Abstract
Stomatal closure is an inducible form of defense that plants exert upon activation of pattern-triggered immunity (PTI). Arabidopsis long-chain base kinase 1 (LCBK1) phosphorylates phytosphingosine, which is essential for PTI-induced stomatal closure. Impairment of stomatal closure of lcbk1 mutants can be rescued by exogenous application of phosphorylated phytosphingosine. PTI-induced stomatal closure also requires salicylic acid (SA). However, the role of LCBK1 in SA-mediated stomatal closure was not known. Here, we have shown that lcbk1 mutants are defective in pathogen-induced SA accumulation and show a reduced level of expression of SA biosynthesis genes such as ICS1, PAD4, and APD1. Interestingly, the exogenous application of SA does not entirely restore the loss of immunity against pathogens in lcbk1 mutants. The lcbk1 mutants are also partially defective in SA-mediated stomatal closure. Application of phytosphingosine-phosphate activate stomatal closure in WT but not in SA biosynthetic mutant sid2. LCBK1 interacts with polycomb-group repressor complex 2 protein MEDEA, which functions as an attenuator of SA-mediated defense. However, MEDEA is not involved in SA-mediated stomatal closure. Results altogether suggest that LCBK1 functions at the upstream of SA biosynthesis as well as at the downstream for SA-mediated stomatal immunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have evolved diverse strategies to combat invading pathogens. Pattern-recognition receptors (PRRs) present on the cell surface recognize conserved molecular patterns present on the pathogens and activate pattern-triggered immunity (PTI) (Jones and Dangl 2006; Dodds and Rathjen 2010). The specific non-host type structural motifs present in microbial pathogens, such as cell walls and flagella components, are examples of microbe/pathogen-associated molecular patterns (M/PAMPs). Successful pathogens suppress PTI by releasing effector molecules. During evolution, plants also developed systems of recognition of specific effectors comprising one or more resistance (R) proteins to activate a higher level of defense, known as effector-triggered immunity (ETI). Pathogen invasion results a series of responses, such as accumulation of reactive oxygen species, over-production of several phytohormones, other phytochemicals, and defense-related proteins, which are common for both PTI and ETI (Spoel and Dong 2012). Plant hormones salicylic acid (SA), ethylene (ET), and jasmonic acid (JA) play crucial roles in mounting defense responses. Plants also protect themselves by restricting the entry of pathogens. Plants strengthen the cell wall barrier by lignin or callose deposition. Bacterial pathogens often enter the host plant through stomatal pores. Stomatal closure is one of the PTI responses that restricts bacterial entry (Melotto et al. 2006, 2017).
Hormonal crosstalk is essential for stomatal immunity. Abscisic acid (ABA)-mediated stomatal closure during drought stress has been well studied. ABA functions as both positive and negative regulators of plant defense, depending on the stage of infection. At the early stage of infection, ABA promotes defense by enhancing stomatal closure (Ton et al. 2009). However, ABA also functions antagonistically with SA signaling and suppresses SA-mediated plant defense and callose deposition (Ton et al. 2009). In addition to ABA, SA has also been reported for the closure of stomata. SA deficient Arabidopsis mutants such as SA induction deficient 2 (sid2; allelic to ics1- isochorismate synthase 1) or transgenic plants expressing SA hydroxylase coded by nahG are impaired for PTI-mediated stomatal closure (Melotto et al. 2006). SA-mediated stomatal closure also requires NO and ROS production, similar to ABA (Melotto et al. 2017). Ethylene (ET) can also close the stomata on intact leaves but inhibits ABA-induced closure of stomata on epidermal peels (Tanaka et al. 2005; Desikan et al. 2006). Antagonistically, jasmonic acid (JA) signaling acts negatively during the stomatal defense. Bacterial pathogens also evolved mechanisms to reinstate the opening of stomata. Experiments showed that plants close stomata within an hour of the virulent pathogen Pseudomonas syringae pv. tomato DC3000 (Pst DC3000) inoculation (Melotto et al. 2006). However, Pst is capable of reopening stomata within three hours with the help of secreted toxin coronatine (COR) (Melotto et al. 2006; Gupta et al. 2020). COR is a structural analog of JA-isoleucine, a biologically active form of JA. COR activates the JA signaling pathway and suppresses SA signaling pathway. Activation of JA/COR responsive genes, like NAC transcription factors, inhibits the SA accumulation and promotes COR-induced stomatal reopening and bacterial multiplication in plant tissues (Robert-Seilaniantz et al. 2011; Zheng et al. 2012). There are many effectors which can commonly target JA signaling response to conquer stomatal defense in plants. For example, P. syringae effector AvrB requires the JA signaling pathway to induce stomatal opening in a RIN4-dependent manner (Zhou et al. 2015).
