Abstract
Circular RNAs (circRNAs) exert pivotal effects on regulating the progression of osteosarcoma (OS). It was found through microarray analysis that circ-0002052 is abnormally expressed in OS, but the role of circ-0002052 in OS remains obscure. The results of this research manifested that relative to that in non-tumor controls, circ-0002052 level was raised in OS tissues. Up-regulated circ-0002052 was associated with advanced stage, tumor size, and metastasis. Additionally, circ-0002052 elevation indicated a low survival rate in OS patients and silencing of circ-0002052 suppressed proliferation, migration, and invasion of OS cells. It was proved that circ-0002052 sponged miR-382 and stimulated STX6 expression, thus activating Wnt/β-catenin. The function of circ-0002052 reduction in OS cells was effectively reversed by miR-382 suppression. To sum up, it can be concluded that circ-0002052, functioning as a sponge for miR-382, enhances the activation of Wnt/β-catenin mediated by STX6 to stimulate the progression of OS, and circ-0002052 may be an underlying treatment target and a biomarker for prognosis of osteosarcoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
OS is an invasive cancer of the bone and usually occurs among teenagers [1]. In adolescent malignant tumors, the incidence rate of OS ranks only second to lymphoma [2]. OS has a high degree of malignancy, early hematogenous metastasis, high incidence of lung metastasis, rapid development of the disease, and very high mortality rate [3,4,5]. OS cases have a rather low 5-year survival rate [6]. Up to now, there are no efficacious regimens for radically curing OS. Particularly, the growth mechanism of OS still needs to explore. Today, there is an urgent need to investigate novel targets for OS therapy and improve patients’ outcomes.
CircRNAs are a type of ncRNAs that are extensively expressed in numerous cell species [7] and their 3′ and 5′ ends join to one another forming a circle-like structure [8]. Increasing studies have manifested that circRNAs exert crucial effects in varying processes related to pathology, including those in cancer [9,10,11]. For example, Ma et al. found that circ-000284 facilitates cells to proliferate and invade in cervical cancer [12]. Li et al. discovered that circDDX17 acts as a tumor suppressor in colorectal cancer [13]. Li et al. found that circRNA BCRC4 overexpression modulates miR-101/EZH2 signaling and cell apoptosis in bladder cancer [14]. In recent years, research has denoted that circRNAs are intimately associated with the pathogenesis of osteosarcoma [15, 16]. So, the microarray GSE96964 was reviewed and it was found that circ-0002052 is abnormally expressed in OS.
The role of circRNA in regulation of miRNAs has always been of concern [17]. MiRNAs, with a length of about 22–25 nt, are a kind of non-coding RNA. It is able to recognize the target gene mRNA by base complementary matching, and directly degrade or inhibit translation, so as to exert post-transcriptional regulation of genes [18, 19]. Studies have shown that hsa-miR-19-3p and hsa-miR106b-3p may be reliable markers for predicting and treating OS [20]. Besides, Xu et al. found that miR-382-suppressing tumors and that miR-382 overexpression is a new strategy to inhibit tumor metastasis and prevent CSC-induced relapse in OS [21].
In conclusion, this study clarified the effect of circ-0002052 on the proliferation, invasion, and migration of OS cells, and verified that circ-0002052 exerts its biological function by binding to miR-382 to regulate STX6 expression. In addition, it may provide us with a new perspective for the in-depth study of the diagnosis of OS.
Materials and methods
Human samples
OS and non-cancer tissues were harvested from the OS patients of the First People’s Hospital of Qujing and kept in liquid nitrogen following operation. All cases received no chemoradiotherapy prior to operation. They singed the informed content. This research received the approval from the Ethic Committee of the First People’s Hospital of Qujing.
Culture and transfection of cells
The American Type Culture Collection (USA) supplied OS cell lines, non-cancer cell line hFOB 1.19 and 293T cell line, which were then raised in DMEM with 10% FBS provided by Gibco (USA), streptomycin (100 mg/mL), and penicillin (100 U/mL). The circ-0002052 overexpressing plasmids, shRNAs targeting circ-0002052, and negative control were purchased from RiboBio (Guangzhou, China). miR-382 mimics and inhibitors were obtained from GeneCopoeia (Guangzhou, China). Transfection was carried out using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocols.
