carrier genotype were observed (GA/AA) in precancer cases [22 % (11/50)] and in oral cancer cases [21 % (21/100)] when compared to controls [15 % (22/150)], and the differences were not statistically significant between oral cancer cases and precancer cases compared with controls. The frequency of the −238A allele was found to be 1.5 times greater in precancer cases (14 %) and cancer cases (14.5 %) than in controls (9.3 %).
3.2.2 TNFA −308G/A Polymorphism (rs1800629) and Risk of Oral Cancer
The distribution of the carrier genotype (GA/AA) at −308G/A was found to be significant for both precancer (P = 0.0236, odds ratio [OR] 3.159, 95 % confidence interval [CI] 1.251–7.975) and cancer (P = 0.0174, OR 2.774, 95 % CI 1.248–6.163) in comparison with the control group. The frequency of polymorphic allele A was found to be higher in precancer cases (15 [15 %]) and oral cancer cases (29 [14.5 %]) than in the control group [16 (5.3 %)].
3.2.3 TNFA −857C/T Polymorphism (rs1799724) and Risk of Oral Cancer
In TNFA −857C/T, the frequency of the carrier genotype was 48 % (24/50) in precancer patients and 54 % (54/100) in cancer patients, and it was found to be significantly associated with the risk of precancerous lesions (P < 0.0001, OR 7.222, 95 % CI 3.410–15.293) and cancerous lesions (P < 0.0001, OR 9.184, 95 % CI 4.842–17.419) in comparison with controls. We also observed a fivefold higher minor T allele frequency in precancer cases (31 %) and cancer cases (30 %) in comparison with controls (6.3 %).
3.2.4 TNFA −863C/A Polymorphism (rs1800630) and Risk of Oral Cancer
The carrier genotypes (CA/AA) at the −863 locus were significantly related to susceptibility to, or development of, oral cancer, with frequencies of 30 % (15/50), 53 % (53/100), and 18 % (27/150) in precancer cases, cancer cases, and controls, respectively. A minor allele frequency (−863A) was also found to be higher in precancer (21 %) and cancer (33 %) patients than in controls (11 %).
3.2.5 TNFA −1031T/C Polymorphism (rs1799964) and Risk of Oral Cancer
The percentages of carrier genotype (CT/CC) distribution at the −1031 locus of TNFA were found to be twofold higher in precancer cases (42 %) and threefold higher in cancer cases (63 %) than in controls (22 %). The minor allelic C frequency was also revealed to be highly significant in association with cancer (P < 0.0001, OR 6.0, 95 % CI 3.4–10.5) and precancer (P = 0.0006, OR 2.6, 95 % CI 1.5–4.6) in comparison with controls.
3.2.6 LTA +252A/G Polymorphism (rs909253) and Risk of Oral Cancer
With regard to the frequency of carrier genotypes at LTA +252 (AG/GG), a nonsignificant association was observed with precancerous lesions (P = 0.1190, OR 2.053, 95 % CI 9205–4.577) in comparison with controls, while a significant difference in this SNP was detected in the cancer group (P < 0.0001, OR 3.500, 95 % CI 1.873–6.539) in comparison with controls. The minor allelic frequencies of +252G were found to be 30 % (15/50) in precancer cases, 19.5 % (39/100) in cancer cases, and 8 % (24/150) in controls, which revealed a significant association after analysis. Higher minor allelic frequencies in precancerous lesions may be a possible way to detect initiation of oral cancer at an early stage (Table 2).
3.3 Effects of Lifestyle Habits on Risk of Oral Cancer
Further analysis was performed to determine any correlation between SNPs, tobacco and alcohol use, and the socioeconomic status of the subjects. During the analysis, we found that five SNPs (i.e., TNFA −308, −857, −863, −1031, and LTA +252) were significantly associated with the risks of oral cancer and precancer in tobacco users (P < 0.05). For the habit of smoking, only three SNPs (i.e., TNFA −857, −863, and −1031) were found to be significantly associated with the risk of oral cancer, but when we tried to assess the combined effect of SNPs and smoking on the development of precancerous lesions, we could find only one SNP (i.e., TNFA −857) that was significantly associated with development of precancerous lesions. With regard to alcohol drinking, interestingly, this showed a significant association with cancer in TNFA −857, −863, −1031, and LTA +252, but no association with precancerous lesions.
