Abstract
Phenotyping and genetic assessment of germplasm provide information about trait variability, which helps for effective breeding programs. In the present study, 12 indigenous finger millet genotypes from Koraput, India and three high-yielding improved genotypes were used for elucidation of genotypic variability of photosynthetic traits and genetic diversity using 36 SCoT (Start codon targeted polymorphism) markers. Significant variations were noticed in the CO2 photosynthetic rate (PN), stomatal conductance (gs), transpiration rate (E), internal CO2 concentration (Ci), water use efficiency and carboxylation efficiency among the genotypes. Significant variations of stomatal traits, pigment content, PS II activity and dry matter accumulation were also observed. The major morpho-physiological traits such as stomatal conductance, dry matter accumulation, shoot length and stomata per leaf area are played a pivotal role and are the major determinants of phenotypic diversity. The positive association of photosynthesis with dry matter accumulation indicates that some of the genotypes remarkably have more photosynthetic rate along with better plant biomass accumulation and can be used in future crop improvement program. Further, SCoT markers were polymorphic and revealed moderately high level of genetic diversity and provided information on population structure among the finger millet genotypes. The SCoT primers, SCoT-14, SCoT-18 SCoT-20 and SCoT-23 showed the higher PIC value and marker index, and potentiality for exploring the genetic diversity of studied millet genotypes. Based on the genetic similarity analysis it is revealed that some of the indigenous finger millet genotypes such as Jhana, Lala, Kurkuti, Ladu, Bhadi and Taya showed highest genetic disimilarity with modern high yielding genotype and can be considered as the potential genetic resources for breeding program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Millets are a diverse group of small-grained cereal and one of the important staple food crops grown around the world (Singh and Sharma 2018). Unique attributes of millet such as ability to adapt under severe and adverse climate condition, least agro-input requirement and with the exceptional nutritional quality makes them crucial plant genetic resources for agriculture (Gupta et al. 2017). Finger millet is [Eleusine coracana (L.) Gaertn.] widely cultivated in arid and semi-arid regions of Africa and South Asia under rainfed conditions (Mukami et al. 2019). Finger millet is one of the important small millet belongs to Poaceae family and commonly known as “Ragi” and “Mandia” in India (Babu et al. 2017; Sakamma et al. 2018). It has been estimated that about 3.38 million hectares of global area are under finger millet cultivation with a production of 3.76 million tons of grain (FAO 2007; Babu et al. 2017). In India, finger millet ranks third next to sorghum and pearl millet as most important millet crop is grown on 1.27 million hectares (Mha) of cultivated land with an annual production of 1.89 million tons (Mt) (FAO 2007). Finger millet grains are rich in methionine and tryptophan, total aromatic amino acids, and high calcium content with superior nutritional qualities, therefore plays a significant role in nutritional security (Panda et al. 2019; Mukami et al. 2019). The crop cultivation under diverse eco-geographical zones globally signifies high genetic variations of finger millet, which could be the basis for future breeding program for climate resilient crop improvement (Assefa et al. 2013). Morphologically finger millet showed huge variations, which need to be captured at DNA level for detection of genetic diversity. Despite its economic and nutritional importance, breeding and selection of finger millet with improved traits are currently limited by the lack of adequately characterized germplasm both at in the physiological and molecular level. The dearth of knowledge regarding population structure has significantly contributed to the genetic erosion of millets. Extensive genetic diversity has been observed, especially among the genotypes for many agronomic traits. However, this diversity has not yet been tapped for crop improvement. Therefore, information on genetic diversity of finger millet is essential for their efficient utilization in the crop improvement programs.
Photosynthesis forms an integral part of the plant metabolism and balance sheet of growth and development, which is known to be regulated by internal and environmental parameters (Gupta et al. 2002; Panda et al. 2018). Previous reports suggested that the global crop production requires alteration of the primary photosynthetic process of the crops to cope up with the changing climatic scenario (Kondamudi et al. 2016; Panda et al. 2018). The increase in photosynthetic efficiency and growth in crop plants led to an increase in grain yield. Extensive studies on variations of traits associated with photosynthesis have been reported in rice, wheat and other cereal crops and were used successfully in the breeding programs (Kondamudi et al. 2016; Haritha et al. 2017; Panda et al. 2018). However, the photosynthetic efficiency in finger millets has been little studied and there is a dearth of knowledge on photosynthetic variants among traditional and modern finger millet genotypes.
