Abstract
Here, we describe a new species of genus Bombus Latreille, 1802 from northeastern Brazil, Bombus (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. Molecular analysis was initially performed to confirm the new species placement within the genus Bombus. Afterward, we performed an integrative approach combining molecular data (DNA barcoding and two nuclear regions) and morphology to confirm its taxonomic status. The genetic and morphological data were very consistent and congruent supporting this group as a new species. B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. has the body totally covered by black pilosity and can be distinguished from closer and/or sympatric species by clypeus shape, which has a distinct flattened dorsal platform, by length of malar area and the length and shape of the hairs, shorter and aligned as if it was trimmed. In addition, brief taxonomic notes on Bombus (Thoracobombus) brevivillus Franklin (1913) and Bombus (Thoracobombus) morio Swederus (1787), the sympatric species, are provided. A key for identification of Brazilian Bombus species, including the new species, was elaborated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Bumblebees belong to the genus Bombus (Hymenoptera: Apidae, Bombini) that encompasses about 250 species. These bees provide important ecosystem services by their extraordinary ability to pollinate (Corbet et al. 1991; Kevan 1991; Memmott et al. 2004; Pywell et al. 2006; Goulson et al. 2008). They occur primarily in cold climate zones in Eurasia and North America (Michener 2007). In Brazil, six species are currently recognized: Bombus bellicosus Smith (1879), Bombus brasiliensis Lepeletier (1836), Bombus brevivillus Franklin (1913), Bombus morio Swederus (1787), Bombus pauloensis Friese (1913) (also known as Bombus atratus Franklin; 1913), and Bombus transversalis Olivier (1789), all belonging to the subgenus Thoracobombus (Williams et al. 2008). The low number of species in Brazil, despite their wide distribution, is very contrasting to the high number found in other world regions as the Palaearctic (120 species), Orient (108 species), and Japan (23 species) (Williams 1996, 1998).
In general, species of Bombus present homogeneity in morphology, an unusual feature for bees. This uniformity is an obstacle to their identification, and consequently, cryptic species may be still unrecognized (Moure and Sakagami 1962; Williams 1998; Michener 2007; Murray et al. 2008). Conversely, Brazilian species present high intra-specific variability, and parallelism in color patterns is observed in sympatric species. As stated by Moure and Sakagami (1962), Bombus is a real “puzzles” for taxonomists and the “most difficult group to classify”.
The DNA barcode has been proposed to supplement current and traditional methods based on morphological traits (Hebert et al. 2004). For animals in general, this method is based on the sequencing of the mitochondrial gene cytochrome C oxidase I (COI) region. Recent works have employed this molecular method with success in the identification of a great variety of taxa, including the genus Bombus (Murray et al. 2008; Williams et al. 2011, 2012). Nevertheless, subsequent to an initial DNA barcoding, an integrative approach using multilocus analysis and traditional taxonomy is still necessary for species delimitation accuracy (Sites and Marshall 2004; Collins and Cruickshank 2012).
To develop a comprehensive phylogeography study based on molecular data, we have been collecting Bombus specimens in Brazil during these past years. Also, we had access to samples kept in collections for DNA extraction (Table I). The preliminary analysis revealed that some specimens from northeastern Brazil were genetically “different” from all other samples. In addition of being more robust, these distinctive specimens were slightly different morphologically from the sympatric species (B. brevivillus and B. morio), according to the morphological key proposed by Moure and Sakagami (1962).
Considering these molecular and morphological evidences, the goals of this research were as follows: (a) verify the placement of these specimens within the genus Bombus (whether they are closely related to the Brazilian species) and ensure that these specimens do not belong to a species already described; (b) test if these specimens constitute a new species by gathering molecular data from the DNA barcode and nuclear regions for all Brazilian Bombus species and performing phylogenetic analysis; (c) characterize these specimens morphologically; (d) describe the new species; and (e) provide a key for identification of Brazilian Bombus species, including the new species.
2 Materials and methods
2.1 Samples and molecular markers
Table I summarizes the Brazilian species studied and the general information for each specimen. We were able to obtain 17 specimens of the suspected species either from nature or from collections. The specimens were identified according to the morphological key proposed by Moure and Sakagami (1962). One middle leg of frozen specimens was used for DNA extraction following the Chelex® 100 (Bio-Rad, UK) method (Walsh et al. 1991). For pinned specimens, we also used one middle leg for DNA extraction using the DNeasy Tissue Kit (Qiagen, Germany) following the supplier’s recommendations.
