Abstract
During the past decade, a series of discoveries has established the potential of the so-called terminally differentiated cells to transition to more primitive progenitor cells. The dramatic demonstration of the ability to reprogram differentiated somatic cells to induced pluripotent stem cells (iPSC) that can then give rise to cells of all three germ layers has opened the possibility of generating virtually any cell type in culture, from any given individual. Taking advantage of these concepts, researchers have generated iPSC by reprogramming a wide variety of somatic cells. In addition to their practical implications, these studies have provided crucial insights into the mechanism of cell plasticity that underlies the transition from one cell type to another. Using concepts derived from research on embryological development, investigators have differentiated iPSC to cells resembling hepatocytes in many ways. Such hepatocyte-like cells could be of enormous value in disease modeling, drug discovery and regenerative medicine. However, the currently available methods do not yield cells that fully reproduce the characteristics of adult primary hepatocytes. Thus, generating hepatocytes from iPSC is very much a work in progress. In addition to chronicling these exciting developments, this review will discuss the emergent new approaches to generating iPSC, improving their differentiation to hepatocyte-like cells, and maintaining the hepatocyte-like cells in culture for longer survival and better function.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In mammals, the liver plays a pivotal role for diverse functions, including protein synthesis, metabolism, detoxification and excretion. Reproducing all or most of these functions in isolated liver cells is a major challenge. Availability of viable, functional hepatocytes would have been highly beneficial for pharmacological evaluation, creating cellular models for pathophysiological analysis of diseases, generating bioartificial liver support and regenerative therapy of the liver. Orthotopic liver transplantation can replace virtually all liver functions and rescue patients with acute and chronic liver failure, as well as monogenic liver diseases, such as Crigler–Najjar Syndrome type 1, alpha-1 antitrypsin deficiency, primary hyperoxaluria, etc. Because liver transplantation is a formidable and expensive procedure, and is dependent on the immediate availability of livers, hepatocyte transplantation is being explored as a minimally invasive alternative to organ transplantation for many of these disorders. However, the severe shortage of donor livers, which are normally prioritized for organ transplantation, drastically limits the availability of usable livers for isolating primary hepatocytes. The problem is compounded by the fact that primary hepatocytes rapidly deteriorate in function in culture and their viability after cryopreservation is extremely variable. Therefore, there is a great need for alternative renewable sources of human hepatocytes. Tissue stem cells, such as mesenchymal and hematopoietic stem cells, liver progenitor cells and pluripotent stem cells (PSC), are being evaluated as sources of human hepatocytes. This review will focus on the different approaches for generating induced pluripotent stem cells (iPSC) from normal or patient-specific somatic cells and their differentiation into hepatocyte-like cells.
Pluripotent Stem Cells for Disease Modeling and Regenerative Medicine
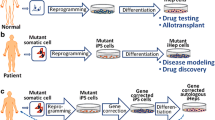
PSC can give rise to all cell types of the body, and therefore offer great promise in disease modeling, drug development and regenerative medicine (Fig. 1). Much progress has been made taking advantage of the unlimited proliferation capacity, plasticity and pluripotency of human embryonic stem cells (hESC) [1•]. The landmark work of Takahashi and Yamanaka [1•, 2•] that led to the reprogramming of somatic cells to iPSC has provided several potential advantages over using ESC. First, ethical concerns are mitigated, as no embryo needs to be destroyed for generation of these cells. Second, iPSC can be generated from individual patients, permitting the development of ‘personalized’ cellular models of genetic diseases. Finally, autologous cells differentiated from iPSC from patients with monogenic disorders could be used for regenerating organs after correction of the genetic defect, thereby circumventing the need for immune suppression.
Generation of iPSC and differentiation to hepatocyte-like cells (iHeps). Somatic cells are collected from normal subjects or patients with liver diseases with known or unknown genetic basis by biopsy, blood drawing, hair plucking or urine collection. The somatic cells are reprogrammed to iPSC by approaches summarized in Fig. 2. The iPSC are differentiated to iHeps with or without gene correction, as indicated, by methods summarized in Fig. 3. The procedures and putative uses are shown in blue font (Color figure online)
Reprogramming Somatic Cells to iPSC
Since Yamanaka and associates generated the first iPSC from mouse fibroblasts in 2006, various additional methods have been reported for reprogramming somatic cells to iPSC. Typically, these methods include expression of the four so-called Yamanaka pluripotency factors, Oct3/4, Sox2, Klf4 and c-Myc (OSKM). As described later in this section, recently an alternative approach has been described based on expression of specific microRNAs that may affect the transcription of multiple genes simultaneously.
