Abstract
Continuous excessive irrigation and industrial activities increase the salinity in the environment which poses a threat to the current technology used in the Methyl Red degradation process. Hence it is quite essential to use a microbe which can be used in very high saline conditions. The present study focuses on the kinetic and isotherm studies on the novel bacterium Lactiplantibacillus plantarum SS-AU1 for Methyl Red bio-degradation under high salinity.he degradation was assessed based on the UV-Spectrophotometer studies. Kinetics and Isotherm studies were also performed to find the mechanism involved. The findings were also justified by Scanning Electron Microscope images and Fourier Transformed Infrared spectra.The strain can degrade up to 400 mg L−1 dye concentration. Kinetic studies revealed chemisorption involvement in the process where maximum adsorption was observed at 2500 mg g−1 and the formation of multilayer on the cell surface was confirmed. It followed the pseudo-second-order at low dye concentration while the pseudo-first-order at high dye concentration. The high degradation efficiency, having Km = 0.092 mMol−1 L and Vm = 0.761 mmol min−1, can be attributed to intra-particle diffusion predicted by the Weber-Morris and Elovich-Model. The weak bonding (E = 28 J/mol) is a key-finding behind the efficient degradation. Infrared spectroscopy study revealed the breakdown of the azo-bond. In addition, a Biodegradation mechanism has been suggested.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
A plethora of synthetic dyes (e.g. direct dyes, reactive dyes etc.) derived from organic compounds are available nowadays, widely employed for more repeatable and consistent dyeing applications in both industries (e.g., textile, paint, tannery, leather, paper, cosmetics, food and pharmaceutical industries etc.) and scientific laboratories (e.g., BODIPYFL (Boron-Dyepyrromethene), fluoresceins, oxazines, phenothiazines etc.) (Bhattacharya et al. 2017; Vinoda et al. 2015). However, the production of inexpensive dyes with long-lasting coloring effects comes at the expense of extremely robust organic structures and inert characteristics towards degradation (Singh 2022). Moreover, degradation products of various dyes have been reported to be recalcitrant, cytotoxic and carcinogenic for living beings (Chen et al. 2009; Mahajan and Kaushal 2020). Approximately 20–30 various dye groups may be characterized based on the chemical structure or chromophore responsible for the dye hue, such as azo, anthraquinone, methine, nitro, aryl methane, carbonyl, and others. Among the most popular synthetic dyes, azo dyes which have the azo (R − N = N − R′) functional group, represent the largest and most versatile class in the manufacturing industry (Cui et al. 2014; Mawad et al. 2020). Among azo dyes, methyl red (MR) is mutagenic and its persistence have major environmental and human health consequences, as reported by many researchers (Gupte et al. 2013; Sharma et al. 2003, 2009). Several physicochemical approaches have been investigated to treat dye-contaminated wastewater; however, the methodologies have substantial disadvantages including high operational cost, low efficiency, secondary contamination, and inapplicability to a wide range of dyes (Singh 2022). Furthermore, the presence of significant levels of salts (up to 100 g L−1 NaCl) in dye-containing wastewater also hampers physicochemical treatment (Lellis et al. 2019). High salts are used in the fixation process of dying which as a result increases the salt concentration in the effluent (Montañez-Barragán et al. 2020; Tian et al. 2021). These issues, along with severe environmental regulations, have sparked interest in an ecologically sound and cost-effective option for dye degradation. Conversely, microbial degradation of azo dyes seems to be a promising strategy and almost no harm to the environment. These tiny microbial isolates possess inherent metabolic machinery to mineralize and decolorize the dye completely (Oliveira et al. 2023; Tripathi et al. 2016). There are several researches available which have used the potential of biological components to reduce the MR. In these studies, microorganisms such as Pseudomonas aeruginosa, Zhihengliuella sp. ISTPL4, Aspergillus sp. etc. have been used to degrade the azo dye (Abd El-Rahim et al. 2021; Ikram et al. 2022; Takkar et al. 2022). Others have used the capabilities of the Phyto-based approaches by using Trachyspermum ammi, Sargassum latifolium, Chara vulgaris to remove the azo dyes efficiently (Mahajan and Kaushal 2020; Mansour et al. 2022; Kaur et al. 2023). The textile industries are the biggest azo dye consumers, releasing tonnes of textile effluent every day. The effluents are characterized by high biological oxygen demand (80–6000 mg L−1 BOD), high chemical oxygen demand (150–12,000 mg L−1 COD), high salinity (> 10 g L−1 NaCl), high color (50–2500 Pt/Co units), and alkaline pH (7–10), leads to detrimental effects on the environment (Takkar et al. 2022). Due to the high solubility of Azo dyes in water, excess dyes used in processing can accumulate on lands and in water reservoirs, making removal complicated through conventional methods (Kgatle et al. 2021). Finding a salt-tolerant and effective bio-treatment system for dye degradation in high-saline wastewater is a challenge. With this objective in mind, a potential salt-tolerant bacterium (Lactiplantibacillus plantarum strain SS-AU1) was isolated from the River Ganges and examined as a potential decolourizer of methyl red. The sampling point was situated close to an effluent dumping site in Prayagraj, Uttar Pradesh, India. The Ganges River is one of India's major rivers, providing not only potable water to 40% of India's population across eleven states but also a source of livelihood for the people who live along its banks. Lactiplantibacillus species are generally present in the gastrointestinal (GI) tract, Fermented food, water reservoirs etc. hence they are called ‘nomadic’ microbes (Jeong et al. 2021; Yu et al. 2021). These species have been involved in a variety of novel applications such as probiotics production and antibiotic production and it is a microbe which can withstand very high salt concentration (Bu et al. 2022; Echegaray et al. 2023; Rocchetti et al. 2021). Moreover, these bacteria are non-pathogenic and non-harmful to the environment as well as it can withstand multiple heavy metals toxicity which is the major reason behind using them in the treatment of industrial effluent in this study (Mostafidi et al. 2023; Sundaramanickam et al. 2023; Yilmaz et al. 2022).
To the best of our knowledge, this research is the first attempt, to use the river Ganges’ self-cleaning capacity to deal with high salinity effluents. Additionally, this may also be the first study of the Lactiplantibacillus species for methyl red degradation.
Materials and methods
Chemicals and media
For the bacterial culturing and other degradation processes, Luria–Bertani media (LB) (Composition: Tryptone 10 g/L; NaCl 10 g/L; Yeast Extract 5 g/L; Agar 15 g/L) was purchased from Himedia. Methyl red, DMSO, Acetonitrile, Ethanol, NaCl, KOH, Glycerol, Indole Reagent, Hydrogen Peroxide, Urea, Simmons Citrate Agar were purchased from Himedia. OsO4, Glutaraldehyde, HCl, Nutrient Agar, Glucose, Gelatin, Lactose, Sucrose, Mannitol, Pectin, Crystal Violet, Iodin solution, Safranin, K2HPO4, KH2PO4, MgSO4, ZnCl2, CaCl2 Acetone were purchased from Himedia. All the chemicals were analytical grade. A stock solution of MR (5000 mg L−1) was prepared by dissolving 0.5 g MR in 100 mL of absolute ethanol and stored in a 250 mL amber glass bottle.
