Abstract
An investigation was carried out to compare the ability of two bacteria Pseudomonas aeruginosa PSA5 and Rhodococcus sp. NJ2 isolated from petroleum sludge for degradation of benzo(a)pyrene [B(a)P], a HMW PAH compound in MSM. During 25 days of incubation, 50 ppm B(a)P was degraded by 88 and 47 % by P. aeruginosa PSA5 and Rhodococcus sp. NJ2, respectively. Besides, involvement of different catabolic enzymes, that is, salicylate hydroxylase, 2-carboxybenzaldehyde dehydrogenase, catechol 1,2-dioxygenase and catechol 2,3-dioxygenase, was also examined to identify their differential role in B(a)P degradation. Among these enzymes, the highest induction of 2-carboxybenzaldehyde dehydrogenase (773.5 nmol mg−1 protein) was recorded in P. aeruginosa PSA5, while salicylate hydroxylase was highly expressed (839.6 nmol mg−1 protein) in Rhodococcus sp. NJ2. Both the bacteria were found biosurfactant (glycolipid) producing, and role of biosurfactant in PAH degradation was also ascertained by reduced surface tension, higher emulsification index and increased cell surface hydrophobicity.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Polycyclic aromatic hydrocarbons (PAHs) are released from various anthropogenic sources like crude oil spillage, waste solid incineration, incomplete combustion of fossil fuels, etc. (Juhasz and Naidu 2000). Hence, they are ubiquitous in the soil environment and tend to accumulate in the top soil mainly around industrial and urban activities (Johnsen and Karlson 2007). Persistence of high molecular weight (HMW) PAHs in the environment mainly due to dense clouds of π-electrons on both sides of the ring structures is a serious environmental concern which needs to be addressed for ensuring human health (Long et al. 2008).
Among PAHs, benzo[a]pyrene is known for carcinogenic, genotoxic and cytotoxic properties (Hsu et al. 2005). Although several reports are available for microbial biodegradation of HMW PAH compounds (Sayara et al. 2010), their degradation still continues to be unreliable due to non-availability of potential soil microbes with appropriate catabolic pathways and their low bioavailability (Cerniglia 1992). However, benzo(a)pyrene [B(a)P] degradation has been reported by Sphingomonas yanoikuyae JAR02 and white rot fungus Armillaria sp. F022 (Rentz et al. 2008; Hadibarata and Kristani 2012).
At times, degradation of HMW PAHs by microbe is inhibited by the presence of more water-soluble PAHs via competition at active sites of oxygenases or through the accumulation of toxic products following co-metabolism of other PAHs (Bouchez et al. 1995). Initial dioxygenase is a key enzyme for attacking the aromatic ring structure of PAHs under aerobic condition (Juhasz and Naidu 2000). Dioxygenase genes, such as NahAc, phnAc, NidA and pdoB, were used to detect PAH-degrading Gram-negative and Gram-positive bacteria (Widada et al. 2002; Sho et al. 2004). Bacteria secrete many other degradative enzymes, growth factors and biosurfactants for fast utilization of PAHs (Mukherjee and Das 2005). Biosurfactants produced by microorganisms are amphiphilic in nature which reduce surface tension, critical micelle concentration (CMC) and interfacial tension (IFT) in both aqueous solutions and hydrocarbon mixtures and thus facilitate PAH degradation (Maier 2003; Desai and Banat 1997).
In the present investigation, a comparison of intrinsic ability of two bacterial strains PSA5 and NJ2 isolated from petroleum sludge was made for degradation of B(a)P in MSM. Besides, differential expression of catabolic enzymes such as salicylate hydroxylase, 2-carboxybenzaldehyde dehydrogenase, catechol 1,2-dioxygenase and catechol 2,3-dioxygenase was also studied to investigate their role in B(a)P degradation. A link between nature and extent of biosurfactant production was explored with cell surface hydrophobicity, surface tension and emulsification index which affect bioavailability of PAHs. This study was carried out at NBRI during June–July, 2012.
Materials and methods
Chemicals and media
Benzo(a)pyrene (with >97.0 % purity) and HPLC grade Acetonitrile were procured from (Sigma Aldrich, USA), while mineral salt medium (MSM) was obtained from Hi media.
Bacterial isolation and screening
Bacteria were isolated from oily sludge collected from Barauni oil refinery, Barauni (Bihar, India), through enrichment in 20 ml MSM supplemented with 1 g of oily sludge in 100 ml Erlenmeyer flask incubated in an orbital shaker for 7 days set at 37 °C and 150 rpm. Subsequently, isolated bacterial strains were screened on the basis of degradation of B(a)P (25–200 ppm) in MSM as a sole carbon source after incubation for 7 days in an orbital shaker set at 37 °C and 150 rpm. Out of 12 bacterial strains, two were selected as potential degraders of B(a)P and designated as PSA5 and NJ2.
