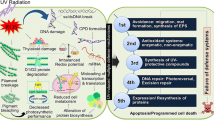
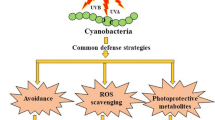
Abstract
Scytonemin is a lipid-soluble, highly stable, yellow–brown-coloured secondary metabolite that is accumulated in the extracellular polysaccharide sheath of several but not all members of cyanobacteria. Chemically, scytonemin is an indole alkaloid composed of two heterocyclic units symmetrically connected through a carbon–carbon bond. Thus, scytonemin is unique among natural products due to its special structure, location in a cell, as well as strong absorption maxima in UV-A in addition to the violet–blue region. Traditionally, scytonemin is a well-established photoprotective compound against ultraviolet radiation. Its accumulation in the cyanobacterial sheath has been suggested to be a strategy adopted by several cyanobacteria to protect their cellular components against damaging effects of UVR. Additionally, recent studies have also established the importance of scytonemin in reactive oxygen species scavenging as well as in controlling the growth of cancerous cells. Thus, scytonemin is both ecologically as well as pharmaceutically important metabolite. Recent developments made in the biochemistry and genetics of this compound have paved the way for its application and commercialization for human welfare. This review aims to present a brief history of the compound with chronological developments made in the study of scytonemin and emphasizes its physiochemistry, analytical chemistry, biochemistry, and genetics. We provide a separate section for metabolic engineering and potential applications of scytonemin, mainly as sunscreen and anti-cancerous drugs. We also discuss the future research directions which need to be worked out.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Scytonemin is exclusively biosynthesized by several but not all cyanobacteria in response to mainly ultraviolet-A (UV-A; 315–400 nm) radiation stress. It is a lipid-soluble and yellow–brown-coloured secondary metabolite that acts as a photoprotective compound [1, 2]. Several other stress factors such as salinity, oxidative stress, desiccation, nitrogen limitation, and high temperature have been also found to influence the biosynthesis of scytonemin in cyanobacteria [3, 4]. Thus, the biosynthesis of scytonemin in cyanobacteria may be regulated as a part of a complex stress response where more than one stress factors can affect its biosynthesis. The well-known habitats for scytonemin-biosynthesizing cyanobacteria are intertidal mats, biofilms attached to rocks, monuments, the bark of trees, and soil surfaces with cyanobacterial crusts in arid or semi-arid ecosystems [2, 5, 6]. Thus, cyanobacteria growing in brightly lit habitats with a recurring cycle of desiccation, nutrient limitation, and high temperature are the good source of scytonemin. Table 1 shows the concentration of scytonemin found in different environmental habitats. Scytonemin accumulates in the extracellular matrix secreted by cyanobacterial cells where it forms a stable protective layer that absorbs incoming UV-A radiation and therefore provides passive protection to the cellular components [1, 7,8,9]. Thus, scytonemin acts as a barrier for incoming UV-A radiation without any further energy investment after its synthesis.
The dry cyanobacterial mats survive under harsh conditions by minimizing their biological activities while accumulating very high levels of scytonemin [11]. The accumulation of high levels of scytonemin has been proposed to be responsible for the survival and reversal of normal biological activities, including photosynthesis and growth, on rehydration of cyanobacterial mats [11]. Thus, scytonemin supports long-term survival at the expenditure of a minimal amount of cellular energy under extreme stress conditions. The production of scytonemin has been proposed to be one of the ancient and crucial photoprotective mechanisms developed by cyanobacteria to protect their cellular components when there was no ozone layer in the Earth’s atmosphere to filter the lethal dose of incoming ultraviolet radiation (UVR) [5, 7]. Thus, scytonemins are evolutionary important molecules that supported life in the absence of the ozone layer and led to the development of oxygenic photosynthesis and aerobic life by protecting the ancestors of cyanobacteria [12]. Also, the antioxidant activity of scytonemin provides additional protection to cellular components by neutralizing the reactive oxygen species (ROS) which are produced in the presence of various abiotic stressors, including UV-A radiation [10].
The importance of scytonemin in cyanobacterial survival and its role in the development of existing life is well documented. However, recent researches have shown that this ancient secondary metabolite is also important to humans as a drug. Scytonemin is well known for its anti-inflammatory and anti-proliferative activities, which necessitate its large-scale production in the pharmacy industry for its application as a potent anti-cancerous drug [13, 14]. The recent advancements in the genetics of scytonemin have led to the development of genetically engineered Escherichia coli (E. coli) strain expressing genes of cyanobacteria for scytonemin biosynthesis [15]. The large-scale production of scytonemin using the above-mentioned metabolically engineered production strain or cyanobacteria can be combined with biofuel production to support the production cost and economic sustainability of such ventures [16]. The commercial production of scytonemin in heterotrophic mode could be economically challenging. Therefore, the production of scytonemin in phototrophic and/or mixotrophic mode together with other valuable chemicals and biofuel can be an option for the economic viability of biorefinery [17]. Additionally, the presence of scytonemin in stromatolitic mats and other extremophilic cyanobacterial mats has led to the development of the proposition that scytonemin can be used as a biomarker for the detection of life in scientific projects aiming astrobiological exploration of life on other planets [18, 19].
Thus, scytonemin has both ecological and economic importance in addition to its potential to be utilized in various branches of biology, including pharmacology and astrobiology. In this review, we have summarized and discussed the findings related to the discovery, physiochemistry, biosynthetic pathway, and commercial applications of scytonemins while giving comprehensive information on recent developments made in the genetics and biochemistry of scytonemin biosynthesis.
