Abstract
Wastewater contaminated with the antibiotic-resistant bacteria, Pseudomonas aeruginosa, can contribute to human community-acquired infections when released into receiving waters. This study outlines a novel process of phage application that can reduce the reservoir of P. aeruginosa in both primary wastewater (PWW) and secondary wastewater (SWW). The phage PA25 was first successfully isolated from SWW and is a double-stranded DNA phage, classified as a Siphoviridae family as defined by plaque morphology, electron microscopy and host range. Bacteria such as Pseudomonas are the natural host of this virus; the addition of Siphoviridae PA25 has resulted in the greatest reduction of bacteria from unsterilized PWW compared to unsterilized SWW. Experimental results showed a bacterial reduction of 5ULog discharge in PWW compared only 3ULog in SWW. The addition of PA25 to wastewater can also eliminate streptomycin resistance in P. aeruginosa ATCC strain 27853. Infected P. aeruginosa showed decreased resistance to the antibiotics gentamicin and rifampicin.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Untreated wastewaters that are discharged into the Mediterranean Sea represent a serious threat to human health, both by swimming in polluted water and by the consumption of contaminated seafood (Prieto et al. 2001; Danovaro et al. 2009a, b). Mandated closures of beaches for swimming and closing shellfish harvesting have occurred after wastewater discharge in response to this public health threat, especially along the coasts of North Africa (Kamizoulis and Saliba, 2004).
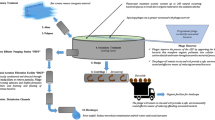
Wastewater treatment processes include three stages aimed to improve the environmental quality of effluents; these stages are primary, secondary and tertiary treatment. Primary treatment involves mechanical scrapers that separate greases and oils from the wastewater streams. Secondary wastewater treatment engages microorganisms to digest particulate organic matter in the wastewater stream. Tertiary wastewater treatment is a final stage to remove pathogens such as coliforms, nutrients and other pollutants to meet treatment performance criteria as determined by regulations for effluent water quality. Tertiary wastewater treatment is essential for producing clean and high-quality effluent that can be discharged into receiving waters. Effluent standards are addressed in the Sustainable Development Goals. Nevertheless, in some cases, environmental regulations and public health concerns require further performance standards in tertiary treatment. Appropriate choices of all stages of wastewater treatment affect the overall wastewater stream cost and efficacy. Community-degrading microorganisms present in the wastewater stream can be affected by the presence of antibiotics naturally present in wastewater. Although wastewater systems are not specifically designed to remove pharmaceuticals, pharmaceuticals as pollutants are increasingly recognized by the negative environmental impacts on receiving waters. Advanced tertiary treatment methods like activated carbon, ozonation and advanced oxidation have shown efficiency in pharmaceutical removal (Rivera-Utrilla et al. 2013).
Biological treatments in most wastewater plants address the removal of some types of pharmaceuticals; however, antibiotics persist in treated wastewater, inducing bacteria resistance. Growing environmental regulations and public health concerns implemented to achieve the Sustainable Development Goals1 have called for tertiary treatment of wastewater before discharge into receiving waters to remove both pathogens and nutrients that contribute to eutrophication. In the past, chlorination was the most common method to disinfect wastewater. However, it has been shown that chlorine disinfection creates by-products that negatively impact both the environmental and human health (Al-Abri et al. 2019; Tsamba et al. 2020). Tunisia wastewater treatment plants (WWTPs) managers wish to initiate more sustainable and effective practices for advanced wastewater treatment that may include ultra-violet (UV) disinfection to reduce antibiotic-resistant bacteria (Mansour 2018). Some harmful microorganisms can escape UV treatment. For example, P. aeruginosa is a ubiquitous opportunistic pathogen of both plants and humans (Ebrahimi et al. 2017). This pathogen causes respiratory, urinary tract and bloodstream infections in humans (Brizou 1972; Slekovec et al. 2012). P. aeruginosa strains are antibiotic resistance for many common antibiotics, disinfectants and biocides (Pukatzki et al. 2002; Schwartz et al. 2015). P. aeruginosa may produce persistent biofilms (Saidi et al. 2011) as well as show resistance to common antibiotics (Abedon 2020).
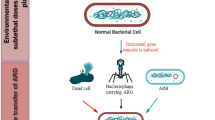
Phage therapy is well established in clinical settings to remove pathogenic bacteria; however, the use of phages to remove pathogenic bacteria in a WWTP setting could be a viable alternative to both UV disinfection and chlorination. The culture of phage was accomplished in simple culture medium including only host cells. The easy, fast and economical methods available for phage multiplication are feasible both in industrialized and developing countries. Phages could kill bacteria or/and to impact the dissemination of genes encoding antibiotic resistance. Phage therapy can be used as a preventive measurement against community infection from exposure to wastewater (Cisek et al. 2017). Methods to reduce community infections that may not be treatable with antibiotics are critical moving forward. The cost of WWTP phage therapy is likely significantly lower than the public health costs from any novel community infections from wastewater effluent exposure (Bradbury 2004). Phages are an efficient alternative to eliminate pathogenic bacteria in specific environmental settings (Ackermann and DuBow, 1987; Summers, 2005). Phages present an interesting tool for scientific research given the ease and speed of laboratory phage cultivation (Ackermann 2001; Merril et al. 2003; Labrie et al. 2009). Two categories of bacteriophage are recognized: temperate and virulent. During lytic infection, virulent phages inject their nucleic acid into the host cell following attachment to the cell membrane, and then, they exploit the cellular machinery of the host to synthesize new phage capsule material to complete the lytic cycle with new host cells infections (Withey et al. 2005). In contrast, during lysogenic infection, temperate phage nucleic acid recombines with the host cell genome forming a dormant prophage; this prophage does not induce the lytic cycle in the bacteria cell. In wastewater treatment, only lytic phages were considered for improving water quality since the goal was to kill the bacteria host cell.