Recently, we showed that Arabidopsis LCBK1 interacts with the PRC2 complex member protein MEDEA (MEA) (Gupta et al. 2020). LCBK1 phosphorylates phytosphingosine (PHS) to phytosphingosine-1-phosphate (PHS-P), a process that is required for stomatal immunity (Gupta et al. 2020). Though LCBK1 localization has not been established, it’s interaction with MEA suggests it be a plasma-membrane localized protein (Gupta et al. 2020). MEA helps in attenuating PTI and ETI in Arabidopsis (Roy et al. 2018). LCBK1 and MEA oppositely regulate plant defense. Whereas MEA is a negative regulator, LCBK1 is a positive regulator of defense against bacterial pathogens (Roy et al. 2018; Gupta et al. 2020). However, the role of LCBK1 is SA-mediated stomatal immunity was not known. Here, we report the positive regulatory role of LCBK1 in SA biosynthesis and a cooperative role of PHS-P and SA in stomatal immunity.
Material and method
Plant and pathogens materials, growth condition, SA treatment and infection experiments
The T-DNA insertion mutants lcbk1-2 (Salk_152371C) and lcbk1-3 (SAIL_529_H04), and bacterial pathogen Pseudomonas syringae pv tomato DC3000 (Pst) have been described previously (Gupta et al. 2020). MEA over-expression and mutant lines were also described earlier (Roy et al. 2018). Plants were grown in a growth room at 22 °C and 65% relative humidity with an alternate light /dark period of 12 h each (Roy et al. 2018). Pathogen inoculation and method of determination of bacterial load were followed exactly as described previously (Singh et al. 2014; Roy et al. 2018; Gupta et al. 2020). In brief, overnight grown bacterial cultures were resuspended in 10 mM MgCl2. Inoculation was carried out by spraying or infiltrating with a needless syringe through the abaxial surface of leaves. For SA treatment, 4-week-old plants were sprayed with only water as the control or a solution of 500 µM SA (Sigma-Aldrich) made in distilled water, and plants were covered overnight with a plastic dome. The pathogen was sprayed after 24 h of SA treatment.
Stomatal aperture measurement
Stomatal aperture size was measured as described previously (Gupta et al. 2020). In brief, abaxial epidermal peels were excised and floated on the stomatal opening buffer (MES-KOH-10 mM, KCl-30 mM; pH 6.15) with or without SA and PHS-P (Avanti polar lipids Cat# 860491). SA and PHS-P stock solutions were prepared in ethanol and ethanol:DMSO (2:1), respectively, and diluted in the stomatal opening buffer. Diluted SA (10 µM) or PHS-P (10 µM) were applied to the peels and incubated under light from 1 to 3 h, and observed under a light microscope. Stomatal apertures was calculated using ImageJ software using microscopic images (Gupta et al. 2020).