It is well established that cancer is a multifactorial disease, with genetic as well as environmental factors. Socioeconomic status, as an environmental factor, plays an important role in maintaining an individual’s lifestyle, which also contributes to progression of disease. The data from our study showed that the association between the risk of cancer and socioeconomic status was more significant in low-socioeconomic-group subjects than in high-/medium-socioeconomic-group subjects (see Tables 2–7 in the Electronic Supplementary Material).
The data revealed higher carrier genotypic frequencies of the studied SNPs among tobacco chewers who developed oral cancer. The data were also significant for smokers and alcohol drinkers but comparatively less significant than those for tobacco chewers. It was concluded that tobacco chewing, smoking, and alcohol drinking in association with TNFA polymorphisms contribute to development of oral cancer with the progression of disease (see Tables 2–7 in the Electronic Supplementary Material).
3.4 Haplotypes and Progression of Oral Carcinoma
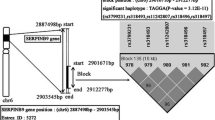
Haplotype analysis using the statistical software Haploview and Plink showed the presence of 11 haplotypes, which are shown in Table 3. The haplotypes presented in the table were present in more than 1 % of the total population studied. When we analysed our data for the cancer and precancer groups, we observed that four haplotypes in the cancer group (ATCCGG, ACACGG, ACATGG, and GCCTGG) and two haplotypes in the precancer group (ATCCGG and ATCTGG) were statistically significant (P < 0.05).
The associations of minor allele SNPs in haplotypes with precancer, cancer, and total cases were found to be more interesting. The minor +252G allele was found in two haplotypes (i.e., GTCCGG and GCCTGG) out of 11. The GCCTGG haplotype showed significant values for cancer cases (P = 0.023, OR 50.6).
The minor −1031C allele was found in five haplotypes (ACCCGG, ACACGG, ACATGG, GCCTGG, and ACCTGG). In cancer, out of a total of five haplotypes, three haplotypes (ACACGG, ACATGG, and GCCTGG) were statistically significant (P = 0.0029, OR 9.44; P = 0.0034, OR 69.9; P = 0.0234, OR 50.6, respectively) and positively associated with the risk of oral carcinoma. However, we could not find any association of these haplotypes with precancerous lesions.
The minor −863A allele was found in three haplotypes (ATACGG, ACACGG, and ACATGG). In cancer, out of three haplotypes, two haplotypes (ACACGG and ACATGG) were statistically significant and positively associated with the risk of oral carcinoma (P = 0.0029, OR 9.44; P = 0.0034, OR 69.9, respectively). Interestingly, for the precancer group, we could not find any significant association for any of the haplotypes (Table 4).
The minor −857T allele was found in four haplotypes (ATCTGG, ACATGG, GCCTGG, and ACCTGG). In cancer, out of a total of four haplotypes, two (ACATGG and GCCTGG) were observed to be statistically significant (P = 0.0034, OR 69.9; P = 0.02, OR 50.6, respectively) and positively associated with the risk of oral carcinoma. Interestingly, the ATCTGG haplotype was found to be significantly and positively (P = 0.003, OR 4.2) associated with precancerous lesions.
The minor −1031C, −863A, and −857T alleles were concurrently present in the ACATGG haplotype, which was statistically significant and positively associated with cancer (P = 0.0034, OR 69.9). However, it was not present in precancer, possibly because of the smaller number of precancer cases. In addition, the minor −1031C allele was also linked with the −863A minor allele in ACACGG haplotypes with a P = 0.0029 (OR 9.44) level of significance and a positive association with cancer, while in precancer, there was a nonsignificant (P = 0.12, OR 3.7) and positive association with oral cancer development. The minor −1031C allele was also linked with the −857T and +252G minor alleles in G CCTGG with statistical significance in cancer (P = 0.023, OR 50.6), but it was not present in precancer, possibly because of the small number of precancer cases. Apart from the above, the minor −1031C allele was also linked with −857T in ACCTGG haplotypes, with a nonsignificant result in cancer (P = 0.172, OR 15.4), but in precancer, it was not found.