Somatic Cells for Reprogramming
Investigators have used many different somatic cell sources for reprogramming into iPSC (Fig. 2). Skin fibroblasts have been used most commonly [2•], but other cells such as peripheral blood cells [3, 4], chord blood endothelia [5], adipose-derived stem cells [6], neural stem cells [7], hepatocytes [8], keratinocytes [9], pancreatic β-cells [10•], amniotic cells [11] or renal tubular epithelial cells shed in the urine [12] can also be used for reprogramming into iPSC. Issues to be considered in selecting the somatic cell source include the ease of collecting the cells, epigenetic memory that may lead to retention of some characteristics of the somatic cells in the derived iPSC, and the possibility of acquired genetic mutations or rearrangement in the somatic cells. For example, umbilical chord blood epithelial cells would be a highly desirable cell type for reprogramming; however, such cells are usually not available in many patients by the time the diagnosis of an inherited disease is made. Tissue stem cells derived from adipose tissue and bone marrow are also excellent cells to reprogram, but invasive procedures are required to obtain them. A less invasive source is venous blood. However, somatic genetic recombination, such as the random combinations of the variable, diverse, and joining gene segments [V(D)J recombination] in T and B lymphocytes remains a concern. Therefore, investigators are exploring the use of peripheral blood hematopoietic stem cells, which are normally present in very small numbers. Skin fibroblasts are the most common cells used for iPSC generation. These cells are obtained by skin biopsy, which is a micro invasive procedure. An additional concern relates to the life-long exposure of the skin cells to environmental pollutants, chemicals and UV rays that may potentially cause somatic cell mutations. The incidence of this has not been determined systematically. Recently, renal tubular epithelial cells shed in the urine have been utilized for iPSC generation. Collection of these cells is completely non-invasive. It should be noted that all somatic cells can acquire somatic mutations, which can persist in the derived iPSC, even after removing the epigenetic marks. To what extent this may affect the characteristics of the iPSC, in terms of their ability to be differentiated to desired target cells, remains to be determined.
The different types of somatic cells reprogrammed to iPSC by various investigators are listed. It appears that any dividing cell has the potential to be reprogrammed. The original reprogramming method is based on delivery of the four Yamanaka pluripotency factors, OCT3/4, SOX2, KLF4 and MYC. The means to achieve this has been listed. An alternative approach is based on the expression of microRNA 316/302 cluster (see text). Once colonies with morphological characteristics and cell surface phenotype of pluripotent cells are obtained, these are passaged multiple rounds and tested for expression of marker genes and teratoma formation
Approaches to Reprogramming
Two basically different approaches have been used by a number of investigators for reprogramming somatic cells to iPSC (Fig. 2). The original approach is based on the landmark work by Takahashi and Yamanaka [2•], which showed that expression of only four transcription factors, OCT3/4, KLF4, SOX2 and MYC, which are now known as Yamanaka factors (Y4), can reprogram somatic cells to iPSC. A number of strategies have been used to achieve the expression of these factors. A completely different approach is based on expression of specific microRNAs, which affect the expression of multiple genes simultaneously. These approaches are discussed in brief below.
Reprogramming Based on Expression of Yamanaka Factors
In their original study, the Yamanaka laboratory used Moloney’s murine leukemia retroviral vectors, which integrate into the cellular genome. Gamma-retroviruses, including the Moloney Leukemia virus, are replication-defective because of the deletion of genes encoding proteins for virus replication and packaging. These vectors infect only dividing cells [13], which reduces the efficiency of reprogramming. Subsequently, expression of the exogenous transgenes is silenced, but in the meantime the reprogrammed cells continue to express endogenous pluripotency factors. However, the transgenes remain integrated into the host cellular genome at random sites. In contrast, vectors based on a different class of retroviruses, termed lentiviruses, can infect non-dividing cells and have been shown to successfully generate iPSC [14]. To reduce the possibility of activation of oncogenes by the randomly integrated proviral genome, vectors have been designed to permit the removal of the exogenous genes using cre recombinase [15]. However, this system may result in incomplete recombination, owing to insufficient expression of Cre recombinase. Another highly efficient integration-based method utilizes the piggyback (PB) transposons. The advantage of this system is that it requires only an active transposase and 13 bp inverted terminal repeats (ITR) for insertion and excision of the reprogramming cassette. The transgenes can be eliminated seamlessly after reprogramming, without leaving any residual nucleic acid sequences in the genome [16].
Despite these design improvements, random integration of proviruses is considered to increase the risk of tumor development in future clinical applications [17]. Therefore, to improve the safety, other methods have been designed that should provide transgene-free iPSC. These methods include transfection of modified mRNAs, delivery of recombinant transcription factor proteins, infection with recombinant episomal viruses (e.g. adenoviruses and Sendaï viruses), and transfection of “minicircles” from which the bacterial component of plasmids is removed [18] or episomal plasmids containing Epstein–Barr viral sequences.
Plews et al. [19] first achieved reprogramming by transfecting in vitro transcribed modified mRNA encoding five pluripotency factors. To increase the half-life of the transfected mRNA by avoiding the cellular interferon response that normally results after RNA transfection, cytidine and uridine residues are replaced with 5-methylcytidine and pseudouridine, respectively. In addition, an interferon receptor mimetic, B18R/B19R, is expressed to reduce interferon binding to its receptor. To further improve the efficiency of this system, other groups have added IRES sequences and strong translational initiation signals in the 5′UTR, and a polyA signal at the 3′UTR [20]. However, despite the mRNA modification to inhibit RNA degradation, this method requires repeated transfection, which may be harmful for more sensitive primary cells.
Reprogramming has also been achieved by direct delivery of reprogramming proteins (OSKM). The major hurdle in this strategy is delivering the proteins across the cell membrane. Peptides rich in arginine or lysine, termed cell penetrating peptides (CPP) [21, 22], such as a peptide fragment of the human immunodeficiency virus transactivator of transcription (HIV-TAT), have been tagged on to the transcription factors to achieve transmembrane delivery. The reprogramming efficiency of this method was low [7], probably because of the need to transfer large amounts of the recombinant transcription factors and a relatively short dwell time of the proteins in the dividing cells.