Primers (Online resource 1) and PCR conditions for amplification of the COI barcode region were as described by Françoso and Arias (2013). The COI sequences for B. bellicosus, B. brevivillus, and B. transversalis were obtained previously in our laboratory, using primers described by Simon et al. (1994; online resource 1). Also, the large ribosomal RNA subunit (16S) region was amplified according to the same authors. We amplified exons and introns from two nuclear genes, arginine kinase (ArgK) (Kawakita et al. 2003) and elongation factor-1 α (EF-1 α) (Hines et al. 2006), to test for mitochondrial introgression and to improve our phylogenetic analyses. All PCR amplifications were according to the articles cited above and performed in a Mastercycler Pro (Eppendorf, Germany). For amplifications using the primers described by Simon et al. (1994), we used the same PCR cycling conditions as for the DNA barcode region. PCR products were separated on a 0.8 % agarose gel, stained with Gel Red 10.000X (Biotium, USA), and visualized under UV light. All PCR products were purified with 0.5 μl of ExoSAP-IT® (USB, USA) following the manufacturer’s recommendations and were sequenced from the forward direction by the Macrogen Company (South Korea). The program Muscle (Edgar 2004) included in Geneious Pro 7.0.3 software (http://www.geneious.com) was used to align the sequences. The COI sequences were edited and translated into amino acid sequences. These DNA sequences were compared with GenBank using BLAST tool (Altschul et al. 1990) and Bold databases (Ratnasingham and Hebert 2007). These two procedures were employed to verify inadvertent amplification of homologous COI from Wolbachia or numts. The DNA sequences obtained were deposited in GenBank (Accession numbers: KT187861–KT187933).
2.2 Phylogenetic analysis
To confirm the placement within the genus and ensure that these specimens could not be a species already known from another region, we selected one Bombus sp. nov. (EF204) COI sequence to compare with the COI sequences deposited at BOLD and GenBank databases. Also, we performed phylogenetic analysis using the sequence alignment of 218 taxa (Hines 2008) already published for the entire genus (Cameron et al. 2007) including the suspect group sequences. Three out of five molecular markers used by Cameron et al. (2007) were used in the new species: 16S, ArgK, and EF-1α. We added the suspect specimen sequences to the data matrix (Study ID: S1927) available in TreeBASE (http://www.treebase.org) and applied the same models of nucleotide substitution and the same phylogenetic reconstruction parameters used in Hines (2008) for Bayesian analysis.
Phylogenetic reconstruction for the Brazilian group of bumblebees was performed by Bayesian analysis through MrBayes 3.2.2 (Huelsenbeck and Ronquist 2001). The program JModelTest 2.1.4 (Darriba et al. 2012) selected the models GTR + I + R for the mitochondrial COI and TIM1 + G for the nuclear ArgK and EF-1 α genes. The analysis was run with 10 million generations with a burn-in of 25 %. Bombus (Bombus) ignitus Smith (1869), Bombus (Bombus) lucorum Linnaeus (1761), and Bombus (Cullumanobombus) volucelloides Gribodo (1891) (GenBank accession numbers: 010967.1, AY267133, and AY739522, respectively) were used as outgroups.
The genetic distance between the new species clade and the other Brazilian species of bumblebee was taken through the sum of the length of branches of a UPGMA tree obtained in Geneious Pro 5.6.3 software.
2.3 Traditional morphology
Four female (f) specimens (queens) were morphologically analyzed and deposited at Coleção Entomológica do Museu de História Natural da Universidade Federal da Bahia (MHNBA-MZUFBA), Salvador, Bahia, Brazil. The other 13 specimens were analyzed only molecularly and were included as paratypes (Table I). General morphological terminology is according to Michener (2007), and the standard abbreviations are as follows: antennal flagellomeres (F1, F2…), metasomal terga (T1 to T6), and puncture diameter (PD). The upper and lower interocular distances were measured using the shortest distance between the compound eyes in frontal view. All measurements are given in millimeters (mm). Label information from separate labels is separated by double slashes, “//.” Photomicrographs were prepared using a Leica M165C stereomicroscope coupled with a Leica DFC295 and a Leica Application Suite V4.1 Interactive Measurements, Montage. All the observation and measurements for all species cited in here were made based in female queen specimens. A key for identification of Brazilian Bombus species was prepared, adapted from Moure and Sakagami (1962), including the new species (see below).
3 Results
3.1 Molecular markers and phylogeny
The following sequence sizes were generated: 376 base pairs (bp) for 16S, 430 bp for COI (with primers MtD7 and MtD9), 631 bp for COI (with primers BarbeeF and MtD9), 820 bp for ArgK, and 733 bp for EF-1 α.
The Bombus sp. nov. (EF204) COI sequence was used for comparisons to our Bombus COI sequence, to Bombus COI sequences homologous from GenBank (1607 sequences) and Bold (1269 sequences), totaling 2893 sequences (about 130 species). The sequence sizes were not identical, which could compromise the accuracy of the genetic distance between them. However, the 20 highest hits (sequence similarities) obtained for Bombus sp. nov. (EF204) encompass two Brazilian species, B. brasiliensis and B. pauloensis, and one North American, Bombus (Thoracobombus) fervidus Fabricius, 1798 (Online resource 2). These species are all united in Thoracobombus subgenus. No Wolbachia or other contaminants were verified.
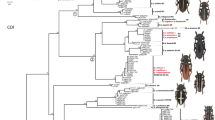
According to the phylogeny of the entire genus (Figure 1a), Bombus sp. nov. is related to the following Brazilian bumblebee species: B. pauloensis, B. brasiliensis, and B. transversalis, with high values of posterior probabilities. For DNA barcode and nuclear regions, in all topologies, samples of Bombus sp. nov. constituted a clade strongly supported by posterior probabilities and are well separated from the other Brazilian bumblebee species (Figure 1). The DNA barcode data revealed a 6.11 % genetic distance between the new lineage and its sister group, comprised of B. bellicosus, B. brasiliensis, B. brevivillus, and B. transversalis.