This research explores methods to improve wastewater quality via bacteriophage use to reduce antibiotic-resistant pathogens. This study aims (i) to isolate and to identify lytic bacteriophages from wastewater; (ii) to use the phage as a biocontrol agent for P. aeruginosa strain ATCC27853 when added in PWW and SWW; and (iii) to document the impact of phage on the P. aeruginosa strain ATCC 27853 antibiotic resistance for selected antibiotics (e.g., gentamicin, streptomycin, rifamycin, spiramycin and tetracycline).
Material and methods
Bacteriophage host range
The strain P. aeruginosa ATCC 27853 was used in this research. PA25 was isolated from domestic SWW from Charguia pilot, Tunisia. The physicochemical characteristics of the PWW and SWW effluents are presented in Table 1.
Phages isolation followed the protocols outlines in Petrovski et al. (2011). A 50 mL sample of wastewater was added to 50 mL Tryptic Soy Broth (TSB, Difco, medium). One mL of an overnight culture of P. aeruginosa strain ATCC 27853 added to the assay mixture was incubated at 37 °C for 2 h for PA25 multiplication. Following incubation, 2 mL of chloroform was added to the assay mixture to lyse bacteria cells. Vacuum evaporation of solvent was followed by centrifugation at 1800 × g for 15 min to collect cell debris, and then, the supernatant was filtered through 0.22 µm pore-size filters to extract PA25. PA25 suspension was stored at – 20 °C. A volume of 100 µl of the suspension was added to 5 mL of TSB medium previously inoculated with 100 µl of overnight P. aeruginosa ATCC strain 27853 culture and then poured over agar plates and incubated 12 h at 37 °C for plaque formation (Kropinski et al. 2009). A single plaque was picked from the plate and eluted into1 mL of fresh culture of P. aeruginosa ATCC strain 27853. Optical density at 600 nm (OD600) was determined for samples incubated with and without PA25. The mixture was then layered on the agar for plaque formation. Titers of PA25 were determined by plaque-forming units PFU/mL, using the agar overlay method on TSA (Armon and Kott 1993). Serial dilutions were performed, followed by the drop count method (Adams, 1959).
Transmission electron microscopy (TEM)
Electron microscopy (TEM) is an important tool for high-resolution structure determination in applications ranging from condensed matter to biology. Particularly, TEM has provided the fastest diagnostic technique in virology and in phage identification. The major contribution of electron microscopy is the demonstration of the capsid of tailed phages shape and dimension that is very specific to establish differences in phage taxonomy. Methods used are those described by Ackermann (2001). In summary, a volume of 1.5 mL of PA25 suspension was centrifuged at 17,000 × g (4 °C) for one hour (High Speed Brushless Centrifuge MPW-350R). After discarding the supernatant, 1.5 mL of 0.1 M ammonium acetate was added to 100 µl of the pellet, and the mixture was urged again at 17,000 × g (4 °C) for one hour. The pellet was prepared for the microscopic observation with coloration by uranyl acetate 2%, on a carbon grid, mixed with a drop of the PA25 particle. Excess liquid was removed with filter paper, then left to dry in room temperature, and examined at 80 kV by using the JEOL JEM-1230 transmission electron microscope.
One-step growth curve
One-step growth curve was constructed to analyze the life cycle of PA25. PA25 was added to 1 mL of log-phase P. aeruginosa ATCC strain 27853 at a multiplicity of infections of 0.01 and then incubated for 2 h at 37 °C. The unabsorbed PA25 particles were removed by centrifugation at 10 000 × g for 5 min. After suspending in 50 mL of TSA medium, the mixture was incubated at 37 °C with continuous shaking. Samples were collected at intervals of 30 min, and viral abundance was immediately quantified with a double-agar plaque assay.