Recombinant adenoviruses can transduce a large variety of cells from various species and can be generated at high transduction efficiency [23]. Adenoviral vectors are episomal, and integration of the transgene into the host genome is extremely infrequent, but not inexistent [24]. Being episomal, adenoviral vectors are rapidly lost in dividing cells, and repeated infection is needed. Unfortunately, the efficiency for generating iPS cells from primary human cells is much lower compared to mouse fibroblasts [25], which may be related in part to the species difference in the cell surface expression of the adenoviral receptor (Coxsackie adenovirus receptor) [26].
Another non-integrative strategy using recombinant Sendaï virus (aka Hemagglutinating Virus of Japan, HVJ) was first reported by Li et al. [27]. The Sendaï virus is a single-stranded RNA virus of the paramyxovirus family, which differs from other viral vectors in that its entire replication cycle occurs within the cytosol, virtually eliminating the possibility of integration into the genome. A single infection with recombinant Sendaï viruses expressing the pluripotency factors results in a high frequency of reprogramming of human primary somatic cells [28, 29, 30, 31•].
Episomal plasmid vectors offer an efficient integration-free method of somatic cell reprogramming. Conventional plasmids are diluted and lost from dividing cells after transfection, requiring repeated transfection and resulting in a low efficiency of reprogramming [32, 33•, 34]. To overcome this shortcoming, episomal vectors containing oriP/EBNA1 (Epstein–Barr nuclear-antigen 1) have been developed that can replicate during cell division for about six cycles [35]. The Yamanaka laboratory has refined the episomal vector system by developing a set of three plasmids that express OCT3/4, SOX2, KLF4, L-Myc and Lin28, in addition to a shRNA that suppresses p53 expression [33•].
Reprogramming Methods Based on Overexpression of microRNAs
MicroRNAs (miRNAs) can affect the expression of multiple genes in a coordinated manner, and are emerging as important regulators of cell function. Somatic cell reprogramming with miRNAs represent the first alternative to overexpression of transcription factors for generating iPSC. miRNAs that are expressed preferentially in ESC are thought to help maintain the pluripotent cell phenotype [36•, 37•]. Several microRNAs (miRNAs), including the miR302/367 cluster, have been reported to enhance the efficiency of transcription factor-based reprogramming of somatic cells [38•]. The miR302/367 cluster, which is highly expressed in pluripotent cells, consists of five miRNAs located in intron 8 of the Larp 7 gene on chromosome 3 that are transcribed as a single polycistronic transcript [39•]. Four of these microRNAs (miR301a, b, c and d) have identical seed sequences. The miR302/367 cluster is highly conserved across species, and its expression is induced by the pluripotency transcription factors, OCT3/4 and SOX2. In turn, expression of the mir302/367 cluster activates endogenous OCT3/4 expression following reprogramming. Pathway analysis highlighted potential relevant effectors, including mesenchymal-to-epithelial transition, cell cycle, and epigenetic regulators. The miR302-367 targets TGFβ receptor 2, promotes E-cadherin expression, accelerates mesenchymal-to-epithelial transition and promotes cell division [40•]. In fact, retrovirus-mediated expression of this gene cluster alone has been reported to be sufficient in reprogramming both mouse and human somatic cells [41•]. It is now well recognized that chromatin remodeling is essential in reprogramming. In this context, several laboratories have reported a potent and cooperative role of the inhibition of histone deacetylase 2 by valproic acid in miR302/367-mediated reprogramming [38•, 40•, 41•].
Differentiation of iPS into Hepatocytes
Generation of iPS-derived hepatocytes not only serves as a source for potential therapeutic application in human liver diseases, but also enables the understanding of inherited liver diseases by providing cell-based pathophysiological models in vitro. iPSC generated from individual patients with several monogenic liver diseases have been shown to reflect several aspects of the pathologic phenotype of patients, and can potentially provide deeper insights of disease processes and expose new therapeutic targets [42].
Comprehension of the molecular correlates of liver development during embryogenesis has contributed tremendously to the understanding of the differentiation processes [43, 44]. In mammalian embryos, the ventral foregut endoderm is the tissue from which the liver originates. Thus, stepwise induction of definitive endoderm, followed by hepatic progenitors, and finally mature hepatocyte-like cells are the three essential consecutive processes through which (PSC) must pass in order to attain hepatocyte-like phenotypes (Fig. 3). A number of protocols have been established to produce hepatocyte-like cells from both human ESC and iPSC [45•]. The initial step of generating with the phenotype of definitive endoderm has turned out to be crucial in hepatocyte differentiation. Critical requirement for this step is exposure to the transforming growth factor β (TGF-β) superfamily members activin A and bone morphogenic protein 4 (BMP4) [46–49]. A short exposure to Wnt3a, which is expressed at critical stages of human liver development and specifically interacts with activin A [50], enhances the production of definitive and hepatic endoderm. Fibroblast growth factors (FGF), in combination with BMP4, contribute to the definitive endoderm cell commitment at a later stage of embryonic development [51, 52]. An additional important discovery was that the effect of all factors that stimulate early mammalian development is inhibited in the presence of fetal bovine serum in culture medium [53]. It is now clear that the interplay of activin A, Wnt3a, FGF2 and BMP4 plays a major role in determining the early cell fate of pluripotent stem cells toward definitive endoderm. Specific marker genes that are expressed during embryonic development of definitive endoderm include SRY (sex determining region Y)-box 17 (Sox17) and forkhead box A2 (Foxa2). Expression of these genes specifies foregut endoderm, which subsequently gives rise to pancreatic and hepatic cells [54].