Isolation, characterization, and maintenance dye degrading microbes
The bacteria used in the present study were isolated from the river Ganges water, Prayagraj, U.P., India. Initially, five bacterial isolates with methyl red decolorizing activity were isolated and purified by repeated sub-culturing on nutritional agar (NA) (Composition: Peptone 5 g/L; NaCl 5 g/L; Yeast Extract 1.5 g/L; Agar 15 g/L) plates, and labelled S-1, S-2, S-3, S-4, and S-5. The isolates were kept at 4 °C and subcultured weekly on new nutrient agar slants. The pure culture was also stored at − 20 °C in stocks of 20% (v/v) glycerol. Based on their MR decolorizing capacity in saline conditions, isolates were screened and S-1 was selected for further experimentation. The S1 isolate was grown in a 250 mL Erlenmeyer flask containing 50 mL of LB with salt (100 g L−1) and incubated on a rotary shaker at 180 rpm at 35 °C temperature to reactivate the cell growth for inoculum preparation.
Morphological characterization
The colony morphological characteristics of the isolate were examined microscopically as prescribed by Cheesbrough (1981) and Gram staining method (Cheesbrough 1981; Cappucino and Sherman 2005) using a trinocular microscope at 100× (Olympus-CX33RTFS2).
Biochemical characterization
The freshly-prepared culture was inoculated in Mineral Salt Media (MSM) (Composition: K2HPO4 1.73 g/L, KH2PO4 0.68 g/L, MgSO4.7H2O 0.1 g/L, NaCl 100 g/L, FeSO4.7H2O 0.03 g/L CaCl2.2 H2O 0.02 g/L, NH4NO3 1 g/L) containing different types of sugar sources such as Glucose, Lactose, Sucrose, Manitol, Pectin and Gelatin for sugar utilization and fermentation tests. After the proper growth at 37.4 ℃ in the MSM, 2–3 drops of MR were added to the solution where color change from yellow to red indicates the fermentation positive results. The culture was also evaluated for other biochemical activities such as urease, citrate, indole and catalase using standard protocols (Sahoo and Prelot 2020).
Genomic characterization
The DNA of the bacterial isolate was extracted using a standard protocol and subjected to molecular characterization by 16S rRNA sequencing. The 16S rRNA gene PCR amplifications were performed with initial denaturation at 94 °C for 2 min, 35 amplification cycles at 94 °C for 45 s, 55 °C for 60 s, 72 °C for 60 s, and a final extension at 72 °C for 10 min using forward (5′-GGATGAGCCCGCGGCCTA-3′) and reverse (5′-CGGTGTGTACAAGGCCCGG-3′) universal primers. PCR products were excised from a 1% agarose gel after electrophoresis and purified by using a QIA quick gel extraction kit (Qiagen) and were sequenced directly with a Biotech Diagnostics Big-Dye sequencing kit on an ABI 377 sequencer (Applied Biosystems, Foster City, Calif.). Isolates were assigned a putative taxonomy based on 16S rDNA sequence analysis using the NCBI 16S BLAST database.
Dye degradation studies
Bacterial cells were grown as shake cultures at 37.4 °C in 100 mL of LB medium supplemented with methyl red (100 mg L−1) and sodium chloride (NaCl, 100 g L−1) at pH 7.1 in 250 mL Erlenmeyer flasks at 100 r min−1. The uninoculated media was used as control. 3 mL of test samples were collected and centrifuged (10,000 rpm for 15 min) for analysis. The disappearance of color was recorded at 491.8 nm by UV–VIS spectrophotometer. The supernatant was taken for the spectrophotometer examination against the control.
The percentage of degradation was calculated by the following formula (Kang et al. 2018);
where % D is the extent of percentage degradation, Ai & Af are initial absorbance and final absorbance respectively.
Optimization of dye degradation conditions
The Correlation of the kinetic properties with dye concentration and other rate-dependent environmental parameters (temperature, pH, and dye concentration) was characterized. The impact of different factors on the degradation capacity of the S1 was studied. The different factors were pH (5 to 11), temperature (20 to 50 °C), salinity (0.5–200 g/L NaCl), inoculum size (1 to 10%), Dye concentration (10 to 60%) and growth condition (stationery and shaking). Growth was monitored by measuring the turbidity of the culture at A600. The reusability efficiency was also investigated, with a new batch of dye supplied to the media after decolorization until the seventh cycle. The corresponding second-order polynomial equation, which the degradation process was following, was fitted to the data by Microsoft Excel software with a significance R2 value. All the equations were in the form of,
where y is the degradation and x are independent variables and denote different parameters. δ represents the error, a represents the quadratic coefficient, b represents the linear coefficient and c represents the intercepts.
Fourier transform infrared spectroscopy and scanning electron microscopy analysis
For identification of change in the functional groups of MR metabolites, FTIR spectrum (4000–400 cm−1) of cell-free aqueous samples was performed before and after biodegradation of 600 mg L−1. The tested samples were F-1 (untreated MR), F-2 (red to colourless), and F-3 (red to black). Scanning Electron Microscopy (SEM) characterization was carried out further to investigate the surface alteration of the bacterium cell wall (prescribed by Hartmann et al. 2010) using SEM-JOEL(JXA-8100) (Li et al. 2018). The samples were included; control ABS-1 (without MR, 5 g/L NaCl concentration), control ABS-2 (without MR, 100 g/L NaCl concentration) & ABS-3 (MR degradation, 100 g/L NaCl).
Degradation kinetics studies of Lactiplantibacillus paraplantarum
For the study of degradation kinetics, freshly prepared 2% inoculum was added in 100 g L−1 NaCl, pH 7, LB media. After every 30 min interval, samples were taken out and centrifuged at 10,000 RPM for 15 min. The supernatant was then characterized by the spectrophotometer at 491.8 nm, and the percentage degradation was calculated by Eq. (1). The percentage of degradation vs. time gives the degradation kinetic curve from where the rate constant can be calculated.
Adsorption isotherm studies
Adsorption efficiency was calculated at 10% MR concentration, 2% inoculum size, 100 g.L−1 NaCl concentration, 37.4 °C, 180 rpm and pH 7. The samples were taken out at regular intervals of 30 min. Samples were centrifuged at 10,000 rpm for 15 min and the supernatant was used to calculate the difference in the concentration of MR using Eq. (1). The pellet was dried and weighed and the adsorption efficiency was calculated by the following Equation (Piccin et al. 2011).
where qe is the adsorption coefficient, Ci & Cf are initial and final concentration respectively, V is volume of sample and W dried biomass.
Langmuir isotherm and Freundlich isotherm were checked for the adsorption efficiency process (Takkar et al. 2022).
The equation of Langmuir isotherm is as follows;
where qeq is adsorption efficiency at time equilibrium, qm is maximum adsorption efficiency which can be calculated from the intercept of the graph. Ceq is the concentration of MR at equilibrium and b (L/mg) is the Langmuir constant, which can be calculated from the slope of the graph. It represents the affinity of sorbate with sorbent where a higher value of b indicates higher affinity.