Bayesian phylogenetic trees obtained through molecular markers analysis (a, b, d, e). Values in node represent posterior probabilities, and dashes represent posterior probabilities lower than 90 %. Letters beside the species name match to the index in Table I. a Phylogeny of the genus Bombus (Cameron et al. 2007; Hines 2008) obtained by analyzing 218 taxa (16S, ArgK, EF-1α, opsin, and PEPCK) with the placement of Bombus (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. b Phylogeny of the Brazilian species obtained by cytochrome C oxidase I sequences. c Geographic distribution of samples in b. d Phylogeny of the Brazilian species obtained by elongation factor-1 α sequences. e Phylogeny of the Brazilian species obtained by arginine kinase sequences. Only posterior probabilities lower than 90 % are represented.
The clades obtained from COI sequences were well resolved, for all species (Figure 1b, c). The nuclear genetic markers gave support for the Bombus sp. nov. as monophyletic group but did not provide enough signal to solve all of the other Brazilian species as distinct from each other, except for B. morio, the most distant in the Brazilian bumblebee group (Cameron et al. 2007; Figure 1d, e). The DNA barcode data also revealed that B. brasiliensis is polyphyletic and apparently represents a species complex.
3.2 Morphology
Morphological characters allowed the recognition of these specimens as a new species, as the flattened structure of clypeus on the middle portion (Figure 2 (1)), its general aligned body pubescence (uniform in length) with straight hair (as if they had been trimmed) (Figure 3 (11)), and also the relatively shorter pubescence on head (Figure 2 (2)). Bombus sp. nov. is quite similar to B. brevivillus and B. morio, mainly by similar integument color (black) and a very dense pubescence and punctures in general. Nevertheless, these three species can be distinguished. The malar area is shorter and wider in the new species in comparison to B. morio. B. brevivillus has the shortest malar area among the three species, and it is narrower compared to the new species. The discal glabrous area of mesoscutum is smooth and shiny in B. brevivillus and Bombus sp. nov., but narrower in B. brevivillus, and microreticulated and dull in B. morio (Figures 2 and 3), as described below in the taxonomic treatment.
Frontal view of the head (1, 4, 7), head in profile (2, 5, 8), and malar area (3, 6, 9) of species of Bombus Latreille, 1802 (Hymenoptera: Apidae: Bombini). 1–3: Bombus (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (Holotype); 4–6: Bombus (Thoracobombus) brevivillus Franklin, 1913; 7–9: Bombus (Thoracobombus) morio (Swederus, 1787).
Dorsal view of the head (vertex) (10, 13, 16), mesoscutum (11, 14, 17), and hind tibia (12, 15, 18) of species of Bombus Latreille, 1802 (Hymenoptera: Apidae: Bombini). 10–12: Bombus (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (Holotype); 13–15: Bombus (Thoracobombus) brevivillus Franklin, 1913; 16–18: Bombus (Thoracobombus) morioSwederus (1787).
4 Discussion
Specimens of the genus Bombus are not easily identified by morphology (Murray et al. 2008). This is an impediment to understanding the taxonomy and systematics of bumblebees, such as revealing new species. For example, Bombus (Bombus) cryptarum Fabricius, 1775, a common and widespread species, remained undetected until 2005 due to the high morphological similarity to B. lucorum (Bertsch et al. 2005; Murray et al. 2008). According to Williams et al. (2012), Bombus s. str. is likely to require additional characters beyond morphology in order to add substantial progress toward resolving cryptic species.
The low number of Bombus species found in Brazil may be a consequence of ecological and environmental features, among other factors. The tropical climate may facilitate dispersion allowing gene flow even in large areas; thus, the species integrity can be maintained. However, the low number of Bombus species presenting large geographic distribution in Brazil may be unreal. The lack of morphological characters seems to be the major constraint to species discrimination in this bee group. Even for well-known taxa, the existence of morphologically unrecognized (or cryptic) species indicates that there are likely to be many more species than has been currently estimated (Frankham et al. 2002; Blaxter 2004; Bickford et al. 2007; Murray et al. 2008). Williams et al. (2011), studying the subgenus Subterraneobombus Vogt (1911), suggested that COI barcodes are a cost-effective source of additional characters where morphological information has been insufficient. Here, after confirming the placement in the subgenus, the DNA barcoding approach enabled the unveiling of a new species of Bombus endemic to Brazil and also suggests that B. brasiliensis is a non-monophyletic species, likely a cryptic species complex. Subsequently, nuclear markers corroborate the mtDNA findings for the new species. The clade comprising the new species lineage was distinct and well supported. It is worth mentioning that although the nuclear markers supported the new species as a distinctive clade, those markers could not discriminate several other Brazilians species, probably due to recent divergence time among B. bellicosus, B. brasiliensis, B. brevivillus, and B. transversalis, estimated in less than 3 Mya (Hines 2008).
The COI divergence between the new species and its nearest neighbor was relatively high (6.11 %) when comparing to other Bombus species. Among the Brazilian species that share a common ancestor (Cameron et al. 2007), the lowest value of the minimum interspecific sequence divergence was 0.8 % (between B. transversalis and B. brevivillus), and the highest value was 4.7 % (between B. pauloensis and B. transversalis). In the subgenus Bombus, the average was 3.2 % (minimum of 1.57 % and maximum of 6.07 %; Williams et al. 2012). Species boundaries signaled by deep COI divergence are generally congruent with those established through classical taxonomic work (Sheffield et al. 2009). In fact, herein, the new species can be distinguished morphologically from their sympatric species on the basis of shape of the clypeus, having a distinct dorsal platform and lateral area, length of malar area, and the length and shape of the hairs (shorter and aligned).