Experimental setup
Preparation of bacterial inoculum
Bacteria were grown in 30 ml NB (nutrient broth) with 50 ppm B(a)P to attain the exponential growth phase. Cell density of the suspension at this stage was found to be OD = 1 at 600 nm as measured by UV–Vis spectrophotometer (Perkin Elmer Lambda 35). After this, the cells were harvested by centrifugation at 10,000 rpm for 10 min, washed with sterile MSM and concentrated in 10 ml of MSM to be used as inoculum.
Biodegradation of benzo(a)pyrene [B(a)P]
In order to measure the potential of bacterial isolates (PSA5 and NJ2) for B(a)P degradation in MSM, 200 μl inoculum of each isolate was added separately to 10 ml MSM (pH 7.2) with 50 ppm B(a)P as a substrate in 100 ml flasks. For reference, a control was also set up in the same way without bacterial inoculum. These flasks were incubated at 37 °C in an orbital shaker at 150 rpm for 25 days. After every 5 days, residual B(a)P in MSM was extracted thrice with 10 ml DCM to ensure complete extraction. The extracts were concentrated to 2 ml by evaporation and quantified by injecting 20 μl of the samples to HPLC (Dionex ultimate 3000) with a reverse-phase ODS-C18 column (5 μm, 250 × 4.6 mm) and PDA detector (254 nm) using a gradient mixture of H2O and acetonitrile (50:50–85:50) in mobile phase at a flow rate of 1.5 ml min−1. The biodegradation efficiency (BE) of the bacterial strains was calculated by using the following formula:
where C 0 is initial concentration of PAH (μg g−1) and C e is equilibrium concentration of PAH (μg g−1).
Bacterial growth and protein analysis
The growth of bacterial strains (PSA5 and NJ2) was monitored at 600 nm by UV–Vis spectrophotometer during the degradation of B(a)P in sterile MSM after every 5 days interval of incubation. For the estimation of protein, cells of the bacterial isolates (PSA5 and NJ2) grown in MSM with 50 ppm B(a)P were harvested, suspended and washed in potassium phosphate buffer, pH 7. Subsequently, cells were sonicated and centrifuged at 20,000 rpm at 4 °C for 25 min to remove the cell debris. The supernatant was then stored at 2 °C, and the protein estimation of the cell extract was carried out following the Lowry et al. (1951) method using bovine serum albumin (BSA) as a standard at 660 nm using UV–Vis spectrophotometer.
Enzyme extraction
For the extraction of enzymes after every 5 days intervals, the cells were harvested and washed twice with 2 ml of 50 mM sodium phosphate buffer (pH-7) and resuspended in the same buffer (pH-7) and then homogenized by sonicator for 5 min. Sonicated sample was centrifuged for 25 min at 20,000 rpm at 4 °C. After removal of the cell debris by centrifugation, the supernatant was stored at 2 °C and was further used for different enzyme assays.
Enzyme assays
Activities of ortho-cleaving catechol 1,2-dioxygenase (decycling) (EC 1.13.11.1) and meta-cleaving enzyme catechol 2,3-dioxygenase (decycling) (EC 1.13.11.2) were measured by following the methods of Ngai et al. (1990), Sala-Trepat and Evans (1971), respectively. Further, the activity of 2-carboxybenzaldehyde dehydrogenase was determined by the method of Kiyohara and Nagao (1978) while salicylate hydroxylase [EC 1.14.13.1] activity was measured as described by Yamamoto et al. (1965).
Screening of bacteria for biosurfactant
Bacterial isolates were tested for glycolipid production using the method of Siegmund and Wagner (1991). Subsequently, biosurfactant was extracted in chloroform and methanol (2:1) following the method of Rahman et al. (2003) and then quantified.
Presence of moiety in biosurfactant
Sugar moiety of biosurfactant was tested by molish reagent using the method described by Nelson and Cox (2005) while protein moiety in biosurfactant was tested by ninhydrin, following the method Noudeh et al. (2005). Similarly, for the presence of lipid moiety in biosurfactant, the culture supernatant and water was mixed in 1:1 ratio in a test tube and shaken gently. Then, 3 drops of 2 % Sudan III stain solution (in ethanol) were added, and formation of red layer on the upper surface of the supernatant indicated presence of lipid moiety.