Bacterial challenge test
Two types of wastewater were used from Charguia Pilot Tunis: Upstream PWW samples and Downstream SWW samples. PA25 bacteriolytic activity was determined in (a) distilled water (DW), (b) sterilized wastewater (StWW) and (c) unsterilized wastewater (UstWW) following the method of Wang et al. (2016). P. aeruginosa ATCC 27853 was incubated into 500 mL TSB medium and grown overnight at 37 °C. Bacteria were harvest from 250 mL saturated TSB culture by centrifugation at 18 000 × g during 30 mn. The pellet was resuspended in three 5-L containers with DW, UstPWW and StPWW, to a density expressed in Colony-forming Unit (CFU) per ml of 6.0 × 103, 1.4 × 105 and 3.0 × 108, respectively. The same experiment was repeated using sterilized secondary wastewater StSWW and unsterilized secondary wastewater UstSWW with initial load of P. aerugnosa of 1.58 × 104 and 2.0 × 105 CFU/mL, respectively. 10 mL of PA25 suspension was added to each 5-L container, corresponding to 105 PFU/mL. All experiments were performed at room temperature around 25 °C during 48 h. Bacteria enumeration with or without PA25 treatment from both PWW and SWW were assayed via serial dilution total plate count (TPC) method using TSA medium (Cabral (2010). All experiments were performed in triplicates. Statistical significance was determined using student-t-test with a P-value threshold of < 0.05.
Effect of PA25 in antibiotic resistance of P. aeruginosa ATCC strain 27853
P. aeruginosa ATCC strain 27853 antibiotic resistance was assayed using two treatments: (1) experimental containers considering 90 mL of sterile wastewater to which was added 10 mL of pellet overnight P. aeruginosa ATCC strain 27853 culture in TSB medium, along with 1 mL of PA25 suspension corresponding to 105 PFU/mL and (2) Control containers with no PA25. Both treatments were incubated for 48 h at 37 °C.
Antibiotics susceptibility of P. aeruginosa ATCC strain 27853 subsequent PA25 contact
Antimicrobial susceptibility was assayed via the agar diffusion method with a disk diffusion test (Acar 2000) using TSA medium and inoculums of 108 CFU/mL (Control treatment) and 104 CFU/mL (Experimental treatment). The following disks for susceptibility testing were used: (a) gentamicin (GM), 15 µg/disk, (b) streptomycin (S10 10 µg/disk); (c) tetracycline (TE), 30 µg/disk; (d) rifampicin (RIF), 30 µg/disk and (e) spiramycin (SP) 100 µg/disk (Oxoid). P. aeruginosa ATCC strain 27853 was plated in Mueller Hinton agar (Biokar, Diagnostic), and the antibiotic disks were placed over the bacteria layer (Nassar et al. 2019). Plates were incubated at 37 °C overnight. After this period, the inhibition zone diameter was measured, and strains classified according to the sensitivity to each antibiotic.
Statistical analysis
All experiments were performed in triplicate. The Student–Newman–Keuls analysis (SPSS 20) software was used to determine the significant differences of removal efficiencies of phage treatment to check the influence of load bacteria variation with a P-value threshold of < 0.05.
Results and discussion
Results
Characterization of bacteriophages PA25 by plaque morphology, and transmission electron Microscopy TEM
Plaque assay of PA25 on P. aeruginosa showed the presence of medium plaques (0.7–0.8 cm) that determined the titer expressed by plaque-forming units PFU/ mL, estimated to be of 105 PFU/mL. (b) Electron micrographs of TEM IBIS, JEOL JEM 1230 of PA25 showed clearly that PA25 appeared as a doubled stranded DNA phage, with Isometric head 61 and 91 nm and Thin, long, noncontractile, and flexible tail with a base plate 110 and 170 nm, classified to belong to the Siphoviridae family, PA25 successfully isolated from SWW was able to undergo the lytic life cycle in P. aeruginosa ATCC27853. The results of OD600 of bacteria culture evolution showed that a 3-h contacting time was necessary to reduce OD600 from 1.6 to 0.6. The titer was determined, by PFU/ mL, estimated to be of 105 PFU/mL (Fig. 1a). PA25 was first characterized by plaque morphology showing medium sized plaques and gave clear-plaques expressing lytic cycles. Based on JEM-1230 transmission electron micrographs (JEOL, Tokyo, Japan), isolated PA25 appeared as a doubled stranded DNA phage, with a head and long tail, classified to belong to the Siphoviridae family (Ackermann 2007; 2009; 2012; Aprea 2015. 2018), similar to lytic Pseudomonas phages isolated previously from central Mexico, reported by Sepúlveda-Robles et al. (2012) describing a novel phage in the Siphoviridae family (Fig. 1b).
Characterization of bacteriophages PA25 by plaque morphology, and transmission electron microscopy (TEM): a Plaque assay of PA25 showing medium plaques (0.7–0.8 cm) on P. aeruginosa that determined the titer expressed by plaque-forming units PFU/ mL, estimated to be of 105 PFU/mL b TEM micrographs of IBIS, JEOL JEM 1230 of PA25 as a doubled stranded DNA phage, with isometric head 61 to 91 nm; a thin, long, noncontractile, and flexible tail with a base plate 110–170 nm, classified as the Siphoviridae family (Ackermann 2007; 2009; 2012; Aprea 2017;2018), Bar length = 200 nm
One-step growth of PA25 lytic of P. aeruginosa ATCC strain 27853
A constructed one-step growth of PA25 showed exponential curves with a short latent period less than 20 min (Fig. 2). The latent PA25 period is defined as the period required between PA25 adsorption and the beginning of the cell disruption.