A Brief History of Scytonemin
In 1849, Swiss botanist Carl Nägeli observed yellowish-brown cyanobacterial sheath colouration [20] and coined the name “scytonemin” in 1877 for this coloured pigment [21]. Garcia-Pichel and Castenholz [1] were the pioneer workers who studied the scytonemin in several cyanobacterial species from laboratory cultures as well as natural populations, including crusts, mats, and colonies. After more than 100 years of its discovery, Proteau et al. [8] for the first time solved the chemical structure of scytonemin which was found to be a novel dimeric molecule of indolic and phenolic subunits. The structure of scytonemin is unique among all known natural organic substances. Proteau et al. [8] gave the name “the scytoneman skeleton” to the carbon skeleton of this unique secondary metabolite. Based on their findings, it was proposed that scytonemin is formed by condensation of tryptophan and phenylpropanoid-derived subunits. Another breakthrough in the field of scytonemin study was made when Balskus and Walsh [22] cracked the genetics of this valuable compound. However, the mechanistic insight of abiotic stressors sensing and signalling pathway(s) involved in the biosynthesis of scytonemin is still lacking.
Physiochemistry of Scytonemin
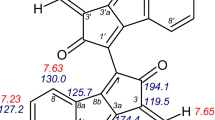
Scytonemin, the most common member of aromatic indole alkaloids, is a relatively small molecule (molecular formula C36H20N2O4; molecular weight 544.566 g/mol). Scytonemin is consisting of two identical condensation products of tryptophanyl and tyrosyl-derived subunits which are linked through a carbon–carbon bond [8]. Its IUPAC name is 3-[(4-oxocyclohexa-2, 5-dien-1-ylidene) methyl]-1-[2-oxo-3-[(4-oxocyclohexa-2, 5-dien-1-ylidene) methyl]-4H-cyclopenta [b] indol-1-yl]-4H-cyclopenta [b] indol-2-one [23]. Depending on the redox state, scytonemin is found in two inter-convertible forms. A predominant oxidized yellowish-brown form of scytonemin is insoluble in water and less soluble in organic solvents such as pyridine and tetrahydrofuran. However, the reduced form is comparatively more soluble in organic solvents and characterized by a bright red colour [1, 8]. In addition to oxidized and reduced forms, several other derivatives of scytonemin have been also reported. Dimethoxy and tetramethoxy scytonemins are derivatives of reduced scytonemin where one or both of the ethenyl groups in the molecule have been saturated by two or four methoxy groups, respectively [24, 25]. Another derivative of the scytoneman skeleton is scytonemin-3a-imine in which C-3a atom of scytonemin is attached with a 2-imino-propyl radical [26]. Only the structure of scytonine deviates substantially from the dimeric scytoneman skeleton due to the loss of one para-substituted phenol group and ring openings of both cyclopentenones where successive methoxylation and secondary cyclizations take place [24]. The spectral and structural characteristics of scytonemin derivatives known so far are shown in Table 2 [9, 26,27,28].
Scytonemin is exclusively produced by cyanobacteria which are capable of producing extracellular polysaccharide (EPS) (Fig. 1a). The complex ring structure of scytonemin permits strong absorption broadly across the UV-C, UV-B, and UV-A-violet–blue spectral regions with a maximum of around 370 nm in vivo [8]. In addition to significant absorbance at 252, 278, and 300 nm, purified scytonemin absorbs maximally at 386 nm (Fig. 1c). The large extinction coefficient (ε = 250 l g−1 cm−1 at 384 nm) of scytonemin makes it an efficient photoprotective compound [8]. Furthermore, the long-term stability of scytonemin in cyanobacterial biocrusts or dried mats, which are exposed to intense solar radiation, suggests its exceptionally high photo-stability and utilization in the cosmetic industry [3, 7, 29].
Scytonemin biosynthesis in cyanobacteria. Photograph (40 ×) showing the false branching in the filament of Scytonema sp. with sheath containing brown-coloured UV-protective pigment scytonemin (indicated by black arrow) (a). HPLC chromatogram of partially purified scytonemin obtained from Scytonema sp. showing retention time (RT) of 1.7 min (indicated above peak) for this compound (b). The absorption spectrum of purified scytonemin extracted from Scytonema sp. showing absorption maxima at 252, 278 and 386 nm (c)
Factors Affecting the Biosynthesis of Scytonemin
A few cyanobacteria produce indole-alkaloid scytonemin on exposure to long wavelengths of UVR [1, 30]. In addition to UVR, nutrient limitation, mainly nitrogen, iron, or magnesium limitation, is also known to induce the biosynthesis of scytonemin in cyanobacteria [4, 31]. Several other stress factors such as heat, salinity, osmotic, and oxidative stress alone or in combination with UVR can also affect the biosynthesis of scytonemin in cyanobacteria [4, 32, 33]. Fleming and Castenholz [3] showed that hydration of Nostoc punctiforme ATCC 29133/PCC 73102 (hereafter, N. punctiforme) filaments for 2 days followed by intermittent desiccation periods results in higher scytonemin biosynthesis as compared to the filaments which were hydrated for 1 day. However, the contradictory results were found in Chroococcidiopsis where periodic desiccation inhibited scytonemin biosynthesis. In conclusion, scytonemin biosynthesis is regulated by several abiotic factors, and its accumulation in a cell helps in maximizing the fitness of organisms under various stressors. Soule et al. [34] investigated the relationship between scytonemin and EPS production under UV-A stress. EPS production was measured following exposure of wild-type (WT) N. punctiforme and non-scytonemin-producing strain SCY59 to UV-A and oxidative stress. Under UV-A exposure, SCY59 produces significantly more EPS than the unstressed controls and the WT, while both SCY59 and WT strains produce more EPS under oxidative stress compared to the controls [34]. These findings suggested that EPS secretion occurs in response to oxidative stress which is a by-product of UV-A irradiation rather than as a direct response to UV-A radiation. The exact mechanism(s) of scytonemin induction are still unclear, but multiple environmental signals could act together to determine the level of this compound in selected species of cyanobacteria.
The biosynthesis of scytonemin in different environmental samples can vary greatly indicating the metabolic status of organisms under different environmental conditions (Table 1). Generally, photosynthetically active cells have the carbon pool to divert it towards the biosynthesis of scytonemin in response to environmental signals. The allocation of carbon pool towards l-tryptophan and l-tyrosine synthesis increases the biosynthesis of scytonemin [15]. Once synthesized, highly stable scytonemin can protect an organism for a long time without further requirement for its biosynthesis. Thus, metabolically active cells with a sufficient amount of nutrients and carbon pools are required for the production of scytonemin. As shown in Table 1, intertidal microbial mats or terrestrial cyanobacterial colonies possess the highest amount of scytonemin. This could be explained by the fact that these mats possess several layers of metabolically active and inactive cells. The upper metabolically active layer of cells performs active photosynthesis and accumulates the highest amount of scytonemin in comparison with the lower layer of cells which is metabolically and photosynthetically inactive [5, 10]. Therefore, in addition to the metabolic status of cells, the above-mentioned inducers of scytonemin biosynthesis are required for the maximum accumulation/production of scytonemin.