The adsorption characteristics of the Langmuir model can be understood by a dimensionless constant RL,
The values of RL show whether the adsorption is favorable or not. If 0 < RL < 1, the reaction is favorable, RL > 1, the reaction is unfavorable, RL = 0, the reaction is irreversible and RL = 1, the reaction is linear. This model is applicable for monolayer adsorption (Sahoo and Prelot 2020).
The equation of Freundlich Isotherm for multilayer is as follows:
where n and Kf denote Freundlich constants. ‘n’ denotes the intensity factor for adsorption and Kf represents adsorbed ions concentration at equilibrium (Takkar et al. 2022). The 1/n value gives the characteristics of the adsorption process. If 1/n < 1, ordinary Langmuir or chemisorption is followed and if 1/n > 1, cooper adsorption is followed (Sahoo and Prelot 2020).
Redlich-Peterson Isotherm (Eq. 6) and SIPS Isotherm (Eq.7) are the composites of Langmuir and Freundlich isotherm. They are designed to overcome the limitations of these isotherms. Freundlich Isotherm cannot calculate maximum adsorption coefficient, which is being solved by the SIPS isotherm, which has similar assumptions to Freundlich, but maximum adsorption coefficient can also be calculated from Kumara et al. (2014). The equations are as follows:
where Ce and qe are equilibrium dye concentration and equilibrium adsorption coefficient, respectively. Β (exponent lies between 0 and 1) and A (L mg−1) are R–P constants. Whereas Ks (L mg−1) is SIPS equilibrium constant (Kumara et al. 2014; Ayawei et al. 2017).
The Dubinin–Radushkevich (D–R) isotherm postulates a Gaussian-type distribution of pores on the multilayer surface. From this isotherm, the mean binding energy of adsorbate with adsorbent can be calculated (Ayawei et al. 2017).
where \(\varepsilon =RTln\left(1+\frac{1}{{C}_{e}}\right)\)
qDR(mg/g)is adsorbed dye in the pores of the formed layer, (mol2/KJ2) is constant of adsorption energy and ε (J/g) is adsorbent Polanyi potential in equation (8). R is gas constant (8.314 J/K mol) and T is Kelvin temperature of the system. The mean adsorption energy E (KJ/mol) can be calculated by using following formula [37. 38, 39]:
Harkin–Jura (Eq. 9) & Halsey Isotherm (Eq. 10) express the possibilities of the heterogeneous pore distribution on the surface of the adsorbent (Ayawei et al. 2017; Amin et al. 2015).
Bhj & Ahj are Harkin–Jura constants whereas nH& KH are Halsey constants in Eq. (9) and Eq. (10) respectively (Ayawei et al. 2017; Amin et al. 2015).
Assumption of Temkin isotherm is that the heat of adsorption of a system decreases linearly rather than logarithmically with increasing coverage of the system. The heat of the system is directly related to the binding energy of adsorbate with adsorbent. The graph can be obtained by plotting qe versus ln Ce. The equation is as follows,
where \({B}_{T}=\frac{RT}{{b}_{T}}\)
R is gas constant, T is system kelvin temperature, KT equilibrium binding constant related to the maximum binding energy and bT is adsorption heat.
Jovanovic isotherm postulates about the mechanical contact between adsorbate and adsorbent (Ayawei et al. 2017).
qm is Jovanovic maximum adsorption which is in mechanical contact between adsorbate and adsorbent. Kj (L/g) is Jovanovic constant related to the fraction of adsorbates involved in mechanical adsorption.
Elovich isotherm postulates that the adsorption sites exponentially increase with increasing adsorption. The graph can be obtained by plotting ln (qe/Ce) vs qe (Ayawei et al. 2017). The equation is as follows,
where qm is maximum adsorption capacity and Ke is related to the adsorption of adsorbate on the adsorbent.
Adsorption kinetics studies
The Lineweaver–Burk plot was plotted to investigate the possible enzymes involved in the process. The equation is as follows
where V is reaction velocity, [S] is dye concentration, Km is Michaelis constant and Vm is maximum reaction velocity.
To check the kinetics of adsorption of MR with time, Lagergren models were explored. There are two types of Lagergren model: pseudo-first-order model and pseudo-second-order model. The Lagergren pseudo-first-order model (PFO) is as follows:
where k1 is the pseudo-first order rate constant and can be calculated from the slope of log (qeq–qt) vs time plot.
The pseudo-second-order model (PSO) is as follows:
where h0 is the second order rate constant and can be calculated by the following equation (Feng et al. 2012).
h0 = k2qe2
Weber and Morris model (IP), also known as intra-particle mass diffusion model, was also explored for the adsorption kinetics. The equation of Weber and Morris model is as follows (Cappucino and Sherman 2005):
where ki is inter-particle diffusion rate constant and Ci is thickness of the boundary layer and can be calculated from qt versus t(1/2) plot.
The adsorption & desorption rate of the MR was calculated by the Elovich kinetic model whose equation is as follows (Ayawei et al. 2017).
where α is the adsorption rate and β is the desorption rate constant and they can be calculated by plotting qt versus ln t graph.
Statistical validation
All tests were carried out in triplicate. Along with their determinations, experimental errors were illustrated and assessed using standard error (error bars). The data from the earlier experiments were collected and subjected to an analysis of variance ANOVA test with a 95% confidence interval on SPSS software for statistical analysis.
Results and discussion
Morphological, biochemical and molecular identification
The morphological characteristics, colony characteristics, and other biochemical features of the isolate S-1 were summarized in Supplementary Table S1 and shown in Figure 1. According to microscopic investigations, S-1cells were slightly curved, rods and occur singly, in pairs, and sometimes in short chains. However, under stressful conditions, active S-1 cells can produce dormant spherical endospores that might be located centrally or terminally (Fig 1a). These growing endospores may aid in the S-1’s ability to survive in low salt conditions to sustain for extended periods of time. Similar findings were reported by many authors. Sari et al. (2019) reported a round-shaped structure at the terminal region of heat-killed cells of Lysinibacillus fusiformis W1B6 (Tripathi et al. 2016). Li et al. (2018) have also reported a somewhat similar structure in the endospore formation by rod-shaped bacteria (Li et al. 2018). Hence the endospore-like modification in the S-1 morphology indicates the stress condition due to the low salt level in the media (0.5% NaCl) which will be justified further in this study by the growth studies on various salt concentrations. 10% NaCl concentration which is optimum for the S-1, shows a different type of structure as compared to 0.5% NaCl. Majorly, it is almost similar to the conventional rod-shaped structure with a slight curve (Fig 1b). Moreover, the cells are single in 0.5 % NaCl as compared to chains in 10% NaCl. The major change in the morphology post-MR degradation is a longer chain with a slightly uneven shape as compared to even shaped structure in pre-MR degradation (Fig. 1c–e), as also reported by Takkar et al. (2022) (Takkar et al. 2022). A number of carbohydrate utilization assays were also carried out with Glucose, Gelatin, Lactose, Sucrose, Mannitol, and Pectin. Results showed that S-1 was able to utilize all carbon sources supplied, except Gelatin (see Supplementary Table S1). It has been proven that S-1 followed the fermentation route and verified methyl red test. Yu et al. (2021) showed that Lactiplantibacillus sp. can efficiently thrive on several carbon sources such as Glucose, Maltose, Sucrose, Galactose and Fructose while Diao et al. (2022) also reported that Mannitol was utilized by the same species (Yu et al. 2021; Diao et al. 2022) (Table 1).