In this study, the DNA barcode methodology revealed a new bumblebee species from northeastern Brazil, with agreement between mitochondrial, nuclear, and morphological data. The large ranges of bumblebee species in Brazil, in addition to the lack of basic biological studies, constitute an auspicious scenario for further unveiling new species currently overlooked in species complexes. The recent reports about the decline in range or abundance of some of these valuable pollinators mean that there is a pressing need to monitor all bumblebee species (Williams and Osborne 2009). Hence, there is a need for revision of the taxonomy and nomenclature of the Brazilian species to give reliable support to further research (Moure and Sakagami 1962) and to guide conservation efforts.
Taxonomic treatment
Genus Bombus Latreille, 1802
B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (Figures 2, 3, and 4)
Head in profile (19), frontal view of the head (vertex) (20), mesosoma in dorsal view (21, 22, 26, 30, 34, 39), lateral view of the body (23, 27, 31, 35, 36), dorsal view of the head (vertex) (24, 28, 32, 37), and mesoscutum (discal glabrous area) (25, 29, 33, 38) of species of Bombus Latreille (1802) (Hymenoptera: Apidae, Bombini). 19–21, 24–27: Bombus (Thoracobombus) pauloensis Friese (1913); 22–23: Bombus (Thoracobombus) brevivillus Franklin (1913); 28–31: Bombus (Thoracobombus) bellicosus Smith (1879); 32–35: Bombus (Thoracobombus) transversalis Olivier (1789); 36–39: Bombus (Thoracobombus) brasiliensis Lepeletier (1836).
Holotype. f (Queen): Holótipo // Favizia 005263 // Brasil, Bahia, Igrapiúna, BR 101-BA001, 12.X.2011, n. 216, E. Françoso leg. // B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. The specimen is in excellent condition, only with the right middle leg removed from the body for the molecular analysis, and is deposited in MZUFBA.
Paratypes. (3f, Queen): Parátipo // Favízia 005264 // Brasil, Bahia, Igrapiúna, Reserva Ecológica da Michelin, 11.X.2011, n. 204, E. Françoso leg. // B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (1f, MZUFBA); Parátipo // Favízia 005265 // Brasil, Bahia, Igrapiúna, Reserva Ecológica da Michelin, 11.X.2011, n. 205, E. Françoso leg. // B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (1f, MZUFBA); Parátipo // Favízia 005266 // Brasil, Bahia, Igrapiúna, Reserva Ecológica da Michelin, 11.X.2011, n. 206, E. Françoso leg. // B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. (1f, MZUFBA). Other specimens (13), studied only for molecular biology analysis, where also included here as Paratypes: Paratype: Parátipo // Brasil, Pará, Abel Figueiredo, 1.VII.2002, E.A.B.. Almeida // 7855–23796 (EF129) (1f, UFMG, LBEA); Parátipo // Brasil, Distrito Federal, Brasília, 27.VIII.1999, F. A. Silveira // 4900–13443 (EF116) (1f, UFMG, LBEA); Parátipo // Brasil, Tocantins, Itacajá, 17.I.1993, J.M.F. Camargo, J.A. Tavares, S.R.M. Pedro // 930450 (EF148) (1f, RPSP); Parátipo // Brasil, Paraíba, João Pessoa, IV.2009, S.S. Neto 6464 (EF20) (1f, ESALQ); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF202 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF203 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF207 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF209 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF210 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF212 (1f, LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF213 (1 f. LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF214 (1 f. LGEA); Parátipo // Brasil, Bahia, Igrapiúna, 10.XI.2011, E. Françoso leg. // EF215 (1 f. LGEA).
Diagnosis: Integument predominantly black, except by a very small spot on mandibular condyle and the light brown strigilis, plumose setae across the body brownish and simple setae black; pubescence on clypeus and labrum more sparse, on clypeus leaving a large area without pubescence; general pubescence of body shorter and aligned, especially on the head, with the tip of hair straight as if trimmed (Figures 2 (2) and 3 (11)). Clypeus flattened on the middle portion longitudinally, with a large subtriangular dorsal area contrasting with an almost vertical lateral area (Figure 2 (1)); discal glabrous area of mesoscutum wide, polished, and shiny (Figure 3 (11)).
Description: f: Structure: Total body length 27.6; forewing length 20.0; head length 6.8, width 6.1; clypeus length 2.2, width 2.9 (clypeus dorsal area length 1.8, larger width 1.9, smaller width 0.66; lateral area width 0.7); malar area length 1.2; malar area width: greatest width 1.9, mandible basis 1.4; scape length 2.7, width 0.3 (apex 0.4); length of compound eye 4.1, width 1.9; ocellus diameter: median 0.43, lateral 0.44; ocellocular distance 0.8 (1.82× lateral ocellar diameter); interocellar distance: lateral to median 0.2, between lateral 0.74; alveolocular distance 1.0; interalveolar distance 0.88; interocular distance: upper 3.4, median 3.6, lower 4.3; metatibia length 7.8, width 2.4; metabasitarsus length 5.2, width 2.0. Clypeus flattened on the middle portion longitudinally, what provide it a large flat dorsal area contrasting with an almost vertical lateral area (Figure 2 (1)).