Quantitative estimation of carbohydrate (anthrone method)
Quantitative analysis of carbohydrate is based on the reaction of glucose with anthrone (9,10-dihydro-9-oxoanthracene) in the presence of a strong acid. The carbohydrate content of the isolated biosurfactant was determined by the anthrone reagent by UV–Vis spectrophotometer at 620 nm as described by Spiro (1966).
Surface tension and surface activity
Surface tension of the MSM was measured by stalagmometer before and after biodegradation of B(a)P. The following formula was used for the calculation of surface tension and surface activity.
where σ = surface tension of MSM with bacterial culture, \( \sigma_{{{\text{H}}_{ 2} {\text{O}}}} \) = surface tension of water, m = the mass of MSM with bacterial culture, \( m_{{{\text{H}}_{ 2} {\text{O}}}} \) = mass of the water and surface activity = surface tension of uninoculated medium − surface tension of inoculated medium.
Cell surface hydrophobicity (CSH)
The BATH test was used to determine changes in CSH during bacterial growth in liquid MSM media with benzene as a carbon source. This test was carried out using the method of Rosenberg et al. (1980).
Emulsification index
Emulsification index of culture samples with kerosene, hexadecane, xylene, olive oil, petrol, diesel and benzene was measured using the method as described by Cooper and Goldenberg (1987).
Estimation of emulsification activity
Emulsification activity of biosurfactant with different hydrocarbons like phenanthrene, anthracene, fluoranthene, pyrene, fluorene, B(a)P and chrysene was measured as described by Rosenberg et al. (1979).
Detection of meta-dioxygenase assay
In order to know whether meta-cleavage pathway was involved in B(a)P degradation, a method of Kim and Zylstra (1995) was followed.
Identification of bacterial strains
These strains were identified as Pseudomonas aeruginosa PSA5 and Rhodococcus sp. NJ2 on the basis of their homology (>99 %) of DNA sequence with NCBI databases of bacteria following 16s ribosomal DNA technology (Chromous Biotech, Bangalore, India) which have been reflected as supplementary data.
Results and discussion
Biodegradation of B(a)P
Degradation study reveals that P. aeruginosa PSA5 and Rhodococcus sp. NJ2 degraded B(a)P by 19 and 16 %, respectively, in 5 days of incubation despite its recalcitrant nature. Beyond this period also, degradation continued to increase with the incubation period. After 25 days of incubation, P. aeruginosa PSA5 degraded B(a)P by 88 %, while Rhodococcus sp. NJ2 could degrade only 47 % (Fig. 1a). This clearly indicated an intrinsic difference in the ability of two bacteria in B(a)P degradation. However, in control, 13 % degradation of B(a)P was observed due to abiotic factor. Thus, P. aeruginosa PSA5 showed more BE than Rhodococcus sp. NJ2.
Luo et al. (2009) have observed 20.98 % of B(a)P degradation by Ochrobactrum sp. BL01 strain and 44.07 % by Pseudomonas fluorescens BL03 strain after 14 days of incubation. When a comparison was made about the degradation ability of our strains with Mycobacterium vanbaalenii PYR-1, it was found that B(a)P (50 μg ml−1) was degraded by 88 and 47 % by P. aeruginosa PSA5 and Rhodococcus sp. NJ2, respectively, during 25 days of incubation against 24.7 % degradation by M. vanbaalenii PYR-1 in 14 days of incubation period. This clearly indicated that PSA5 and NJ2 strains have higher degradation ability than M. vanbaalenii PYR-1 as reported by Heitkamp and Cerniglia (1988).
Moody et al. (2004) have elucidated the mechanism of B(a)P degradation by M. vanbaalenii PYR-1 which initiates its attack on B(a)P by dioxygenation or monooxygenation at C-4, 5, C-9, 10 and C-11, 12. Different metabolites, formed during B(a)P degradation by M. vanbaalenii PYR-1, have been identified as benzo(a)pyrene-11,12-epoxide, benzo(a)pyrene-cis-4,5-dihydrodiol, benzo(a)pyrene-cis-11,12-dihydrodiol, benzo(a)pyrene-trans-11,12-dihydrodiol, 11,12-dihydroxybenzo(a)pyrene, hydroxymethoxybenzo(a)pyrene, dimethoxybenzo(a)pyrene, 4,5-dihydroxybenzo(a)pyrene, 4-formylchrysene-5-carboxylic acid, 4,5-chrysene-dicarboxylic acid, chrysene-4 or 5 carboxylic acid, cis-4-(8-hydroxypyren-7yl)-2-oxobut-3-enoic acid and 10-oxabenzo(def)chrysene-9-one.