One-step growth of PA25 lytic phage on P. aeruginosa ATCC strain 27853. A constructed one-step growth of PA25 showed exponential curves with a short latent period less than 20 min (Fig. 2). The latent phage PA25 period is defined as the period required between phage PA25 adsorption and the beginning of the cell disruption
Bacterial challenge test
PA25 was added to treatment of PWW, SWW and sterile distilled water (StDW); all treatments artificially contaminated by P. aeruginosa ATCC 27853. A control without PA25 addition was included. The results showed that bacterial load remain unchanged with PA25 addition to StDW; the discharge was weak, and the bacteria load was 7.00 × 103 and 3.00 × 103, for StDW and StDW + Pg, respectively (Fig. 3). This result may be due to the limited medium for bacteria growth that limited phage load. With the treatments of StSWW and StSWW + Pg, the PA25 does not induce cell lysis and bacteria load was 1.5 × 104 and 1.00 × 104, respectively. However, bacteria density in SWW effluent was reduced when PA25 was added to UStSWW + Pg, P. aeruginosa ATCC strain 27853 density was reduced from 2.00 × 105 to 6.00 × 103 CUF/mL. The effluent bacteria density was greatly reduced when PA25 was added to UStPWW; P. aeruginosa ATCC strain 27853 density to 3.00 × 108 to 2.00 × 103 CUF/mL, for untreated control (no phage) and treatment with PA25, respectively. In contrast, in StPWW the P. aeruginosa ATCC strain 27853 density deceased only slightly from to 1.40 × 105 to 5.00 × 104. The obtained results showed clearly that the sterilization of the sample and artificially inoculation of P. aeruginosa ATCC strain 27853 induced a less efficient reduction of bacteria cells by PA25. This result was consistent for both PWW and SWW. These results showed that PA25 may be added to both treated and untreated wastewater. The greatest reduction in P. aeruginosa ATCC strain 27853 density in effluents comes from adding PA25 to UStPWW compared to UStSWW. This difference in efficiency of PA25 when added to PWW compared to that of SWW may be related to the high organic content of PWW, higher at upstream of the pilot wastewater treatment stream. PA25 appears to thrive in the endogenous microbial community and chemistry of the wastewater samples; P. aeruginosa ATCC strain 27853 lyses easily occurs at the level of PWW. Other methods such chlorination or UV light are limited in their application to PWW, and require additional levels of pre-treatment to be effective. The wastewater disinfection was effective in this study, and PA25 application deserves attention to reduce effluent pollution.

taken from the primary domestic and from the secondary wastewater according to the addition or not of the phage PA25. (Pg): phage PA25, StDW: Sterilized distilled water + P. aeruginosa. StDW + Pg: Sterilized distilled water + P. aeruginosa + Phage PA25 StPWW = Sterilized Primary wastewater + P. aeruginosa StPWW + Pg: Sterilized Primary wastewater + P. aeruginosa + Phage PA25. UstPWW: Unsterilized primary wastewater + P. aeruginosa UstPWW + Pg: Unsterilized primary wastewater + P. aeruginosa + Phage PA25. StSWW = Sterilized Secondary wastewater + P. aeruginosa StSWW + Pg = Sterilized Secondary wastewater + P. aeruginosa + Phage PA25. UstSWW:Unsterilized secondary wastewater + P. aeruginosa UstSWW + Pg Unsterilized secondary wastewater + P. aeruginosa + Phage PA25
Load of P. aeruginosa ATCC strain 27853 over 48 h, in sample
PA25 infection significantly impacted P. aeruginosa ATCC27853 regression. The bacterial density reduction was most dramatic in unsterilized wastewater. It can be explained by the presence of endogenous P. aeruginosa present previously in wastewater associated with P. aeruginosa ATCC27853 strain artificially inoculated in wastewater. Several studies reported linear relationship between phages and bacteria load (Leverentz et al. 2004; Wong et al. 2020). Burmeister et al. (2020) highlighted the abundance and the significant roles played by phage thereby modifying bacterial population structure, composition and dynamic as well as density. Untreated wastewater has a high organic content, thus supports high microbial growth (Al-Rekabi et al. 2007). However, SWW that has aerobic conditions can more easily be treated with phages (Gerba and Pepper 2019; Tan et al. 2020). In fact, phage requires aerated and suspended conditions to effectively attach bacteria hosts; suspended solids and high organic content of wastewater can remove phage from suspension, thus decreasing its efficiency in bacteria removal (Watts et al. 2020). This suggests that phage act on the endogenous P. aeruginosa present normally in wastewater microbial communities. Indeed, bacteria and phages are in a perpetual competition in the environment and continually evolve by modification of cell receptors (bacteria) and binding proteins (phages) (Van Houte et al. 2016; Broniewski, et al. 2020). In fact, phage can influence the abundance, diversity and evolution of bacterial communities (León and Bastías 2015). Several phages have been reported to add virulence factors to their host and to increase bacterial virulence (Brussow et al. 2004). However, lytic phage can also exert a selective pressure allowing the proliferation of strains with reduced virulence (Yuan et al. 2019). In reality, phage use structures present on the bacterial surface as receptors, which can be virulence factors in different bacterial species. Therefore, some strains may modify in these receptors and will be resistant to phage infection and may also exhibit reduced virulence. Silva et al. (2014) showed that the efficiency of phage therapy is highly dependent on different environmental factors such as salinity and the availability of organic matter. In this study, PWW with high organic content showed better performance of the phage in decreasing P. aeruginosa ATCC strain 27853 cells suggesting the use of phage in PWW. Phage is highly specific for bacterial species and multiplies at the expense of the cell, eventually reducing the number of viable bacterial (Sulakvelidze and Morris 2001). Even though the major advantage of phage treatment is phage specificity, since the non-target bacterial populations should remain unaffected, phages should be able of lysing the majority of strains of a given bacterial species.