RNA isolation, cDNA synthesis, and expression analysis
Extraction of total RNA, synthesis of cDNA and quantitative real-time PCR (qPCR) were carried out as described earlier (Roy et al. 2018). cDNA was made from 1.0 µg of DNAse-treated RNA by using the Bio-Rad cDNA synthesis kit (catalog # 170–8891). Each sample consisted of three biological replicates. qPCR was carried out for each sample with two technical replicates. The average Ct values of two technical replicates was taken for the calculation. We used BioRad (CFX connect) system for qPCR with gene-specific primers and 2 × SYBR Green master mix (Bio-Rad; catalog no. #172–5124). Samples were normalized with TUBULIN2 (At5g62690) mRNA level for quantification. The following primers were used for qPCR. TUB2: AGCAATACCAAGATGCAACTGCG and TAACTAAATTATTCTCAG TAC; PAD4: TTCTTTTCCCCGGCTTAT and GTTATCACCACCAGCTTTTTACC; APD1: TCAAAACATATGCATTTCAGGTCT and GTGGATTTGTTTCTTCTCGACTTT; ICS1: CTAATCTCCGCCGTCTCTGAACT and TTGGAACCTGTAACCGAACGA; PR2-ATCGTTGGAAATCGTGGTGTC and TAGCTTTCCCTGGCCTTCTC.
SA estimation
SA content was determined by luminescence assay using Acinetobacter sp. ADPWH_lux (Huang et al. 2005; DeFraia et al. 2008). Overnight-grown Acinetobacter sp. ADPWH_lux culture was diluted 20 times in LB media and grown further for 2 to 3 hours (~ 0.4OD) at 28 °C, before using in the assay. Leaf samples (100 mg) were frozen in liquid N2 and homogenized in 250 µl acetate buffer (0.1M, pH 5.6). This crude extract was centrifuged at 12,000 rpm for 15 min. The supernatant was divided equally and transferred into two tubes. One tube was treated with β-glucosidase (Sigma, USA Cat # G0395) was incubated for 1.5 h at 37 °C, for total SA. The other tube was kept on ice for free SA. All samples were centrifuged for 5 min at 12,000 rpm. 20 µl of extract + 60 µl LB + 50 µl Acinetobacter (0.4OD) was added in a microplate and incubated for 1hr at 37 °C. Luminescence was observed in POLAR star Omega (BMG Labtech) luminometer.
Results
lcbk1 mutant plants are defective in SA-mediated stomatal immunity
ABA-mediated stomatal closure during abiotic and biotic stress has been well documented. Besides ABA, stomatal closure is also influenced by SA (Melotto et al. 2006). Our earlier studies demonstrated lcbk1 mutants are also defective in pathogen-induced stomatal closure (Gupta et al. 2020). In the current study, we examined the effect of SA on stomatal closure of lcbk1 mutant. We exogenously applied SA on epidermal peels of WT and lcbk1-2 and observed stomatal aperture. As expected, SA closes the stomata in WT (Fig. 1a). SA treatment also induced stomatal closure in lcbk1-2 plants, but to a lower extent than WT (Fig. 1a). Results suggested that the SA-mediated stomatal closure is partly dependent on the LCBK1 function.
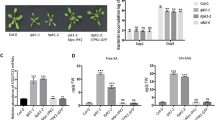
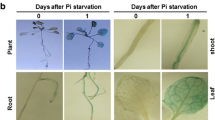
Stomatal aperture measurement and disease defense response after SA treatment in WT, lcbk1-2 and lcbk1-3. a Stomatal aperture after 10 µM SA treatment for 2 h in WT and lcbk1-2. Each bar represents the mean ± S.D. (n ~ 80). b Bacterial load and disease symptoms after spraying SA (500 µM) or water in indicated genotypic plants. Pst at 5 × 108 CFU/ml was sprayed after 24 h of SA or water treatment and colonies were counted after 4 days of spray inoculation. Each bar represents the mean ± S.D. (n = 04). c Relative level of PR2 mRNA after Pst inoculation. *(P < 0.05) and **(P < 0.001) indicate the mean values of lcbk1-2, lcbk1-3 or npr1-1 plants are significantly different from mock or SA-treated WT samples as determined by student’s t-test
To further investigate whether LCBK1 is also required for SA-mediated stomatal immunity, we examined the effect of exogenous SA application on disease resistance in WT, lcbk1-2, lcbk1-3 and npr1-1 plants. The npr1-1 mutant is defective in SA response and thus was taken as a control for the study (Pieterse and Van Loon 2004; Singh et al. 2018). Five-week-old soil-grown plants were sprayed with water or SA (500 µM). After 24 h of SA spray, all plants were spray-inoculated with Pst. As expected, SA treatment provided resistance to WT but not in npr1 plants. The lcbk1 mutant plants responded to SA application but to a significantly lower extent than WT plants (Fig. 1b). Previously we showed that lcbk1-2 plants are impaired in pathogen-induced PR1 gene expression (Gupta et al. 2020), which is in agreement with impaired SA biosynthesis. To further confirm, we investigated accumulation of PR2 after pathogen inoculation. We inoculated WT and lcbk1-2 plants with either 10 mM MgCl2 (mock) or by Pst and relative PR2 mRNA was determined at 0, 12 and 24 hours-post-inoculation. We observed significantly lower level of PR2 mRNA in lcbk1-2 compared to WT plants (Fig. 1c). Together, these results demonstrated that LCBK1 promotes SA-mediated stomatal closure and stomatal immunity in Arabidopsis.
