Abstract
Titanium dioxide nanoparticles are widely used in consumer products, paints and pharmaceutical preparations. They have been shown to induce cytotoxicity, genotoxicity and carcinogenicity—in vitro and in vivo. So far there is lack of standardized protocol for in silico analysis of nano-toxicological evaluations. In the present study, it was attempted to analyze the titanium dioxide nanoparticles-protein interaction through docking using AutoDock 4.0.5 software. Titanium dioxide nanoparticles with particle size of 1.09 nm were docked with different cellular proteins. Binding site area and volume has been determined by using CastP online server and docking has been performed at the active site as a pocket. The docked structures were analyzed for the most efficient binding with amino acids. It is the first study to report the interaction of titanium dioxide nanoparticles without any surface modification with proteins using docking analysis. The negative binding and docking energy inferred that the interaction of titanium dioxide nanoparticles with certain proteins is significant. Titanium dioxide nanoparticle shows significant interaction with intercellular adhesion molecule-1, P38 mitogen-activated protein kinases (P-38), placental growth factor and nuclear factor kappa-light-chain-enhancer of activated B cell proteins. Further, it has been observed that titanium dioxide nanoparticles show frequent interaction with proline, lysine as well as leusine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Titanium dioxide nanoparticles (TNP) have a number of applications in which they can directly or indirectly gain access to human body. They have applications ranging from cosmetics to medicine, food and tissue engineering. The route of TNP is mainly through ingestion, dermal or inhalation. The toxic behavior of TNP, its mechanisms of action, its effect on different cell lines and protein is still unexplored. There is a need to develop in silico, in vitro and in vivo protocols for toxicological analysis of nanomaterials and to understand the mechanism of action [1–4].
TNP interactions with several cellular proteins have been studied earlier by in vitro method. Some of the important chemokines and other cytological proteins studied for interaction with TNP are intercellular adhesion molecule 1 (ICAM-1), C–C-motif chemokine ligand 20 (CCL-20), interleukin 8 (IL-8), nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB), P38 mitogen-activated protein kinases (P-38), placental growth factor (PlGF), C–X–C motif chemokine ligand 1 (CXCL1), C–X–C motif chemokine ligand 5 (CXCL-5), cluster of differentiation 35 (CD35) and matrix metallopeptidase 9 (MMP-9). The chemical composition, size, shape and surface characteristics of nanoparticles affect the way proteins bind to these particles, and this in turn influences the way in which nanoparticles interact with cells and tissues. Nanomaterials bound with proteins can result in physiological and pathological changes including protein aggregation and complement activation, but the mechanisms that lead to these changes remain poorly understood. Kruger et al. [5] have identified by quantitative PCR that TNP transiently induces the expression of ICAM-1, CCL-20, COX-2 and IL8 proteins. Further, using nuclear factor (NF)-κB reporter gene assays, they have shown that TNP-induced IL-8 mRNA expression occurs, in part, through activation of NF-κB and p38 mitogen-activated protein kinase pathways. Through the series of in vitro and in vivo analysis Chen et al. [6] concluded that TNP induces some of the chemokine expressions such as PlGF, CXCL1, CXCL3 and CXCL5 that may cause pulmonary emphysema and alveolar epithelial cell apoptosis. Additionally, flow cytometry analysis done by Babin et al. [7] concluded that TNP slightly down-regulated cell surface expression of the granule marker CD35 and the gelatin zymography studies suggests that TNP markedly increased the enzymatic activity of MMP-9.
To understand the TNP interaction with protein a protocol can be developed for in silico analysis for binding of nanoparticle (NP) and protein, which will give an insight to understand the mechanism of toxicological behavior of nanoparticle [8]. Due to the smaller size of the nanomaterial, dispersivity and diffusivity of the nanomaterials is higher in animal models [9, 10]. Additionally, after a number of researches in vitro and in vivo the mechanism of action of nanomaterial is not clear as it is dependent on the size variation, charge density variation, shape variation and other surface property variations [11]. Various reports are present for assessing toxicological evaluation of these NPs but yet a standardized protocol is not present. Moreover, very few reports are present for assessing the toxicity by computational method.
In the present study using computational analysis this has been attempted for the first time to understand the mechanism of TNP-protein interaction by using PDB file of NP in docking. In this work only those proteins have been selected for which interaction with TNP was well established in the earlier works using in vitro techniques and the in silico results have been discussed comparatively. Through this in silico analysis, the basic analysis for the effect of nanomaterial toxicity has been provided and an insight into the regularity of interaction has been attempted between nanomaterials with specific amino acids. It can be noted that, the cellular proteins for which interaction has been studied earlier have been shortlisted by using in vitro approach and the earlier studied findings have been validated through in silico method.