Antibiotics susceptibility of P. aeruginosa ATCC strain 27853 subsequent to PA25 contact
The results showed that P. aeruginosa ATCC 27853 untreated with PA25 was resistant to tetracycline (TE), streptomycin (S10) and spiramycin (SP), and showed limited sensitivity to gentamicin and rifampicin. However, when PA25 was added to the bacteria suspensions, bacterial resistance to streptomycin (S10) declined. The inhibition zone diameter increased with addition of PA25; indeed, diameter of sensitivity of gentamicin increased from 19 to 21 mm, and from 11.3 to 20 mm for rifampicin. Streptomycin inhibition zone diameter increased to 20 mm (Fig. 4). Antibiotics resistance results are shown in Table 2; PA25 appears to have effectively attacked P. aeruginosa ATCC strain 27853 and the stressed bacteria became more sensitive to antibiotics (considered separately gentamicin and rifampicin).
Photographs of Petri dishes of P. aeruginosa ATCC27853 treated with 5 antibiotics a in the absence of phage PA25 and b in the presence of phage PA25, (G): gentamicin (S): streptomycin; (T):tetracycline (R): rifampicin (S1): spiramycin all discs (Oxoid). P. aeruginosa ATCC 27853 untreated with PA25 phage was resistant to tetracycline: (TE), streptomycin: (S10) and spiramycin: (SP), and showed limited sensitivity to gentamicin and rifampicin. However, when phage PA25 was added to the bacteria suspensions, bacterial resistance to streptomycin (S10) declined. The inhibition zone diameter increased with addition of phage PA25
The effect of phage strains on changing bacterial virulence is still poorly understood (León and Bastías 2015). Recently, there has been an interest in research of the use of phage therapy against pathogenic bacteria (Moghadam et al. 2020). Current study indicates clearly that bacteria associated with phage have altered their sensitivity to streptomycin (S10) compared to resistance without phage treatment. However, Olivares et al. (2020) showed that P. aeruginosa resistant to β-lactams antibiotic may lose this resistance by another way by increasing doses of antibiotics. P. aeruginosa show low resistance to aminoglycoside; studies have found gentamicin (an aminoglycoside) resistance rates ranging from 12 to 22%. Gentamicin is the least active of the aminoglycosides, with lower rates of resistance being reported for tobramycin and amikacin (Lister et al. 2009). But, this kind of resistance is usually enzymatic, thus bacteria do not respond to dose increases. However, Gene PA14_40260-40,230 is part of an operon that encodes a novel efflux pump, and deletion of this operon in P. aeruginosa resulted in a decrease of the resistance of the bacterium to gentamicin and ciprofloxacin in biofilm (Zhang and Mah 2008). This may suggest that the selected phage may contribute to Gene PA14_40260-40,230 deletion. Further research is needed to confirm this hypothesis. Concerning streptomycin antibiotic, it has a unique fixation site which is the ribosomal protein S12. The addition of the phage affects the pathogen load (Levin and Bull 1996) and also limits bacteria resistance to antibiotics (Torres-Barceló et al. 2014). The effect of the phage on bacteria is likely a genetic modification that induces a loss of antibiotic resistance (Levin and Bull 1996). Rifampicine is one of the most potent and broad spectrum antibiotics used against bacterial pathogens and is a key component of anti-tuberculosis therapy. Rifampicine effectiveness stems from its inhibition of the bacterial RNA polymerase (RNAP) (Campbell et al. 2001). P. aeruginosa has been found to be resistant to tetracycline (Dean et al. 2003), streptomycin (Cervantes-Vega et al. 1986) and spiramycin and has a limited sensitivity to rifamycin (Yee et al. 1996), and gentamycin (Kadurugamuwa et al. 1993). In addition, the apparition of strain sensitivity to streptomycin could be due to bacterial response to phage infection (Levin and Bull 1996). Bacterial resistance against these antibiotics has been associated with their overuse in livestock veterinary practices; phages could be a new tool to mitigate antibiotic resistance in P. aeruginosa found in wastewater.