The general strategy for directed differentiation of iPSC to hepatocyte-like cells is to emulate the stages of ontogenic development of the liver. Pluripotent cells (hESC and hiPSC) may be differentiated in monolayer cultures or after conversion to embryoid bodies. The first critical step is differentiation to definitive endoderm-like cells. The next step is to achieve hepatocyte specification, which gives rise to hepatoblast-like cells. Finally, attempt is made to induce maturation to cells as close to primary hepatocytes as possible. Factors that are considered to be critical by multiple investigators are bulleted and bold faced. Other factors that have been reported to be helpful are also listed. LY294002 is an inhibitor of phosphatidylinositide 3-kinases (PI3-kinase)
The next step toward hepatocyte generation is the developmental induction of hepatic progenitor cells or hepatoblasts from the definitive endoderm. In cell culture, this is accomplished by adding specific growth factors, of which hepatocyte growth factor (HGF) appears to be the most important [55, 56]. Hepatocyte nuclear factor 4 alpha (HNF4α) is a transcription factor that is expressed initially in the developing hepatic diverticulum. HNF4α expression increases during liver development, and marks the differentiation toward hepatocyte lineage during in vitro differentiation [57]. Another important marker of hepatic progenitor cells is the serum glycoprotein alpha-fetoprotein (AFP), which is expressed in primitive hepatocytes [58].
The final step of the differentiation process is the induction of hepatocyte maturation. In culture this has been achieved by adding oncostatin M (OSM), an interleukin-6 family cytokine, in combination with glucocorticoids [59] to the cell culture medium. Differentiation of hepatocyte-like cells is associated with assumption of hepatocyte-like morphology and intracellular glycogen accumulation. The strategy for changing the liver lineage cells from a hepatoblast/fetal hepatocyte phenotype to cells with adult primary hepatocyte-like characteristics, involves mimicking the molecular/physiological changes that occur during perinatal and neonatal life. Some hepatocellular functions, such as albumin and apolipoprotein synthesis, and urea production, are already near the adult level during late fetal life. Other functions, such as bilirubin glucuronidation and alpha-1 antitrypsin synthesis, are at a very low level at birth, and are stimulated after birth by various factors, including perinatal hormonal changes, increase of portal blood flow and blood oxygen tension after the closure of the ductus venosus, elevation of plasma glucagon and catecholamine levels, and possibly many other changes. Epigenetic modification of DNA during perinatal life plays an important role in the expression of adult hepatocyte-specific genes [60]. Some forms of UDP-glucuronosyltransferase and sulfotransferases reach adult levels only at adolescence, coincident with the surge of sex hormones. As the underlying mechanisms of hepatocyte maturation are not fully known, simulating the perinatal changes in vitro is challenging. On the other hand, the empirically gained knowledge of differentiating cells from the hepatoblast stage to the final hepatocyte-like cells offers a model for progression of hepatocyte maturation.
Expression of marker genes that are generally evaluated at this stage includes albumin, CK18, cytochrome p450 enzymes (CyP), α1-antitrypsin (ATT), asialoglycoprotein receptor 1 (ASGPR), C/EBPα, UGT1A1 and Prox1. In addition to the marker gene expression profile, functional in vitro assays are carried out to further evaluate the features of hepatocyte-like cells; for example, urea production, the uptake of indocyanin green and low-density lipoprotein (LDL), inducible cytochrome P450 activity, and secretion of albumin and alpha-1 antitrypsin into the medium. Primary hepatocytes are usually used for comparison in these in vitro assays. However, a high quality of primary human hepatocytes needs to be assured for this purpose [61].
For a comprehensive list of cell culture components used by various investigators for the stepwise directed differentiation of iPSC to hepatocyte-like cells, see the review by Han et al. [45•].
Improving Differentiation of iPSC-Derived Hepatocyte-Like Cells
Despite the effort of many investigators, differentiation of human iPSC to cells equivalent to primary hepatocytes have not been achieved yet. It is generally stated in literature that the iPSC-derived hepatocytes have fetal hepatocyte-like characteristics. However, although these cells have some characteristics of fetal hepatocytes, they also express some genes, such as uridinediphosphoglucuronate glucuronosyltransferase-1 (UGT1A1) and alpha 1-antitrypsin (SERPINA1). Most hepatocyte-preferred genes are expressed at much lower levels than in primary hepatocytes. Upon transplantation into the liver of immunodeficient or immunosuppressed hosts, these cells engraft with much lower efficiency than the primary hepatocytes. Moreover, as in cultured primary hepatocytes, expression of liver-specific genes in iPSC-derived hepatocyte-like cells declines rapidly. Therefore, much effort is underway to improve the differentiation and maintenance of iPSC-derived hepatocyte-like cells, some of which are discussed below.
Forced Expression of Transcription Factors
In addition to exposure to cytokines and other chemicals, developmental stage-specific transcription factors have been expressed in the cells at various steps of differentiation. Thus, adenovector-mediated expression of SOX17 [62], HEX [63] and HNF4 [64] has been used to improve the differentiation to definitive endoderm, hepatoblasts and hepatocyte-like cells, respectively. Therefore, it may be possible to use sequential transient expression of transcription factors, or perhaps microRNAs, to achieve a more desirable phenotype of hepatocyte-like cells. Until iPSC-derived hepatocytes with morphology and gene expression patterns very similar to adult primary hepatocytes are achieved, it may be necessary to pay special attention to characteristics specifically required for a given application. For example, the nuclear receptor, constitutive androstane receptor (CAR), regulates the expression of multiple gene products involved in the detoxification of endogenous metabolites, drugs and other xenobiotics. It has been reported that permanent transduction of the iPSC-derived cells at a late stage of differentiation with CAR yielded hepatocyte-like cells that exhibited high levels of xenobiotic detoxification functions [65].