Growth kinetics
Cells of P. aeruginosa PSA5 and Rhodococcus sp. NJ2 continued to multiply in MSM with 50 ppm B(a)P during incubation period. However, the growth of Rhodococcus sp. NJ2 was recorded faster than P. aeruginosa PSA5 (Fig. 1b). However, in control, the bacterial growth was not significant due to nonavailability of B(a)P as a carbon source.
Cell protein in both the bacteria continued to increase with cell multiplication, attaining maximum value of protein 3.62 and 3.2 mg ml−1 in Rhodococcus sp. NJ2 and P. aeruginosa PSA5, respectively, after 15 days of incubation (Fig. 1c).
Degradative enzymes
In B(a)P degradation, many enzymes may be involved, but in this study activities of only four enzymes salicylate hydroxylase, 2-carboxybenzaldehyde dehydrogenase, catechol 1,2-dioxygenase and catechol 2,3-dioxygenase actually involved in degradation process were investigated.
Salicylate hydroxylase
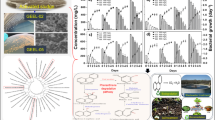
It was observed that salicylate hydroxylase was induced during B(a)P degradation in both the bacteria, but at different incubation periods. Rhodococcus sp. NJ2 exhibited maximum induction of salicylate hydroxylase (839 nmol mg−1 protein) after 10 days of incubation, while P. aeruginosa PSA5 showed a peak induction of this enzyme (662.28 nmol mg−1 protein) after 15 days of incubation period (Fig. 2a). This indicates that salicylate hydroxylase was induced more in Rhodococcus sp. NJ2, than in P. aeruginosa PSA5 during B(a)P degradation.
Salicylate hydroxylase is mainly involved in the conversion of salicylate to catechol in B(a)P degradation and forms a reduced enzyme substrate complex in the presence of NADH. Subsequently, molecular oxygen binds to the complex for production of catechol, CO2 and H2O.
There are two pathways for the microbial degradation of naphthalene, in which salicylate is formed as an intermediate. In the classical pathway, as studied extensively in Pseudomonas sp., salicylate undergoes oxidative decarboxylation by salicylate 1-hydroxylase to produce catechol (Yen and Serdar 1998). However, an alternative pathway was reported in Rhodococcus sp., which involves a salicylate 5-hydroxylase yielding gentisate for naphthalene degradation (Allen et al. 1997).
2-Carboxybenzaldehyde dehydrogenase (2-CBD)
This enzyme was more expressed in P. aeruginosa PSA5 than in Rhodococcus sp. NJ2. The activity of 2-CBD continued to increase with incubation period attaining a peak (973.5 nmol mg−1 protein) after 15 days of incubation in P. aeruginosa PSA5, while in Rhodococcus sp. NJ2, highest 2-CBD activity was recorded 775.9 nmol mg−1 protein after 20 days (Fig. 2b).
Induction of the 2-CBD activity indicates the formation of phenanthrene in degradation pathway of B(a)P. Krishnan et al. (2004) reported two major pathways for phenanthrene degradation in Pseudomonas PP2 strain. Both pathways generate 1-hydroxy 2-naphthoic acid (1-H-2-NA) via 3,4-dihydroxyphenanthrene which is further oxidized by two different routes. Route 1 leads to complete mineralization of 1-H-2-NA to TCA cycle intermediates via 1,2-dihydroxynaphthalene, salicylate and catechol, while route 2 partially degrades 1-H-2-NA to 2-carboxybenzapyruvate and 2-carboxybenzaldehyde and finally to o-phthalic acid.
Catechol 1,2-dioxygenase (C12O)
As compared to salicylate hydroxylase and 2-CBD, C12O was less induced, and its peak induction was recorded after 15 days of incubation period in both the bacteria. The highest induction of this enzyme was found to be 280.9 nmol mg−1 protein in P. aeruginosa PSA5 and 174.1 nmol mg−1 protein in Rhodococcus sp. NJ2 (Fig. 2c).
Ring-cleaving dioxygenases play an important role in the degradation of aromatic compounds by incorporating O2 into the aromatic nucleus which results oxidation of aromatic ring (Goyal and Zylstra 1996). A terminal dioxygenase has been found to be an important factor in PAH degradation in M. vanbaalenii’s pathway to transform B(a)P into benzo[a]pyrene-cis-4R,5S-dihydrodiol and benzo[a]pyrene cis-4S,5R-dihydrodiol.