The phage therapy was shown as a potential solution for the antibiotic resistance crisis in clinical medium (Golkar et al. 2014; Berglund, 2015). In wastewater treatment systems, microbial communities may increase genetic diversity linked to antibiotic resistance, and wastewater effluents may consequently impact microorganisms entering the environment and ecological systems (Sorum, 2006), thus impacting pathogen transmission to humans and animals (Brüssow and Kutter, 2005; Cabello, 2006). Inadequate treatment of municipal wastewater may increase the conditions that promote microbial antibiotic resistance through new mutations (Courvalin, 2001). Synergy between judicious antibiotic use generally and phage treatment of wastewater specially could address the threat of bacterial pathogens released in wastewater effluent to improve environmental and human health (Sulakvelidze and Kutter, 2005; Comeau et al. 2008), and that PA25 may be added at the level of PWW and SWW indicating efficiency of PA25 as tool in primary or secondary plant treatment. This study supports the use of PA25 as a tool to fight bacterial infections and should serve as a baseline for additional research for broader wastewater treatment applications to prevent pathogen spreading (Alisky et al. 1998).
Conclusion
In this manuscript, the phage PA25 was isolated from secondary domestic wastewater to remove P. aeruginosa ATCC strain 27853. The effect of PA25 on reducing P. aeruginosa ATCC strain 27853 discharge was more effective when the phage was added in PWW. Also, the isolated PA25 was able to reduce antibiotic resistance in the target bacteria. The result provided should be useful for limiting pathogens spread in environment and should be used as new tool for both wastewater primary treatment and wastewater disinfection.
References
Abedon ST (2020) Phage-phage, phage-bacteria, and phage-environment communication in biocommunication of phages. Springer, pp 23–70
Acar JF (2000) Antibiotic synergy and antagonism. Med Clin North Am 84(6):1391–1406
Ackermann HW (2001) Frequency of morphological phage descriptions in the year 2000. AdvVirol 146(5):843–857
Ackermann HW (2007) 5500 Phages examined in the electron microscope. Arch Virol 152:227–243. https://doi.org/10.1007/s00705-006-0849-1
Ackermann HW (2009) Basic phage electron microscopy. Humana press, pp 113–126
Ackermann HW (2012) Bacteriophage electron microscopy. Adv Virus Res 82:1–32
Ackermann HW, DuBow MS (1987) Viruses of prokaryotes. CRC Press, General properties bacteriophages 1:86
Adams MH (1959) Methods of study of bacterial viruses Bacteriophages. Interscience Publishers
Al-Abri M, Al-Ghafri B, Bora T, Dobretsov S, Dutta J, Castelletto S, Boretti A (2019) Chlorination disadvantages and alternative routes for biofouling control in reverse osmosis desalination. NPJ Clean Water 2(1):1–16
Alisky J, Iczkowski K, Rapoport A, Troitsky N (1998) Bacteriophages show promise as antimicrobial agents. J Infect 36(1):5–15
Aprea G, ARDPrencipe VA, Migliorati G (2015) Bacteriophage morphological characterization by using transmission electron microscopy. J Life Sci 9:214–220
Aprea G, D’Angelantonio D, Boni A, Connerton P, Connerton I, Scattolini S, Marotta F, Pomilio F, Migliorati G, D’Alterio N, Di Giannatale E (2018) Isolation and morphological characterization of new bacteriophages active against campylobacter jejuni. Am J ClinMicrobiolAntimicrob 1(1):1004
Al-Rekabi WS, Qiang H, Qiang WW (2007) Improvements in wastewater treatment technology. Pak J Nutr 6(2):104–110
Armon R, Kott Y (1993) A simple, rapid and sensitive presence/absence detection test for bacteriophage in drinking water. J ApplBacteriol 74(4):490–496
Berglund B (2015) Environmental dissemination of antibiotic resistance genes and correlation to anthropogenic contamination with antibiotics. Infect EcolEpidemiol 5(1):1–28564
Bradbury J (2004) My enemy’s enemy is my friend Using phages to fight bacteria. Lancet 363:624–625
Brizou F (1972) Famille de Pseudomonacae dans « bactériologie médicale ». Editions médicales Flammarison, 469–1523.