Maintenance of the iPSC-Derived Hepatocyte-Like Cells in Culture
Within the liver, hepatocytes form chords with other hepatocytes and exist in the close vicinity of non-parenchymal cells, such as hepatic sinusoidal endothelial cells, stellate cells and Kupffer cells. Clearly, the native liver matrix, the three-dimensional structure of the liver and cross-talk with non-parenchymal cells all play important roles in maintaining the viability and gene expression characteristics of hepatocytes in situ. Therefore, it is unrealistic to expect either primary hepatocytes or iPSC-derived hepatocyte-like cells to retain their function in “minimalistic” monolayer cultures. Based on this consideration, investigators are attempting to partially recreate the spatial organization of the liver to support the function and viability of iPSC-derived hepatocytes [66]. Primary hepatocytes and hepatic sinusoidal epithelial cells appear to support each other in three-dimensional co-culture [67]. These principles have been applied to iPSC-derived hepatocyte-like cells, showing that co-culturing with endothelial and stromal cells favor the maturation of hepatocyte-like cells by cell–cell contact or via paracrine factors [68]. Recently, hepatocyte-specific definitive endoderm was cultured with human umbilical vein endothelial cells and mesenchymal stem cells (MSC), resulting in three-dimensional cellular clusters in which the iPSC-derived cells expressed alpha-fetoprotein, albumin and other hepatocyte-preferred genes, indicating that cluster formation promoted maturation toward a hepatocyte phenotype. When implanted intracranially, in the small bowel mesentery, or under the renal capsule of immune-deficient mice, the clusters became vascularized and proliferated for 2 months. The engrafted cell clusters secreted human albumin and alpha-1-antitrypsin into the host plasma. Furthermore, the cells exhibited cytochrome P450 activity, and improved the survival of mice subjected to toxic hepatic injury. This model partially recreates the liver environment, but does not provide a drainage system, namely bile ducts. Other investigators have attempted to provide a more complete liver-like scaffold to isolated hepatocytes by decellularizing whole livers by detergent perfusion, and then populating the scaffold with primary human hepatocytes and endothelial cells [69, 70]. Although these scaffolds were generated with the purpose of transplanting primary hepatocytes, a similar environment could potentially enhance the differentiation and survival of iPSC-derived hepatocyte-like cells.
In summary, the advent of reprogramming somatic cells to iPSC and directed differentiation of these cells to hepatocyte-like cells offers great promise for applications in pathophysiological studies, pharmacological testing and regenerative medicine. Differentiation of the iPSC toward hepatocyte phenotype can be considered a “work in progress” at this time, but research by many groups worldwide is producing creative and original solutions for overcoming a myriad of existing hurdles, and to eventually translate these exciting developments for pharmacological, pathophysiological and therapeutic applications.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
• Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872. doi:10.1016/j.cell.2007.11.019. Identification of the four pluripotency factors, now termed Yamanaka factors, that can reprogram somatic cells to induced pluripotent cells. Work represented in these seminal papers created the field of iPS cell technology and led to a Nobel prize for Medicine and Physiology in 2012
• Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676. doi:10.1016/j.cell.2006.07.024. Identification of the four pluripotency factors, now termed Yamanaka factors, that can reprogram somatic cells to induced pluripotent cells. Work represented in these seminal papers created the field of iPS cell technology and led to a Nobel prize for Medicine and Physiology in 2012
Loh YH, Hartung O, Li H, Guo C, Sahalie JM, Manos PD et al (2009) Reprogramming of T cells from human peripheral blood. Cell Stem Cell 7:15–19. doi:10.1016/j.stem.2010.06.004
Staerk J, Dawlaty MM, Gao Q, Maetzel D, Hanna J, Sommer CA et al (2010) Reprogramming of human peripheral blood cells to induced pluripotent stem cells. Cell Stem Cell 7:20–24. doi:10.1016/j.stem.2010.06.002
Haase A, Olmer R, Schwanke K, Wunderlich S, Merkert S, Hess C et al (2009) Generation of induced pluripotent stem cells from human cord blood. Cell Stem Cell 5:434–441. doi:10.1016/j.stem.2009.08.021
Sun N, Panetta NJ, Gupta DM, Wilson KD, Lee A, Jia F et al (2009) Feeder-free derivation of induced pluripotent stem cells from adult human adipose stem cells. Proc Natl Acad Sci USA 106:15720–15725. doi:10.1073/pnas.0908450106
Kim JB, Greber B, Arauzo-Bravo MJ, Meyer J, Park KI, Zaehres H et al (2009) Direct reprogramming of human neural stem cells by OCT4. Nature 461:643–649. doi:10.1038/nature08436