Quantitative real-time PCR (qRT-PCR)
Based on the TRIzol total RNA manual, centrifugation was performed at 4 °C, and isopropanol precipitates in the upper aqueous phase were harvested, rinsed, and dried at room temperature. Subsequently, 20–30 mL DEPC-treated water was added, RNA concentration was calculated, and RNAs were preserved in a refrigerator at − 80 °C. With reference to the Takara OneStep PrimeScript® miRNA cDNA Synthesis Kit instructions, RT was conducted, followed by PCR detection using SYBR Green I fluorescence method. STX6 primer sequences: F: 5′-CACCAACGAGCTGAGAAATAACC-3′, R: 5′-CCCTGACAACTTGCCGAGT-3′. U6 primer sequences: F: 5′-CTCGCTTCGGCAGCACA-3′, R: 5′-AACGCTTCACGAATTTGCGT-3′. GAPDH primer sequences: F: 5′-GTCAACGGATTTGGTCTGTATT-3′, R: 5′-AGTCTTCTGGGTGGCAGTGAT-3′. 2−△△Ct was utilized to calculate the relative concentration of the samples to be examined. Each experiment was repeated thrice for averaging.
Examination of proliferation
To implement CCK-8 assay, 2 × 103 cells were raised for specified time following inoculation into a ninety-six-well plate, followed by incubation with CCK-8 solution acquired from Dojindo Laboratories (Japan) for 2 h. After that, a microplate reader supplied by TECAN (Austria) was employed to measure absorbance at 450 nm. EdU experiment was implemented to test cell proliferation. Each assay was implemented thrice one by one.
Wound healing assay
Following 24 hours of transfection, a micropipette tip was used to make a straight scratch in each well center. In each well, cells migrated to the scratch, which was observed to analyze cell migration. 24 hours later, the speed of wound closure was measured and normalized to the length at 0 h. Each assay was conducted in triplicate.
Transwell invasion assay
Transwell chambers (Corning, NY, USA) coated with Matrigel (BD Biosciences) were applied to test cell invasion. The upper Transwell chamber was inoculated with the transfected cells (2.0 × 104 per well) in medium without serum, whereas the lower Transwell chamber was added with DMEM (10% FBS) as a chemoattractant. Following 48 h of incubation at 37 °C, the cells not subjected to invasion on the upper chamber were rubbed away using cotton swabs, but those undergoing invasion on the lower chamber underwent fixation in 4% paraformaldehyde for 20 min, followed by 15 min of 1% crystal violet (Sigma) staining. Cells undergoing invasion were photographed and calculated in five fields of view randomly selected in each filter using an optical microscope (Olympus).
Luciferase reporter assays
Circ-0002052 WT, circ-0002052 MUT, STX6 WT or STX6 MUT sequence were put into pmiR-reporter attained from Promega (USA). Subsequently, OS cells were treated with miR-382 mimics or miR-NC, at 48 h after which, the dual-luciferase reporter assay kit supplied by Promega (USA) was utilized to test the relative activity of luciferases.
Fractionation in the nucleus and cytoplasm
Nuclear and cytoplasmic fractions were picked with the use of PARIS Kit obtained from Life Technologies (USA) by reference to the regimen of the manufacturer. Then qRT-PCR was carried out. U6 was considered to be the nuclear control, whereas GAPDH to be the cytoplasmic control.
Statistical analysis
GraphPad Prism 6.0 from GraphPad Inc. (USA) was utilized for data processing. Experimental results were represented as mean ± standard deviation (SD). ANOVA or Student’s t test was carried out for the assessment of differences. Pearson’s correlation coefficient was used to test associations among circ-0002052, miR-382, and STX6. Besides, the log-rank test and Kaplan–Meier method were adopted for the assessment of survival rates. p < 0.05 denoted a statistically significant difference.