Other biochemical assays showed that S-1 is a Gram-positive non-motile bacterium with catalase, urease, and citrase activity, summarized in Supplementary Table 2. Saleh et al. (2022) have reported similar observations except for the urease test, which is negative in their results (Saleh et al. 2022). According to colony morphology, physiology, and biochemical characterization, those isolates were presumptively identified as Bacillus. The S-1 isolate seems to be alkaliphilic and moderately halo-tolerant, growing optimally at a pH range of 7–11 and NaCl concentration range of 1–16 %.
The 16S rRNA gene sequences were submitted via ‘BANKIT’ option of Gen Bank on NCBI webpage to get Gen Bank accession number(s) for nucleotide sequence(s) (OQ459850). The novel strain was subjected to genomic relatedness analysis to confirm membership to the genus from a public database. Phylogenetic studies of S-1, based on core 16S r RNA genome phylogeny confirmed its similarity > 95.5% with the Lactobacillaceae family.
Effect of physico-chemical parameters on dye decolourization
The optimization of physico-chemical conditions of MR degradation by the novel Lactiplantibacillus plantarumSS-AU1 isolate was carried out on different salinity, pH, initial dye concentration, inoculum size, and temperature under aerobic shaking conditions (180 rpm).
Effect of inoculum size on the rate of degradation
The effect of inoculum size was investigated while maintaining pH, salinity, temperature and dye concentration constant at 7.0, 100 g L−1, 35 °C and 100 mg L−1, respectively. Results showed that the dye removal capacity was affected significantly by the inoculum size used (Fig. 2).
The degradation rate increased with the increasing inoculum sizes from 1 to 10 % (Fig 3a). The most significant degradation rate of 21.54 mg/h was obtained when 10% inoculum was used. Whereas 8%, 6%, 4%, 2% and 1% of inoculum sizes have degradation rates of 19.84 mg/h, 16.20 mg/h, 14.15 mg/h, 13.21 mg/h and 6.63 mg/h, respectively (p = 3.5E–03). Results showed that degradation rate doubled sharply from 6.63 to 13.21 mg/h at 1% and 2% of inoculum sizes. Furthermore, thresholds for degradation products were expressed as a percentage of the methyl red dye. It was observed that with increasing inoculum size from 1% to 10%, time required to achieve maximum degradation reduces (Fig 3b). The threshold degradation of inoculum size of 10 % and 8 % were achieved within 4.5 and 5 h, respectively. Furthermore, threshold degradation was attained in 6.5 h and 7 h with 6% and 2%, respectively. While inoculum size of 1% was shown to have longer threshold degradation after 24 h. Sari et al. (2019) showed a similar trend, indicating that the dye removal capability of the method varies dramatically with increase in inoculum size. They demonstrated that raising the initial inoculum size to 10% (v/v) improved the degrading efficiency of Lysinibacillus fusiformis W1B6 (Tripathi et al. 2016).
a Variation in Dye degradation rate (Y-axis) against initial inoculum size (X-axis). This graph shows that the degradation rate is not constant but changing b Percentage degradation (Z-axis) with time (Y-axis) on varying inoculum size (X-axis). All the values were significant at p < 0.05 on ANOVA analysis
Effect of salinity
The effect of varying salinity (0.5–180 g L−1) was investigated while maintaining pH, inoculum size, temperature and dye concentration constant at 7.0, 2 % (v/v), 35 °C and 100 mg L−1, respectively. As a result, shown in Figure 4a, percentage degradation was time-dependent which increased significantly with an increase in salinity from 0.5 to 120 g L−1, thereafter, decreasing sharply. The isolated strain was capable of degrading up to maximum of 94.8 % methyl red within 24 h after inoculation in the presence of 120 g.L−1 NaCl. The maximum degradation decreased to 77.26%, 81.46% and 29.36% in the presence of 140, 160 and 180 g L−1 salinity, respectively (p = 3.3E–04). Similarly, growth analysis of bacterial isolates showed that OD650 values were increased with increased salinity from 0.5 to 120 g L−1, thereafter decreased (Fig 4b). The highest degradation percentage was observed in between 10 and 120 g L−1 salinity, where OD650 values were greater. However, samples grown in very low or high salinity concentrations could not show good performance in decolorization. Our finding suggests that activity of azo reductase enzyme of isolates may be affected by the surrounding concentrations of salt. As our strain has the ability to grow in both lower (0.5 g L−1) as well as higher saline concentrations (180 g L−1), it could be effectively used for large-scale bio-decolorization. The moderate decolorization ability with lower growth rates at lower and higher saline concentrations may cause plasmolysis and/or loss of activity of cells. Many studies discovered that bacteria had a remarkable capacity to adapt to a wide variety of salt concentrations. Recently Diao et al. (2022) shown that Lactiplantibacillus sp. can grow up to 5 % NaCl concentration (Yu et al. 2021) Bhattacharya et al. (2017) observed that the Nesterenkonia lacusekhoensis EMLA3 can efficiently decompose up to 96% MR at 48 h in the presence of 5.0, 20, 40, and 60 g L−1 NaCl. At higher salt concentrations of 80 and 100 g L−1, however, deterioration was reduced by up to 60 and 48%, respectively, after 48 h. (Bhattacharya et al. 2017). In this investigation, however, a maximum of 94.8% MR degradation was reached after 24 h of inoculation in the presence of 120 g L−1 NaCl. In another study, Cui et al. (2014) showed that Klebsiella sp. strain Y3 can degrade the MR efficiently in a 1–4% saline environment (Cui et al. 2014). Our findings reveal that isolated L. plantarum SS-AU1 not only survives in extremely high salinity but also degrades at a rate of more than 90%.
Effect of initial dye concentration
A series of experiments was performed while maintaining pH, salinity, inoculum size, and temperature i.e. 7.0, 100 g L−1, 2 % (v/v), and 35 °C, respectively but varying initial dye concentrations (50, 100, 150, 200, 250, 300, and 400 mg L−1). In the present study, the L. plantarum SS-AU1 showed a good performance with the different concentrations of methyl red. The results showed over 90 % degradation up to 400 mg L−1 of dye concentration. It seems that initial dye concentration affected the degradation capability of the bacterium (see Fig 5b). The degradation rate initially increased from 20.3 to 32. mg h−1 with an increase in dye concentration up to 200 mg.L-1. Following this, there was constant reduction reaching 15.2 mg h−1 for 400 mg L−1 initial dye concentration. Furthermore, the period to attain over 90 % of degradation was also varied among initial dye concentration. It was obvious that more than 90 % decolorization could be achieved between 4 and 6 h. Incubation for the initial dye concentrations of 50–200 mg L−1. Decolourization reached a maximum of 95.4 % at 100 mg L−1, while 300 mg L−1 achieved 93.5 % decolourization followed by 400 mg L−1 showing 93.6 % of decolourization in 24 h of incubation (Fig 5a) (p = 3.7E−06). At concentrations higher than 400 mg L−1MR a black product was formed which could not be characterized spectroscopically. This study showed that when the initial dye concentration was increased, the percentage of decolorization reduced. Bhattacharya et al. (2017) reported similar findings, demonstrating that initial dye concentration has a significant impact on the degradation process. They observed that if the dye concentration is more than 400 mg L−1, the degradation is greatly impeded (Bhattacharya et al. 2017).