Liu H, Ye Z, Kim Y, Sharkis S, Jang YY (2010) Generation of endoderm-derived human induced pluripotent stem cells from primary hepatocytes. Hepatology 51:1810–1819. doi:10.1002/hep.23626
Aasen T, Raya A, Barrero MJ, Garreta E, Consiglio A, Gonzalez F et al (2008) Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nat Biotechnol 26:1276–1284. doi:10.1038/nbt.1503
• Bar-Nur O, Russ HA, Efrat S, Benvenisty N (2011) Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet beta cells. Cell Stem Cell 9:17–23. doi:10.1016/j.stem.2011.06.007. Reprogramming of somatic cells requires removal of epigenetic marks that regulates gene expression, leading to characteristic gene expression pattern of lineage-specified and differentiated cells. Incomplete removal of these epigenic marks may result in the retention of some characteristics of the somatic cells from which the iPS cells were generated. This may reduce the efficiency of reprogramming and the ability to differentiate the iPS cells to desired cells (such as hepatocytes), An implication of this is that maneuvers to fully remove the epigenetic marks could result in more effcient and complete reprogramming of somatic cells
Li H, Collado M, Villasante A, Strati K, Ortega S, Canamero M et al (2009) The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature 460:1136–1139. doi:10.1038/nature08290
Zhou T, Benda C, Duzinger S, Huang Y, Li X, Li Y et al (2011) Generation of induced pluripotent stem cells from urine. J Am Soc Nephrol 22:1221–1228. doi:10.1681/ASN.2011010106
Kitamura T, Koshino Y, Shibata F, Oki T, Nakajima H, Nosaka T et al (2003) Retrovirus-mediated gene transfer and expression cloning: powerful tools in functional genomics. Exp Hematol 31:1007–1014
Picanco-Castro V, de Sousa Russo-Carbolante EM, Tadeu Covas D (2012) Advances in lentiviral vectors: a patent review. Recent Pat DNA Gene Seq 6:82–90
Soldner F, Hockemeyer D, Beard C, Gao Q, Bell GW, Cook EG et al (2009) Parkinson’s disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 136:964–977. doi:10.1016/j.cell.2009.02.013
Woltjen K, Michael IP, Mohseni P, Desai R, Mileikovsky M, Hamalainen R et al (2009) piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458:766–770. doi:10.1038/nature07863
Mikkers H, Berns A (2003) Retroviral insertional mutagenesis: tagging cancer pathways. Adv Cancer Res 88:53–99
Jia F, Wilson KD, Sun N, Gupta DM, Huang M, Li Z et al (2010) A nonviral minicircle vector for deriving human iPS cells. Nat Methods 7:197–199. doi:10.1038/nmeth.1426
Plews JR, Li J, Jones M, Moore HD, Mason C, Andrews PW et al (2010) Activation of pluripotency genes in human fibroblast cells by a novel mRNA based approach. PLoS One 5:e14397. doi:10.1371/journal.pone.0014397
Warren L, Manos PD, Ahfeldt T, Loh YH, Li H, Lau F et al (2010) Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7:618–630. doi:10.1016/j.stem.2010.08.012
Frankel AD, Pabo CO (1988) Cellular uptake of the tat protein from human immunodeficiency virus. Cell 55:1189–1193
Ziegler A, Nervi P, Durrenberger M, Seelig J (2005) The cationic cell-penetrating peptide CPP(TAT) derived from the HIV-1 protein TAT is rapidly transported into living fibroblasts: optical, biophysical, and metabolic evidence. Biochemistry 44:138–148. doi:10.1021/bi0491604
Graham FL, Prevec L (1992) Adenovirus-based expression vectors and recombinant vaccines. Biotechnology 20:363–390
Stephen SL, Sivanandam VG, Kochanek S (2008) Homologous and heterologous recombination between adenovirus vector DNA and chromosomal DNA. J Gene Med 10:1176–1189. doi:10.1002/jgm.1246
Zhou W, Freed CR (2009) Adenoviral gene delivery can reprogram human fibroblasts to induced pluripotent stem cells. Stem Cells 27:2667–2674. doi:10.1002/stem.201
Roy-Chowdhury J, Horwitz MS (2002) Evolution of adenoviruses as gene therapy vectors. Mol Ther 5:340–344. doi:10.1006/mthe.2001.0575S1525001601905751
Li HO, Zhu YF, Asakawa M, Kuma H, Hirata T, Ueda Y et al (2000) A cytoplasmic RNA vector derived from nontransmissible Sendai virus with efficient gene transfer and expression. J Virol 74:6564–6569
Tokusumi T, Iida A, Hirata T, Kato A, Nagai Y, Hasegawa M (2002) Recombinant Sendai viruses expressing different levels of a foreign reporter gene. Virus Res 86:33–38
Fusaki N, Ban H, Nishiyama A, Saeki K, Hasegawa M (2009) Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc Jpn Acad Ser B Phys Biol Sci 85:348–362
Macarthur CC, Fontes A, Ravinder N, Kuninger D, Kaur J, Bailey M et al (2012) Generation of human-induced pluripotent stem cells by a nonintegrating RNA Sendai virus vector in feeder-free or xeno-free conditions. Stem Cells Int 2012:564612. doi:10.1155/2012/564612
• Nishimura K, Sano M, Ohtaka M, Furuta B, Umemura Y, Nakajima Y et al (2011) Development of defective and persistent Sendai virus vector: a unique gene delivery/expression system ideal for cell reprogramming. J Biol Chem 286:4760–4771. doi:10.1074/jbc.M110.183780. In contrast to other recombinant viral vectors, the Sendai virus life cycle does not include a nuclear phase. This virtually eliminates the possibility of integration of the pluripotency genes into the host cell genome. In addition, infection with recombinant sendai viruses is a highly efficient method of expressing the Yamanak factors in most cell types