Genetics and Biochemistry of Scytonemin Biosynthesis
Elucidation of the genetic and biochemical basis of scytonemin biosynthesis is of great interest because of its unique structure, place of partial biosynthesis, and accumulation in a cell, i.e., in the periplasmic space of Gram-negative cell wall. The genetics of scytonemin biosynthesis was solved after a long period of its discovery in the cyanobacterium N. punctiforme using a random transposon mutagenesis approach [35]. In N. punctiforme, the genomic region (Table 3) associated with scytonemin biosynthesis contains an 18-gene cluster (Npun_R1276 to Npun_R1259) [35]. The chemical structure of scytonemin and initial feedback inhibition, as well as radiotracer studies, had earlier suggested tyrosine and tryptophan as the biosynthetic precursors of the scytonemin [8]. Interestingly, the identified gene cluster possesses genes encoding proteins involved in the shikimic acid pathway (aroB and aroG) as well as proteins involved in the biosynthesis of tryptophan (trpE, trpC, trpA, trpB, and trpD) and tyrosine (tyrP) [35, 36]. Further genomic analysis revealed the presence of an additional copy of genes encoding AroB, AroG, TrpE, TrpC, TrpA, TrpB, and TrpD and TyrP proteins at different places in the genome of N. punctiforme. This suggested that identified genes (Table 3) are exclusively dedicated to the biosynthesis of scytonemin by providing the precursor amino acids [35, 36]. Thus, it appears that N. punctiforme has duplicated certain genes of the shikimic acid pathway in its genome to produce enough amount of raw materials needed for the essential amino acids and scytonemin biosynthesis. Furthermore, during the evolution of this strain, these duplicated genes managed to cluster with other genes exclusively involved in the synthesis of amino acids or scytonemin.
The comparative genomic analysis revealed the presence of genes associated with scytonemin biosynthesis in other cyanobacterial species such as Lyngbya sp. PCC 8106, Anabaena sp. PCC 7120, Nodularia sp. CCY 9414, and Chlorogloeopsis sp. CCMEE 5094 [35, 36]. Table 4 gives an overview of the occurrence of scytonemin and its derivatives in different cyanobacteria from diverse habitats. Additionally, six putative genes of the scytonemin gene cluster in N. punctiforme (Npun_R1276 to Npun_R1271) annotated as scyA, scyB, scyC, scyD, scyE, and scyF were proposed to be involved in the assembly of scytonemin from central metabolites [36]. Among newly identified proteins, ScyA, TyrP, and NPR1259 possess transmembrane domain, while ScyD, ScyE, and ScyF possess export signal domains, which suggest that biosynthesis of scytonemin is compartmentalized between the cytoplasm and periplasm [36]. A two-component regulatory system (Npun_F1277/Npun_F1278) upstream of the scytonemin biosynthetic genes has been identified in N. punctiforme genome [36]. Based on the comparative genomic analysis, these gene products have been proposed to be involved in the regulation of scytonemin biosynthesis [36]. The gene expression analysis of the identified 18-gene cluster suggests that all 18 genes are upregulated by UV-A exposure, confirming the involvement of UV-A radiation in the induction of scytonemin biosynthesis. Furthermore, all 18 genes are co-transcribed as a single transcriptional unit in N. punctiforme [37]. The in vitro characterization of ScyA, ScyB, and ScyC proteins for their involvement in the early stages of scytonemin assembly was demonstrated by overexpression of corresponding genes in E. coli [38, 39]. ScyB (Npun_R1275), a leucine dehydrogenase homolog which has been recently functionally characterized and renamed as l-tryptophan dehydrogenase [40], catalyzes the first biosynthetic step by oxidative deamination of l-tryptophan to provide indole-3-pyruvic acid. TyrA (Npun_R1269), a putative prephenate dehydrogenase, is responsible for the oxidation of prephenate to p-hydroxyphenylpyruvic acid [41]. Subsequently, ScyA (Npun_R1276), a thiamine-dependent enzyme, mediates the acyloin coupling of indole-3-pyruvic acid and p-hydroxyphenylpyruvic acid, producing the labile β-ketoacid precursor [38]. Thereafter, ScyC (Npun_R1274) catalyzes the cyclization and decarboxylation of β-ketoacid precursor to form a ketone. The compound yielded by ScyC further undergoes the oxidation process to form a scytonemin monomeric precursor [39]. The detailed model for the genetic and biochemical bases of scytonemin biosynthesis in cyanobacteria is shown in Fig. 2.
