Abstract
Repetitive DNA sequences have been related to the maintenance of heterochromatin, evolution, and speciation, mainly due to the variations to which they are subject. Bees of the Meliponini tribe generally possess large amounts of heterochromatin that occupy the long arm of the chromosomes or even almost the entire chromosome. Although widely studied by conventional methods, little is known about the composition of these regions. Thus, this study aimed to analyze a repetitive sequence of Tetragonisca angustula and compare it with other genera in order to broaden the understanding of the evolution of these sequences. The results indicate that the isolated repeating DNA sequence of Tetragonisca angustula is present in the heterochromatin of all chromosomes of this species and in Tetragonisca fiebrigi. However, the same was not observed for the other evaluated species, which suggests that the isolated sequence is conserved within the genus Tetragonisca and that, in the other genera, the sequences that compose the heterochromatin may have undergone different processes of diversification or have arisen independently.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Repetitive sequences represent large fractions of the eukaryotic genome (Bostock 1980; Charlesworth et al. 1994) and perform different functional roles, such as the assembly of kinetochores (Huang et al. 2016), maintenance of heterochromatin (Grewal and Jia 2007), and regulation of gene expression (Koo et al. 2016). Due to their high variation rates, these sequences are used in evolutionary and speciation studies (Plohl et al. 2012; Garrido-Ramos 2017). In turn, variations that occur in the sequence can be spread throughout the genome by the homogenization process and be fixed between individuals of a species by sexual reproduction, which is known as the concerted evolution (Dover 2002). Despite the rapid evolution of these sequences, related species may share the same family of repetitive DNA (Lorite et al. 2017).
Species of the Meliponini tribe have a large amount of heterochromatin and their distribution in the chromosomes may vary. In Tetragonisca angustula Latreille, 1811, for example, heterochromatin predominates in the short arm of the chromosomes, while in Plebeia phrynostoma Moure, 2004, it is mainly found in the centromeric and terminal regions of the chromosomes (Barth et al. 2011; Godoy et al. 2013). The genus Melipona Illiger, 1806 presents a peculiar pattern of heterochromatin distribution that is found exclusively in the pericentromeric region of some species, and in almost all the chromosome extension in others (Rocha and Pompolo 1998; Cunha et al. 2018).
Studies with repetitive DNA sequences in the heterochromatin of Meliponini tribe chromosomes are scarce (Lopes et al. 2014; Piccoli et al. 2018; Santos et al. 2018), so documented results together with those obtained from additional studies may enhance the understanding of the evolution of the heterochromatin in this group of bees. Our objective was to analyze the repetitive DNA fragment of Tetragonisca angustula and compare this region in other different genera of this tribe in order to broaden the understanding of the evolution of these regions in these bees. For such, we isolated a repetitive DNA sequence and compared its distribution in seven species of different genera, including more related species and phylogenetically distant species.
2 Materials and methods
2.1 Biological samples and chromosome preparation
For this work, we selected the species Tetragonisca angustula, Tetragonisca fiebrigi Schwarz, 1938; Melipona mondury Smith, 1863; Nannotrigona testaceicornis, Lepeletier, 1836; Frieseomelitta varia Lepeletier, 1836; Partamona helleri Friese, 1900; and Austroplebeia australis Friese, 1898 (Table I). To obtain mitotic chromosomes, brain ganglia of post-defecating larvae were used (Imai et al. 1988).
2.2 Obtaining repetitive DNA by restriction enzyme
Total T. angustula DNA was extracted as Waldschmidt et al. (1997) and tested with restriction enzymes MspI, EcoRI, HinfI, BcnI, BgIII, Bsp68I, and HinfI e VspI. The tests took into account enzyme concentration and digestion time. For the reaction, we used 2.5 μl DNA (100 ng/μl), 0.5 μl enzyme (10 U/μl), and 2.0 μl Buffer (10X). DNA cleavage was conducted in a water bath at 37 °C for 6 h. The enzyme digestion product was evaluated on 1.2% agarose gel. The fragment corresponding to the repetitive sequence was purified using NucleoSpin ® Gel and PCR Clean-up kit (Neumann-Neander-Str. 6–852,355 Düren Germany) according to manufacturer’s instructions.
2.3 Fluorescent in situ hybridization (FISH)
The isolated repetitive sequence was used as a probe for FISH according to Pinkel et al. (1986) with modifications. Denaturation of mitotic chromosomes was achieved with 70% formamide at 75 °C for 5 min. The isolated sequence was labeled with Dig-Nick Translation (Roche Applied Science) following manufacturer’s instructions and the obtained probe used to hybridize the chromosomes at 37 °C in a humid chamber overnight. The signal was detected with anti-digoxigenin-rhodamine (Roche Applied Science). The chromosomes were counterstained with Fluoroshield with DAPI (Sigma). The images were captured on an Olympus BX53F microscope equipped with an Olympus MX10 camera and the CellSens imaging software.