Material and Methods
PDB File of Proteins and Titanium Dioxide Nanoparticles
The PDB file of TNP was downloaded from the University of Sydney website [12]. Further, PDB file of proteins was obtained from RCSB protein data bank [13] of whose PDB id is given in Table 1.
Dimensions of Titanium Dioxide Nanoparticles
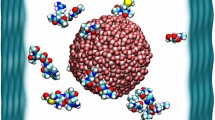
The dimension of the lattice was determined by using Pymol software (Ver 1.1, DeLano Scientific LLC, US) and the size of the nanoparticle was identified [14]. All the water molecules were removed using a plain text editor. The polar hydrogen, partial charge, and solvation parameters for use with the AutoDock 4.0.5 simulation were applied using AutoDock-Tools.
Determination of Binding Site
Computed Atlas of Surface Topography of Proteins (CASTp) [15–17] is used for visualization of the annotated functional residues, with emphasis on mapping to surface pockets and interior voids. CASTp gives a prediction of active site and the number of amino acids involved in it [18]. It has the ability to locate functionally important residues and to obtain a comprehensive understanding of the structural basis of protein function. The best binding pockets with high area and volume were predicted by CASTp as active sites for each protein. Depending on CASTp prediction, these binding pockets seem to be the potential active sites for further docking analysis. Thus, the present model could further be explored for in silico docking studies with TNP.
Docking Studies of Titanium Dioxide Nanoparticles with Proteins
AutoDock 4.0 docking program was applied to study the interaction between the TNP and the proteins. The PDB file id of the proteins has been listed in Table 1. The docking simulations were carried out with the rigid TNP and the proteins. Using AutoGrid tools, the grid maps were generated adequately large to include the binding site(s) of protein as depicted from CASTp. The points of the grids were thus with a grid spacing of 0.375 Å. The grid parameter file and the docking parameter file were set up by the AutoDock Tools program. The binding mode was predicted by using the Lamarckian genetic algorithm search engine implemented in AutoDock software package. Population size was 150, number of GA runs is 20 and maximum number of generation has been set as 27,000. Default settings were used for all other parameters. The interaction of amino acid of each protein with TNP has been analyzed by visualizing the docked structure (Fig. 1).
Results and Discussion
It is a novel approach to determine the nanoparticle–protein interaction by using docking without any surface modification of nanoparticle. The size of the TNP has been determined using pymol as 1.09 nm (Fig. 2). The best binding pockets with high area and volume were predicted by CASTp as active sites for each protein. The best binding pockets with area and volume are as under: ICAM (area-748.5; volume-1823.9), CCL-20 (area-632.4; volume-1120.4), IL8 (area-510.7; volume-588.7), NF-κB (area-212.7; volume-249), P-38 (area-866.8; volume-1377.8), PlGF(area-291.1; volume-321.4), CXCL-1 (area-293.5; volume-431.4), CXCL-5 (area-478.9; volume-441.9), CD-35 (area-213.8; volume-256.1), and MMP-9 (area-1164.1; volume-2603.4).
The negative binding energy and intermolecular energy of TNP-protein complexes show the efficient formation of protein-nanoparticle complexes. TiO2 makes most stable docked complex with ICAM as it has most binding energy and intermolecular energy as −1163 and −12.73 respectively. The TNP makes hydrogen bonds in the active pockets of protein (Fig. 1). It shows good binding energy and intermolecular energy with ICAM and P-38 as it has negative binding energy and intermolecular energy as −11.63 and 12.73; 11.73 and 12.83 respectively for each protein but does not show any stable complex with CXCL-1, CXCL-5 and CD-35 because of the positive binding and intermolecular energy (Table 1). All other proteins selected for this study showed good binding energy ranging from −11.63 to −4.04 which confirm the binding. The binding energy, intermolecular energy, torsion energy and the interacting amino acids have been tabulated in Table 1.
Docking studies suggests that TNP binds more efficiently with nonpolar R-group containing amino acids and positively charged R-group containing amino acids. Mainly, it shows more efficient interaction with proline and lysine. Further it also shows frequent bindings with polar uncharged R-groups and negatively charged R-group containing amino acids. Additionally, it shows least binding with aromatic R-group containing amino acids. TNP shows least frequent binding with alanine, serine, aspargine, phenylalanine and tryptophan (Table 2).