Results
Circ-0002052 elevation is related to poor prognosis of OS
Through analyzing the microarray GSE96964 from platform GPL19978, circ-0002052 was found to be highly expressed in OS cells (Fig. 1a). To ascertain the possible functions of circ-0002052 in OS, qRT-PCR was implemented to test its levels in OS tissues and matching non-tumor controls collected. Circ-0002052 expression was increased in OS tissues (Fig. 1b). Obviously, the levels of circ-0002052 in OS tissues at the late stage (Fig. 1c) and those subjected to metastasis (Fig. 1d) were higher. Additionally, the association between clinical characteristics and circ-0002052 was assessed. It was affirmed that circ-0002052 expression was positively correlated with late tumor stage as well as tumor size and metastasis in a positive manner, and circ-0002052 level was not overtly associated with sex or age (Table 1). Identically, circ-0002052 expression went up in OS cells (Fig. 1e). The assessment of the overall survival rate unraveled that the higher circ-0002052 expression was accompanied with a lower survival rate (Fig. 1f), suggesting that circ-0002052 is a possible marker for the prognosis of OS cases. As circ-0002052 is resistant to RNase R digestion, it was verified to possess the characteristics of circRNAs (Fig. 1g).
Circ-0002052 elevation is related to poor prognosis of OS. a Analysis of microarray GSE96964 from platform GPL19978 containing even human OS cell lines (U2OS, U2OS/MTX300, HOS, MG63, 143B, ZOS, ZOSM) and the human osteoblast hFOB1.19. CircRNAs with varying expressions are identified on the basis of the criteria of fold change > 2 and p < 0.05. bCirc-0002052 expression is tested in forty OS tissues and forty non-cancer controls. c Assessment of circ-0002052 expression in OS tissues at denoted stages. d Assessment of circ-0002052 expression in OS tissues undergoing distant metastasis or not. e Assessment of circ-0002052 expression in OS cells. f Kaplan–Meier curve is utilized to determine overall survival rate. gCirc-0002052′s resistance to RNase R digestion in OS cells. Each value represents mean ± SD in three independent experiments. *p < 0.05, **p < 0.01, **p < 0.001
Circ-0002052 sponges miR-382 in OS
To date, whether circ-0002052 could be a miRNA sponge remains unknown. We performed nuclear/cytoplasmic fractionation assay and found that circ-0002052 was mainly located in the cytoplasm of OS cells (Fig. 2a). Then, we designed shRNAs targeting circ-0002052 and confirmed their efficiency (Fig. 2b). Bioinformatic analysis using circinteractome (https://circinteractome.nia.nih.gov/) denoted that miR-382 might be a target. Moreover, circ-0002052 reduction in OS cells led to a remarkable rise in miR-382 level (Fig. 2c), and circ-0002052 expression had an inverse relationship to miR-382 in OS tissues (Fig. 2d), implying the regulatory correlation between them. Since sh-circ-0002052#2 showed the greatest knockdown efficiency, sh-circ-0002052#2 (named as sh-circ-0002052) was used for subsequent experiments. Then circ-0002052-WT or -MUT reporter was utilized for luciferase reporter assay, so as to ascertain the interaction between the two. MiR-382 mimics in HOS and MG63 cells repressed the activity of circ-0002052-WT reporter (Fig. 2e, f). What’s more, reduction assay manifested that miR-382 reduced circ-0002052 in OS cell lysates in a direct manner (Fig. 2g). Hence, circ-0002052 sponges miR-382 in OS in a direct way.
Circ-0002052 sponges miR-382 in OS. a Fractionation assays are implemented in the nucleus and cytoplasm to detect circ-0002052 expression. bCirc-0002052 expression is lowered with the use of two shRNAs specific to it. cCirc-0002052 reduction raises miR-382 expression. d Inverse relationship between circ-0002052 and miR-382 in OS tissues (r = − 0.4082). e, fCirc-0002052-WT activity is weakened by miR-382 displayed in luciferase reporter assay. g Biotin-miR-382-WT mutually acts with circ-0002052 by reduction assay. Each value represents mean ± SD in three independent experiments. *p < 0.05, ***p < 0.001; ns no significance
Circ-0002052-miR-382 regulatory loop plays a vital role in cellular functions
Afterwards, the potential roles of circ-0002052 in OS were investigated. CCK-8 and EdU incorporation assays were carried out and indicated that circ-0002052 knockdown suppressed the proliferation of HOS and MG63 cells (Fig. 3a, b). To explore whether circ-0002052 regulates metastasis and invasion, wound healing assay and Transwell assay was performed. As shown, decreased expression of circ-0002052 led to attenuated migration and invasion of HOS and MG63 cells (Fig. 3c, d). More importantly, transfection with miR-382 inhibitors reversed the effects of circ-0002052 knockdown on proliferation, migration, and invasion (Fig. 3a–d).