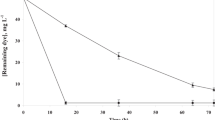
The degradation kinetic curve in Figure 5b follows a second-order reaction with 0.9697 R2 (Supplementary Table 3), as also reported by Takkar et al. (2022). They demonstrated the second-order degradation of MR by novel Actinobacterium Zhihengliuella sp. ISTPL4 (Takkar et al. 2022). It is apparent that the deterioration takes place in two stages. Degradation is quite slow in the initial phase, around 10% within the first 3.5 h. In the second phase, the degradation rate was more than 90 % in the additional 3.3 h. Within 24 h, more than 95 % deterioration was accomplished. The half-life of MR in the reaction is 6.08 h, which suggests that 50% of the MR is degraded in approximately 6 h. According to Lellis et al. (2019), azo dyes have a half-life in the environment ranging from 2 to 13 years (Lellis et al. 2019). This study clearly emphasizes the potential of L. plantarum SS-AU1 as an efficient bioremediating tool.
Effect of pH and temperature
To investigate the influence of both pH and temperature on MR degradation, a wide range of pH (5–12) with a constant temperature of 35 °C, and temperature (20–50 °C) with constant pH of 7 was studied while maintaining constant salinity, dye concentration, inoculum size and incubation time i.e. 100 g L−1, 2 % (v/v), 100 mg L−1, and 24 h., respectively.
The results revealed that the isolate L. plantarum SS-AU1could not only survive in a pH range from 7 to 11, but also degraded the methyl red (Fig 6a) However, at pH 5, 6, and 12, growth of the bacterium was significantly suppressed, and dye degradation potential was also diminished after 24 h of incubation. At pH 7, 8, 9, 10, and 11, the percentages of decolorization were 92.2%, 88.9%, 92%, 97.24%, and 94.4%, respectively (1.7E−07). Diao et al. (2022) reported a similar observation for the MR degradation in Lactiplantibacillus sp at pH ranges 2–9 (Yu et al. 2021).
The studies on varying temperatures revealed that L. plantarum SS-AU1 efficiently decolorized the dye with the maximum degradation of 92.2% at 35 °C. The percentage degradation at 20, 25, and 30 °C was 18.2, 53.5, and 87.9 %, respectively (Fig 6b). At temperatures higher than 35 °C, the percentage degradation was 89.4, 78.9, and 4.8% at 40, 45, and 50 °C, respectively (5.2E−03). Similar results have been reported by Bhattcharya et al. (2017) with N. lacusekhoensis EMLA3 (Bhattacharya et al. 2017). The MR precipitates out at less than or equal to pH 6 which may be a reason behind the negligible degradation due to aggregation of the solute generating a localized high concentration.
Effect of cycle time on biodegradation of dye
The effects of cycle time on the biodegradation of the methyl red dye were investigated in a batch experiment using stationary and shaking modes. The experiment was repeated seven times with the addition of MR (100 mg L−1) each time. All experiments were performed at constant initial pH, salinity, temperature, and inoculum size i.e. 7.0, 100 g L−1, 35 °C and 10 % (v/v). The time required for wastewater treatment at the industrial scale is an important criterion for the effective adoption of any bioremediating system. The higher inoculum size of 10% (v/v) was used to make MR degradation process more rapid.
The result showed that after the first cycle of degradation, the second cycle takes only 2.5 h, and the seventh cycle 1 h (Supplementary Fig. S1). The degradation under stationary conditions was 87.23 % and 92.38 % in the shaking conditions of the first cycle. This subsequently decreased to 79.3 and 74.3 %, respectively in the seventh cycle. The shift in the percentage degradation pattern, after the fourth cycle from stationary to shaking conditions was due to enhanced cell density. The shaking phase leads to a greater cell mass, thereby rapidly depleting the media nutrients. A study has shown a decline in dye degradation capabilities of the bacteria due to cell density. A higher cell density and nutrient depletion are directly proportional (Tripathi et al. 2016). It is noteworthy that after the seventh cycle, a black color appears which might be related to another byproduct build-up. Mahajan et al. (2020) have also studied the reusability of the inoculum over different cycles and they have concluded that the microbe can degrade the dye up to 6th cycle after that the efficiency starts to diminish drastically (Mahajan and Kaushal 2020).
Metabolite characterization by FTIR
The wave number corresponding to the Hydroxyl group (3200–2700), Carboxylic group(1730), Azo group (1570–1640), Aromatic C–N (1240–1370), C–N amine (1030–1180), C=C (950) were prominently visible in the control sample of MR (Supplementary Fig. 5a). In this study, there were two steps of degradation of MR. It first converts into a decolorized sample that has only stretched hydroxyl group, azo bond, C–N amine and C=C with new bond appearance corresponding to the Phenol group (1350) and Aldehyde group (1380) (Supplementary Fig. S5b). The second step of degradation where a black product was formed may be due to absence of aromatic amine (Supplementary Fig. S5c). Moreover, there is a substantial change in the azo bond wave number of metabolites as compared to the control which suggests a change in the azo bond structure in the metabolites. Sari et al. (2019) and Ikram et al. (2022) have also reported substantial changes in the hydroxyl, amine and azo function groups after microbial treatment (Cappucino and Sherman 2005; Sari and Simarani 2019). Based on the above observations, following mechanism can be postulated for MR degradation (Supplementary Fig. S5d). The MR first breaks down into phenol, an aldehyde and an intermediate benzyl hydrazine derivative. The final steps show the conversion of benzyl hydrazine derivative into benzyl hydrazine. Hence in the overall reaction, Methyl Red is degraded in two steps by SS-AU1 into an aldehyde, a phenol and a benzyl hydrazine.
Adsorption kinetics models
The Lineweaver–Burk plot was assessed to identify the theoretical maximum velocity of reaction and possible enzymes involved in the process. The Vm value was 0.761 mmol min−1 whereas Km value was 9.1 µmol−1 L which shows quite high degradation efficiency. The similar levels were reported by Cong et al. (2022) where they reported that novel azo reductase BVU5 can remove MR with Km = 0.092 mMol−1 L and Vm = 0.067 mmol L−1 min−1 (Brdarić et al. 2021).
The Pseudo-first order (PFO), pseudo-second-order (PSO), Elovich kinetic models, and Weber–Morris intra-particle diffusion model were employed to appraise the experimental adsorption data of MR dye biodegradation by L. plantarum SS-AU1. The kinetic parameters at different initial MR concentrations were determined based on the corresponding linear equations and are summarized in Tables 1 and 2.