Negative binding energy and negative intermolecular energy indicates the more interaction of TNP and protein. Based on the binding energy obtained in this analysis, P-38, ICAM, PlGF, MMP-9, NF-κB, CCL-20 and IL-8 show interaction with TNP and the data supports the earlier findings [5–7]. Although, Chen et al. [6] have reported that TNP up regulates PlGF, CXCL-1 and CXCL-5 but the docking results thus obtained i.e. binding energy and intermolecular energy of TNP-protein complex for CXCL-1 and CXCL-5 contradict their findings, so, more in vitro investigation is needed to understand the TNP interaction with CXCL-1 and CXCL-5. Additionally, research is required to find out the reason as to why TNP activates CXCL-1 and CXCL-5 without having the stable complexes. This approach of in silico analysis for TNP-protein interaction supports the earlier findings of Frauke et al. [19] since they got binding energy in the range of −3.9 to −7.6 for the material science flexible ligands.
Docking results show that TNP binds more frequently with nonpolar aliphatic R-group amino acids and positively charged R-group containing amino acids of the proteins. It binds with proline and lysine more often than other amino acids. Interestingly, TNP does not bind with methinone; this suggests that the sulfur containing heavy chain R group prevents it to form stable hydrogen bond. Apart from the nonpolar aliphatic R-group and positively charged R-group amino acids; it binds with polar uncharged R-groups amino acids, negatively charged R-group and aromatic R-group containing amino acids but with less frequency. It shows least binding with alanine, serine, aspargine, phenylalanine and tyrosine. TNP shows least affinity with amino acids containing aromatic R-group which may be attributed to the fact that TNP cannot form a stable hydrogen bond because of the resonance energy stabilization of the amino acid. A detailed result for TNP binding with specific amino acids during the interaction has been tabulated (Table 2).
All the cellular proteins selected for the analysis has specific roles which can be affected when they come in contact with TNP. ICAM-1 also known as CD54 (Cluster of Differentiation 54), has important role in cell–cell signaling as stabilizing cell–cell interactions and facilitating leukocyte endothelial transmigration [20, 21]. ICAM-1 has antagonistic effect on the tight junctions forming the blood-testis barrier, thus playing a major role in spermatogenesis [22]. P-38 has important role in cell signaling against stress such as it is responsive to stress stimuli, such as cytokines, ultraviolet irradiation, heat shock, and osmotic shock, and is involved in cell differentiation, apoptosis and autophagy [23, 24]. Similarly, PIGF and NF-κB are having very important role against several cellular functions which have been described below. PlGF is a key molecule in angiogenesis and vasculogenesis, in particular during embryogenesis. It is a potent angiogenic factor involved in extravillous trophoblast proliferation, migration and invasiveness [25–27]. NF-κB is important in regulating cellular responses because it belongs to the category of “rapid-acting” primary transcription factor, i.e., transcription factor that is present in cells in an inactive state and do not require new protein synthesis in order to become activated [28, 29]. NF-κB is a major transcription factor that regulates genes responsible for both the innate and adaptive immune response [30–32]. Because of the efficient interaction of TNP with these proteins, it can be said that functioning of ICAM-1, P-38, PLGF and NF-κB might be altered. In short it can be said that TNP interaction with these proteins may affect stabilization of cell–cell interactions, leukocyte endothelial transmigration, formation the blood-testis barrier, cellular response to stress stimuli, anchorage of the proteins to the cell surface, trophoblast proliferation- migration- invasiveness, innate immune response and adaptive immune response.
Future Perspective
TNP-protein interaction highly depends upon the size [33, 34] so the PDB file need to be developed from ‘single crystal X-ray diffraction’ analysis of TNP so that TNP-protein can be analyzed with respect to size and mechanism. It can be noted that most of the inorganic nanoparticles are toxic in nature other than TNP mainly of gold [35] and silver [36] etc. for which the PDB-file should be developed to explore the in silico nanotoxicology. Apart from this, there is a need of PDB file for other NPs as well to have a detailed analysis of NP-protein interaction. Computational approaches like molecular docking could be adopted for screening of all the NPs, which can bind to the target with experimental or modeled structures. Further, molecular simulation studies could be the next approach for researchers to understand the TNP-protein interaction.