Circ-0002052 promotes OS progression through suppressing miR-382. a CCK-8 and b EdU assays are conducted to test cell proliferation. c Wound healing assay is implemented to assess HOS and MG63 cell migration. d Invasion of HOS and MG63 cells is evaluated via Transwell assay. HOS and MG63 cells are treated with NC shRNA, sh-circ-0002052 or co-treated with sh-circ-0002052 and miR-382 inhibitor. Each value represents mean ± SD in three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001
We next examined whether miR-382 affected cell proliferation, migration, and invasion in OS cells. Compared with negative controls, overexpression of miR-382 obviously inhibited capability of cell proliferation, migration, and invasion, and circ-0002052 up-regulation could partially reverse miR-382 suppression effect (Fig. 4a–d). Thus, circ-0002052 promotes OS progression by sponging miR-382.
Circ-0002052-miR-382 regulatory loop plays a vital role in cellular functions. a CCK-8 and b EdU assays are conducted to test cell proliferation. c Wound healing assay is implemented to assess HOS and MG63 cell migration. d Invasion of HOS and MG63 cells is evaluated via Transwell assay. HOS and MG63 cells are treated with negative controls, miR-382 mimics or co-treated with circ-0002052 overexpressing plasmids and miR-382 mimics. Each value represents mean ± SD in three independent experiments. *p < 0.05, **p < 0.01
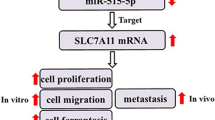
STX6 up-regulation by circ-0002052-caused miR-382 suppression induces activation of Wnt/β-catenin signaling
To further explore the molecular mechanism of circ-0002052/miR-382 axis, we further investigated downstream signaling. Through bioinformatic analysis (TargetScan7) and literature review, we selected STX6 as a potential target of miR-382. To validate it, luciferase reporter assay was conducted. MiR-382 mimics markedly reduced the activity of WT-STX6-reporter (Fig. 5a, b). Furthermore, STX6 expression had an inverse association with miR-382, but was positively related to circ-0002052 in OS tissues (Fig. 5c). Moreover, miR-382 inhibited STX6 expression in OS cells (Fig. 5d). Also, circ-0002052 reduction caused the lowered STX6 expression, but treatment with miR-382 inhibitor abolished this effect (Fig. 5d). Hence, circ-0002052 facilitates STX6 expression through impeding miR-382. Numerous studies have shown that Wnt/β-catenin is closely related to OS [22,23,24]. Hence, we sought to explore whether circ-0002052 eventually regulates activation of Wnt/β-catenin signaling and promotes OS progression. The effects of circ-0002052 on expressions of the target genes (MYC, SOX4, and CCND1) of Wnt/β-catenin signaling were tested. It was discovered that circ-0002052 reduction lessened their expressions in a manner dependent on STX6 (Fig. 5e). To sum up, circ-0002052 mediates miR-382/STX6 axis in OS to modulate the activation of Wnt/β-catenin signaling.