Gonzalez F, Barragan Monasterio M, Tiscornia G, Montserrat Pulido N, Vassena R, Batlle Morera L et al (2009) Generation of mouse-induced pluripotent stem cells by transient expression of a single nonviral polycistronic vector. Proc Natl Acad Sci USA 106:8918–8922. doi:10.1073/pnas.0901471106
• Okita K, Nakagawa M, Hyenjong H, Ichisaka T, Yamanaka S (2008) Generation of mouse induced pluripotent stem cells without viral vectors. Science 322:949–953. doi:10.1126/science.1164270. Transfection of self-replicating episomal plasmids is an effective and economical method of expressing pluripotency factors. Therefore, this method is being used by an increasing number of laboratories
Si-Tayeb K, Noto FK, Sepac A, Sedlic F, Bosnjak ZJ, Lough JW et al (2010) Generation of human induced pluripotent stem cells by simple transient transfection of plasmid DNA encoding reprogramming factors. BMC Dev Biol 10:81. doi:10.1186/1471-213X-10-81
Yu J, Hu K, Smuga-Otto K, Tian S, Stewart R, Slukvin II et al (2009) Human induced pluripotent stem cells free of vector and transgene sequences. Science 324:797–801. doi:10.1126/science.1172482
• Wang Y, Baskerville S, Shenoy A, Babiarz JE, Baehner L, Blelloch R (2008) Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat Genet 40:1478–1483. doi:10.1038/ng.250. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
• Wang Y, Blelloch R (2009) Cell cycle regulation by MicroRNAs in embryonic stem cells. Cancer Res 69:4093–4096. doi:10.1158/0008-5472.CAN-09-0309. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
• Judson RL, Babiarz JE, Venere M, Blelloch R (2009) Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat Biotechnol 27:459–461. doi:10.1038/nbt.1535. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
• Card DA, Hebbar PB, Li L, Trotter KW, Komatsu Y, Mishina Y et al (2008) Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol Cell Biol 28:6426–6438. doi:10.1128/MCB.00359-08. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
• Liao B, Bao X, Liu L, Feng S, Zovoilis A, Liu W et al (2011) MicroRNA cluster 302–367 enhances somatic cell reprogramming by accelerating a mesenchymal-to-epithelial transition. J Biol Chem 286:17359–17364. doi:10.1074/jbc.C111.235960. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
• Anokye-Danso F, Trivedi CM, Juhr D, Gupta M, Cui Z, Tian Y et al (2011) Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell 8:376–388. doi:10.1016/j.stem.2011.03.001. Reprogramming approach that does not utilize direct forced expression of pluripotency factors. In this approach reprogramming is orchestrated by expressing a single microRNA cluster
Park IH, Arora N, Huo H, Maherali N, Ahfeldt T, Shimamura A et al (2008) Disease-specific induced pluripotent stem cells. Cell 134:877–886. doi:10.1016/j.cell.2008.07.041
Irion S, Nostro MC, Kattman SJ, Keller GM (2008) Directed differentiation of pluripotent stem cells: from developmental biology to therapeutic applications. Cold Spring Harb Symp Quant Biol 73:101–110. doi:10.1101/sqb.2008.73.065
Murry CE, Keller G (2008) Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell 132:661–680. doi:10.1016/j.cell.2008.02.008
• Han S, Bourdon A, Hamou W, Dziedzic N, Goldman O, Gouon-Evans V (2012) Generation of functional hepatic cells from pluripotent stem cells. J Stem Cell Res Ther. doi:10.4172/2157-7633.S10-008. This paper lists and summarizes the different methods used by various investigators for differentiating pluripotent stem cells to hepatocyte-like cells
Touboul T, Hannan NR, Corbineau S, Martinez A, Martinet C, Branchereau S et al (2010) Generation of functional hepatocytes from human embryonic stem cells under chemically defined conditions that recapitulate liver development. Hepatology 51:1754–1765. doi:10.1002/hep.23506
Yamanaka Y, Ralston A (2010) Early embryonic cell fate decisions in the mouse. Adv Exp Med Biol 695:1–13. doi:10.1007/978-1-4419-7037-4_1
Vallier L, Touboul T, Chng Z, Brimpari M, Hannan N, Millan E et al (2009) Early cell fate decisions of human embryonic stem cells and mouse epiblast stem cells are controlled by the same signalling pathways. PLoS One 4:e6082. doi:10.1371/journal.pone.0006082
Teo AK, Ali Y, Wong KY, Chipperfield H, Sadasivam A, Poobalan Y et al (2012) Activin and BMP4 synergistically promote formation of definitive endoderm in human embryonic stem cells. Stem Cells 30:631–642. doi:10.1002/stem.1022
Toivonen S, Lundin K, Balboa D, Ustinov J, Tamminen K, Palgi J et al (2013) Activin A and Wnt-dependent specification of human definitive endoderm cells. Exp Cell Res 319:2535–2544. doi:10.1016/j.yexcr.2013.07.007
Hansson M, Olesen DR, Peterslund JM, Engberg N, Kahn M, Winzi M et al (2009) A late requirement for Wnt and FGF signaling during activin-induced formation of foregut endoderm from mouse embryonic stem cells. Dev Biol 330:286–304. doi:10.1016/j.ydbio.2009.03.026