LCBK1 promotes pathogen-induced SA accumulation
LCBK1 is a positive regulator of defense and pathogen-induced PR1 transcript accumulation (Gupta et al. 2020). To examine whether LCBK1 influences SA biosynthesis, we measured basal and pathogen-induced SA accumulation in WT, lcbk1-2, and lcbk1-3 plants. Plants were inoculated with Pst suspended in 10 mM MgCl2 or only 10 mM MgCl2 as mock. Samples were harvested at 12 h-post-inoculation for SA estimation. We observed pathogen-induced SA accumulation in WT plants but not in lcbk1 mutant plants (Fig. 2), suggesting that LCBK1 function is essential for SA accumulation in Arabidopsis.
Total SA (SA + SAG) or free SA content in mock or Pst-inoculated WT, lcbk1-2, and lcbk1-3 plants. Plants were infiltrated with Pst, and samples were harvested at 12 hours post-inoculation (hpi) for SA quantification. Each bar represents the mean ± S.D. (n = 03). **Indicated a statistically significant difference (P < 0.001) in Pst-treated samples with corresponding mock-treated samples, as obtained by student’s t-test
LCBK1 positively regulates expression of SA biosynthetic genes
To further investigate whether SA biosynthesis is regulated by LCBK1 at the transcriptional level, we monitored the expression of a few genes that positively regulate SA biosynthesis. Arabidopsis ISOCHORISMATE SYNTHASE1 (ICS1) converts chorismate to isochorismate, which is the rate-limiting step of pathogen-induced SA biosynthesis (Wildermuth et al. 2001). The AP2 FAMILY PROTEIN INVOLVED IN DISEASE DEFENSE (APD1) is the AP2/ERF family transcription factor that positively regulates ICS1 expression, SA biosynthesis, and SA-mediated defense in Arabidopsis (Giri et al. 2014). The PHYTOALEXIN DEFICIENT 4 (PAD4) codes for a lipase-like gene and positively regulates expression of APD1 (Jirage et al. 1999; Giri et al. 2014). We observed pathogen-induced expression of all the three SA biosynthesis genes were compromised in lcbk1-2 mutant compared to WT plants (Fig. 3). Results altogether demonstrated that LCBK1 function is essential for pathogen-induced SA biosynthesis in Arabidopsis.