The Lagergren model for first and second-order reactions (PFO & PSO) was investigated to study the kinetics of the process. Supplementary Fig S2a and b show the fitting plots of the PFO and PSO models, in which the kinetic data fitted well based on the relatively high R2 values. The PSO model was exhibited at lower dye concentrations (50–100 mg L−1), while PFO model was observed at higher concentrations. At lower concentrations, the R2 value for the PFO model is non-significant (0.50), however the R2 value for the PSO model is strongly significant (> 0.80) (Table 1). Furthermore, theoretical qe values reveal a substantial divergence (> 80%) from experimental qe in the PFO model, but a strong similarity with the PSO model for this range of dye concentration. In contrast, higher MR concentrations (> 100 mg L−1) exhibited extremely significant R2 (Table 2) values with the PFO model, but not with the PSO model. The matching qe-values confirm this. In this range, there is relatively little difference between experimental and theoretical qe for the first order and extremely significant divergence for the second order. This is explained by the fact that at low dye concentrations, there is a high surface area accessible for adsorption, but at high concentrations, the free sites accessibility for the dye molecules is limited. As a result, at lower concentrations, accessible sites as well as dye concentrations play important roles in the PSO model. However, at higher concentrations, only accessible sites are important because there is a constant equilibrium state between the occupied site and available sites for adsorption over a significant period of time. As a result, dye concentration becomes an insignificant component for adsorption, and the PFO model is used in this range. Sari et al. (2019) and Mahajans et al. (2020) have also studied the validity of PFO and PSO. Sari et al. (2019) reported that Lysinibacillus fusiformis W1B6 followed the PFO at 100 mg/L dye concentration whereas Mahajans et al. (2020) reported that the C. vulgaris follows the PSO at 20 mg/L dye concentration in MR degradation process (Mahajan and Kaushal 2020; Tripathi et al. 2016)
The Elovich and Weber–Morris models were investigated for the study of the mechanism after dye adsorption on the cell surface (Supplementary Fig. S2c and d). The 100 mg/L dye concentration clearly follows these models (R2 > 0.50). (See supplementary Table S4). The negative values of Ci depict multilayer corrosion induced by the continuous degradation process. The positive Ki values imply that there is a particularly rapid intra-particle diffusion phenomenon within the multilayer. This could be largely attributable to the constant degradation process, which restricts the saturation level in the multilayer and leads to a high Ki value. The Elovich model also suggests this phenomenon. There are clear indications of a simultaneous adsorption-desorption process, with both constants α & β being positive. A higher value of α than β indicates the desorption phenomenon. As a result, available adsorption sites are increasing, explaining the positive value of α. The continuous decrease of all kinetic model constants may be due to the hindrance phenomenon, in which an increased amount of dye prevents interaction with available adsorption sites on the cell surface. As a result, the MR is continuously adsorbing on the surface while also being degraded by the L.plantarum SS-AU1. Mahajan et al. (2020) concluded that the degradation is due to the interaction between the functional groups and dye rather than the surface of the cell (Mahajan and Kaushal 2020).
Adsorption isotherm studies
The Freundlich, Langmuir, R–P, SIPS and Elovich isotherms (Supplementary Fig. S3a–e respectively) were studied to assess the formation of the layer type, e.g., monolayer or multilayer. The experimental R2 values imply that a multilayer is growing on the cell surface, as the multilayer-suggesting isotherms (Freundlich, Sips, and R–P) have larger R2 values than Langmuir (Table 3). The high R2 values for the Freundlich and Sips isotherms indicate the formation of multiple layers. The value of n is between 1 and 10, indicating that multilayer formation is favorable, but 1/n = 0.968 indicates that the system's heterogeneity is very high and the binding energy is quite random. Based on the Sips isotherm, the qmax value of the L. plantarum SS-AU1 is 2500 mg g−1, which is very high, and hence the effectiveness of L. plantarumSS-AU1 in MR degradation is likewise quite high. As 1-β is equivalent to 1/n, the R-P model validates the formation of a multilayer in the system. This has been proven strongly earlier in this article by D–R isotherm studies. The Elovich isotherm postulates that the number of adsorption sites induced in the presence of adsorbent facilitates the formation of multilayer. The adsorption is clearly following multilayer formation in the above graph with lower R2 (0.69). Thus, the adsorption coefficient value is low (i.e. 1.7 × 103 mg/g), compared to the Sips Model (2500 mg/g). The observations of the model are justified by the facts observed in the SEM characterization. The formation of multilayer indicates the degradation process is a surface phenomenon which may be the reason behind the modified/elongated surface of the cell. Sari et al. (2019) reported that Lysinibacillus fusiformis W1B6 follows the Langmuir isotherm rather than Freundlich isotherm which predicts the formation of a monolayer structure on the surface of the microbe (Tripathi et al. 2016). Mahajan et al. (2020) reported that the C. vulgaris follows the Freundlich model (Mahajan and Kaushal 2020).
Harkin–Jura, Halsey, Temkin and D–R isotherms (Supplementary Fig. S4a–d) were investigated for heterogeneous pore distribution on a multilayer system. Significant R2 (0.8063) values of the D–R isotherm reflect the formation of a porous structure on the cell’s surface, which may be analogous to numerous ligands. Bradaric et al. (2021) also demonstrated that biomolecules (such as Lipopolysaccharide) on the cell surfaces of Lactoplantibacilli are responsible for the adsorption of foreign entities, implying that these molecules are required for degradation processes (Liu et al. 2019). The maximum adsorption coefficient was 109.18 mg g−1, implying that a large amount of dye may be accommodated and saturated in the pores, simultaneously. The system’s mean free activation energy (activity coefficient) is exceptionally low, 6.0 × 10−4 g2/J2, suggesting that very little energy is required to initiate the process. Furthermore, the binding energy is relatively low, 28.9 J/mol, which is substantially less than 8 KJ/mol, confirming the presence of physical forces. The energy is beyond the range of Van der Waal forces (0.4–4 KJ/mol), implying that other weaker forces are involved in the process, and so relatively little energy is necessary to adsorb the dye on the cell's surface (Amin et al. 2015). Furthermore, the H–J and Halsey isotherms validate the presence of heterogeneous porous structures on the multilayer system (Kecili and Hussain 2018). The uneven surface of the cell morphology observed in the SEM characterization also validates the above mathematical model. The almost perfect fitting of the experimental data to the Halsey model (R2 = 0.9988) indicates a highly packed porous structure on the surface, implying that the system is particularly efficient in the degrading process. According to the Temkin isotherm, increasing the coverage of the adsorbate surface leads to the stabilization of the binding energy, and hence the heat of the system. The preceding Fig. 4b clearly shows no signs of saturation; contrary, it follows an exponential trajectory. This indicates that the surface's covering is not achieving saturation, but that there are always residual regions on the surface that are free of any coupling (Ayawei et al. 2017) and thereby accessible for adsorption. This suggests that adsorption and degradation are occurring simultaneously on the cell's surface, with the degradation rate being greater than the adsorption rate. This could also explain S-1’s efficient degrading capabilities even on the seventh cycle of examination (Supplementary Fig. S1).
The Jovanovic isotherm proposes that, in addition to physical contact, there is mechanical interaction between adsorbate and adsorbent. In our study (Supplementary Fig. S4e), the R2 value of Jovanovic isotherms is quite high i.e., 0.8896. The saturation adsorption coefficient is 24.58 mg/g, compared to the maximal adsorption efficiency (2500 mg/g), and the Kj is quite low, − 0.0057 L/g (Ayawei et al. 2017).