Schneider et al. (1996) found Mycobacterium sp. Strain RJGII-135 highly capable of transforming benzo[a]pyrene to initial ring oxidation and ring cleavage products by initial enzymatic attack at C-4, 5, C-7, 8 and/or C-9, 10 of benzo[a]pyrene. However, only benzo[a]pyrene-cis-7,8-dihydrodiol was identified. It was suggested that the formation of 4,5-chrysene-dicarboxylic acid is the ortho-cleavage ring fission product of benzo[a]pyrene-cis-4,5-dihydrodiol.
Catechol 2,3-dioxygenase (C23O)
During B(a)P degradation, an induction of C23O was not as high as C12O in both the bacteria. In P. aeruginosa PSA5, a maximum induction of this enzyme was found to be 172.9 nmol mg−1 protein after 20 days of incubation period, while in Rhodococcus sp. NJ2, its highest activity (116.3 nmol mg−1 protein) was observed after 15 days of incubation (Fig. 2d).
Schneider et al. (1996) observed the transformation of B(a)P through meta-cleavage by Mycobacterium sp. Strain RJGII-135. During meta-cleavage of B(a)P, they suggested formation of either cis-4-(8-hydroxypyren-7-yl)-2-oxobut-3-enoic acid or cis-4-(7-hydroxypyren-8-yl)-2-oxobut-3-enoic acid and either 7,8-dihydropyrene-7-carboxylic acid or 7,8-dihydropyrene-8-carboxylic acid as intermediates. The meta-fission products were formed after initial dioxygenation and subsequent dehydrogenation at C-7, 8 and/or C-9, 10.
It was observed from the activity of degradative enzymes that out of four, three enzymes, 2-CBD, C12O and C23O, were highly expressed in P. aeruginosa PSA5 and salicylate hydroxylase in Rhodococcus sp. NJ2, in B(a)P degradation after 25 days of incubation. The genes for degradation of HMW PAH such as pyrene and chrysene were found to be inducible and chromosomally located (Kim et al. 2006, 2007), while genes responsible for degradation of low molecular weight PAH such as naphthalene, phenanthrene and anthracene occur on plasmids (Mallick et al. 2007). This observation was further confirmed by Lily et al. (2010) in relation to B(a)P degradation by Bacillus subtilis BMT4i (MTCC 9447).
pH of the media
No significant change in pH of the media was recorded during B(a)P degradation by Rhodococcus sp. NJ2. However, in P. aeruginosa PSA5, the medium pH was changed to acidic side (6.0) (Fig. 1d).
Role of biosurfactant
Both bacteria were found glycolipid producing. Although P. aeruginosa PSA5 produced rhamnolipid, but Rhodococcus sp. NJ2 did not produce rhamnolipid as evidenced by the formation of a blue halo around the colony of P. aeruginosa which indicated the presence of an anionic surfactant, that is, rhamnolipid. When moieties of biosurfactant produced by bacteria were examined, both indicated sugar and lipid moieties confirming glycolipid production. The carbohydrate content was found to be 650.63 mg ml−1 in P. aeruginosa PSA5 and 215.06 mg ml−1 in Rhodococcus sp. NJ2. It is possible that higher degradability of B(a)P by P. aeruginosa PSA5 is linked to high production of rhamnolipid evidenced by enhanced production of carbohydrates.
Several species of P. aeruginosa have been reported to produce rhamnolipid, which increases the solubility and bioavailability of petroleum hydrocarbons for enhanced biodegradation of diesel contamination in soil (Kojima et al. 1961). Different trehalose containing glycolipids are produced by several other microorganisms belonging to mycolates group, such as Arthrobacter, Nocardia, Rhodococcus and Gordonia.
Surface tension and surface activity
When surface tension and surface activity of the MSM inoculated with two bacteria were measured, it was observed that surface tension decreased with the incubation period, while the surface activity increased. As evident from Fig. 3a, b, the surface tension was decreased from 90.8 to 36 mN m−1 in P. aeruginosa PSA5, while in Rhodococcus sp. NJ2, it declined from 90.7 to 58.2 mN m−1 as compared to control (82.3 mN m−1). Similarly, Noudeh et al. (2010) also observed a sharp decline in surface tension from 60 to 35 mN m−1 in 24 h fermentation by P. aeruginosa PTCC 1561. However, surface activity increased with incubation period in both the bacteria, showing an inverse relationship with surface tension.
Low molecular weight biosurfactants (e.g., glycolipids and lipopeptides) having high surface activity are able to reduce the surface tension of water to 25–30 mN m−1, which facilitates the uptake of hydrocarbons into bacterial cells for enzymatic degradation (Franzetti et al. 2010).