The relative abundance of transcripts of ICS1, PAD4, and APD1 after mock or Pst inoculation in WT and lcbk1-2 plants. Expression analysis was done by qRT-PCR. Plants were inoculated with Pst at 106 CFU/ml suspended in 10 mM MgCl2 or 10 mM MgCl2 as mock control. Samples were harvested after 12 h post-inoculation. Each bar represents the mean ± S.D. (n = 03). * (P < 0.05) and ** (P < 0.001) indicate the mean values of lcbk1-2 plants are significantly different from respective WT samples as determined by student’s t-test
SA is essential for phytosphingosine-1-phosphate (PHS-P)-mediated stomatal closure
Previous observation suggested LCBK1 phosphorylates phytosphingosine, which leads to PTI-induced stomatal closure (Gupta et al. 2020). Exogenous application of PHS-P alone can induce stomatal closure in the absence of pathogen or PTI inducers such as flg22, suggesting that LCBK1 functions downstream of PTI activation (Gupta et al. 2020). Since LCBK1 also positively regulates SA-mediated stomatal closure (Fig. 1), we hypothesized phosphorylation of PHS is a downstream event of SA accumulation. To that end, we examined PHS-P mediated stomatal closure in WT and SA deficient mutant ics1/sid2. As expected, we found significant reduction in the stomatal aperture size in WT after 1 h and 3 h post PHS-P inoculation (Fig. 4). However, in contrary to the expectation, PHS-P application failed to close stomata in sid2 plants (Fig. 4). This observation is interesting, as it suggests that PHS-P alone is not sufficient to close stomata and SA is an essential component for PHS-P-mediated stomatal closure.
PHS-P induced stomatal closure in WT and sid2 plants. Stomatal aperture sizes were observed through microscopy at 1 h and 3 h post-inoculation in WT and sid2. Each bar represents the mean ± SD (n = ~ 80). Different letters above the bars indicated a statistically significant difference (P < 0.001) as obtained by one-way ANOVA (Holm–Sidak method)
MEA does not influence SA-mediated stomatal closure
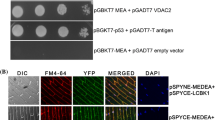
Previously, we observed that MEA over-expression lines were defective in PTI and PTI-induced stomatal closure (Roy et al. 2018; Gupta et al. 2020). Since LCBK1 interacts with MEA and promotes SA-mediated stomatal immunity, we wanted to investigate the role of MEA in this process. We exogenously applied SA on WT, MEA over-expressing, and mea mutant plants and recorded stomatal aperture. We observed no significant difference between WT and MEA Oex plants (Fig. 5a) or between WT and mea mutant plants (Fig. 5b) in terms of SA-mediated stomatal closure. The results showed that SA-induced stomatal closure defect in lcbk1 mutants (Fig. 1a) is unlikely to be influenced by MEA.
Stomatal aperture measurement after SA treatment in WT, MEA Oex and mea-6. Stomatal aperture size was observed through microscopy after 10 µM SA treatment for 2 h in a WT and MEA Oex b WT and mea-6. Each bar represents mean ± S.D. (n ~ 80). Each bar represents the mean ± SD (n = ~ 80). Different letters above the bars indicated a statistically significant difference (P < 0.001) as obtained by one-way ANOVA (Holm–Sidak method)
Discussion
Our data revealed a complex interaction of SA and LCBK1 in stomatal immunity. LCBK1 codes for an enzyme that phosphorylates PHS to PHS-P (Imai and Nishiura 2005). LCBK1 plays crucial roles in ABA and PTI-induced stomatal closure. However, the mechanisms by which LCBK1 or PHS-P contributes to stomatal closure is still not known. Our results show a dual role of LCBK1 in stomatal immunity (Fig. 6). LCBK1 transcriptionally upregulates SA biosynthesis genes such as ICS1, PAD4, and APD1. Unfortunately, the regulatory mechanisms of expression of these genes are not much known. Both PAD4 and APD1 promote ICS1 expression, and conversely, expressions of PAD4 and APD1 are upregulated by SA in a positive feedback loop (Giri et al. 2014). Thus, it is possible that LCBK1 may directly promote any of these genes, and other genes may be upregulated in the feedforward mechanism (Fig. 6). Nevertheless, our data reveal that pathogen-induced SA accumulation, at least in the early hours of inoculation, requires LCBK1 (Fig. 1a). Intriguingly, the exogenous application of SA does not completely rescue the loss-of-immunity of lcbk1 plants (Fig. 1b). This result suggested that role of LCBK1 is not limited to SA biosynthesis only.