Conclusion
Methyl red is one of the most popular chemical azo dye colourants used in textile, printing, and paper industries, and its toxicity trigger deleterious effects on human health. To the best of our abilities, Lactiplantibacillus plantarum SS-AU1, an alkalaphilic and halotolerant bacterium isolated from the Ganges River, may have used for the first time in this investigation as a critical tool to breakdown MR in a high saline environment. The study demonstrated that several physicochemical conditions, such as pH (7–10), initial dye concentrations (50, 100, 150, 200, 250, 300, and 400 mg L−1), temperature (20–45 °C), and salinity (0.5–180 g L−1), might influence MR degradation ability of bacterium and should be regulated for optimal commercial use. Furthermore, after the first cycle of MR degradation (92.38 %) in 3 h, the second cycle reduces degradation time to 2.5 h, and the 7th cycle requires only 1 h for the 74.3 % degradation. The kinetics and isotherm analysis of dye adsorption on the bacterial surface and the generation of less hazardous metabolites confirmed by FTIR reveal that isolate is capable of efficient degradation under high salinity. Thus, the L. plantarum SS-AU1 strain has the potential to be a cost-effective and efficient MR dyes removal tool via biological routes, particularly from high salinity industrial effluents.
References
Abd El-Rahim WM, Moawad H, Azeiz AZA, Sadowsky MJ (2021) Biodegradation of azo dyes by bacterial or fungal consortium and identification of the biodegradation products. Egypt J Aquat Res 47(3):269–276
Amin MT, Alazba AA, Shafiq M (2015) Adsorptive removal of reactive black 5 from wastewater using bentonite clay: isotherms, kinetics and thermodynamics. Sustainability 7(11):15302–15318
Ayawei N, Ebelegi AN, Wankasi D (2017) Modelling and interpretation of adsorption isotherms. J Chem. https://doi.org/10.1155/2017/3039817
Bhattacharya A, Goyal N, Gupta A (2017) Degradation of azo dye methyl red by alkaliphilic, halotolerant Nesterenkonia lacusekhoensis EMLA3: application in alkaline and salt-rich dyeing effluent treatment. Extremophiles 21:479–490
Brdarić E, SokovićBajić S, Đokić J, Đurđić S, Ruas-Madiedo P, Stevanović M, Tolinački M, Dinić M, Mutić J, Golić N, Živković M (2021) Protective effect of an exopolysaccharide produced by Lactiplantibacillus plantarum BGAN8 against cadmium-induced toxicity in Caco-2 Cells. Front Microbiol 12:759378
Bu Y, Liu Y, Liu Y, Wang S, Liu Q, Hao H, Yi H (2022) Screening and probiotic potential evaluation of bacteriocin-producing Lactiplantibacillus plantarum in vitro. Foods 11(11):1575
Cappucino JG, Sherman N (2005) Microbiology a laboratory manual, 7th edn. Aurora, Torrance
Cheesbrough M (1981) Medical laboratory manual for tropical countries (vol 1). M. Cheesbrough, 14 Bevills Close, Doddington, Cambridgeshire, PE15 OTT
Chen H, Xu H, Heinze TM, Cerniglia CE (2009) Decolorization of water and oil-soluble azo dyes by Lactobacillus acidophilus and Lactobacillus fermentum. J Ind Microbiol Biotechnol 36(12):1459
Cong J, Xie X, Liu Y, Qin Y, Fan J, Fang Y, Liu N, Zhang Q, Song X, Sand W (2022) Biochemical characterization of a novel azo reductase named BVU5 from the bacterial flora DDMZ1: application for decolorization of azo dyes. RSC Adv 12(4):1968–1981
Cui D, Li G, Zhao M, Han S (2014) Decolourization of azo dyes by a newly isolated Klebsiella sp. strain Y3, and effects of various factors on biodegradation. Biotechnol Biotechnol Equip 28(3):478–486
Diao X, Yamada K, Shibata Y, Imada C (2022) Metabolites produced by a new Lactiplantibacillus plantarum strain BF1-13 isolated from deep seawater of Izu-Akazawa protect the intestinal epithelial barrier from the dysfunction induced by hydrogen peroxide. Mar Drugs 20(2):87
Echegaray N, Yilmaz B, Sharma H, Kumar M, Pateiro M, Ozogul F, Lorenzo JM (2023) A novel approach to Lactiplantibacillus plantarum: from probiotic properties to the omics insights. Microbiol Res 268:127289
Feng J, Cerniglia CE, Chen H (2012) Toxicological significance of azo dye metabolism by human intestinal microbiota. Front Biosci (elite Ed) 4:568
Gupte S, Keharia H, Gupte A (2013) Toxicity analysis of azo Red BS and methyl Red dye solutions on earthworm (Pheretima phosthuma), micro-organisms, and plants. Desalin Water Treat 51(22–24):4556–4565
Hartmann M, Berditsch M, Hawecker J, Ardakani MF, Gerthsen D, Ulrich AS (2010) Damage of the bacterial cell envelope by antimicrobial peptides gramicidin S and PGLa as revealed by transmission and scanning electron microscopy. Antimicrob Agents Chemother 54(8):3132–3142
Ikram M, Naeem M, Zahoor M, Rahim A, Hanafiah MM, Oyekanmi AA, Shah AB, Mahnashi MH, Al Ali A, Jalal NA, Bantun F (2022) Biodegradation of azo dye methyl red by Pseudomonas aeruginosa: optimization of process conditions. Int J Environ Res Public Health 19(16):9962
Jakobek L, Matić P, Kraljević Š, Ukić Š, Benšić M, Barron AR (2020) Adsorption between quercetin derivatives and β-glucan studied with a novel approach to modeling adsorption isotherms. Appl Sci 10(5):1637
Jeong CH, Sohn H, Hwang H, Lee HJ, Kim TW, Kim DS, Kim CS, Han SG, Hong SW (2021) Comparison of the probiotic potential between Lactiplantibacillus plantarum isolated from kimchi and standard probiotic strains isolated from different sources. Foods 10(9):2125
Kang Y, Xu X, Pan H, Tian J, Tang W, Liu S (2018) Decolorization of mordant yellow 1 using Aspergillus sp. TS-A CGMCC 12964 by biosorption and biodegradation. Bioengineered 9(1):222–232
Kaur N, Kaushal J, Mahajan P, Mantri A (2023) Phytoremediation of methylene blue dye (triarylmethane) and Congo red (diazo) by T. ammi L.: kinetic studies. Int J Environ Sci Technol. https://doi.org/10.1007/s13762-023-05037-x
Kecili R, Hussain CM (2018) Mechanism of adsorption on nanomaterials. In: Hussain CM (ed) Nanomaterials in chromatography. Elsevier, Amsterdam, pp 89–115
Kgatle M, Sikhwivhilu K, Ndlovu G, Moloto N (2021) Degradation kinetics of methyl orange dye in water using trimetallic Fe/Cu/Ag nanoparticles. Catalysts 11(4):428
Kumara NTRN, Hamdan N, Petra MI, Tennakoon KU, Ekanayake P (2014) Equilibrium isotherm studies of adsorption of pigments extracted from Kuduk-kuduk (Melastoma malabathricum L.) pulp onto TiO2 nanoparticles. Journal of Chem. https://doi.org/10.1155/2014/468975