Fascinatingly, the minor −308A and −238A alleles were also found in ATCCAG and ATCCGA, respectively, without positive associations and statistical significance in haplotype analysis by Plink. In multiple testing studies, the TNFA −238 SNP (rs361525) showed a nonsignificant P value, while other SNPs showed significant P values at the 0.05 level. Different multiple testing corrections are listed in Table 8 in the Electronic Supplementary Material.
3.5 Risks of Oral Cancer with Lifestyle Habits and Socioeconomic Status
For a better understanding of oral cancer etiology, we examined associations between haplotype findings, lifestyle habits (tobacco chewing, smoking, and alcohol drinking), socioeconomic status, and clinical aggressiveness. All 11 haplotypes were divided into three groups on the basis of their frequencies and significance in the studied population for analysis. These three groups were designated as wild haplotypes (ATCCGG), risk haplotypes (ATCTGG + ACACGG), and other haplotypes (ACCCGG, ATACGG, GTCCGG, ATCCGA, ACA TGG, ATCCAG, GCCTGG, and ACCTGG). After analysis, risk haplotypes were found to be significantly associated with tobacco chewing (P = 0.04) and low socioeconomic status (P = 0.01). Further, we also analyzed the effects of risk haplotypes on progression of oral cancer (Table 5). After the analysis, we found an increased percentage of risk haplotypes associated with aggressiveness of oral cancer. The percentage of risk haplotypes was 3-fold higher in precancer cases than in controls, while for advanced stages of cancer, it was increased fourfold.
4 Discussion
Oral cancer cases constitute approximately 40 % of total cancers in the Indian population, and the number of cases is continuously increasing from both the global and Indian perspective. The etiology of oral cancer is quite complex, where a host of genetic factors play a key role. Therefore, it is important to evaluate the roles of different biomarkers in oral cancer susceptibility for better understanding of the disease etiology, which may contribute toward treatment and early detection of cancer. TNFA and TNFB/LTA polymorphisms have previously been reported to be associated with inflammatory and immunomodulatory diseases, including cancer, but the results were inconsistent. Multiple SNPs of the functionally active TNF-LTA locus are responsible for impaired function of the gene and have been implicated in many cancers [17–19]. Some studies have reported data suggesting that TNF polymorphisms alter expression of the gene. For example, cells containing the −308A allele have been reported to produce up to six times more messenger RNA than those containing the −308G allele [20, 21]. Similarly the TNFB G/A polymorphism located at position 252 affects expression of the gene and concentrations of TNF-α and TNF-β proteins in plasma [10, 22]. Previously, our group has reported the role of TNF-LTA SNPs/haplotype associations in the risks of cervical and breast cancer [7, 23]. So, our interest was to study the possible contribution of these TNFA and LTA gene polymorphisms, either individually or in combination, in the development of oral cancer, in association with lifestyle-associated factors.
Individually, we observed a significant association between the TNFA −308G/A SNP and a threefold increased risk of oral cancer, as well as precancer, in comparison with the minor allelic frequency in controls, while the TNFA −238G/A SNP showed a nonsignificant association with the risks of oral cancer and precancer. Similar types of studies in Korean [24] and Taiwanese [8] populations are also in agreement with our findings. Polymorphism of TNFA −308 is associated with HLA haplotypes, which play an important role in immune responses [25]. In the Indian population, two other studies have showed associations between the above two SNPs and the risk of oral carcinoma, one of which was in agreement with our findings [6], while the other conflicted with our data [26]; this difference may have been due to different habitats and lifestyle factors. Like oral cancer, other cancers have also shown almost similar patterns of association [7, 27].