Broniewski JM, Meaden S, Paterson S, Buckling A, Westra ER (2020) The effect of phage genetic diversity on bacterial resistance evolution. ISME J 14(3):828–836
Brüssow H, Kutter E (2005) Phage ecology. Bacteriophages BiolAppl 70:129–164
Brussow H, Canchaya C, Hardt WD (2004) Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. MicrobiolMolBiol 68(3):560–602
Burmeister AR, Fortier A, Roush C, Lessing AJ, Bender RG, Barahman R et al (2020) Pleiotropy complicates a trade-off between phage resistance and antibiotic resistance. Proc Nat Acad Sci 117(21):11207–11216
Cabello FC (2006) Heavy use of prophylactic antibiotics in aquaculture: a growing problem for human and animal health and for the environment. Environ Microbiol 8(7):1137–1144
Cabral JP (2010) Water microbiology bacterial pathogens and water. Int J Environ Res Publ Health 7(10):3657–3703
Campbell EA, Korzheva N, Mustaev A, Murakami K, Nair S, Goldfarb A, Darst SA (2001) Structural mechanism for rifampicin inhibition of bacterial RNA polymerase. Cell 104(6):901–912
Cervantes-Vega C, Chávez J, Rodríguez MG (1986) Antibiotic susceptibility of clinical isolates of Pseudomonas aeruginosa. Antonie Van Leeuwenhoek 52:319–324
Cisek AA, Dąbrowska I, Gregorczyk KP, Wyżewski Z (2017) Phage therapy in bacterial infections treatment: one hundred years after the discovery of bacteriophages. CurrMicrobiol 74(2):277–283
Comeau AM, Tétart F, Trojet SN, Prère MF, Krisch HM (2008) La « synergiephages-antibiotiques » Un enjeu pour la phagothérapie. Médecine/sciences 24(5):449–451
Courvalin P (2001) Resistance aux antibiotiques. SocMicrobiol 45:1007–1031
Danovaro R, Fonda US, Pusceddu A (2009a) Climate change and the potential spreading of marine mucilage and microbial pathogens in the Mediterranean Sea. PLoS ONE 4:e7006. https://doi.org/10.1371/journal.pone.0007006
Danovaro R, Umani SF, Pusceddu A (2009) Climate change and the potential spreading of marine mucilage and microbial pathogens in the Mediterranean Sea. Plos One 4(9):256
Dean CR, Visalli MA, Projan SJ, Sum PE, Bradford PA (2003) Efflux mediated resistance to tigecycline (GAR-936) in Pseudomonas aeruginosa PAO1. Antimicrob Agents Chemother 47(3):972–978
Ebrahimi A, Arvaneh Z, Mahzounieh M, Lotfalian S (2017) Antibiotic resistance induction by benzalkonium chloride exposure in nosocomial pathogens. Int J Infect 4(2):e40296. https://doi.org/10.5812/iji.40296
Gerba CP, Pepper IL (2019) Municipal wastewater treatment in environmental and pollution science. Academic Press
Golkar Z, Bagasra O, Pace DG (2014) Bacteriophage therapy: a potential solution for the antibiotic resistance crisis. J Infect Develop Ctries 8(02):129–136
Kadurugamuwa JL, Clarke AJ, Beveridge TJ (1993) Surface action of gentamicin on Pseudomonas aeruginosa. J Bacteriol 175(18):5798–5805
Kamizoulis G, Saliba L (2004) Development of coastal recreational water quality standards in the Mediterranean. Environ Int 30(6):841–854
Kropinski AM, Mazzocco A, Waddell TE, Lingohr E, Johnson RP (2009) Enumeration of bacteriophages by double agar overlay plaque assay in bacteriophages. Humana Press, pp 69–76
Labrie M, Reguiba N, Tremblay N (2009) ManuelleApprentissage par problèmes à l’Université du Québec à Montréal (UQAM). Point Biolo 3:1–16
León M, Bastías R (2015) Virulence reduction in bacteriophage resistant bacteria. Front Microbiol 6:1–343
Leverentz B, Conway WS, Janisiewicz W, Camp MJ (2004) Optimizing concentration and timing of a phage spray application to reduce Listeria monocytogenes on honeydew melon tissue. J Food Prot 67(8):1682–1686
Levin BR, Bull JJ (1996) Phage therapy revisited: the population biology of a bacterial infection and its treatment with bacteriophage and antibiotics. Am Nat 147(6):881–898
Lister PD, Wolter DJ, Hanson ND (2009) Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. ClinMicrobiol Rev 22(4):582–610
Mansour W (2018) Tunisian antibiotic resistance problems: three contexts but one health. Afr Health Sci 18(4):1202–1203
Merril CR, Scholl D, Adhya SL (2003) The prospect for bacteriophage therapy in Western medicine. Nat Rev Drug Discovery 2(6):489–497
Moghadam MT, Amirmozafari N, Shariati A, Hallajzadeh M, Mirkalantari S, Khoshbayan A, Jazi FM (2020) How phages overcome the challenges of drug resistant bacteria in clinical infections. Infection and Drug Resistance 13:1–45
Nassar MS, Hazzah WA, Bakr WM (2019) Evaluation of antibiotic susceptibility test results: how guilty a laboratory could be? J Egypt Public Health Assoc 94(1):4
Olivares E, Badel-Berchoux S, Provot C, Prévost G, Bernardi T, Jehl F (2020) Clinical impact of antibiotics for the treatment of Pseudomonas aeruginosa biofilm infections. Front Microbiol 10:2894
Petrovski S, Seviour RJ, Tillett D (2011) Genome sequence and characterization of the Tsukamurella bacteriophage TPA2. Appl Environ Microbiol 77(4):1389–1398
Prieto MD, Lopez B, Juanes JA, Revilla JA, Llorca J, Delgado-Rodriguez M (2001) Recreation in coastal waters: health risks associated with bathing in sea water. J Epidemiol Community Health 55(6):442–447