Lellis B, Fávaro-Polonio CZ, Pamphile JA, Polonio JC (2019) Effects of textile dyes on health and the environment and bioremediation potential of living organisms. Biotechnol Res Innov 3(2):275–290
Li SL, Wang YH, Chao YC, Bai MD, Cheng SS (2018) Amylolysis is predominated by cell-surface-bound hydrolase during anaerobic fermentation under mesophilic conditions. J Biosci Bioeng 125(4):432–438
Liu L, Luo XB, Ding L, Luo SL (2019) Application of nanotechnology in the removal of heavy metal from water. In: Luo X, Deng F (eds) Nanomaterials for the removal of pollutants and resource reutilization. Elsevier, Amsterdam, pp 83–147
Mahajan P, Kaushal J (2020) Phytoremediation of azo dye methyl red by macroalgae Chara vulgaris L.: kinetic and equilibrium studies. Environ Sci Pollut Res 27(21):26406–26418
Mansour AT, Alprol AE, Abualnaja KM, El-Beltagi HS, Ramadan KM, Ashour M (2022) Dried brown seaweed’s phytoremediation potential for methylene blue dye removal from aquatic environments. Polymers 14(7):1375
Mawad AM, AbdHesham EL, Yousef NM, Shoreit AA, Gathergood N, Gupta VK (2020) Role of bacterial-fungal consortium for enhancement in the degradation of industrial dyes. Curr Genom 21(4):283–294
Montañez-Barragán B, Sanz-Martín JL, Gutiérrez-Macías P, Morato-Cerro A, Rodríguez-Vázquez R, Barragán-Huerta BE (2020) Azo dyes decolorization under high alkalinity and salinity conditions by Halomonas sp. in batch and packed bed reactor. Extremophiles 24:239–247
Mostafidi M, Sanjabi MR, Mojgani N, Eskandari S, ArbabiBidgoli S (2023) Heavy metal bioremediation potential of autochthonous lactic acid bacteria for use in edible leafy vegetables. J Food Qual. https://doi.org/10.1155/2023/8730676
Oliveira JMS, Poulsen JS, Foresti E, Nielsen JL (2023) Microbial communities and metabolic pathways involved in reductive decolorization of an azo dye in a two-stage AD system. Chemosphere 310:136731
Piccin JS, Dotto GL, Pinto LAA (2011) Adsorption isotherms and thermochemical data of FD&C Red n 40 binding by chitosan. Braz J Chem Eng 28:295–304
Rocchetti MT, Russo P, Capozzi V, Drider D, Spano G, Fiocco D (2021) Bioprospecting antimicrobials from Lactiplantibacillus plantarum: key factors underlying its probiotic action. Int J Mol Sci 22(21):12076
Rudzinski W, Plazinski W (2008) Kinetics of solute adsorption at solid/solution interfaces: on the special features of the initial adsorption kinetics. Langmuir 24(13):6738–6744
Sahoo TR, Prelot B (2020) Adsorption processes for the removal of contaminants from wastewater: the perspective role of nanomaterials and nanotechnology. In: Bonelli B, Freyria FS, Rossetti I, Sethi R (eds) Nanomaterials for the detection and removal of wastewater pollutants. Elsevier, Amsterdam, pp 161–222
Saleh AK, El-Gendi H, Soliman NA, El-Zawawy WK, Abdel-Fattah YR (2022) Bioprocess development for bacterial cellulose biosynthesis by novel Lactiplantibacillus plantarum isolate along with characterization and antimicrobial assessment of fabricated membrane. Sci Rep 12(1):2181
Sari IP, Simarani K (2019) Comparative static and shaking culture of metabolite derived from methyl red degradation by Lysinibacillus fusiformis strain W1B6. R Soc Open Sci 6(7):190152
Sharma S, Pathak S, Sharma KP (2003) Toxicity of the azo dye methyl red to the organisms in microcosms, with special reference to the guppy (Poecilia reticulata Peters). Bull Environ Contam Toxicol 70:753–760
Sharma S, Sharma S, Upreti N, Sharma KP (2009) Monitoring toxicity of an azo dye methyl red and a heavy metal Cu, using plant and animal bioassays. Toxicol Environ Chem 91(1):109–120
Singh P (2022) Bioremediation of hazardous azo dye methyl red by a newly isolated Enterobacter asburiae strain JCM6051 from industrial effluent of Uttarakhand regions. J Appl Biol Biotechnol 10(2):64–72
Sundaramanickam A, Awasthi S, Patel V (2023) Bioremediation of heavy metals from aquatic environments by lactic acid bacteria. In: Montet D, Ray RC, Azevedo VA, Paramithiotis S (eds) Lactic acid bacteria as cell factories. Woodhead Publishing, Sawston, pp 389–399
Takkar S, Tyagi B, Kumar N, Kumari T, Iqbal K, Varma A, Thakur IS, Mishra A (2022) Biodegradation of methyl red dye by a novel actinobacterium Zhihengliuella sp. ISTPL4: kinetic studies, isotherm and biodegradation pathway. Environ Technol Innov 26:102348
Tian F, Wang Y, Guo G, Ding K, Yang F, Wang H, Cao Y, Liu C (2021) Enhanced azo dye biodegradation at high salinity by a halophilic bacterial consortium. Biores Technol 326:124749
Tripathi A, Singh Y, Verma DK, Ranjan MR, Srivastava SK (2016) Bioremediation of hazardous azo dye methyl red by a newly isolated Bacillus megaterium ITBHU01: process improvement through ANN-GA based synergistic approach
Vinoda BM, Vinuth M, Bodke YD, Manjanna J (2015) Photocatalytic degradation of toxic methyl red dye using silica nanoparticles synthesized from rice husk ash. J Environ Anal Toxicol 5(1000336):2161–2525
Yilmaz B, Bangar SP, Echegaray N, Suri S, Tomasevic I, Manuel Lorenzo J, Melekoglu E, Rocha JM, Ozogul F (2022) The impacts of Lactiplantibacillus plantarum on the functional properties of fermented foods: a review of current knowledge. Microorganisms 10(4):826
Yu AO, Goldman EA, Brooks JT, Golomb BL, Yim IS, Gotcheva V, Angelov A, Kim EB, Marco ML (2021) Strain diversity of plant-associated Lactiplantibacillus plantarum. Microb Biotechnol 14(5):1990–2008
Acknowledgements
Authors are thankful to the Centre of Material Science and National Centre of Experimental Petrology and Mineralogy, University of Allahabad, Prayagraj, India for providing necessary laboratory facilities. Mr. Abhijeet Sharma is grateful to the Department of Biotechnology, New Delhi for providing him with the Junior Research Fellowship for his work under the sanction order BT/PR31340/BCE/8/1505/2019.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Editorial responsibility: Nabin Aryal.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sharma, A., Singh, S.K., Nath, A. et al. Methyl red biodegradation by novel halophilic Lactiplantibacillus plantarum SS-AU1 isolated from river Ganges. Int. J. Environ. Sci. Technol. 21, 7191–7206 (2024). https://doi.org/10.1007/s13762-024-05468-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-024-05468-0