Model for the biosynthesis of scytonemin in Nostoc punctiforme ATCC 29133 showing different biochemical steps and corresponding gene products involved in the synthesis of photoprotective compound. ScyA, ScyB, and ScyC proteins encoded by scyA, scyB, and scyC genes, respectively, lead to the accumulation of monomer moieties of scytonemin in the cytoplasm. Monomers are translocated into the periplasm by Ebo complex, encoded by eboF, eboE, eboC, eboB, and eboA genes, where ScyE encoded by scyE gene dimerizes the monomers to produce the final product scytonemin [35, 38, 61]
Recently, heterologous expression of only three N. punctiforme genes, i.e., scyABC, in E. coli, resulted in the production of monomer moiety of scytonemin upon supplementation of tryptophan and tyrosine [15]. This finding further confirms the involvement of scyABC gene products in scytonemin biosynthesis and suggests that only ScyA, ScyB, and ScyC proteins are required for the heterologous production of scytonemin monomer in E. coli. The additional reaction required for monomer biosynthesis can be carried out by the endogenous enzymes of the E. coli [15]. Furthermore, Ferreira and Garcia-Pichel [59] characterized the role of ScyD, ScyE, and ScyF proteins in the biosynthesis of scytonemin in N. punctiforme. The ∆scyE strain was found to lack scytonemin production; however, this strain did not accumulate any predicted intermediates of scytonemin biosynthetic pathway. ∆scyD and ∆scyF strains produce scytonemin similar to the WT; however, scytonemin production in ∆scyF strain is slightly hampered in comparison with WT [59]. These findings collectively suggest that ScyD and ScyF are not essential for the last steps of scytonemin biosynthesis in N. punctiforme and could be redundant. The transcriptional analysis of two-component regulatory system genes, i.e., Npun_F1277 and Npun_F1278, suggested that these two genes are co-transcribed and upregulated by scytonemin inducing condition, i.e., UV-A exposure [60]. This finding further strengthens the proposition that Npun_F1277 and Npun_F1278 products regulate the scytonemin biosynthesis. Naurin et al. [30] studied the role of response regulator (RR) protein encoded by Npun_F1278, which was earlier identified by the comparative genomic study and proposed to regulate the biosynthesis of scytonemin. The mutant strain of Npun_F1278 of N. punctiforme completely lacks scytonemin even after induction by UV-A [30]. This suggests that RR Npun_F1278 is essential for scytonemin biosynthesis and participates in the signal transduction pathway at least for the UV-A-mediated biosynthesis of scytonemin.
Taken together, the above-mentioned findings suggest that scyA, scyB, and scyC gene products together with the two-component regulatory system are essential for UV-A-dependent biosynthesis of monomer moiety of scytonemin in the cytoplasm (Fig. 2). The protein encoded by scyE is essential for catalyzing the oxidative dimerization of monomer moiety in the periplasm to finally produce the scytonemin molecules (Fig. 2). However, how monomer moieties of scytonemin are translocated into the periplasm for their final assembly by ScyE was unknown until the recent study by Klicki et al. [61]. They reported that the Ebo complex is responsible for the transportation of monomer moieties of scytonemin. The group of five highly conserved genes (Npun_F5232 to Npun_F5236) are located at a different location from that of the scytonemin operon in the genome of N. punctiforme. However, the expression of this gene cluster is coordinated with the expression of scytonemin operon after induction with UV-A radiation [36, 37, 61]. This highly conserved five-gene cluster is also found in several bacteria as well as in the plastid genome of eustigmatophyte algae and therefore named as eustigmatophyte/bacterial operon or ebo [61, 62]. Interestingly, ebo genes are found within scytonemin operons of several scytonemin-producing cyanobacteria [61]. The products encoded by ebo gene cluster helps in transportation of monomer moiety of the scytonemin from the cytoplasm to periplasm. The knockout of any individual gene of ebo cluster or removal of whole ebo cluster from the genome of N. punctiforme results in the accumulation of monomer moiety of scytonemin in the cytoplasm. However, in ∆scyE strain, monomer moieties are accumulated in the periplasm [61]. These findings collectively suggest that Ebo complex encoded by ebo gene cluster plays a crucial role in the export of scytonemin monomers to the periplasm for their final dimerization by ScyE (Fig. 2). In summary, research in the field of scytonemin in the last decade has significantly advanced our knowledge about the genetics, biochemistry, and regulators of scytonemin biosynthesis. This information can be useful for the large-scale production of scytonemin for its commercial application in different areas by fine-tuning its biosynthesis using metabolically engineered strains as well as in understanding the evolutionary significance of scytonemin. It can also be utilized to produce non-cyanobacterial commercial strain for scytonemin production by installing the missing genes in the strain using synthetic biology tools.
Metabolic Engineering of Scytonemin Biosynthesis
Recent advancements in the field of genetics of scytonemin biosynthesis have paved the way for the development of metabolically engineered strains producing scytonemin for commercialization. All the essential enzymes and substrates required for the production of scytonemin are now well known (Fig. 3). Scytonemin biosynthesis requires products of the shikimate pathway, i.e., l-tryptophan and l-tyrosine, which enters the scytonemin biosynthesis pathway through the action of cyanobacterial genes coding ScyB and ScyA proteins, respectively. These steps are rate limiting for the synthesis of scytonemin as exogenous addition of l-tryptophan and l-tyrosine substrates increases the production of monomer moieties of scytonemin in a metabolically engineered strain of E. coli [15]. Therefore, diversion of more carbon flux towards l-tryptophan and l-tyrosine biosynthesis from chorismic acid could potentially increase the scytonemin biosynthesis by providing the two substrates in enough quantity. ScyB product is a tryptophan dehydrogenase that catalyzes the conversion of l-tryptophan to indole-3-pyruvic acid [40]. However, ScyB of N. punctiforme is catalytically inefficient and protein engineering using error-prone PCR method has been suggested to increase its efficiency by directed evolution [40]. In this approach, different mutant versions of ScyB are selected against higher efficiency of indole-3-pyruvic acid production. A similar approach can be taken for other enzymes to increase the stability of enzymes and scytonemin production. Installation of cyanobacterial genes encoding ScyA, ScyB, ScyC, and ScyE proteins in E. coli resulted in the production of monomer moieties of scytonemin [15]. However, the production of scytonemin was not achieved even after providing all essential proteins which are required for the synthesis of scytonemin in N. punctiforme [15]. This could be explained by the fact that ScyE protein of N. punctiforme, which catalyzes the dimerization of monomer moieties to give scytonemin, contains a target peptide sequence for targeting this enzyme to periplasm [36]. Thus, monomer moieties of scytonemin, which are produced in the cytoplasm in engineered strain, might not get transported to periplasm for their dimerization by ScyE enzyme [15]. A recent discovery of the Ebo FECBA complex could solve this problem as this complex is responsible for the transportation of monomer moieties of scytonemin to periplasmic space [61]. Therefore, the installation of Ebo FECBA complex in a metabolically engineered strain of E. coli, which accumulates ScyA, ScyB, ScyC, and ScyE proteins, could be required for the production of scytonemin. In summary, ScyA, ScyB, ScyC, ScyE, and Ebo FECBA, which are cyanobacterial in origin (shown in the green box in Fig. 3), could be installed in any organism possessing shikimate pathway (shown in a white box in Fig. 3) to give rise to engineered strain. Once scytonemin production is achieved, substrate supplementation and/or protein engineering approach can be used for maximizing its production.