Cell surface hydrophobicity (CSH)
Cell surface hydrophobicity is an important factor in determining hydrocarbon adhesion to cell surface in place of surfaces. When cell surface hydrophobicity of bacterial cultures was measured, P. aeruginosa PSA5 showed a maximum hydrophobicity of 51 %, while Rhodococcus sp. NJ2 exhibited hydrophobicity of 47 % for benzene.
The ability of different microorganisms to access hydrocarbons depends on their cell surface hydrophobicity. Microorganisms with high CSH directly contact oil drops and solid hydrocarbons, while low CSH permits the adhesion of microbial cells to the micelles or emulsified oils, formed due to the presence of extracellular biosurfactants or bioemulsifiers (Van Hamme et al. 2003).
Chang et al. (2009) demonstrated that the cell surface hydrophobicity was enhanced by the accumulation of different fatty acids on the cell surface during growth of Rhodococcus erythropolis NTU-1 on hydrocarbon. Zhong et al. (2007) showed that the adsorption of dirhamnolipid biosurfactants on cells of Bacillus subtilis, P. aeruginosa and Candida lipolytica affected the cell surface hydrophobicity depending on the rhamnolipid concentration and the physiological state of the cell.
Emulsification activity and emulsification index
When emulsification activity of 7 different solid hydrocarbons phenanthrene, fluoranthene, fluorene, anthracene, pyrene, chrysene and B(a)P and emulsification index of hexadecane, xylene, olive oil, kerosene, petrol and diesel were measured, it was observed that P. aeruginosa PSA5 showed higher emulsification activity and emulsification index than Rhodococcus sp. NJ2 (Fig. 3c, d).
Emulsification of the hydrocarbons by biosurfactant helps in microbial degradation. Van Hamme and Ward (2001) observed that hydrocarbon-grown Rhodococcus sp. Strain F9-D79 cells had high surface-active and emulsification properties. Anyanwu and Chukwudi (2010) reported that biosurfactants produced by P. aeruginosa LS1 were effective in producing good emulsification with kerosene. Obayori et al. (2009) have also observed high emulsification activity of engine oil, crude oil and diesel by Pseudomonas strain sp. LP1.
Thus, both the bacteria were found to be potential degraders of B(a)P as mediated by catabolic enzymes and facilitated by the biosurfactant production. Hence, these bacteria can be recommended for use in in situ bioremediation of soil contaminated with PAHs.
Conclusion
It may be concluded from this investigation that both the bacteria P. aeruginosa PSA5 and Rhodococcus sp. NJ2 showed differentiated ability to degrade B(a)P in MSM, mediated by catabolic enzymes, that is, salicylate hydroxylase, 2-CBD, C12O and C23O. These enzymes were differentially expressed in both the bacteria, showing their involvement in B(a)P degradation at different stages. A clear link between B(a)P degradation and glycolipid production was also found.
References
Allen CCR, Boyd DR, Larkin MJ, Reid KA, Sharma ND, Wilson K (1997) Metabolism of naphthalene, 1-naphthol, indene, and indole by Rhodococcus sp. strain NCIMB 12038. Appl Environ Microbiol 63:151–155
Anyanwu, Chukwudi U (2010) Surface activity of extracellular products of a Pseudomonas aeruginosa isolated from petroleum contaminated soil. Int J Environ Sci 1(2):225–235
Bouchez M, Blanchet D, Vandecasteele JP (1995) Degradation of polycyclic aromatic hydrocarbons by pure strains and by defined strain associations: inhibition phenomena and cometabolism. Appl Microbiol Biotechnol 43:156–164
Cerniglia CE (1992) Biodegradation of polycyclic aromatic hydrocarbons. Biodegradation 3:351–368
Chang WN, Liu CW, Liu HS (2009) Hydrophobic cell surface and bioflocculation behavior of Rhodococcus erythropolis. Process Biochem 44:955–962
Cooper DG, Goldenberg BG (1987) Surface-active agents from two Bacillus species. Appl Environ Microbiol 53(2):224–229
Desai JD, Banat IM (1997) Microbial production of surfactants and their commercial potential. Microbiol Mol Revol 61:47–64