SNPs at −238(G/A) and −308(G/A) have been well studied not only in oral cancer but also in other cancers [7]; it is tempting to speculate on the role of other SNPs [TNFA −857(C/T), −863(C/A), and −1031(T/C)] in the pathogenesis of oral cancer. We observed that −857T was present fivefold more frequently in both oral cancer and precancer patients, whereas the presence of TNFA −863A and −1031C was noticed to be twofold higher in precancer and threefold higher in oral cancer patients, respectively, in comparison with the control group. Previous studies have also shown that TNF promoter activity is increased in TNFA −857T, −863A and −1031C genotypes [28], although contradictory findings have also been reported [29, 30].
In addition to SNPs in TNFA, the +252(A/G) SNP in LTA has been found to be associated with oral cancer in the Indian population, with twofold increased risks in both cancer and precancer patients compared with controls. Our findings are in accordance with the heterogeneity in the LTA +252(A/G) polymorphism shown in studies of oral cancer in Greek and German populations [10] and also in gastric cancer cases in a Korean population [31].
The strength of the present study was not only genotypic but also haplotypic analysis, which showed the linkage between minor alleles of multiple SNPs of the TNF gene. Individual haplotype pairs, including TNFA −1031, −863, and −857, were consistently associated with a significantly increased risk of oral cancer. ATCTGG (with the minor allele TNFA −857T) and ACACGG (with the minor alleles −1031C and TNFA −863A) were found to be risk haplotype pairs for oral cancer and precancer in comparison with controls. Our consistent findings from the different statistical methodologies are quite meaningful and are supported by other Asian population studies in different cancers [32]. Similar findings were observed in a sarcoidosis disease study, which found a significant association with the haplotype TCTgg (with the minor allele TNFA −857T) and a nonsignificant association with CACgg (with the minor alleles −1031C and −863A) haplotype pairs [33].
We found a synergistic relationship, in our combined study of TNFA and LTA genotypes, with lifestyle habits (smoking, tobacco chewing and alcohol drinking), which personalized the risk of oral cancer. Smoking, alcohol drinking, and tobacco chewing may play important roles in the etiology of oral submucous fibrosis (OSF) and OSCC, and may act in synergy [34–37]. In our study, the habit of tobacco chewing was more significantly associated with oral cancer and precancer than smoking or alcohol drinking. Risk haplotypes (ATCTGG and ACACGG) showed positive associations, with significant P values, in oral cancer and precancer cases among people with low socioeconomic status. It is possible that use of low-quality products contributes to the pathogenesis of OSCC through suppression of immune action by affecting the production of inflammatory cytokines such as TNF-α [38–43]. Some studies have reported that cigarette smoking suppresses production of TNF-α, adversely affecting the function of human natural killer (NK) cells and contributing to a higher incidence of cancer [39, 42, 44].
Increasing trend of risk haplotype pairs with aggressiveness of disease has showed the correlation between multiple SNPs and oral cancer development. Multiple testing analyses also supported our findings. Bonferroni calculations showed significant values for TNFA −308, −857, −863, and −1031 and for LTA +252, while TNFA −238 had a nonsignificant value. No such study has been conducted previously in oral cancer in India.
The major limitation of our study was its small sample size. On the other hand, this type of small-scale case–control study may be useful for planning of future investigations, providing information for large-scale studies with proper study designs, proper sample sizes and relevant clinical information, or for meta-analyses.
5 Conclusion
Out of six studied SNPs, five SNPs (i.e., −308 G>A, −857 C>T, −1031 T>C, −863 C>A, and +252 A>G) were found to be associated with development of oral cancer. In combination, ATCTGG and ACACGG haplotypes emerged as major risk haplotypes for oral carcinoma progression. In addition, we also observed a positive association between risk haplotypes and tobacco use, socioeconomic status and aggressiveness of tumors. Taking together all of the above findings, we were able to demonstrate that people with a history of tobacco use; the ATCTGG and ACACGG risk haplotypes; and low socioeconomic status are at higher risk of oral cancer progression. This information could be used for early detection of the disease, improving treatment efficacy and effectiveness, which will ultimately contribute to the discovery of personalized therapy.