Coloration: Integument predominantly black (Figures 2 (1–3) and 3 (10–12)), except by a very small spot on mandibular condyle and the anterior tibial spur (strigilis), light brown; antennal flagellum grayish from F3 to apex on ventral surface; tegula black; wing membrane dark amber brownish, a little darker in the anterior border and apex of the marginal and on basis of first submarginal cell, the veins blackish.
Pubescence: General setae on body long, but comparatively shorter than other similar species, and extremely dense, except on head, relatively uniform in length on the regions of body, especially on the head and mesoscutum and posterior edge of the posterior tibia, aligned with the tip of hair straight like they had been trimmed (Figures 2 (2, 3) and 3 (11, 12)). Body entirely covered by long black thick branched setae (the branches are extremely short and compact, almost imperceptible), interspersed by shorter plumose brown hairs, easily visible on the parocular area near the alveoli, on pronotal lobes, on sides of propodeum, apical border of the sternum, and on the last segment of the metasoma (dorsal and ventral sides); the longer bristles on sides of clypeus (0.61), vertex (0.8), mesoscutum (1.5), scutellum (2.0), and mesepisternum (1.8); setae of legs long, thicker, black, longer, and denser in the femurs than in the tibias, on posterior border of metatibia a little shorter (1.5; 1.0 on the anterior border), uniform in length and directed downward (0.8 on the posterior border of the metabasitarsus) (Figure 3 (12)); pilosity even longer on terga (3.2), except T1, where is extremely short and thin on disk and longer on the sides of the apical borer (0.8); microtrichia of forewing relatively long and dense in the basal two thirds of wing (0.2). A pale-yellowish-golden decumbent velvety microvilli, resembling that which covers the flagellum but longer and denser, is also observed on the external surface of the mandibles (denser in apical half), the apex of the coxae and first sternum, and ventral face of femora (denser in apical half), covering the apical third of basitarsus and entirely the other tarsomeres, especially the latter which is denser.
Sculpturing: Punctures extremely dense and marked, interspaces polished and mostly smaller than 0.3PD, the space between the points mostly represented by only the edge of the points, on vertex, mesepisternum, mesoscutum, and scutellum; some punctures a little larger on clypeus and labrum, but remarkable less marked; punctures very sparse on head, especially on clypeus (reach 3PD) (Figure 2 (1–3)), fore tibiae and corbicula and malar area where there is a large smooth area. Clypeus flattened on the middle portion longitudinally, with a large subtriangular dorsal area contrasting with an almost vertical lateral area (Figures 2 (1) and 3 (10)); discal glabrous area of mesoscutum relatively wide, polished and shiny (Figure 3 (11)).
Male: unknown.
Geographical distribution: Pará (Cabo Branco Beach, Abel Figueiredo), Distrito Federal (Brasília), Tocantins (Itacajá), Paraíba (João Pessoa), and Bahia (Igrapiúna). Basically in areas of sub-evergreen Equatorial forest, Brazilian savannah and Atlantic forest.
Comments. Among the Brazilian species of Bombus, B. brevivillus and B. morio are totally black, making them difficult to identify, especially in the field. B. brasiliensis has yellow bands on tergal hairiness, and B. pauloensis can have both morphotypes (totally black or with yellow stripes).
Although B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. is quite similar to B. brevivillus and B. morio in the color of integument, very dense pubescence and punctures, in general, it can be easily differentiated from both species by the flattened structure of clypeus on the middle portion and by its general pubescence of body aligned (uniform in length), with the tip of hair straight like they had been trimmed, and also by the relatively shorter pubescence on head (Figures 2 and 3). Although the malar area is a little shorter in B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. than in B. morio, it is wider in the new species; B. brevivillus has the shorter malar area among the three species, and it is narrower than the new species. Comparing the discal glabrous area of mesoscutum in the three species, it is smooth and shiny in B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. and in B. brevivillus (narrower in the new species) and microreticulated and dull in B. morio.
Etymology: The specific epithet, from Latin (applanatis = flattened), is a reference to flatten clypeus, main character distinguishing B. (Thoracobombus) applanatus Oliveira, Françoso & Arias, sp. nov. from the other similar black species.