Conclusion
Docking studies have been done to analyze the interaction of TNP with different cellular proteins and cytokines. Further, efficiency and frequency of amino acid has been analyzed from the docked structure. It can be concluded that TNP shows maximum interaction with proline, lysine and leusine. Interestingly, TNP does not show any interaction with methionine for the docked structure of protein. Further, it can be concluded that the interaction of TNP is more with amino acid containing aliphatic R-group and amino acid with positively charged R-group. Additionally it shows least interaction with amino acid containing aromatic R-group which can be attributed by the fact that due to the resonance energy stabilization and the heavier R-group inhibiting the hydrogen bond formation between TNP and protein. Since, TNP binds most efficiently with ICAM-1, P-38, PlGF and NF-κB, so, it can be said that many cellular or non cellular responses like cell–cell interaction, spermatogenesis, blood-testis barrier formation and innate as well as adaptive immunity can be altered. To consider the future aspect of this work it can be stated that simulation is the serious threat for the in silico nanotoxicological analysis to understand the mechanism. Further, the PDB file for each NPs can be developed for different size so that the mechanism and size dependence of NP-protein interaction can be analyzed.
References
Shukla RK, Kumar A, Naga VSV, Kumar AP, Dhawan A (2014) Titanium dioxide nanoparticle-induced oxidative stress triggers DNA damage and hepatic injury in mice. Nanomedicine 9:1423–1434. doi:10.2217/nnm.13.100
Ranjan S, Dasgupta N, Chakraborty AR, Samuel SM, Ramalingam C, Kumar A, Shanker R (2014) Nanoscience and nanotechnologies in food industries: opportunities and research trends. J Nanopart Res 16(6):1–23. doi:10.1007/s11051-014-2464-5
Dasgupta N, Ranjan S, Deepa M, Chidambaram R, Shanker R, Kumar A (2015) Nanotechnology in agro-food: from field to plate. Food Res Int 69:381–400. doi:10.1016/j.foodres.2015.01.005
Kingsley DJ, Ranjan S, Dasgupta N, Saha P (2013) Nanotechnology for tissue engineering: need, techniques and applications. J Pharm Res 7(2):200–204. doi:10.1016/j.jopr.2013.02.021
Kruger K, Cossais F, Horst N, Martin K (2014) Titanium dioxide nanoparticles activate IL8-related inflammatory pathways in human colonic epithelial Caco-2 cells. J Nanopart Res 16(5):1–12. doi:10.1007/s11051-014-2402-6
Chen HW, Su SF, Chien CT, Lin WH, Yu SL, Chou CC, Chen JJ, Yang PC (2006) Titanium dioxide nanoparticles induce emphysema-like lung injury in mice. FASEB J 20(13):2393–2395
Babin K, Francis A, David MG, Denis G (2013) TiO2, CeO2 and ZnO nanoparticles and modulation of the degranulation process in human neutrophils. Toxicol Lett 221:57–63
O’Shaughnessy PT (2013) Occupational health risk to nanoparticulate exposure. Environ Sci: Process Impacts 15:49–62
Feliu N, Fadeel B (2010) Nanotoxicology: no small matter. Nanoscale 2:2514–2520
Zuzana M, Andrew C, Ashutosh K, Alok D, Vicki S, Maria D (2014) Mechanisms of genotoxicity. A review of in vitro and in vivo studies with engineered nanoparticles. Nanotoxicology 8(3):233–278. doi:10.3109/17435390.2013.773464
Kumar A, Alok D, Shanker R (2011) The need for novel approaches in ecotoxicity of engineered nanomaterials. J Biomed Nanotechnol 7:79–80
University of Sydney (2014) TiO2.PDB 19-May-2007. Weblink: http://firstyear.chem.usyd.edu.au/jmol/ (Retrieved on: 10th April 2014)
RCSB PDB (2014) Weblink: http://www.rcsb.org/pdb/home/home.do (Retrieved on: 25th April, 2014)
TPMGS (2014) ThePyMOL Molecular Graphics System, Version 1.5.0.4 Schrödinger, LLC
Dundas J, Zheng O, Jeffery T, Andrew B, Yaron T, Jie L (2006) CASTp: computed atas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucl Acids Res 34:W116–W118
Binkowski TA, Naghibzadeh S, Liang J (2003) CASTp: computed atlas of surface topography of proteins. Nucleic Acids Res 3:3352–3355
Liang J, Edelsbrunner H, Woodward C (1998) Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci 7:1884–1897
Singh S, Kumar A, Patel A, Tripathi A, Kumar D et al (2010) In silico 3D structure prediction and comparison of nucleocapsid protein of H1N1. J Model Simul Syst 1:108–111