STX6 up-regulation by circ-0002052-caused miR-382 suppression induces activation of Wnt/β-catenin signaling. a, b MiR-382 targets STX6 in a direct way demonstrated by luciferase reporter assay. cSTX6 in OS is negatively correlated with miR-382 (r = − 0.4355), but positively related to circ-0002052 (r = 0.2799). d Expression of STX6 was measured by qRT-PCR in OS cells transfected with described plasmids. e Expression of target genes (MYC, SOX4, and CCND1) of Wnt/β-catenin signaling was tested by qRT-PCR transfected with described plasmids. Each value represents mean ± SD in three independent experiments. *p < 0.05, **p < 0.01
Discussion
Elucidating the molecular mechanism of OS development is still urgently required. It was shown in this research that circ-0002052 rose in OS tissues and had associations with late tumor stage as well as tumor size and metastasis. Apart from that, high circ-0002052 expression represented a low survival rate. Reduction of circ-0002052-suppressed OS cells to migrate, proliferate, and invade. Nevertheless, the result of invasion assay only due to proliferation and migration, or also with matrix degradation by MMPs needs further study.
The mutual action between circRNA and miRNA has been extensively researched [25, 26]. Through regulating the activity of miRNAs, circRNA indirectly promotes expression of specific mRNAs [27]. For instance, circRNA_010567 boosts myocardial fibrosis by repressing miR-141 through targeting TGF-β1 [28]. However, whether circ-0002052 has target miRNAs remains to be figured out. In the present research, circ-0002052 was discovered to be present in the OS cell cytoplasm, implying that circ-0002052 may sponge miRNA. After analysis, circ-0002052 was predicted to sponge miR-382. The mutual action between the two was subsequently confirmed by reduction assay and luciferase reporter assay. Apart from that, circ-0002052 impeded miR-382 expression in OS. MiR-382 acts as a tumor suppressor in many cancers [29, 30], including OS [31]. Thus, circ-0002052 sped up the growth of OS via repressing miR-382.
Wnt/β-catenin signaling is a widely accepted pathway in OS, so its correlation with circ-0002052 was examined. According to the findings, circ-0002052 contributed to Wnt/β-catenin signaling activation through elevating STX6 in OS cells. Nevertheless, whether PI3K/AKT, MEK/ERK, JNK, NF-kB, or Notch signaling pathways are involved in circ-0002052/miR-382/STX6 axis needs further study.
To conclude, our work demonstrated circ-0002052 as a novel oncogene in OS. Moreover, we elucidated that circ-0002052 activates Wnt/β-catenin signaling through miR-382/STX6 axis to promote OS progression.
Availability of data and materials
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.
References
Lee YJ, Chung JG, Chien YT, Lin SS, Hsu FT. Suppression of ERK/NF-kappaB activation is associated with amentoflavone-inhibited osteosarcoma progression in vivo. Anticancer Res. 2019;39(7):3669–755. https://doi.org/10.21873/anticanres.13515.
Zhou J, Xiao X, Wang W, Luo Y. Association between PTEN and clinical-pathological features of osteosarcoma. Biosci Rep. 2019;39(7):BSR20190954. https://doi.org/10.1042/BSR20190954.
Yu WC, Chen HH, Qu YY, Xu CW, Yang C, Liu Y. MicroRNA-221 promotes cisplatin resistance in osteosarcoma cells by targeting PPP2R2A. Biosci Rep. 2019;39(7):BSR20190198. https://doi.org/10.1042/BSR20190198.
Mu Y, Zhang L, Chen X, Chen S, Shi Y, Li J. Silencing microRNA-27a inhibits proliferation and invasion of human osteosarcoma cells through the SFRP1-dependent Wnt/beta-catenin signaling pathway. Biosci Rep. 2019;39(6):BSR20182366. https://doi.org/10.1042/BSR20182366.
Dou Y, Zhu K, Sun Z, Geng X, Fang Q. Screening of disorders associated with osteosarcoma by integrated network analysis. Biosci Rep. 2019;39(5):BSR20190235. https://doi.org/10.1042/BSR20190235.
Xi Y, Fowdur M, Liu Y, Wu H, He M, Zhao J. Differential expression and bioinformatics analysis of circRNA in osteosarcoma. Biosci Rep. 2019;39(5):BSR20181514. https://doi.org/10.1042/BSR20181514.
Bian L, Zhi X, Ma L, Zhang J, Chen P, Sun S, Li J, Sun Y, Qin J. Hsa_circRNA_103809 regulated the cell proliferation and migration in colorectal cancer via miR-532-3p/FOXO4 axis. Biochem Biophys Res Commun. 2018;505(2):346–52. https://doi.org/10.1016/j.bbrc.2018.09.073.