Diagnosis: B. (Thoracobombus) brevivillus f (Figures 2 (4–6) and 3 (13–15)): Structure: Total body length 26.5; forewing length 18.2; head length 7.0, width 5.8; clypeus length 2.1, width 2.8; malar area length 1.0; malar area width: greatest width 1.6, mandible basis 1.2; scape length 2.8, width 0.32 (apex 0.56); length of compound eye 3.9, width 1.6; ocellus diameter: median 0.34, lateral 0.33; ocellocular distance 0.8 (2.42× lateral ocellar diameter); interocellar distance: median to lateral 0.24, between lateral 0.72; alveolocular distance 0.9; interalveolar distance 0.8; interocular distance: upper 3.2, median 3.4, lower 3.9; metatibia length 7.4, width 2.3; metabasitarsus length 4.8, width 1.8. Clypeus slightly convex on the middle portion longitudinally, without flat dorsal area (Figures 2 (4) and 3 (13)); discal glabrous area of the mesoscutum relatively narrow, but polished and shiny (Figure 3 (14)). Pubescence: General setae on body long and extremely dense, except on head, not uniform in length on the regions of body, and with the apical extremity of the bristle very thinned in relation to other portions along its length (Figures 2 (5, 6) and 3 (14, 15)); some setae of body 2× longer than others, especially on the head and mesoscutum; the longer bristles on sides of clypeus (0.87), supraclypeal area (0.5–1.7), vertex (1.1), and mesepisternum (1.4); setae of legs long, thicker, black, longer, and denser in the femurs than in the tibias, on posterior border of metatibia a little shorter (1.6; 0.83 on the anterior border), not uniform in length (0.85 on the posterior border of the metabasitarsus and 0.7 on anterior border) (Figure 3 (15)); even longer terga (2.5), except T1, where is extremely short and thin on disk and longer on the sides of the apical borer (0.4).
Diagnosis: Bombus morio f (Figures 2 (7–9) and 3 (16–18)): Structure: Total body length 28.0; forewing length 20.8; head length 7.7, width 6.1; clypeus length 2.6, width 2.8; malar area length 1.45; malar area width: greatest width 1.4, mandible basis 1.3; scape length 2.9, width 0.3 (apex 0.5); length of compound eye 4.1, width 1.8; ocellus diameter: median 0.37, lateral 0.42; ocellocular distance 0.8 (1.9× lateral ocellar diameter); interocellar distance: lateral to median 0.2, between lateral 0.7; alveolocular distance 0.9; interalveolar distance 0.8; interocular distance: upper 3.3, median 3.5, lower 4.2; metatibia length 7.4, width 2.3; metabasitarsus length 3.5, width 1.3. Clypeus convex on the middle portion longitudinally, without flat dorsal area (Figures 2 (7) and 3 (16)); discal glabrous area of the mesoscutum microreticulated and dull, well-defined (Figure 3 (17)). General setae on body long and extremely dense, except on head, not uniform in length on the regions of body, and with the apical extremity of the bristle very thinned in relation to other portions along its length (Figures 2 (8, 9) and 3 (17, 18)); some setae of body 2× longer than others, especially on the head and mesoscutum; some extremely long on lateral of clypeus (1.4); the longer bristles on sides of clypeus (1.51), supraclypeal area (1.53), vertex (1.2), and mesepisternum (1.4); setae of legs long, thicker, black, longer, and denser in the femurs than in the tibias, on posterior border of metatibia a little shorter (1.7; 1.13 on the anterior border), not uniform in length (1.64 on the posterior border of the metabasitarsus and 0.66 on anterior border) (Figure 3 (18)); even longer terga (2.2), except T1, where is extremely short and thin on disk and longer on the sides of the apical borer (1.2).
Morphological identification key for Brazilian species of Bombus Latreille (1802), modified from Moure and Sakagami (1962)
Females
1. Pubescence entirely black, except for some brownish hairs or slightly whitish at the apical end, observed in the terga and ventral half of the body (Figures 2 (1–9), 3 (10–18) and 4 (21-23)).................................................................... 2
1′. Yellow or reddish pubescence on the thorax (mesosoma) and/or abdomen (metasoma), forming stripes for both, or fully covering the dorsal side of the thorax (Figure 4 (19, 20), (24–39))......................................................... 5
2. Clypeus flattened medially, so that features a large flat subtriangular in the central area with a lateral area almost vertical (Figures 2 (1–9) and 3 (10)); pubescence of the head, thorax, and posterior edge of the posterior tibial relatively long and dense, uniform in length, with the tips of hair straight, as if they had been trimmed (Figures 2 (2, 3) and 3 (11, 12)); discal glabrous area of mesoscutum well developed, smooth, and glossy (Figure 3 (11)).............................................................. B. (T.) applanatus Oliveira, Françoso & Arias, sp. nov.