Pathogen or PTI-induced stomatal closure is an early defense response, which functions within an hour of pathogen inoculation. Although stomatal dynamics is dependent on hormonal crosstalk, SA signaling is important among them for PAMP-induced stomatal closure (Melotto et al. 2006, 2017). In addition to SA, ethylene is also known to close the stomata in intact leaves (Desikan et al. 2006). Antagonistically, JA signaling acts negatively during the stomatal defense. PstDC3000 releases COR that can reopen the stomata at 3hr post-inoculation (Melotto et al. 2006). COR activates the JA signaling pathway to antagonize the SA signaling pathway (Robert-Seilaniantz et al. 2011; Zheng et al. 2012). The lcbk1 mutants being defective in SA biosynthesis may also promote JA signaling during PAMP-triggered stomatal immunity. Thus, the partial rescue of disease defense phenotype and stomatal closure after exogenous SA application in lcbk1 plants may have been attributed towards the indirect effect of JA signaling. However, our experiments of PHS-P application in sid2 plants suggested a direct role of SA and LCBK1 in the process of stomatal closure. LCBK1 has been known to phosphorylate PHS, which can induce stomatal closure in WT plants (Imai and Nishiura 2005; Gupta et al. 2020). We observed SA biosynthetic mutant sid2 was insensitive to PHS-P application compared to WT (Fig. 4). Altogether, our results demonstrated that SA and PHS-P both contribute to stomatal closure (Fig. 6). Upon pathogen inoculation, expression of SA biosynthetic genes as well as LCBK1, are upregulated. LCBK1 converts PHS to PHS-P, which together with SA activates stomatal closure (Fig. 6).
Data availability
Data available with both the authors. AKN may be contacted for materials.
Abbreviations
- ABA:
-
Abscisic acid
- APD1:
-
AP2 family protein involved in disease defense 1
- COR:
-
Coronatine
- ET:
-
Ethylene
- ETI:
-
Effector-triggered immunity
- ICS1:
-
Isochorismate synthase 1
- JA:
-
Jasmonic acid
- LCBK1:
-
Long-chain base kinase 1
- MEA:
-
MEDEA
- NPR1:
-
Non-expressor of PR 1
- PAD4:
-
Phytoalexin deficient 4
- PHS:
-
Phytosphingosine
- PHS-P:
-
Phytosphingosine-1-phosphate
- PRC2:
-
Polycomb-group repressor complex 2
- PRR:
-
Pattern recognition receptor
- PTI:
-
Pattern-triggered immunity
- SA:
-
Salicylic acid
- SID2:
-
SA induction deficient 2
References
DeFraia CT, Schmelz EA, Mou Z (2008) A rapid biosensor-based method for quantification of free and glucose-conjugated salicylic acid. Plant Methods 4:28
Desikan R, Last K, Harrett-Williams R, Tagliavia C, Harter K, Hooley R, Hancock JT, Neill SJ (2006) Ethylene-induced stomatal closure in Arabidopsis occurs via AtrbohF-mediated hydrogen peroxide synthesis. Plant J 47:907–916
Dodds P, Rathjen J (2010) Plant immunity: Towards an integrated view of plant-pathogen interactions. Nat Rev Genet 11:539–548
Giri MK, Swain S, Gautam JK, Singh S, Singh N, Bhattacharjee L, Nandi AK (2014) The Arabidopsis thaliana At4g13040 gene, a unique member of the AP2/EREBP family, is a positive regulator for salicylic acid accumulation and basal defense against bacterial pathogens. J Plant Physiol 171:860–867
Gupta P, Roy S, Nandi AK (2020) MEDEA-interacting protein LONG-CHAIN BASE KINASE 1 promotes pattern-triggered immunity in Arabidopsis thaliana. Plant Mol Biol 103:173–184
Huang WE, Wang H, Zheng H, Huang L, Singer AC, Thompson I, Whiteley AS (2005) Chromosomally located gene fusions constructed in Acinetobacter sp. ADP1 for the detection of salicylate. Environ Microbiol 7:1339–1348