3 Results
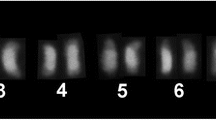
Eight restriction enzymes were tested for total DNA digestion of T. angustula and only Bsp68I produced band pattern with cleavage profile in repetitive regions. The probe obtained from the isolated repetitive DNA sequence hybridized to the heterochromatic region of the short arm of all T. angustula chromosomes (Figure1a). The labeling presented itself as a large block forming a cluster, and no differences in the pattern of labeling between T. angustula chromosomes were observed (Figure 2c, d). In T. fiebrigi, we also observed a similar result with short arm markings of all chromosomes, coinciding with the heterochromatin regions (Figure 1b). In T. fiebrigi, the markings also followed the same cluster pattern as in T. angustula (Figure 2g, h). No markings were observed on the chromosomes of the other species investigated (Figure 1c–g).
Patterns obtained with hybridization of the repeating DNA probe isolated from Tetragonisca angustula. a Tetragonisca angustula. b Tetragonisca fiebrigi. c Frieseomelitta varia. d Partamona helleri. e Nannotrigona testaceicornis. f Melipona mondury. g Austroplebeia australis. On blue chromosomes stained with DAPI and green marking with the probe. Scale bar = 5 μm.
Metaphases of Tetragonisca angustula a C-banding, b DAPI, c repeating DNA probe, and d probe merged with DAPI; metaphases of Tetragonisca fiebrigi e C-banding, f DAPI, g repeating DNA probe, and h probe merged with DAPI; metaphases of Melipona mondury i C-banding, j DAPI, k repeating DNA probe, and l probe merged with DAPI. Scale bar = 5 μm.
4 Discussion
This is the first work reported using restriction enzyme to isolate repetitive DNA sequences in bees. The result was satisfactory for one enzyme among the eight enzymes tested on T. angustula. With this technique, repetitive sequences have already been isolated in other organisms such as ants (Lorite et al. 2004), frogs (Amor et al. 2009), and fish (Vicari et al. 2010). In all these species, the sequences were shown to be satellite DNA, and the results contributed to the understanding of genome evolution and organization (Lorite et al. 2004; Amor et al. 2009; Vicari et al. 2010). Thus, the use of this tool to study these sequences opens the door for further studies in the Meliponini tribe and other Hymenoptera.
Markings observed on the heterochromatin of all T. angustula chromosomes (Figure1a) indicate that there is a type of repetitive DNA that makes up most of the heterochromatin of this species. Lopes et al. (2014), by chromosomal microdissection, constructed probes from the heterochromatin of only one chromosome, using Melipona rufiventris Lepeletier, 1836 and T. fiebrigi as models. Each probe hybridized on all chromosomes of the respective species, which reveals similarity between the sequences that compose the heterochromatin of the species and corroborates our results.
Besides being observed in all T. angustula chromosomes, this sequence was found in another species of the genus, T. fiebrigi (Figure1b). Despite the dynamism and high variability of the repetitive DNA, related species may share repetitive DNA sequences (Lorite et al. 2017). This characteristic can be related to the time of diversification between species, since the propagation of a new variant may depend on time (Pérez-Gutiérrez et al. 2012). The recent time of diversification added to the possibility that T. angustula and T. fiebrigi are subspecies and not species (Francisco et al. 2014), due to the great similarities between them, can explain the conservation of the repetitive DNA sequence among these bees.
Although time may be an important factor in explaining the sharing of repetitive DNA sequences among phylogenetically close species, the long-term evolution rate of these sequences may be influenced by other factors. Among these factors, we can consider the location and organization of these sequences (Navajas-Pérez et al. 2009), biological factors (Lorite et al. 2017), and functional restrictions (Huang et al. 2016). In ants, the conservation of repetitive sequences may be related to the sex determination system for haplodiploidy (Lorite et al. 2017) which could also explain the conservation of this sequence in Tetragonisca. In this case, considering that events, such as crossing-over, are important for new mutations in satellite DNA sequences to be homogenized between chromosomes, the fact that males are haploid and do not present recombination may hinder the transmission of mutations and consequently preserve the composition nucleotide of repeating sequences (Lorite et al. 2017).