Metabolic pathway for the biosynthesis of scytonemin. The white box shows the shikimate pathway which provides raw materials, i.e., l-tryptophan and l-tyrosine, for the biosynthesis of scytonemin. The green box shows scytonemin biosynthetic pathway which is exclusively present in only a few cyanobacteria. Shikimate pathway is present in bacteria, archaea, fungi, algae, cyanobacteria, some protozoans and plants. Genes coding for ScyA and ScyB proteins are required for connecting the two pathways. ScyC, ScyE, and Ebo FECBA proteins are required for the successful production of scytonemin by metabolically engineered strain. Substrates and enzymes shown in bold letters are essential and rate-limiting steps for the biosynthesis of scytonemin [15, 35, 38, 61] (colour figure online)
Extraction and Analysis of Scytonemin
Generally, after the initial screening with a spectrophotometer, high-performance liquid chromatography (HPLC) equipped with a reverse-phase column is utilized for the detection of characteristic UV wavelengths (252, 300 and 384 ± 2 nm). Thus, a particular scytonemin is identified using the distinctive nature of its absorption spectrum and retention time (RT) (Fig. 1b, c). However, the identification of scytonemins using HPLC becomes difficult for unknown compounds having similar RT and absorption maxima. Therefore, techniques such as ESI–MS (Fig. 4) and NMR are commonly utilized to examine the structural diversity in scytonemin. Scytonemin is usually extracted in methanol: ethyl acetate (1:1 v/v) by overnight incubation at 4 °C. Other organic solvents such as 100% acetone, 100% tetrahydrofuran, 100% ethyl acetate, and dimethylsulfoxide have also been used for the extraction of scytonemins [1, 46]. Preliminary spectroscopic analysis is carried out between 200 and 800 nm using a UV–Vis spectrophotometer to observe the characteristic peaks at 384, 300, 278, and 252 ± 2 nm. Further analysis of extract can be carried out using HPLC/TLC, ESI–MS, FTIR, MALDI-TOF MS, Raman spectroscopy, or NMR system [1, 10, 15, 63, 64]. Column chromatography on silica gel, ODS, or Sephadex can also be used to yield different forms of scytonemin [65]. In HPLC analysis, which is commonly used for purification, the flow rate of elution is usually maintained to 1.5 ml min−1 using the mobile phase of solvent A (ultrapure water) and solvent B (acetonitrile: methanol: tetrahydrofuran, 75:15:10 v/v). A 30-min gradient elution program is set with 0–15 min linear increase from 10% solvent A to 100% solvent B and 15–30 min at 100% solvent B. The detection wavelength is set at 380 nm, and the PDA scan wavelength ranges from 200 to 800 nm. The concentration of scytonemin can be determined using a trichromatic equation given by Garcia-Pichel and Castenholz [1]. It can also be quantified in terms of peak area of HPLC chromatograms of purified compound [57]. The commercial standards for various forms of scytonemins are unavailable. Therefore, the mass spectral information provides a measure to identify and characterize various forms of scytonemins based on their fragmentation pattern. Online UV/Vis spectra and mass spectra are available for the identification of UV-protective compounds such as scytonemin and mycosporine-like amino acids [64, 66].
Qualitative identification of different forms of scytonemins can be achieved by Raman spectroscopy, which is based on fundamental vibration modes that can be assigned to specific chemical functional groups. Recently, using Raman spectroscopy, a novel structure of scytonemin has been identified where two scytonemin molecules interact with iron (III) to form sandwich complexes [67]. Thus, Raman spectroscopy can be a useful technique for solving the structure of scytonemin molecules forming a complex with other ions. Therefore, this technique has the potential for the identification of past or present extraterrestrial life on other planets using scytonemin as one of the signature molecules [68]. Similarly, coherent anti-Stokes Raman scattering (CARS) microscopy has been recently used for successful identification and visualization of scytonemin in the sheath of the living cells of Nostoc commune [69]. The intracellular localization of scytonemin and its monomer can also be achieved by fluorescence confocal microscopy using excitation and emission wavelengths of 405 nm and 410 nm, respectively [61].
Application of Scytonemin
The photoprotective role of scytonemin has been studied in the number of cyanobacteria inhabiting various ecological niches [1, 7, 46]. Furthermore, the UV sunscreen role of scytonemin in the terrestrial cyanobacterium Chlorogloeopsis sp. has been well documented [7, 8]. Scytonemin is highly stable and performs its screening activity without further metabolic investment in extreme stressful physiological conditions such as desiccation and high temperature when other photo-repair mechanisms are not operational [46, 52]. The increasing interest of the scytonemin use for pharmaceutical applications might lead to a higher demand for this pigment [70]. Scytonemin shows fairly good radical scavenging activity and hence could be utilized as a natural antioxidant. Scytonemin extracted from N. commune showed quenching of an organic radical in vitro and account for up to 10% of the total activity of an ethanol extract of the cyanobacterium [10]. This demonstrates that scytonemin could be potentially utilized as UV sunscreen and antioxidant in the cosmetic industry. Furthermore, the hydrophobic property of the scytonemin may greatly increase its penetration into the skin where it can protect against UVR-related damages. The maximum absorption of scytonemin in the spectral range of UV-A radiation further strengthens its application as a UV-A sunscreen [8].