Franzetti A, Tamburini E, Banat IM (2010) Application of biological surface active compounds in remediation technologies. In: Sen R (ed) Biosurfactants. Advances in experimental medicine and biology. Springer, Berlin, pp 121–134
Goyal AK, Zylstra GJ (1996) Molecular cloning of novel genes for polycyclic aromatic hydrocarbon degradation from Comamonas testosteroni GZ39. Appl Environ Microbiol 62:230–236
Hadibarata T, Kristani RA (2012) Fate and cometabolic degradation of benzo(a)pyrene by white rot fungus Armillaria sp. F022. Bioresour Technol 107:314–318
Heitkamp MA, Cerniglia CE (1988) Mineralization of polycyclic aromatic hydrocarbons by a bacterium isolated from sediment below an oil field. Appl Environ Microbiol 54:1612–1614
Hsu GW, Huang X, Luneva NP, Geacintov NE, Beese LS (2005) Structure of a high fidelity DNA polymerase bound to benzo(a)pyrene adducts that blocks replication. J Biol Chem 280(50):3764–3770
Johnsen AR, Karlson U (2007) Diffuse PAH contamination of surface soils: environmental occurrence, bioavailability and microbial degradation. Appl Microbiol Biotechnol 76:533–543
Juhasz AL, Naidu R (2000) Bioremediation of high molecular weight polycyclic aromatic hydrocarbons: a review of the microbial degradation of benzo[a] pyrene. Int Biodeterior Biodegradation 45:57–88
Kim E, Zylstra GJ (1995) Molecular and biochemical characterization of two meta-cleavage dioxygenases involved in biphenyl and m-xylene degradation by Beijerinckia sp. strain B1. J Bacteriol 177:3095–3103
Kim SJ, Kweon O, Freeman JP, Jones RC, Adjei MD, Jhoo JW, Edmondson RD, Cerniglia CE (2006) Molecular cloning and expression of genes encoding a novel dioxygenase involved in low- and high-molecular-weight polycyclic aromatic hydrocarbon degradation in Mycobacterium vanbaalenii PYR-1. Appl Environ Microbiol 72:1045–1054
Kim SJ, Kweon O, Jones RC, Freeman JP, Edmondson RD, Cerniglia CE (2007) Complete and integrated pyrene degradation pathway in Mycobacterium vanbaalenii PYR-1 based on systems biology. J Bacteriol 189:464–472
Kiyohara H, Nagao K (1978) The catabolism of phenanthrene and naphthalene by bacteria. J Gen Microbiol 105:69–75
Kojima Y, Itada N, Hayaisbi O (1961) Metapyrocatechase: a new catechol-cleaving enzyme. J Biol Chem 236:2223–2228
Krishnan S, Prabhu Y, Phale PS (2004) O-phthalic acid, a dead-end product in one of the two pathways of phenanthrene degradation in Pseudomonas sp. strain PP2. Ind J Biochem Biophys 41(5):227–232
Lily MK, Bahuguna A, Dangwal K, Garg V (2010) Optimization of an inducible, chromosomally encoded benzo [a] pyrene (BaP) degradation pathway in Bacillus subtilis BMT4i (MTCC 9447). Ann Microbiol 60:51–58
Long C, Lu JD, Li A, Hu D, Liu FQ, Zhang QX (2008) Adsorption of naphthalene onto the carbon adsorbent from waste ion exchange resin, equilibrium and kinetic characteristics. J Hazard Mater 182:771–772
Lowry OH, Roserbrough NT, Farr AL, Randall RJ (1951) Protein measurement with Folin phenol reagent. J Biol Chem 193:265–275
Luo YR, Tian Y, Huang X, Yan CL, Hong HS, Lin GH, Zheng T (2009) Analysis of community structure of a microbial consortium capable of degrading benzo(a)pyrene by DGGE. Mar Pollut Bull 58:1159–1163
Maier RM (2003) Biosurfactant: evolution and diversity in bacteria. Adv Appl Microbiol 52:101–121
Mallick S, Chatterjee S, Dutta TK (2007) A novel degradation pathway in the assimilation of phenanthrene by Staphylococcus sp. Strain PN/Y via meta-cleavage of 2-hydroxy-1-naphthoic acid: formation of trans-2,3-dioxo-5-(29-hydroxyphenyl)-pent-4-enoic acid. Microbiol 153:2104–1215
Moody JD, Freeman JP, Fu PP, Cerniglia CE (2004) Degradation of benzo[a]pyrene by Mycobacterium vanbaalenii PYR-1. Appl Environ Microbiol 70:340–345
Mukherjee AK, Das K (2005) Correlation between diverse cyclic lipopeptides production and regulation of growth and substrate utilization by Bacillus subtilis strains in a particular habitat. FEMS Microbiol Ecol 54:479–489
Nelson DL, Cox MM (2005) Principles of biochemistry, 4th edn. W. H. Freeman and Company, New York