Considering the abundance of heterochromatin in the chromosomes among most Meliponini, the amplification of this region may have occurred in a Meliponini ancestor and later shared among the genera of this tribe. On the other hand, Lopes et al. (2014) reported that the chromosomes of M. rufiventris and T. fiebrigi have heterochromatin with different repetitive sequences. The genus Melipona has recently diverged and presents chromosomal characteristics different from those of the other Meliponini (Rocha and Pompolo 1998; Rocha et al. 2002; Rasmussen and Cameron 2010). Thus, variation in the heterochromatin composition between M. rufiventris and T. fiebrigi may reveal that heterochromatin from Melipona followed different evolutionary paths.
Among the species studied, Austroplebeia australis, (Indo-Malayan/Australasian clade), is phylogenetically the furthest from Tetragonisca (Rasmussen and Cameron 2010). Even showing large blocks of heterochromatin (Travenzoli 2018), no marking signal on the chromosomes of A. australis was observed (Figure1g), which suggests that the repetitive DNA sequence evaluated could have arisen more recently within the Neotropical Meliponini. However, no other evaluated species showed markings, not even Frieseomelitta varia (Figure1c), which, among the species investigated (Figure 1), is phylogenetically closest to Tetragonisca (Rasmussen and Cameron 2010). Thus, the repetitive DNA sequence present in the heterochromatin of the Tetragonisca species possibly arose in the common ancestor among the species of this genus. Since we did not observe this sequence of T. angustula in species of the other genus evaluated, the conservation can be a consequence of the recent time of diversification among the species of this group, without the accumulation of enough mutations to make them divergent. In addition, the repetitive DNA sequence within the Meliponini tribe was not shared, which suggests that these sequences are evolving rapidly and differently in each genus. Heterochromatin may also have appeared at different times throughout the evolution of Meliponini.
In this work, we highlight the importance of the use of molecular cytogenetics for studies on repetitive DNA sequences in bees, and how these sequences can increase the understanding of the evolution of heterochromatin in this group. In addition, we have demonstrated that heterochromatin may evolve independently in each genus. Thus, this study promotes other discussions about the evolution of heterochromatin in Meliponini.
References
Amor, N., Odierna, G., Chinali, G., Said, K., Picariello, O. (2009) Unusual chromosomal distribution of a major satellite DNA from Discoglossus pictus (Amphibia, Anura). Cytogenet. Genome Res. 127 (1), 33-42
Barth, A., Fernandes, A., Pompolo, S. D. G., Costa, M. A. (2011) Occurrence of B chromosomes in Tetragonisca Latreille, 1811 (Hymenoptera, Apidae, Meliponini): a new contribution to the cytotaxonomy of the genus. Genet. Mol. Biol. 34 (1), 77-79
Bostock, C. (1980) A function for satellite DNA? Trends Biochem. Sci. 5 (5), 117-119
Charlesworth, B., Paul, S., Wolfgang, S. (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature. 371 (6494), 215-220
Cunha, M. S., Travenzoli, N. M., Ferreira, R. P., Cassinela, E. K., Silva, H., Salomão, T. M. F., Lopes, D. M. (2018) Comparative cytogenetics in three Melipona species (Hymenoptera: Apidae) with two divergent heterochromatic patterns. Genet. Mol. Biol. 41 (4), 806-813
Dover, G. A. (2002) Molecular drive. Trends Genet. 18 (11), 587-589
Francisco, F. O., Santiago, L. R., Brito, R. M., Oldroyd, B. P., Arias, M. C. (2014) Hybridization and asymmetric introgression between Tetragonisca angustula and Tetragonisca fiebrigi. Apidologie. 45 (1), 1-9
Garrido-Ramos, M.A. (2017) Satellite DNA: An evolving topic. Genes. 8 (9), 230
Godoy, D.C., Ferreira, R.P., Lopes, D.M. (2013) Chromosomal Variation and Cytogenetics of Plebeia lucii and P. phrynostoma (Hymenoptera: Apidae). Fla. Entomol. 96 (4), 1559-1566
Grewal, S. I., Jia, S. (2007) Heterochromatin revisited. Nat. Rev. Genet. 8 (1), 35-47
Huang, Y. C., Lee, C. C., Kao, C. Y., Chang, N. C., Lin, C. C., Shoemaker, D., Wang, J. (2016) Evolution of long centromeres in fire ants. BMC Evol. Biol. 16 (1), 189
Imai, H.T., Taylor, R.W., Crosland, M.W., Crozier, R.H. (1988) Modes of spontaneous chromosomal mutation and karyotype evolution in ants with reference to the minimum interaction hypothesis. Jpn. Soc. Appl. Entomol. Zool. 63 (2), 159-185