Scytonemin is equally important for the treatment of several hyperproliferative disorders, and furthermore, its structural properties such as lack of chirality, multiple dissection points, and phenolic groups easily allow its modification for the development of a new generation of a pharmacophore. The reduced form of scytonemin (R-scytonemin) can induce ROS production and mitochondrial dysfunction, which ultimately results in type II programmed cell death of human T-lymphoid Jurkat cells [65]. R-scytonemin also possesses the anti-tumour activity and checks the nitric oxide production induced by lipopolysaccharide and interferon-γ in murine macrophage RAW264 cells [51]. Scytonemin is known to alter the activity of cell cycle-related kinases such as serine/threonine kinase, protein kinase A, tyrosine kinase, Myt1, checkpoint kinase 1, cyclin-dependent kinase 1/cyclin B, and protein kinase Cbeta2 [13]. Collectively, these reports strongly support the application of scytonemin for the treatment of hyperproliferative and hyper-inflammatory disorders. Scytonemin A possesses calcium antagonistic properties, and therefore, it can also be used as calcium channel blocker [28]. Scytonemin is a potential candidate for the treatment of multiple myeloma as it inhibits the activity of polo-like kinase [14]. Thus, scytonemin is a valuable molecule and its production together with the biofuel production could substantially support economic viability by adding the value to biorefinery [16]. However, screening of cyanobacterial strains capable of producing scytonemin that could also act as a feedstock for biofuel production will be a challenge for the industry. Moreover, synthetic biology tools could be applied for developing novel strains of cyanobacteria combining desired traits of energy and valuable chemical production.
Conclusion and Future Research Directions
The research conducted over the past few decades has significantly contributed to the advancement of our knowledge regarding the biosynthesis, genetics, biochemistry, and importance of scytonemin. Therefore, future research should pave the way to commercialize this valuable compound for various applications. Scytonemin is known to be only produced by some but not all cyanobacterial species, and therefore, identification and development of novel strains are required for scytonemin production at large scale. Significant advancement has been made in the genetics and biochemistry of scytonemin biosynthesis. However, the signal transduction pathway and photoreceptor for UV-A dependent induction of scytonemin, or other signalling molecules or regulators involved in the biosynthesis of scytonemin, are still to be identified. Several studies mentioned above have shown the therapeutic importance of scytonemin. Therefore, these observations need to be strictly verified and taken to the next level for the successful development of scytonemin as a potential anti-cancerous drug. Scytonemin is also important as a biosignature molecule and can be useful in the exploration of life on other planets. Additionally, the question of why only a few cyanobacterial species produce scytonemin still needs to be answered. The revelation of the genetic basis of scytonemin biosynthesis could help in answering this important question.
References
Garcia-Pichel F, Castenholz RW (1991) Characterization and biological implications of scytonemin, a cyanobacterial sheath pigment. J Phycol 27:395–409
Pathak J, Sonker AS, Richa R et al (2017) Screening and partial purification of photoprotective pigment, scytonemin from cyanobacteria dominated biological crusts dwelling on the historical monuments in and around Varanasi, India. Microbiol Res (Pavia) 8:6559
Fleming ED, Castenholz RW (2007) Effects of periodic desiccation on the synthesis of the UV-screening compound, scytonemin, in cyanobacteria. Environ Microbiol 9:1448–1455
Fleming ED, Castenholz RW (2008) Effects of nitrogen source on the synthesis of the UV screening compound, scytonemin, in the cyanobacterium Nostoc punctiforme PCC 73102. FEMS Microbiol Ecol 63:301–308
Balskus EP, Case RJ, Walsh CT (2011) The biosynthesis of cyanobacterial sunscreen scytonemin in microbial mat communities. FEMS Microbiol Ecol 77:322–332
Büdel B, Karsten U, Garcia-Pichel F (1997) Ultraviolet-absorbing scytonemin and mycosporine-like amino acid derivatives in exposed, rock-inhabiting cyanobacterial lichens. Oecologia 112(2):165–172
Garcia-Pichel F, Sherry ND, Castenholz RW (1992) Evidence for an ultraviolet sunscreen role of the extracellular pigment scytonemin in the terrestrial cyanobacterium Chlorogloeopsis sp. Photochem Photobiol 56:17–23
Proteau PJ, Gerwick WH, Garcia-Pichel F et al (1993) The structure of scytonemin, an ultraviolet sunscreen pigment from the sheaths of cyanobacteria. Experientia 49:825–829
Simeonov A, Michaelian K (2017) Properties of cyanobacterial UV-absorbing pigments suggest their evolution was driven by optimizing photon dissipation rather than photoprotection. Biol Phys 1–38. arXiv:1702.03588 [physics.bio-ph]
Matsui K, Nazifi E, Hirai Y et al (2012) The cyanobacterial UV-absorbing pigment scytonemin displays radical scavenging activity. J Gen Appl Microbiol 58:137–144
Richter PR, Sinha RP, Häder D-P (2006) Scytonemin-rich epilithic cyanobacteria survive acetone treatment. Curr Trends Microbiol 2:13–19
Garcia-Pichel F, Lombard J, Soule T et al (2019) Timing the evolutionary advent of cyanobacteria and the later great oxidation event using gene phylogenies of a sunscreen. mBio 10(3):e00561-19
Stevenson CS, Capper EA, Roshak AK et al (2002) The identification and characterization of the marine natural product scytonemin as a novel antiproliferative pharmacophore. J Pharm Exp Ther 303:858–866
Zhang G, Zhang Z, Liu Z (2013) Scytonemin inhibits cell proliferation and arrests cell cycle through down regulating Plk1 activity in multiple myeloma cells. Tumor Biol 34:2241–2247