2′. Clypeus moderately convex or slightly convex, without central flat area (Figures 2 (4, 7) and 4 (19, 20)); pubescence of the head and thorax relatively long, but not uniform in length around the head and thorax, bristles with the apical end quite tapered (Figure 2 (5, 6, 8, 9)); discal glabrous area of the mesoscutum variable..................................... 3
3. Malar area longer than its width of the jaw base (Figure 2 (9)); discal glabrous area of mesoscutum microreticulated and dull, well defined (Figure 3 (17))........................ B. (T.) morio
3′. Malar area shorter than its width of the jaw base (Figure 2 (5, 6)); discal glabrous area of mesoscutum somewhat or very developed, however shiny (Figure 3 (14))................................... 4
4. Superior micropunctuated justaorbital area, at least as wide as the diameter of medium ocellus (Figure 3 (13)); shorter hairiness (1.1 in T3), thick and relatively dense, velvety (Figure 4 (22))......................................... B. (T.) brevivillus
4′. Superior micropunctuated justaorbital area very narrow, generally around one third of the ocelorbital distance; hairiness longer in general (1.4 in T3), thin and looser, especially in T4 and T5, easily allowing easy viewing of the integument through the bristles (Figure 4 (21)). melanocytic form.................... B. (T.) pauloensis
5. Pronotum, mesoscutum, and scutellum covered with yellowish or rusty pubescence, without a black hairy interalar stripe; sometimes, some yellowish hairs on T1 and T2; the last three terga rusty-hairy (Figure 4 (28–31)) ....................................... B. (T.) bellicosus
5′. Pronotum, mesoscutum, and scutellum covered with yellowish or rusty pubescence more or less developed, but always with a black hairy interalar stripe (Figure 4 (25, 33, 38)); usually with a yellow stripe on the third tergum and the last segments black-hairy and black (Figure 4 (26, 34, 39))................................................................ 6
6. Posterior glabrous discal area of mesoscutum undefined, with quite strong punctuation and somewhat more sparsely than in other regions of mesoscutum (Figure 4 (33)); short pubescence (around 0.9 in T3), stripe of pronotum, mesoscutum, scutellum, and third tergum with an intense ocher yellow (Figure 4 (33–35))......................... B. (T.) transversalis
6′. Posterior glabrous discal area of mesoscutum well defined, shiny, more or less infundibuliform, without point in the middle (Figure 4 (25, 38)); the hairiness longer and looser (minimum 1.2 in T3) (Figure 4 (26, 39))................................................................ 7
7. The yellow pubescence fully covering the mesepisternum and propodeum to bottom, beyond the pronotum strips, scutellum, first and third tergum (Figure 4 (36–39)) .......... ............................................... B. (T.) brasiliensis
7′. The yellow pubescence slightly paler, restricted to the pronotum, dorsal portion of mesosoma, and T3 (flavinic form; Figure 4 (24–27)) ......................................... B. (T.) pauloensis
References
Altschul, S.F., Gish, W., Miller, W., Myers, E.W., Lipman, D.J. (1990) Basic local alignment search tool. J. Mol. Biol. 215, 403–410
Bertsch, A., Schweer, H., Titze, A., Tanaka, H. (2005) Male labial gland secretions and mitochondrial DNA markers support species status of Bombus cryptarum and B. magnus (Hymenoptera, Apidae). Insectes Soc 52, 45–54
Bickford, D., Lohman, D.J., Sodhi, N.S., Ng, P.K.L., Meier, R., Winker, K., Ingram, K.K., Das, I. (2007) Cryptic species as a window on diversity and conservation. Trends Ecol. Evol. 22, 148–155
Blaxter, M.L. (2004) The promise of a DNA taxonomy. Philos. T. R. Soc. B 359, 669–679
Cameron, S.A., Hines, H.M., Williams, P.H. (2007) A comprehensive phylogeny of the bumble bees (Bombus). Biol. J. Linn. Soc. 91, 161–188
Collins, R.A., Cruickshank, R.H. (2012) The seven deadly sins of DNA barcoding. Mol. Ecol. Res. 13, 969–975
Corbet, S.A., Williams, I.H., Osborne, J.L. (1991) Bees and the pollination of crops and wildflowers in the European Community. Bee World 72, 47–49
Darriba, D., Taboada, G.L., Doallo, R., Posada, D. (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772
Edgar, R.C. (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl. Acids Res. 32, 1792–1797
Françoso, E., Arias, M.C. (2013) Cytochrome c oxidase I primers for corbiculate bees: DNA barcode and mini-barcode. Mol. Ecol. Res. 13, 844–850
Frankham, R., Ballou, J.D., Briscoe, D.A. (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Goulson, D., Lye, G.C., Darvil, B. (2008) Decline and conservation of bumble bees. Annu. Rev. Entomol. 53, 191–208
Hebert, P.D.N., Penton, E.H., Burns, J.M., Janzen, D.H., Hallwachs, W. (2004) Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci-Biol. 101, 14812–14817
Hines, H.M. (2008) Historical biogeography, divergence times, and diversification patterns of bumble bees (Hymenoptera: Apidae: Bombus). Syst. Biol. 57, 58–75
Hines, H.M., Cameron, S.A., Williams, P.H. (2006) Molecular phylogeny of the bumble bee subgenus Pyrobombus (Hymenoptera: Apidae: Bombus) with insights into gene utility for lower-level analysis. Invertebr. Syst. 20, 289–303
Huelsenbeck, J.P., Ronquist, F. (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755
Kawakita, A., Sota, T., Ascher, J.S., Ito, M., Tanaka, H., Kato, M. (2003) Evolution and phylogenetic utility of alignment gaps within intron sequences of three nuclear genes in bumble bees (Bombus). Mol. Biol. Evol. 20, 87–92
Kevan, P.G. (1991) Pollination: keystone process in sustainable global productivity. Acta Hort. 288, 103–110
Memmott, J., Waser, N.M., Price, M.V. (2004) Tolerance of pollination networks to species extinctions. Proc. R. Soc. B 271, 2605–2611
Michener, C.D. (2007) The bees of the world, 2nd edn. Johns Hopkins University Press, Baltimore
Moure, J.S., Sakagami, S.F. (1962) As mamangabas sociais do Brasil (Bombus Latreille) (Hymenoptera, Apoidea). Studia Entomol. 5, 65–194
Murray, T.E., Fitzpatrick, U., Brown, M.F.F., Paxton, R.J. (2008) Cryptic species diversity in a widespread bumble bee complex revealed using mitochondrial DNA RFLPs. Conserv. Genet. 9, 653–666
Pywell, R.F., Warman, E.A., Hulmes, L., Hulmes, S., Nuttall, P., Sparks, T.H., Critchley, C.N.R., Sherwood, A. (2006) Effectiveness of new agri-environment schemes in providing foraging resources for bumble bees in intensively farmed landscapes. Biol. Conserv. 129, 192–206
Ratnasingham, S., Hebert, P.D.N. (2007) Bold: The Barcode of Life Data System (www.barcodinglife.org). Mol. Ecol. Notes 7, 355–364
Sheffield, C.S., Hebert, P.D.N., Kevan, P.G., Packer, L. (2009) DNA barcoding a regional bee (Hymenoptera: Apoidea) fauna and its potential for ecological studies. Mol. Ecol. Res. (Suppl. 1), 196–207.