References
Fitzmaurice C, Dicker D, Pain A, Hamavid H, Moradi-Lakeh M, MacIntyre MF, et al. The global burden of cancer 2013. JAMA Oncol. 2015;1(4):505–27.
Williams HK. Molecular pathogenesis of oral squamous carcinoma. Mol Pathol. 2000;53(4):165–72.
Jayalekshmi PA, Gangadharan P, Akiba S, Nair RR, Tsuji M, Rajan B. Tobacco chewing and female oral cavity cancer risk in Karunagappally cohort, India. Br J Cancer. 2009;100(5):848–52.
Mehta FS, Bhonsle RB, Murti PR, Daftary DK, Gupta PC, Pindborg JJ. Central papillary atrophy of the tongue among bidi smokers in India: a 10-year study of 182 lesions. J Oral Pathol Med. 1989;18(8):475–80.
Raabe T, Bukrinsky M, Currie RA. Relative contribution of transcription and translation to the induction of tumor necrosis factor-alpha by lipopolysaccharide. J Biol Chem. 1998;273(2):974–80.
Gupta R, Sharma SC, Das SN. Association of TNF-alpha and TNFR1 promoters and 3’ UTR region of TNFR2 gene polymorphisms with genetic susceptibility to tobacco-related oral carcinoma in Asian Indians. Oral Oncol. 2008;44(5):455–63.
Kohaar I, Tiwari P, Kumar R, Nasare V, Thakur N, Das BC, et al. Association of single nucleotide polymorphisms (SNPs) in TNF-LTA locus with breast cancer risk in Indian population. Breast Cancer Res Treat. 2009;114(2):347–55.
Yang CM, Hou YY, Chiu YT, Chen HC, Chu ST, Chi CC, et al. Interaction between tumour necrosis factor-alpha gene polymorphisms and substance use on risk of betel quid-related oral and pharyngeal squamous cell carcinoma in Taiwan. Arch Oral Biol. 2011;56(10):1162–9.
Vairaktaris E, Yapijakis C, Serefoglou Z, Avgoustidis D, Critselis E, Spyridonidou S, et al. Gene expression polymorphisms of interleukins-1 beta, -4, -6, -8, -10, and tumor necrosis factors -alpha, -beta: regression analysis of their effect upon oral squamous cell carcinoma. J Cancer Res Clin Oncol. 2008;134(8):821–32.
Yapijakis C, Serefoglou Z, Vylliotis A, Nkenke E, Derka S, Vassiliou S, et al. Association of polymorphisms in tumor necrosis factor alpha and beta genes with increased risk for oral cancer. Anticancer Res. 2009;29(6):2379–86.
Kramer IR, Lucas RB, Pindborg JJ, Sobin LH. Definition of leukoplakia and related lesions: an aid to studies on oral precancer. Oral Surg Oral Med Oral Pathol. 1978;46(4):518–39.
Bjartveit K, Tverdal A. Health consequences of smoking 1–4 cigarettes per day. Tob Control. 2005;14(5):315–20.
Dufour MC. What is moderate drinking? Defining “drinks” and drinking levels. Alcohol Res Health. 1999;23(1):5–14.
Bairwa M, Rajput M, Sachdeva S. Modified Kuppuswamy’s socioeconomic scale: social researcher should include updated income criteria, 2012. Indian J Community Med. 2012;38(3):185–6.
Sambrook J, Fritsh EF, Maniatis T. Molecular cloning: a laboratory manual. New York: Cold Spring Harbor Laboratory Press; 1989.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21(2):263–5.
Azmy IA, Balasubramanian SP, Wilson AG, Stephenson TJ, Cox A, Brown NJ, et al. Role of tumour necrosis factor gene polymorphisms (−308 and −238) in breast cancer susceptibility and severity. Breast Cancer Res. 2004;6(4):R395–400.