Traditional genotypes possess genetic variability and adapted to local climatic condition and soil quality. Assessment of genetic variation among genotypes might help understand evolutionary relationship and pattern of genetic diversity for conservation of valuable resources as well as efficient use in the crop improvement program (Bashir et al. 2015). Currently, molecular markers have been used as an efficient tool to assess the genetic variation and relatedness among cultivars. Various molecular markers such as RAPD, RFLP, AFLP, ISSR and SSR, etc. have been used in the genetic diversity analysis in different crop species (Ramakrishnan et al. 2016). Each marker system has certain advantages and disadvantages compared to each other and influenced by crop varieties (Collard and Mackill 2009). The start codon targeted polymorphism (SCoT) marker is one of the novel marker systems is based on a short-conserved region in the plant genes flanking the ATG start codon (Collard and Mackill 2009). Earlier some attempts have been made to use molecular markers for assaying genetic diversity to estimate the relationship among African and Asian finger millet germplasm (Dida et al. 2007). For instance, genetic diversity of extensive collection of global finger millet accessions (Babu et al. 2014), genotypes from Indians (Rajendran et al. 2016) and millets of world-wide (Ramakrishnan et al. 2016; Kumar et al. 2016; Wakista et al. 2017) were assessed. However, there is a deficiency of documented information on genetic structure and diversity of indigenous finger millets of India and SCoT marker-based genetic diversity in finger millets is yet to be carried out.
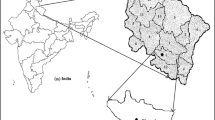
Koraput is one of the tribal-dominated districts of Odisha in India (18° 14′ to 19° 14′ N latitude and 82° 05′ to 83° 25′ E longitude) and is recently declared as one of the agro-biodiversity hot spots in India (Mishra and Chaudhury 2012). Large numbers of finger millet genotypes have been reported from Koraput and conserved in different national and base collection centres. The indigenous finger millet genotypes cultivated by traditional farmers might contain considerable genetic diversity and could serve as potential genetic resources for future crop improvement programs. For proper utilization and incorporation useful traits from these genotypes for future crop improvement program, information on genetic structure and diversity is of utmost importance. Therefore, the present study aims to evaluate genotypic variability in morpho-physiological traits and genetic diversity in selected indigenous finger millets of Koraput.
Materials and methods
Plant material and growth condition
The experiment was conducted by taking 12 indigenous finger millet genotypes from Koraput, India namely Jhana, Lala, Kurkuti, Ladu, Bati, Bhadi, Taya, Limca, Chilli, Biri, Sili janha and Dasara along with three high yielding improved varieties such as Arjuna, Chilika and Bhairabi (Table 1). These indigenous genotypes possess many primitive features and the tribal farmers are being grown to suit their local circumstances. The seeds were obtained from MS Swaminathan Research Foundation (MSSRF), Koraput, India.
Uniformly sized matured seeds of each genotype were selected and kept at 48 ± 2 °C for 5 days to break the seed dormancy. The seeds were directly sown in plastic pots (45 cm in diameter) having two kg of farm soil and farmyard manure in a ratio of 3:1. The seedlings were thinned after 10 days of germination and ten plants per pots were kept for further studies. Each pot was provided with 190 mg single super phosphate (P2O5) and 50 mg murate of potash (K2O). N-fertilizer in the form urea at 1 g per pot was applied after 10, 30 and 50 days of sowing. Plants were regularly irrigated with tap water and subjected to natural solar radiations, with daily maximum photosynthetic photon flux density, air temperature and relative humidity being about 1360 ± 20 µmol m−2 s−1, 31.6 ± 2 °C and 65–70%, respectively throughout the experiment. The experiments were conducted with three replications in each variety.
Measurement of plant growth parameters
The seedling growth parameters of finger millets were estimated by measuring shoot and root length, fresh weight and dry weight of seedling during the flowering stage. The shoot and root dry mass of each replication were measured after drying at 80 °C until a constant weight recorded. The dry matter accumulation (DMA) was determined by the following formula:
The leaf area (LA) was measured in 2nd leaf of each finger millet seedlings by measuring the length and breadth of leaf and the leaf area was calculated by following equation (Yoshida et al. 1976):
Measurement of leaf gas exchange, chlorophyll fluorescence and stomatal traits
Leaf gas exchange was measured in matured leaves during flowering stage using an open system photosynthetic gas analyzer (CI-304, CID, USA). The parameters such as photosynthetic rate (PN), transpiration rate (E), internal CO2 concentration (Ci) and stomatal conductance (gs) were recorded under normal ambient environmental condition. The leaf water use efficiency (WUE) and carboxylation efficiency (CE) were calculated as per the formula previously described by Panda et al. (2018).
The same leaves which were used for gas exchange measurements were used for chlorophyll fluorescence measurement during flowering stage using a portable chlorophyll fluorometer (JUNIOR-PAM, WALZ, Germany). Minimal fluorescence (Fo), maximal fluorescence (Fm), variable fluorescence (Fv = Fm − Fo) and Maximum photochemical efficiency of PSII (Fv/Fm) were measured in 20 min of dark-adapted sample (Maxwell and Johnson 2000). In light-adapted leaves at a PPFD of 400 µmol m−2 s−1, steady-state fluorescence yield (Fs), maximal fluorescence (Fm′) and minimal fluorescence (Fo′) were measured. Quenching value due to non-photochemical dissipation of absorbed light energy (NPQ) and the coefficient for photochemical quenching (qP) was also calculated (Maxwell and Johnson 2000).