Ngai KL, Neidle EL, Ornston LN (1990) Catechol and chlorocatechol 1,2-dioxygenase. Methods Enzymol 188:122–126
Noudeh GD, Housaindokht M, Fazzly Bazzaz BS (2005) Isolation, characterization and investigation of surface and hemolytic activities of a lipopeptide biosurfactant produced by Bacillus subtilis ATCC 6633. Microbiol Soc Korea 43:272–276
Noudeh GD, Noodeh AD, Moshafi MH, Behravan E, Afzadi MA, Sodagar M (2010) Investigation of cellular hydrophobicity and surface activity effects of biosynthesed biosurfactant from broth media of PTCC 1561. Afr J Microbiol Res 4(17):1814–1822
Obayori OS, Adebusoye SA, Adewale AO, Oyetibo GO, Oluyemi OO, Amokun RA, Iiori MO (2009) Differential degradation of crude oil (Bonny light) by four Pseudomonas strains. J Environ Sci 21:243–248
Rahman KSM, Rahman TJ, Kourkoutas Y, Petsas I, Marchant R, Banat IM (2003) Enhanced bioremediation of n-alkane in petroleum sludge using bacterial consortium amended with rhamnolipid and micronutrients. Bioresour Technol 90:159–168
Rentz JA, Alvarez PJJ, Schnoor JL (2008) Benzo[a]pyrene degradation by sphingomonas yanoikuyae JAR02. Environ Pollut 151:669–677
Rosenberg E, Zuckerbera G, Rubinowitz H, Gutnick DL (1979) Emulsifier of Arthrobacter RAG-1: isolation and emulsifying properties. Appl Environ Microbiol 31:402–408
Rosenberg M, Gutnick D, Rosenberg E (1980) The adherence of bacteria to hydrocarbons: a simple method for measuring cell surface hydrophobicity. FEMS Microbiol Lett 9:29–33
Sala-Trepat JM, Evans WC (1971) The meta cleavage pathway of catechol by Azobacter species. Eur J Biochem 20:400–413
Sayara T, Pognani M, Sarra M, Sanchez A (2010) Anaerobic degradation of PAHs in soil: impacts of concentration and amendment stability on the PAHs degradation and biogas production. Int Biodeterior Biodegradation 64:286–292
Schneider J, Grosser R, Jayasimhulu K, Xue W, Warshawsky D (1996) Degradation of pyrene, benz[a]anthracene, and benzo[a]pyrene by Mycobacterium sp. strain RJGII-135, isolated from a former coal gasification site. Appl Environ Microbiol 62:13–19
Sho M, Hame C, Greer CW (2004) Two distinct gene clusters encode pyrene degradation in Mycobacterium sp. strain S65. FEMS Microbiol Ecol 48:209–220
Siegmund I, Wagner F (1991) New method for detecting rhamnolipids excreted by Pseudomonas during growth on mineral agar. Biotechnol Tech 5:265–268
Spiro RG (1966) Analysis of sugars found in glycoproteins. Methods Enzymol 8:7–9
Van Hamme JD, Ward OP (2001) Physical and metabolic interactions of Pseudomonas sp. strain JA5-B45 and Rhodococcus sp. strain F9–D79 during growth on crude oil and effect of a chemical surfactant on them. Appl Environ Microbiol 67:4874–4879
Van Hamme JD, Singh A, Ward OP (2003) Recent advances in petroleum microbiology. Microbiol Mol Biol Rev 67:503–549
Widada J, Nojiri H, Omori T (2002) Recent developments in molecular techniques for identification and monitoring of xenobiotic-degrading bacteria and their catabolic genes in bioremediation. Appl Environ Microbiol 60:45–59
Yamamoto R, Ortmayer HB, Bigland CH, Seely ML, Corstvet RE (1965) Isolation of “N” Mycoplasma from different sites of the turkey. Poult Sci 44:732–736
Yen KM, Serdar CM (1998) Genetics of naphthalene catabolism in Pseudomonas. Crit Rev Microbiol 15:247–268
Zhong H, Zeng GM, Yuan XZ, Fu HY, Huang GH, Ren FY (2007) Adsorption of dirhamnolipid on four microorganisms and the effect on cell surface hydrophobicity. Appl Microbiol Biotechnol 77:447–455
Acknowledgments
Authors are thankful to Director, CSIR-NBRI, Lucknow, for providing us laboratory facilities to carry out investigation and to CSIR, New Delhi, for financial assistance in the form of NWP-19.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mishra, S., Singh, S.N. Biodegradation of benzo(a)pyrene mediated by catabolic enzymes of bacteria. Int. J. Environ. Sci. Technol. 11, 1571–1580 (2014). https://doi.org/10.1007/s13762-013-0300-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-013-0300-6