Luan W, Shi Y, Zhou Z, Xia Y, Wang J. CircRNA_0084043 promote malignant melanoma progression via miR-153-3p/Snail axis. Biochem Biophys Res Commun. 2018;502(1):22–9. https://doi.org/10.1016/j.bbrc.2018.05.114.
Zhan W, Liao X, Chen Z, Li L, Tian T, Yu L, Wang W, Hu Q. Circular RNA hsa_circRNA_103809 promoted hepatocellular carcinoma development by regulating miR-377-3p/FGFR1/ERK axis. J Cell Physiol. 2019. https://doi.org/10.1002/jcp.29092.
Yu J, Yang M, Zhou B, Luo J, Zhang Z, Zhang W, Yan Z. CircRNA-104718 acts as competing endogenous RNA and promotes hepatocellular carcinoma progression through microRNA-218-5p/TXNDC5 signaling pathway. Clin Sci. 2019;133(13):1487–503. https://doi.org/10.1042/CS20190394.
Zhao F, Han Y, Liu Z, Zhao Z, Li Z, Jia K. circFADS2 regulates lung cancer cells proliferation and invasion via acting as a sponge of miR-498. Biosci Rep. 2018;38(4):BSR20180570. https://doi.org/10.1042/BSR20180570.
Ma HB, Yao YN, Yu JJ, Chen XX, Li HF. Extensive profiling of circular RNAs and the potential regulatory role of circRNA-000284 in cell proliferation and invasion of cervical cancer via sponging miR-506. Am J Transl Res. 2018;10(2):592–604.
Li XN, Wang ZJ, Ye CX, Zhao BC, Li ZL, Yang Y. RNA sequencing reveals the expression profiles of circRNA and indicates that circDDX17 acts as a tumor suppressor in colorectal cancer. J Exp Clin Cancer Res. 2018;37(1):325. https://doi.org/10.1186/s13046-018-1006-x.
Li B, Xie F, Zheng FX, Jiang GS, Zeng FQ, Xiao XY. Overexpression of CircRNA BCRC4 regulates cell apoptosis and MicroRNA-101/EZH2 signaling in bladder cancer. J Huazhong Univ Sci Technol Med Sci. 2017;37(6):886–90. https://doi.org/10.1007/s11596-017-1822-9.
Xiao-Long M, Kun-Peng Z, Chun-Lin Z. Circular RNA circ_HIPK3 is down-regulated and suppresses cell proliferation, migration and invasion in osteosarcoma. J Cancer. 2018;9(10):1856–62. https://doi.org/10.7150/jca.24619.
Huang L, Chen M, Pan J, Yu W. Circular RNA circNASP modulates the malignant behaviors in osteosarcoma via miR-1253/FOXF1 pathway. Biochem Biophys Res Commun. 2018;500(2):511–7. https://doi.org/10.1016/j.bbrc.2018.04.131.
Jiang X, Wu X, Chen F, He W, Chen X, Liu L, Tang H. The profiles and networks of miRNA, lncRNA, mRNA, and circRNA in benzo(a)pyrene-transformed bronchial epithelial cells. J Toxicol Sci. 2018;43(4):281–9. https://doi.org/10.2131/jts.43.281.
Li Q, Wang H, Peng H, Huang Q, Huyan T, Huang Q, Yang H, Shi J. MicroRNAs: key players in bladder cancer. Mol Diagn Ther. 2019. https://doi.org/10.1007/s40291-019-00410-4.
Pishkari S, Paryan M, Hashemi M, Baldini E, Mohammadi-Yeganeh S. The role of microRNAs in different types of thyroid carcinoma: a comprehensive analysis to find new miRNA supplementary therapies. J Endocrinol Invest. 2018;41(3):269–83. https://doi.org/10.1007/s40618-017-0735-6.
Wang M, Xie R, Si H, Shen B. Integrated bioinformatics analysis of miRNA expression in osteosarcoma. Artif Cells Nanomed Biotechnol. 2017;45(5):936–43. https://doi.org/10.1080/21691401.2016.1196456.