Simon, C., Frati, F., Becknbach, A., Crespi, B., Liu, H., Flook, P. (1994) Evolution, weighting and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 87, 651–701
Sites, J.W.J., Marshall, J.C. (2004) Operational criteria for delimiting species. Annu. Rev. Ecol. Evol. Syst. 35, 199–227
Walsh, P.S., Metzger, D.A., Higuchi, R. (1991) Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10, 506–513
Williams, P.H. (1996) Mapping variations in the strength and breadth of biogeographic transition zones using species turnover. Proc. R. Soc. B 263, 579–588
Williams, P.H. (1998) An annotated checklist of bumble bees with an analysis of patterns of description (Hymenoptera: Apidae, Bombini). Bull. Br. Mus. Nat. Hist. Entomol. 67, 79–152. Available at http://www.nhm.ac.uk/research-curation/research/projects/bombus/subgenericlist.html
Williams, P.H., Osborne, J.L. (2009) Bumblebee vulnerability and conservation world-wide. Apidologie 40, 367–387
Williams, P.H., Cameron, S.A., Hines, H.M., Cederberg, B., Rasmont, P. (2008) A simplified subgeneric classification of the bumblebees (genus Bombus). Apidologie 39, 46–74
Williams, P.H., An, J., Huang, J. (2011) The bumblebees of the subgenus Subterranobombus: integrating evidence from morphology and DNA barcodes (Hymenoptera, Apidae, Bombus). J. Linn. Soc-Lond. 163, 813–862
Williams, P.H., Brown, M.J.F., Carolan, J.C., An, J., Goulson, D., et al. (2012) Unveiling cryptic species of the bumblebee subgenus Bombus s. str. worldwide with COI barcodes (Hymenoptera: Apidae). Syst. Biodivers. 10, 21–56
Acknowledgments
We wish to thank Flávio de Oliveira Francisco, Kevin M. Flesher (Reserva Ecológica da Michelin), Silvia H. Sofia, Alessandra N. Alves, Antonio Aguiar (Coleção de Insetos da UNB), Sinval Silveira Neto (Museu de Entomologia da Esalq), Fernando A. Silveira (UFMG), Silvia R. M. Pedro (Coleção Camargo–RPSP), Aline C. Martins, Alexandre Zuntini, Jenifer Lopes, and Walmir Augusto for providing samples; Susy Coelho for laboratory maintenance and J. Richard Abbott for English review. We are also thankful to the Natural History Museum of the Federal University of Bahia (MHNBA/MZUFBA), for loaning us material for comparative studies, and to the Brazilian Pollinator Initiative. We are grateful to Tércio Alves de Lima Matos and Mardson Araújo Silva from BIOSIS (UFBA, Brazil) for assistance with photomicrography and, also, to the Food and Agriculture Organization of the United Nations (FAO), Global Environment Facility (GEF), United Nations Environment Programme (UNEP), Fundo Brasileiro para a Biodiversidade (FUNBIO), Ministério do Meio Ambiente (MMA), Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Conselho Nacional de Pesquisa (PIBIC/2008 scholarship to EF), Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP Proc. 10/50597-5 PhD and scholarship to EF 2010/20548-2 and 2013/03961-1), and Research Center on Biodiversity and Computing (BioComp), supported by the USP Provost’s Office for Research.
Author contributions
EF wrote the manuscript and performed the molecular analysis, FFO performed the morphological analysis, and MCA provided support and assistance in preparing the manuscript.
Conflict of interests
The authors declare no potential conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Marina Meixner
Une approche intégrative identifie une nouvelle espèce de bourdon (Hymenoptera: Apidae: Bombini) du nord-est du Brésil
Bombus sp. nov. / barcoding moléculaire / marqueurs nucléaires / taxonomie traditionnelle
Eine integrative Methode identifiziert eine neue Hummelart (Hymenoptera: Apidae: Bombini) in Nordost-Brasilien
Hummel / Bombus sp. nov. / DNA barcoding / nukleare Marker/ traditionelle Taxonomie
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 69 kb)
Rights and permissions
About this article
Cite this article
Françoso, E., de Oliveira, F.F. & Arias, M.C. An integrative approach identifies a new species of bumblebee (Hymenoptera: Apidae: Bombini) from northeastern Brazil. Apidologie 47, 171–185 (2016). https://doi.org/10.1007/s13592-015-0385-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-015-0385-7