Koo, D.H., Zhao, H., Jiang J. (2016) Chromatin-associated transcripts of tandemly repetitive DNA sequences revealed by RNA-FISH. Chromosom. Res. 24 (4), 467-480
Lopes, D.M., Fernandes, A., Diniz, D., Scudeler, P.E.S., Foresti, F., Campos, L.A.O. (2014) Similarity of heterochromatic regions in the stingless bees (Hymenoptera: Meliponini) revealed by chromosome painting. Caryologia. 67 (3), 222-226
Lorite, P., Carrillo, J.A., Tinaut, A., Palomeque, T. (2004) Evolutionary dynamics of satellite DNA in species of the genus Formica (Hymenoptera, Formicidae). Gene. 332, 159-168
Lorite, P., Muñoz-López, M., Carrillo, J.A., Sanllorente, O., Vela, J., Mora, P., Palomeque, T. (2017) Concerted evolution, a slow process for ant satellite DNA: study of the satellite DNA in the Aphaenogaster genus (Hymenoptera, Formicidae). Org. Divers. Evol. 17 (3), 595-606
Navajas-Pérez, R., Quesada del Bosque, M.E., Garrido-Ramos, M.A. (2009) Effect of location, organization and repeat-copy number in satellite-DNA evolution. Mol. Gen. Genomics. 282 (4), 395-406
Pérez-Gutiérrez, M.A., Suárez-Santiago, V.N., López-Flores, I.; Romero, A.T.; Garrido-Ramos, M.A. (2012) Concerted evolution of satellite DNA in Sarcocapnos: A matter of time. Plant Mol. Biol. 78 (1-2), 19-29
Piccoli, M. C. A., Bardella, V. B., Cabral-de-Mello, D.C. (2018) Repetitive DNAs in Melipona scutellaris (Hymenoptera: Apidae: Meliponidae): chromosomal distribution and test of multiple heterochromatin amplification in the genus. Apidologie. 49 (4), 497-504
Pinkel, D. Straume, T. Gray, J.W. (1986). Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc. Natl. Acad. Sci. 83: 2934-2938
Plohl, M., Meštrović, N., Mravinac, B. (2012) Satellite DNA evolution. Genome Dyn. 7, 126-152
Rasmussen, C., Cameron, S.A. (2010) Global stingless bee phylogeny supports ancient divergence, vicariance, and long distance dispersal. Biol. J. Linn. Soc. Lond. 99 (1), 206–232
Rocha, M.P., Pompolo, S.G. (1998) Karyotypes and heterochromatin variation (C-bands) in Melipona species (Hymenoptera, Apidae, Meliponinae). Genet. Mol. Biol. 21 (1), 41–45
Rocha, M.P., Pompolo, S.G., Dergam, J.A., Fernandes, A., Campos, L.A.O. (2002) DNA characterization and karyotypic evolution in the bee genus Melipona (Hymenoptera Meliponini). Hereditas. 21 (1), 19–27
Santos, J.M., Diniz, D., Rodrigues, T.A.S., Cioffi, M.B., Waldschmidt, A.M. (2018) Heterochromatin distribution and chromosomal mapping of microsatellite repeats in the genome of Frieseomelitta stingless bees (Hymenoptera: Apidae: Meliponini). Florida Entomol. 101 (1), 33-39
Travenzoli, N.M. (2018) Citogenética clássica e molecular com ênfase na evolução cromossômica em Meliponini. Dissertation, Universidade Federal de Viçosa
Vicari, M. R., Nogaroto, V., Noleto, R. B., Cestari, M. M., Cioffi, M. B., Almeida, M. C., Artoni, R. F. (2010) Satellite DNA and chromosomes in Neotropical fishes: methods, applications and perspectives. J. Fish Biol. 76 (5) 1094-1116
Waldschmidt, A.M., Salomão, T.M.F., Barros, E.G.D., Campos, L.D.A.O. (1997) Extraction of genomic DNA from Melipona quadrifasciata (Hymenoptera: Apidae, Meliponinae). Braz. J. Genet. 20 (3), 421-423
Acknowledgments
The authors would like to thank the “Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES)” and “Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG)”.
Author information
Authors and Affiliations
Contributions
JAP performed the experiments and analyses and wrote the manuscript. TMFS participated in the revisions of the manuscript. DLM assisted in the analysis and revision of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Manuscript editor: Yves Le Conte
Différentes séquences répétitives d'ADN composent l'hétérochromatine chez les Meliponini.
enzyme de restriction / abeille sans dard / évolution chromosomique / Tetragonisca angustula.
Unterschiedliche repetitive DNA Sequenzen bilden das Heterochromatin bei Meliponini.
Restriktionsenzyme / Stachellose Bienen / Chromosomenevolution / Tetragonisca angustula .
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pereira, J.A., Salomão, T.M.F. & Lopes, D.M. Different repetitive DNA sequences make up heterochromatin in Meliponini. Apidologie 51, 855–860 (2020). https://doi.org/10.1007/s13592-020-00766-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-020-00766-1