Pukatzki S, Kessin RH, Mekalanos JJ (2002) The human pathogen Pseudomonas aeruginosa utilizes conserved virulence pathways to infect the social amoeba Dictyosteliumdiscoideum. ProcNatlAcadSci 99(5):3159–3164
Rivera-Utrilla J, Sánchez-Polo M, Ferro-García MÁ, Prados-Joya G, Ocampo-Pérez R (2013) Pharmaceuticals as emerging contaminants and their removal from water. Rev Chemos 93(7):1268–1287
Saidi N, Kouki S, Mehri I, Benrajeb A, Bellila A, Abdennaceur H, Ouzari HI (2011) Biofilm and siderophore effects on secondary waste water disinfection. CurrMicrobiol 63:337–340
Schwartz T, Armant O, Bretschneider N, Hahn A, Kirchen S, Seifert M, Dötsch A (2015) Whole genome and transcriptome analyses of environmental antibiotic sensitive and multi-resistant Pseudomonas aeruginosa isolates exposed to waste water and tap water. MicrobBiotechnol 8(1):116–130
Sepúlveda-Robles O, Kameyama L, Guarneros G (2012) High diversity and novel species of Pseudomonas aeruginosa bacteriophages. Appl Environ Microbiol 78(12):4510–4515
Silva YJ, Costa L, Pereira C, Cunha Â, Calado R, Gomes NC, Almeida A (2014) Influence of environmental variables in the efficiency of phage therapy in aquaculture. MicrobBiotechnol 7(5):401–413
Slekovec C, Plantin J, Cholley P, Thouverez M, Talon D, Bertrand X, Hocquet D (2012) Tracking down antibiotic-resistant Pseudomonas aeruginosa isolates in a wastewater network. PloS one 7(12):e49300
Sorum H (2006) Antimicrobial drug resistance in fish pathogens. In: Aarestrup FM (ed) Antimicrobial resistance in bacteria of animal origin. American Society for Microbiology Press, pp 213–238
Sulakvelidze A, Kutter E (2005) Bacteriophage therapy in humans bacteriophages biology and applications. CRC Press
Sulakvelidze A, Morris JG (2001) Bacteriophages as therapeutic agents. Ann Med 33:507–509
Summers WC (2005) Bacteriophage research: early history. Bacteriophages: Biology and applications., pp. 5-27
Tan X, Li H, Li X, Sun W, Jin C, Chen L, Sun C (2020) A novel isophorone wastewater treatment technology-wet electrocatalytic oxidation and its degradation mechanism study. J Hazard Mater 1:122035
Torres-Barceló C, Arias-Sánchez FI, Vasse M, Ramsayer J, Kaltz O, Hochberg ME (2014) A window of opportunity to control the bacterial pathogen Pseudomonas aeruginosa combining antibiotics and phages. PloS one 9(9):1056
Tsamba L, Correc O, Couzinet A (2020) Chlorination by-products in indoor swimming pools: development of a pilot pool unit and impact of operating parameters. Environ Int 137:105566
Van Houte S, Ekroth AK, Broniewski JM, Chabas H, Ashby B, Bondy-Denomy J, Westra ER (2016) The diversity-generating benefits of a prokaryotic adaptive immune system. Nature 532(7599):385–388
Wang Z, Zheng P, Ji W, FuQYan WHY, Sun J (2016) SLPW: A virulent bacteriophage targeting methicillin-resistant Staphylococcus aureus in vitro and in vivo. Front Microbiol 7:1–934
Watts S, Julian TR, Maniura-Weber K, Graule T, Salentinig S (2020) Colloidal transformations in MS2 virus particles: driven by pH, influenced by natural organic matter. ACS Nano 14(2):1879–1887
Withey S, Cartmell E, Avery LM, Stephenson T (2005) Bacteriophages—potential for application in wastewater treatment processes. Sci Total Environ 339(1–3):1–18
Wong CW, Delaquis P, Goodridge L, Lévesque RC, Fong K, Wang S (2020) Inactivation of Salmonellaenterica on post-harvest cantaloupe and lettuce by a lytic bacteriophage cocktail. Curr Res Food Sci 2:25–32
Yee Y, Kisslinger B, Yu V, Jin D (1996) A mechanism of rifamycin inhibition and resistance in Pseudomonas aeruginosa. Antimicrob Chemoth 38:133–137
Yuan Y, Qu K, Tan D, Li X, Wang L, Cong C, Xu Y (2019) Isolation and characterization of a bacteriophage and its potential to disrupt multi-drug resistant Pseudomonas aeruginosa biofilms. MicrobPathog 128:329–336
Zhang L, Mah TF (2008) Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J Bacteriol 190(13):4447–4452
Acknowledgments
This study was supported by a grant from Tunisian Ministry of Higher Education and Scientific Research in the ambit of LR19CERTE04 (2o19_2022 Programs). Authors are Grateful for technical team of CERTE, particularly to Miss Nessrine Chourabi. We thank Pr. Sylvain Moineau and Pr Josée Harel and members of their teams for discussion at the beginning of the project and for phage identification. Also, authors are grateful for Pr Steven Aust for revision and English improvement.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Editorial Responsiblity: Parveen Fatemeh Rupani.
Rights and permissions
About this article
Cite this article
Grami, E., Salhi, N., Sealey, K.S. et al. Siphoviridae bacteriophage treatment to reduce abundance and antibiotic resistance of Pseudomonas aeruginosa in wastewater. Int. J. Environ. Sci. Technol. 19, 3145–3154 (2022). https://doi.org/10.1007/s13762-021-03366-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-021-03366-3