Xu M, Jin H, Xu CX, Sun B, Song ZG, Bi WZ, Wang Y. miR-382 inhibits osteosarcoma metastasis and relapse by targeting Y box-binding protein 1. Mol Ther. 2015;23(1):89–988. https://doi.org/10.1038/mt.2014.197.
Xia P, Gao X, Shao L, Chen Q, Li F, Wu C, Zhang W, Sun Y. Down-regulation of RAC2 by small interfering RNA restrains the progression of osteosarcoma by suppressing the Wnt signaling pathway. Int J Biol Macromol. 2019;137:1221–31. https://doi.org/10.1016/j.ijbiomac.2019.07.016.
Tan T, Chen J, Hu Y, Wang N, Chen Y, Yu T, Lin D, Yang S, Luo J, Luo X. Dihydrotanshinone I inhibits the growth of osteosarcoma through the Wnt/beta-catenin signaling pathway. Onco Targets Ther. 2019;12:5111–222. https://doi.org/10.2147/OTT.S204574.
Huang Q, Shi SY, Ji HB, Xing SX. LncRNA BE503655 inhibits osteosarcoma cell proliferation, invasion/migration via Wnt/beta-catenin pathway. Biosci Rep. 2019;39(7):BSR20182200. https://doi.org/10.1042/BSR20182200.
Jin X, Feng CY, Xiang Z, Chen YP, Li YM. CircRNA expression pattern and circRNA-miRNA-mRNA network in the pathogenesis of nonalcoholic steatohepatitis. Oncotarget. 2016;7(41):66455–67. https://doi.org/10.18632/oncotarget.12186.
Su Y, Xu C, Liu Y, Hu Y, Wu H. Circular RNA hsa_circ_0001649 inhibits hepatocellular carcinoma progression via multiple miRNAs sponge. Aging. 2019. https://doi.org/10.18632/aging.101988.
Rong D, Sun H, Li Z, Liu S, Dong C, Fu K, Tang W, Cao H. An emerging function of circRNA-miRNAs-mRNA axis in human diseases. Oncotarget. 2017;8(42):73271–81. https://doi.org/10.18632/oncotarget.19154.
Zhou B, Yu JW. A novel identified circular RNA, circRNA_010567, promotes myocardial fibrosis via suppressing miR-141 by targeting TGF-beta1. Biochem Biophys Res Commun. 2017;487(4):769–75. https://doi.org/10.1016/j.bbrc.2017.04.044.
Feng J, Qi B, Guo L, Chen LY, Wei XF, Liu YZ, Zhao BS. miR-382 functions as a tumor suppressor against esophageal squamous cell carcinoma. World J Gastroenterol. 2017;23(23):4243–51. https://doi.org/10.3748/wjg.v23.i23.4243.
Yao H, Xia D, Li ZL, Ren L, Wang MM, Chen WS, Hu ZC, Yi GP, Xu L. MiR-382 functions as tumor suppressor and chemosensitizer in colorectal cancer. Biosci Rep. 2018. https://doi.org/10.1042/BSR20180441.
Xu M, Jin H, Xu CX, Sun B, Mao Z, Bi WZ, Wang Y. miR-382 inhibits tumor growth and enhance chemosensitivity in osteosarcoma. Oncotarget. 2014;5(19):9472–83. https://doi.org/10.18632/oncotarget.2418.
Author information
Authors and Affiliations
Contributions
PZ made substantial contributions to conception and design. JR, RS, and YL made acquisition of data. PZ and JR performed the experiments. JW and RS wrote the draft manuscript. All authors contributed to the reviewing of the manuscript, and approved the final manuscript for submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no conflict of interest.
Ethical standards
The study was performed in accordance with the Declaration of Helsinki and obtained the approval from the Ethics Committee of the First People’s Hospital of Qujing.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Pr., Ren, J., Wan, Js. et al. Circular RNA hsa_circ_0002052 promotes osteosarcoma via modulating miR-382/STX6 axis. Human Cell 33, 810–818 (2020). https://doi.org/10.1007/s13577-020-00335-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-020-00335-9