Guo W, Wang N, Li Y, Zhang JH. Polymorphisms in tumor necrosis factor genes and susceptibility to esophageal squamous cell carcinoma and gastric cardiac adenocarcinoma in a population of high incidence region of North China. Chin Med J (Engl). 2005;118(22):1870–8.
Saito S, Kasai Y, Nomoto S, Fujiwara M, Akiyama S, Ito K, et al. Polymorphism of tumor necrosis factor in esophageal, gastric or colorectal carcinoma. Hepatogastroenterology. 2001;48(38):468–70.
Kroeger KM, Carville KS, Abraham LJ. The −308 tumor necrosis factor-alpha promoter polymorphism effects transcription. Mol Immunol. 1997;34(5):391–9.
Liu CJ, Wong YK, Chang KW, Chang HC, Liu HF, Lee YJ. Tumor necrosis factor-alpha promoter polymorphism is associated with susceptibility to oral squamous cell carcinoma. J Oral Pathol Med. 2005;34(10):608–12.
Messer G, Spengler U, Jung MC, Honold G, Blomer K, Pape GR, et al. Polymorphic structure of the tumor necrosis factor (TNF) locus: an NcoI polymorphism in the first intron of the human TNF-beta gene correlates with a variant amino acid in position 26 and a reduced level of TNF-beta production. J Exp Med. 1991;173(1):209–19.
Kohaar I, Thakur N, Salhan S, Batra S, Singh V, Sharma A, et al. TNFalpha-308G/A polymorphism as a risk factor for HPV associated cervical cancer in Indian population. Cell Oncol. 2007;29(3):249–56.
Jang WH, Yang YI, Yea SS, Lee YJ, Chun JH, Kim HI, et al. The −238 tumor necrosis factor-alpha promoter polymorphism is associated with decreased susceptibility to cancers. Cancer Lett. 2001;166(1):41–6.
Gallagher G, Eskdale J, Oh HH, Richards SD, Campbell DA, Field M. Polymorphisms in the TNF gene cluster and MHC serotypes in the West of Scotland. Immunogenetics. 1997;45(3):188–94.
Singh PK, Bogra J, Chandra G, Ahmad MK, Gupta R, Kumar V, et al. Association of TNF-alpha (−238 and −308) promoter polymorphisms with susceptibility of oral squamous cell carcinoma in North Indian population. Cancer Biomark. 2014;15(2):125–31.
Liu L, Yang X, Chen X, Kan T, Shen Y, Chen Z, et al. Association between TNF-alpha polymorphisms and cervical cancer risk: a meta-analysis. Mol Biol Rep. 2011;39(3):2683–8.
Higuchi T, Seki N, Kamizono S, Yamada A, Kimura A, Kato H, et al. Polymorphism of the 5′-flanking region of the human tumor necrosis factor (TNF)-alpha gene in Japanese. Tissue Antigens. 1998;51(6):605–12.
Kaijzel EL, Bayley JP, van Krugten MV, Smith L, van de Linde P, Bakker AM, et al. Allele-specific quantification of tumor necrosis factor alpha (TNF) transcription and the role of promoter polymorphisms in rheumatoid arthritis patients and healthy individuals. Genes Immun. 2001;2(3):135–44.
Uglialoro AM, Turbay D, Pesavento PA, Delgado JC, McKenzie FE, Gribben JG, et al. Identification of three new single nucleotide polymorphisms in the human tumor necrosis factor-alpha gene promoter. Tissue Antigens. 1998;52(4):359–67.
Lee SG, Kim B, Yook JH, Oh ST, Lee I, Song K. TNF/LTA polymorphisms and risk for gastric cancer/duodenal ulcer in the Korean population. Cytokine. 2004;28(2):75–82.
Yang JJ, Ko KP, Cho LY, Shin A, Gwack J, Chang SH, et al. The role of TNF genetic variants and the interaction with cigarette smoking for gastric cancer risk: a nested case–control study. BMC Cancer. 2009;9:238.
Grutters JC, Sato H, Pantelidis P, Lagan AL, McGrath DS, Lammers JW, et al. Increased frequency of the uncommon tumor necrosis factor −857T allele in British and Dutch patients with sarcoidosis. Am J Respir Crit Care Med. 2002;165(8):1119–24.