Malla S, Sommer MOA (2014) A sustainable route to produce the scytonemin precursor using Escherichia coli. Green Chem 16:3255–3265
Rajneesh R, Singh SP, Pathak J et al (2017) Cyanobacterial factories for the production of green energy and value-added products: an integrated approach for economic viability. Renew Sustain Energy Rev 69:578–595
Chandra R, Iqbal HM, Vishal G et al (2019) Algal biorefinery: a sustainable approach to valorize algal-based biomass towards multiple product recovery. Bioresour Technol 278:346–359
D’Agostino PM, Woodhouse JN, Liew HT et al (2019) Bioinformatic, phylogenetic and chemical analysis of the UV-absorbing compounds scytonemin and mycosporine-like amino acids from the microbial mat communities of Shark Bay, Australia. Environ Microbiol 21(2):702–715
Edwards HG, Sadooni F, Vítek P et al (2010) Raman spectroscopy of the Dukhan sabkha: identification of geological and biogeological molecules in an extreme environment. Philos Trans R Soc A Math Phys Eng Sci 368:3099–3107
Nägeli C (1849) Gattungen einzelliger algen, physiologisch und systematisch bearbeitet. Neue Denkschr. Allg Schweiz Ges Ges Naturwiss 10:1–138
Nägeli C, Schwenderer S (1877) Das Mikroskop, 2nd edn. Willhelm Engelmann Verlag, Leipzig, p 505
Balskus EP, Walsh CT (2010) The genetic and molecular basis for sunscreen biosynthesis in cyanobacteria. Science 329:1653–1656
Kim S, Thiessen PA, Bolton EE et al (2016) PubChem substance and compound databases. Nucleic Acids Res 44(D1):D1202–D1213
Bultel-Poncé V, Félix-Théodose F, Sarthou C et al (2004) New pigments from the terrestrial cyanobacterium Scytonema sp. collected on the Mitaraka Inselberg, French Guyana. J Nat Prod 67:678–681
Varnali T, Edwards HGM (2010) Ab initio calculations of scytonemin derivatives of relevance to extremophile characterization by Raman spectroscopy. Philos Trans R Soc A Math Phys Eng Sci 368:3099–3107
Grant CS, Louda JW (2013) Scytonemin-imine, a mahogany-colored UV/Vis sunscreen of cyanobacteria exposed to intense solar radiation. Org Geochem 65:29–36
Varnali T, Edwards HGM (2014) Reduced and oxidised scytonemin: theoretical protocol for Raman spectroscopic identification of potential key biomolecules for astrobiology. Spectrochim Acta A 117:72–77
Helms GL, Moore RE, Niemczura WP et al (1988) Scytonemin A, a novel calcium antagonist from a blue-green alga. J Org Chem 53:1298–1307
Lepot K, Compère P, Gérard E et al (2014) Organic and mineral imprints in fossil photosynthetic mats of an East Antarctic lake. Geobiology 12:424–450
Naurin S, Bennett J, Videau P (2016) The response regulator Npun_F1278 is essential for scytonemin biosynthesis in the cyanobacterium Nostoc punctiforme ATCC 29133. J Phycol 52:564–571
Sinclair C, Whitton BA (1977) Influence of nutrient deficiency on hair formation in the rivulariaceae. Br Phycol J 12:297–313
Rath J, Mandal S, Adhikary SP (2012) Salinity induced synthesis of UV-screening compound scytonemin in the cyanobacterium Lyngbya aestuarii. J Photochem Photobiol B Biol 115:5–8
Kokabi M, Yousefzadi M, Soltani M et al (2019) Effects of different UV radiation on photoprotective pigments and antioxidant activity of the hot‐spring cyanobacterium Leptolyngbya cf. fragilis. Phycol Res. https://doi.org/10.1111/pre.12374
Soule T, Shipe D, Lothamer J (2016) Extracellular polysaccharide production in a scytonemin-deficient mutant of Nostoc punctiforme under UV-A and oxidative stress. Curr Microbiol 73:455–462
Soule T, Stout V, Swingley WD et al (2007) Molecular genetics and genomic analysis of scytonemin biosynthesis in Nostoc punctiforme ATCC 29133. J Bacteriol 189:4465–4472
Soule T, Palmer K, Gao Q et al (2009) A comparative genomics approach to understanding the biosynthesis of the sunscreen scytonemin in cyanobacteria. BMC Genom 10:336–345
Soule T, Garcia-Pichel F, Stout V (2009) Gene expression patterns associated with the biosynthesis of the sunscreen scytonemin in Nostoc punctiforme ATCC 29133 in response to UV-A radiation. J Bacteriol 191:4639–4646
Balskus EP, Walsh CT (2008) Investigating the initial steps in the biosynthesis of cyanobacterial sunscreen scytonemin. J Am Chem Soc 130:15260–15261
Balskus EP, Walsh CT (2009) An enzymatic cyclopentyl[b]indole formation involved in scytonemin biosynthesis. J Am Chem Soc 131:14648–14649
Matsui D, Asano Y (2019) Creation of thermostable L-tryptophan dehydrogenase by protein engineering and its application for L-tryptophan quantification. Anal Biochem 579:57–63
Gao Q, Garcia-Pichel F (2011) Microbial ultraviolet sunscreens. Nat Rev Microbiol 9:791–802
Rastogi RP, Sonani RR, Madamwar D (2015) Cyanobacterial sunscreen scytonemin: role in photoprotection and biomedical research. Appl Biochem Biotechnol 176:1551–1563
Mushir M, Fatma T (2012) Monitoring stress responses in cyanobacterial scytonemin-screening and characterization. Environ Technol 33:153–157
Dillon JG, Castenholz RW (1999) Scytonemin, a cyanobacterial sheath pigment, protects against UV-C radiation: implications for early photosynthetic life. J Phycol 35:673–681
Javor B, Castenholz RW (1984) Productivity studies of microbial mats, Laguna Guerrero Negro, Mexico. In: Cohen Y, Castenholz RW, Halvorsan H (eds) Microbial mats: stromatolites. Alan Liss Inc., New York, pp 149–170
Sinha RP, Klisch M, Vaishampayan A et al (1999) Biochemical and spectroscopic characterization of the cyanobacterium Lyngbya sp. inhabiting Mango (Mangifera indica) trees: presence of an ultraviolet-absorbing pigment, scytonemin. Acta Protozool 38:291–298
Rastogi RP, Incharoensakdi A (2014) Characterization of UV screening compounds, mycosporine-like amino acids, and scytonemin in the cyanobacterium Lyngbya sp. CU2555. FEMS Microbiol Ecol 87:244–256
Muehlstein L, Castenholz RW (1983) Sheath pigment formation in a blue green alga, Lyngbya aestuari, as an adaptation to high light. Biol Bull 165:521–522
Dodds WK (1989) Photosynthesis of two morphologies of Nostoc parmelioides as related to current velocities and diffusion patterns. J Phycol 25:258–262