Frauke G, Sonja MS, Annick D, Stefan F, Jeremy CS (2005) Protein/ligand binding free energies calculated with quantum mechanics/molecular mechanics. J Phys Chem B 109:10474–10483
Abraham G, Colonno RJ (1984) Many rhinovirus serotypes share the same cellular receptor. J Virol 51(2):340–345
Rui B, Shaoqiong Y, Xuejie Z, Huiliang L, Xiaohong F (2014) Role of ICAM-1 polymorphisms (G241R, K469E) in mediating its single-molecule binding ability: atomic force microscopy measurements on living cells. Biochem Biophys Res Commun 448(4):372–378
Xiang X, Dolores DM, Yan CC (2013) Intercellular adhesion molecules (ICAMs) and spermatogenesis. Hum Reprod Update 19(2):167–186
Sabzali J, Sehwan J, Bryan A (2013) Crosstalk between mitogen-activated protein kinases and mitochondria in cardiac diseases: therapeutic perspectives. Pharmacol Ther 144(2):202–225. doi:10.1016/j.pharmthera.2014.05.013
Yu-Ting K, Wei-Chi H, Huei-Ting H, Shih-Hsien H, Chang-Shen L, Chien-Chih C, Chi-Yu L, Tzyh-Chyuan H, Yeong-Shiau P, A-Mei H (2014) Involvement of p38 mitogen-activated protein kinase in acquired gemcitabine-resistant human urothelial carcinoma sublines. Kaohsiung J Med Sci 30(7):323–330
Athanassiades A, Lala PK (1998) Role of placenta growth factor (PLGF) in human extravilloustrophoblast proliferation, migration and invasiveness. Placenta 19(7):465–473
Khaliq A, Li X-F, Dunk D, Shams M, Whittle MJ, Ahmed A (1996) Placenta growth factor (PLGF) expression and function in human placenta. Placenta 17(5–6):A43
Lorenzo GD, Ceccarello M, Cecotti V, Ronfani L, Monasta L, Vecchi BL, Montico M, D’Ottavio G (2012) First trimester maternal serum PLGF, free β-hCG, PAPP-A, PP-13, uterine artery Doppler and maternal history for the prediction of preeclampsia. Placenta 33(6):495–501
Susana HM, Carla R, Luísa C, Helena S, Fernando A (2014) Hydrogen peroxide sensing, signaling and regulation of transcription factors. Redox Biol 2:535–562
Silvia P, Karolina K, Dušan V, Luboš R, Abdul HH, Saleh HA, Alexander VS (2013) The involvement of SIRT1 and transcription factor NF-κB (p50/p65) in regulation of porcine ovarian cell function. Anim Reprod Sci 14(3–4):180–188
Hayden MS, West AP, Ghosh S (2006) NF-κB and the immune response. Oncogene 25:6758–6780
Jin J, Hongbo H, Haiyan SL, Jiayi Y, Yichuan X, George CB, Qiang Z, Xuhong C, Frédérick AM, Stephanie SW, Shao-Cong S (2014) Noncanonical NF-κB pathway controls the production of type I interferons in antiviral innate immunity. Immunity 40(3):342–354
Yu L, Hongtao L, Qing-Song X, Yu-Guang D, Jian X (2014) Chitosan oligosaccharides block LPS-induced O-GlcNAcylation of NF-κB and endothelial inflammatory response. Carbohydr Polym 99:568–578
Kumar A, Dhawan A (2013) Genotoxic and carcinogenic potential of engineered nanoparticles: an update. Arch Toxicol 87(11):1883–1900
Kumar A, Sharma V, Dhawan A (2013) Methods for detection of oxidative stress and genotoxicity of engineered nanoparticles. Methods Mol Biol 1028:231–246
Sireesh BD, Badal KM, Ranjan S, Dasgupta N (2015) Diastase assisted green synthesis of size-controllable gold nanoparticles. RSC Adv 5(34):26727–26733. doi:10.1039/C5RA03117F
Dasgupta N, Ranjan S, Bhavapriya R, Venkatraman M, Chidambaram R, Ganesh SA, Kumar A (2015) Thermal co-reduction approach to vary size of silver nanoparticle: its microbial and cellular toxicology. Env Sci Pollut Res. doi:10.1007/s11356-015-4570-z
Acknowledgments
The authors acknowledge VIT University, Vellore, India for providing Research Associateship and RGEMS-VC-Fund to carry out the research for nano-food research group, Veer Kunwar Singh Memorial Trust, Chapra, Bihar, India for partial funding with the Grant Number VKSMT/SN/NFNA/SN/011 and Department of Biotechnology (DBT, India) for the project under consideration with permanent Project Number—BT/PR10414/PFN/20/961/2014.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Shivendu Ranjan and Nandita Dasgupta have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Ranjan, S., Dasgupta, N., Chinnappan, S. et al. A Novel Approach to Evaluate Titanium Dioxide Nanoparticle–Protein Interaction Through Docking: An Insight into Mechanism of Action. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 87, 937–943 (2017). https://doi.org/10.1007/s40011-015-0673-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-015-0673-z