Hashibe M, Brennan P, Chuang SC, Boccia S, Castellsague X, Chen C, et al. Interaction between tobacco and alcohol use and the risk of head and neck cancer: pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. Cancer Epidemiol Biomarkers Prev. 2009;18(2):541–50.
Joshi MS, Verma Y, Gautam AK, Parmar G, Lakkad BC, Kumar S. Cytogenetic alterations in buccal mucosa cells of chewers of areca nut and tobacco. Arch Oral Biol. 2010;56(1):63–7.
Ko YC, Huang YL, Lee CH, Chen MJ, Lin LM, Tsai CC. Betel quid chewing, cigarette smoking and alcohol consumption related to oral cancer in Taiwan. J Oral Pathol Med. 1995;24(10):450–3.
Lee SS, Yang SF, Ho YC, Tsai CH, Chang YC. The upregulation of metallothionein-1 expression in areca quid chewing-associated oral squamous cell carcinomas. Oral Oncol. 2008;44(2):180–6.
Estruch R, Sacanella E, Badia E, Antunez E, Nicolas JM, Fernandez-Sola J, et al. Different effects of red wine and gin consumption on inflammatory biomarkers of atherosclerosis: a prospective randomized crossover trial. Effects of wine on inflammatory markers. Atherosclerosis. 2004;175(1):117–23.
Haddy N, Sass C, Maumus S, Marie B, Droesch S, Siest G, et al. Biological variations, genetic polymorphisms and familial resemblance of TNF-alpha and IL-6 concentrations: STANISLAS cohort. Eur J Hum Genet. 2005;13(1):109–17.
Hsu HJ, Chang KL, Yang YH, Shieh TY. The effects of arecoline on the release of cytokines using cultured peripheral blood mononuclear cells from patients with oral mucous diseases. Kaohsiung J Med Sci. 2001;17(4):175–82.
Jeng JH, Wang YJ, Chiang BL, Lee PH, Chan CP, Ho YS, et al. Roles of keratinocyte inflammation in oral cancer: regulating the prostaglandin E2, interleukin-6 and TNF-alpha production of oral epithelial cells by areca nut extract and arecoline. Carcinogenesis. 2003;24(8):1301–15.
Lambert C, McCue J, Portas M, Ouyang Y, Li J, Rosano TG, et al. Acrolein in cigarette smoke inhibits T-cell responses. J Allergy Clin Immunol. 2005;116(4):916–22.
Volpato S, Pahor M, Ferrucci L, Simonsick EM, Guralnik JM, Kritchevsky SB, et al. Relationship of alcohol intake with inflammatory markers and plasminogen activator inhibitor-1 in well-functioning older adults: the Health, Aging, and Body Composition study. Circulation. 2004;109(5):607–12.
Mian MF, Lauzon NM, Stampfli MR, Mossman KL, Ashkar AA. Impairment of human NK cell cytotoxic activity and cytokine release by cigarette smoke. J Leukoc Biol. 2008;83(3):774–84.
Acknowledgments
We gratefully acknowledge our patients, their relatives, and clinicians for their support and cooperation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
KB, PS, US, SH, SB, AP, VS, AS, AS, PA, MB, BDB, and RM have no conflict of interest to report.
Funding
This study was funded by grant no. DST SR/SO/HS/0041/2011 from the Government of India to MB and core funds from ICPO (ICMR), Noida, India.
Ethical approval and informed consent
The study procedures were approved by the institutional ethical committee (approval no. ICPO/IEC/P-003/2011), Noida, India. Informed consent was obtained from all participating individuals.
Additional information
K. Bandil and P. Singhal contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bandil, K., Singhal, P., Sharma, U. et al. Impacts of TNF-LTA SNPs/Haplotypes and Lifestyle Factors on Oral Carcinoma in an Indian Population. Mol Diagn Ther 20, 469–480 (2016). https://doi.org/10.1007/s40291-016-0215-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-016-0215-2