Chen J, Zhao L, Xu J et al (2013) Determination of oxidized scytonemin in Nostoc commune Vauch cultured on different conditions by high performance liquid chromatography coupled with triple quadrupole mass spectrometry. J Appl Phycol 25:1001–1007
Itoh T, Koketsu M, Yokota N et al (2014) Reduced scytonemin isolated from Nostoc commune suppresses LPS/IFNc-induced NO production in murine macrophage RAW264 cells by inducing hemeoxygenase-1 expression via the Nrf2/ARE pathway. Food Chem Toxicol 69:330–338
Ehling-Schulz M, Bilger W, Scherer S (1997) UV-B induced synthesis of photoprotective pigments and extracellular polysaccharides in the terrestrial cyanobacterium Nostoc commune. J Bacteriol 179:1940–1945
Dodds WK, Castenholz RW (1988) The nitrogen budget of an oligotrophic cold water pond. Arch Hydrobiol Suppl 79:343–362
Rastogi RP, Sinha RP, Incharoensakdi A (2013) Partial characterization, UV-induction and photoprotective function of sunscreen pigment, scytonemin from Rivularia sp. HKAR-4. Chemosphere 93:1874–1878
Asencio AD, Hoffmann L (2013) Chemosystematic evaluation of the genus Scytonema (Cyanobacteria) based on occurrence of phycobiliproteins, scytonemin, carotenoids and mycosporine-like amino acid compounds. Eur J Phycol 48:331–344
Rastogi RP, Kumari S, Richa, Han T et al (2012) Molecular characterization of hot spring cyanobacteria and evaluation of their photoprotective compounds. Can J Microbiol 58:719–727
Rastogi RP, Incharoensakdi A, Madamwar D (2014) Responses of a ricefield cyanobacterium Anabaena siamensis TISTR-8012 upon exposure to PAR and UV radiation. J Plant Physiol 171:1545–1553
Castenholz RW (1984) Composition of hot-spring microbial mats: a summary. In: Cohen Y, Castenholz RW, Halvorsan H (eds) Microbial mats: stromatolites. Alan R. Liss, Inc., New York, pp 101–119
Ferreira D, Garcia-Pichel F (2016) Mutational studies of putative biosynthetic genes for the cyanobacterial sunscreen scytonemin in Nostoc punctiforme ATCC 29133. Front Microbiol 7:735
Janssen J, Soule T (2016) Gene expression of a two-component regulatory system associated with sunscreen biosynthesis in the cyanobacterium Nostoc punctiforme ATCC 29133. FEMS Microbiol Lett 363:1–6
Klicki K, Ferreira D, Hamill D et al (2018) The widely conserved ebo cluster is involved in precursor transport to the periplasm during scytonemin synthesis in Nostoc punctiforme. mBio 9:e02266-18
Yurchenko T, Ševčíková T, Strnad H et al (2016) The plastid genome of some eustigmatophyte algae harbours a bacteria-derived six-gene cluster for biosynthesis of a novel secondary metabolite. Open Biol 6(11):160249
Edwards HGM, Garcia-Pichel F, Newton EM et al (2000) Vibrational Raman spectroscopic study of scytonemin, the UV-protective cyanobacterial pigment. Spectrochim Acta 56:193–200
Squier AH, Airs RL, Hodgson DA et al (2004) Atmospheric pressure chemical ionisation liquid chromatography/mass spectrometry of the ultraviolet screening pigment scytonemin: characteristic fragmentations. Rapid Commun Mass Spectrom 18:2934–2938
Itoh T, Tsuzuki R, Tanaka T et al (2013) Reduced scytonemin isolated from Nostoc commune induces autophagic cell death in human T-lymphoid cell line Jurkat cells. Food Chem Toxicol 60:76–82
Whitehead K, Hedges JI (2002) Analysis of mycosporine-like amino acids (MAAs) in aquatic plankton by liquid chromatography electrospray-ionization mass spectrometry. Mar Chem 80:27–39
Varnali T, Gören B (2018) Two distinct structures of the sandwich complex of scytonemin with iron and their relevance to astrobiology. Struct Chem 29:1565–1572
Edwards HGM, Hutchinson I, Ingley R (2012) The ExoMars Raman spectrometer and the identification of biogeological spectroscopic signatures using a flight-like prototype. Anal Bioanal Chem 404:1723–1731
Venckus P, Paliulis S, Kostkevičiene J et al (2018) CARS microscopy of scytonemin in cyanobacteria Nostoc commune. J Raman Spectrosc 49:1333–1338
Soule T, Garcia-Pichel F (2014) Ultraviolet photoprotective compounds from cyanobacteria in biomedical applications. In: Sharma NK, Rai AK, Stal LJ (eds) Cyanobacteria: an economic perspective. Wiley, Chichester, pp 119–143
Acknowledgements
J. Pathak (09/013/0515/2013-EMR-I) and A. Pandey (09/013/0619/2016-EMR-I) are thankful to the Council of Scientific and Industrial Research, New Delhi, India. Rajneesh is grateful to Department of Biotechnology, Govt. of India, (DBT-JRF/13/AL/143/2158), for the grant in the form of senior research fellowships. P.K. Maurya (3616/NET-DEC2014) is thankful to University Grant commission (UGC), New Delhi, India, for financial support in the form of JRF. S.P. Singh acknowledges the UGC for start-up Grant (F.30-370/2017; BSR) and DST-SERB for early career research Award (ECR/2016/000578).
Author information
Authors and Affiliations
Contributions
JP and SPS conceptualized the idea, did the literature survey, and wrote the paper; Rajneesh, AP, and PM developed the figures and tables; and RPS and SPS edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest to publish this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Significance Statement This review highlights the recent advancements made in the study of a cyanobacterial photoprotective compound called scytonemin. The biochemistry and genetics of scytonemin production have been discussed in detail. The roadmap for scytonemin production using metabolically engineered strains and rate-limiting steps for scytonemin biosynthesis have been presented. Also, it emphasizes the various application of scytonemin in different industries for human welfare.
Rights and permissions
About this article
Cite this article
Pathak, J., Pandey, A., Maurya, P.K. et al. Cyanobacterial Secondary Metabolite Scytonemin: A Potential Photoprotective and Pharmaceutical Compound. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 90, 467–481 (2020). https://doi.org/10.1007/s40011-019-01134-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-019-01134-5