Imai H, Nishiura H (2005) Phosphorylation of sphingoid long-chain bases in Arabidopsis: functional characterization and expression of the first sphingoid long-chain base Kinase gene in plants. Plant Cell Physiol 46:375–380
Jirage D, Tootle TL, Reuber TL, Frost LN, Feys BJ, Parker JE, Ausubel FM, Glazebrook J (1999) Arabidopsis thaliana PAD4 encodes a lipase-like gene that is important for salicylic acid signaling. Proc Natl Acad Sci U S A 96:13583–13588
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Melotto M, Underwood W, Koczan J, Nomura K, He SY (2006) Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–980
Melotto M, Zhang L, Oblessuc PR, He SY (2017) Stomatal defense a decade later. Plant Physiol 174:561–571
Pieterse CM, Van Loon LC (2004) NPR1: the spider in the web of induced resistance signaling pathways. Curr Opin Plant Biol 7:456–464
Robert-Seilaniantz A, Grant M, Jones JD (2011) Hormone crosstalk in plant disease and defense: more than just jasmonate-salicylate antagonism. Annu Rev Phytopathol 49:317–343
Roy S, Gupta P, Rajabhoj MP, Maruthachalam R, Nandi AK (2018) The polycomb-group repressor MEDEA attenuates pathogen defense. Plant Physiol 177:1728–1742
Singh V, Roy S, Singh D, Nandi A (2014) Arabidopsis FLOWERING LOCUS D influences systemic-acquired-resistance-induced expression and histone modifications of WRKY genes. J Biosci (Bangalore) 39:119–126
Singh N, Swain S, Singh A, Nandi AK (2018) AtOZF1 positively regulates defense against bacterial pathogens and NPR1-independent salicylic acid signaling. Mol Plant-microbe Interact 31:323–333
Spoel SH, Dong X (2012) How do plants achieve immunity? Defence without specialized immune cells. Nat Rev Immunol 12:89–100
Tanaka Y, Sano T, Tamaoki M, Nakajima N, Kondo N, Hasezawa S (2005) Ethylene inhibits abscisic acid-induced stomatal closure in Arabidopsis. Plant Physiol 138:2337–2343
Ton J, Flors V, Mauch-Mani B (2009) The multifaceted role of ABA in disease resistance. Trends Plant Sci 14:310–317
Wildermuth MC, Dewdney J, Wu G, Ausubel FM (2001) Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414:562–565
Zheng XY, Spivey NW, Zeng W, Liu PP, Fu ZQ, Klessig DF, He SY, Dong X (2012) Coronatine promotes Pseudomonas syringae virulence in plants by activating a signaling cascade that inhibits salicylic acid accumulation. Cell Host Microbe 11:587–596
Zhou Z, Wu Y, Yang Y, Du M, Zhang X, Guo Y, Li C (2015) An Arabidopsis plasma membrane proton ATPase modulates JA signaling and is exploited by the Pseudomonas syringae effector protein AvrB for stomatal invasion. Plant Cell 27:2032–2041
Acknowledgements
Arabidopsis WT and mutant seeds were obtained from the Arabidopsis Biological Resource Centre, USA. The Acinetobacter sp. ADPWH_lux strain was a kind gift from Prof. Hui Wang NERC/Centre for Ecology and Hydrology Wallingford, UK.
Funding
This work was supported by the Science and Engineering Research Board Project (SERB/SR/SO/PS/150/2012 to AKN), and the CSIR fellowship to PG.
Author information
Authors and Affiliations
Contributions
AKN conceptualized and designed most of the experiments. PG designed some of the experiments, performed all the experiments, analyzed data. Both the authors equally contributed in writing the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gupta, P., Nandi, A.K. Long-chain base kinase1 promotes salicylic acid-mediated stomatal immunity in Arabidopsis thaliana. J. Plant Biochem. Biotechnol. 29, 796–803 (2020). https://doi.org/10.1007/s13562-020-00608-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-020-00608-2