Xu X, Browning VL, Odorico JS (2011) Activin, BMP and FGF pathways cooperate to promote endoderm and pancreatic lineage cell differentiation from human embryonic stem cells. Mech Dev 128:412–427. doi:10.1016/j.mod.2011.08.001
Johansson BM, Wiles MV (1995) Evidence for involvement of activin A and bone morphogenetic protein 4 in mammalian mesoderm and hematopoietic development. Mol Cell Biol 15:141–151
Norrman K, Strombeck A, Semb H, Stahlberg A (2013) Distinct gene expression signatures in human embryonic stem cells differentiated towards definitive endoderm at single-cell level. Methods 59:59–70. doi:10.1016/j.ymeth.2012.03.030
Kamiya A, Kinoshita T, Miyajima A (2001) Oncostatin M and hepatocyte growth factor induce hepatic maturation via distinct signaling pathways. FEBS Lett 492:90–94
Schmidt C, Bladt F, Goedecke S, Brinkmann V, Zschiesche W, Sharpe M et al (1995) Scatter factor/hepatocyte growth factor is essential for liver development. Nature 373:699–702. doi:10.1038/373699a0
Duncan SA, Manova K, Chen WS, Hoodless P, Weinstein DC, Bachvarova RF et al (1994) Expression of transcription factor HNF-4 in the extraembryonic endoderm, gut, and nephrogenic tissue of the developing mouse embryo: HNF-4 is a marker for primary endoderm in the implanting blastocyst. Proc Natl Acad Sci USA 91:7598–7602
Gualdi R, Bossard P, Zheng M, Hamada Y, Coleman JR, Zaret KS (1996) Hepatic specification of the gut endoderm in vitro: cell signaling and transcriptional control. Genes Dev 10:1670–1682
Kamiya A, Kinoshita T, Ito Y, Matsui T, Morikawa Y, Senba E et al (1999) Fetal liver development requires a paracrine action of oncostatin M through the gp130 signal transducer. EMBO J 18:2127–2136. doi:10.1093/emboj/18.8.2127
Thomassin H, Flavin M, Espinas ML, Grange T (2001) Glucocorticoid-induced DNA demethylation and gene memory during development. EMBO J 20:1974–1983. doi:10.1093/emboj/20.8.1974
Hengstler JG, Brulport M, Schormann W, Bauer A, Hermes M, Nussler AK et al (2005) Generation of human hepatocytes by stem cell technology: definition of the hepatocyte. Expert Opin Drug Metab Toxicol 1:61–74. doi:10.1517/17425255.1.1.61
Takayama K, Inamura M, Kawabata K, Tashiro K, Katayama K, Sakurai F et al (2011) Efficient and directive generation of two distinct endoderm lineages from human ESCs and iPSCs by differentiation stage-specific SOX17 transduction. PLoS One 6:e21780. doi:10.1371/journal.pone.0021780PONE-D-11-02464
Inamura M, Kawabata K, Takayama K, Tashiro K, Sakurai F, Katayama K et al (2011) Efficient generation of hepatoblasts from human ES cells and iPS cells by transient overexpression of homeobox gene HEX. Mol Ther 19:400–407. doi:10.1038/mt.2010.241
Takayama K, Inamura M, Kawabata K, Katayama K, Higuchi M, Tashiro K et al (2012) Efficient generation of functional hepatocytes from human embryonic stem cells and induced pluripotent stem cells by HNF4alpha transduction. Mol Ther 20:127–137. doi:10.1038/mt.2011.234
Funakoshi N, Duret C, Pascussi JM, Blanc P, Maurel P, Daujat-Chavanieu M et al (2011) Comparison of hepatic-like cell production from human embryonic stem cells and adult liver progenitor cells: CAR transduction activates a battery of detoxification genes. Stem Cell Rev 7:518–531. doi:10.1007/s12015-010-9225-3
Decaens C, Durand M, Grosse B, Cassio D (2008) Which in vitro models could be best used to study hepatocyte polarity? Biol Cell 100:387–398. doi:10.1042/BC20070127
Kim Y, Rajagopalan P (2010) 3D hepatic cultures simultaneously maintain primary hepatocyte and liver sinusoidal endothelial cell phenotypes. PLoS One 5:e15456. doi:10.1371/journal.pone.0015456
Stevens KR, Ungrin MD, Schwartz RE, Ng S, Carvalho B, Christine KS et al (2013) InVERT molding for scalable control of tissue microarchitecture. Nat Commun 4:1847. doi:10.1038/ncomms2853
Baptista PM, Siddiqui MM, Lozier G, Rodriguez SR, Atala A, Soker S (2011) The use of whole organ decellularization for the generation of a vascularized liver organoid. Hepatology 53:604–617. doi:10.1002/hep.24067
Uygun BE, Soto-Gutierrez A, Yagi H, Izamis ML, Guzzardi MA, Shulman C et al (2010) Organ reengineering through development of a transplantable recellularized liver graft using decellularized liver matrix. Nat Med 16:814–820. doi:10.1038/nm.2170
Acknowledgments
This work was supported in part by NIDDK. 1PO1 DK 096990-01 (to JRC, PD: D. Perlmutter); NIDDK DK092469 (to NRC), New York Stem Cell Foundation CO26440 (to JRC) and German Research Foundation SA 2451/1-1 (to VS).
Compliance with Ethics Guidelines
Conflict of Interest
Vanessa Sauer, Namita Roy-Chowdhury, Chandan Guha and Jayanta Roy-Chowdhury declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sauer, V., Roy-Chowdhury, N., Guha, C. et al. Induced Pluripotent Stem Cells as a Source of Hepatocytes. Curr Pathobiol Rep 2, 11–20 (2014). https://doi.org/10.1007/s40139-013-0039-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40139-013-0039-2