The stomatal traits such as stomatal density (SD), stomatal size (SS) and stomatal index (SI) were measured in the flag leaf of each finger millet seedlings following the method of Radoglou and Jarvis (1990). Briefly, the leaves were collected and abaxial epidermal side of the leaf was cleaned by tissue paper, smeared with nail varnish carefully and after that thin film was peeled off from the leaf surface, mounted on a glass slide. The stomatal traits such as Numbers of stomata (s) and stomatal size (SS) were counted under a trinocular microscope (Olympus model No. MIPS-3 MP CAMERA, Japan).
Measurement of SPAD index photosynthetic pigments
The same leaves which were used for measurement of the gas exchange and chlorophyll fluorescence were used for measurement of SPAD chlorophyll index using the Minolta SPAD 502 chlorophyll meter (Konica Minolta Sensing, Inc., Osaka, Japan) (Shrestha et al. 2012). The photosynthetic pigments such as chlorophyll and carotenoid were estimated by using 100 mg fresh leaves of finger millets dipped in 10 ml of 80% cold acetone, and kept dark inside a refrigerator (4 °C) for 48 h. The total chlorophyll and carotenoid were measured spectrophotometrically by taking absorbance at 665 nm, 445 nm and 470 nm. The chlorophyll a, chlorophyll b, total chlorophyll and carotenoid contents were calculated by using the equation of (Arnon 1949) and (Lichtenthaler and Wellburn 1983), respectively.
Measurement of leaf relative water content (RWC) and membrane stability index (MSI)
For the measurement of RWC the terminal leaflet of each finger millet genotype were collected and immediately weighed, and recorded as leaf fresh weight (LFW). After weighing the leaflet was poured in distilled water and left in dark for 48 h. After 48 h of incubation, the leaves were again weighed to measure the leaf turgid weight (LTW) was recorded and then the leaf was placed in an oven at 70°c for 48 h to know the leaf dry weight (LDW). The relative water content (RWC) of the leaf was calculated according to the equation of González and González-Vilar (2001).
The membrane stability index (MSI) was measured by utilizing, leaf sample (0.1 g) of each finger millet genotype in two sets of test tubes and the leaves were dipped in 10 ml of distilled water in the test tube. Test tubes of one set kept in 40 °C in a water bath for 30 min and another set were incubated at 100 °C in boiling water bath for 15 min. Then the electrical conductivity of the water containing the leaf samples were measured as C1 at 40 °C and C2 at 100 °C separately. After that, MSI was calculated as per the following equation of Mishra et al. (2019):
Molecular profiling of finger millet genotypes through SCoT markers
For molecular marker analysis, 20–30 seeds of each genotype were shown in a Hyco tray at temperature 25 °C. About 3–4 g fresh leaf samples were collected after 25 days of sowing and leaf were stored at − 70 °C. Total genomic DNA was extracted from lyophilized leaves following a modified CTAB-based procedure (Sharma et al. 2011; Singh et al. 1999). The quality of DNA was checked on 1% agarose gel and with a UV–VIS 2450 spectrophotometer (Shimadzu, Japan). The genotyping was carried out by taking 36 SCoT primers. Detailed sequence information of SCoT markers was presented in Table S1. The SCoT primers were used in the study, according to primers developed by Collard and Mackill (2009).
SCoT marker analysis
The typical PCR mix for SCoT marker contained 50 ng genomic DNA in 1 × reaction buffer, 1.5 mM MgCl2, 10 pmol primers and 1 U Taq DNA polymerase (Biotools, Spain) in 20 µl reaction volume. PCR amplification conditions were as follows: 94 °C for 5 min followed by 35 cycles of 94 °C (30 s)/48–50 °C (30 s)/72 °C (60 s) and a final extension at 72 °C for 10 min. All PCR reactions were performed in a Gene Amp PCR 9700 Thermal Cycler (Applied Biosystems, USA). The amplification products were resolved on 1.5% agarose gels. The amplified fragments were scored manually for their presence (denoted as ‘1’) or absence (denoted as ‘0’) for each primer. The Jaccard’s similarity matrix was subjected to the unweighted pair group method of arithmetic averages clustering to construct the phonetic dendrogram using NYSys software (version 2.2). The marker attributes of each SCoT primer combination was assessed by evaluating polymorphism information content (PIC), marker index (MI) and resolving power (RP).
The PIC was computed using the following equation (Roldan-Ruiz et al. 2000)
where PICi is the polymorphic information content of the marker i, fi is the frequency of the amplified allele (band present) and 1 − fi is the frequency of the null allele. PIC was averaged over the fragments for each primer combination.
The MI was calculated using the following formula (Roldan-Ruiz et al. 2000):
where effective multiplex ratio (EMR) is the total number of polymorphic loci/fragments per primer.
The resolving power (RP) of each primer combination was calculated using the following formula (Prevost and Wilkinson 1999):
where Ib represents band informativeness expressed as Ib = 1 − (2 × |5 − p|), where, p is the fraction of the total accessions in which the band is present.
Data analysis
Morpho-physiological traits were analyzed by analysis of variance (ANOVA) using CropStat-7.2 software (International Rice Research Institute, Philippines). The statistical significance of the parameter means was carried out by the least significance difference (LSD) test. The standard deviations and correlation analysis were carried out by Microsoft Excel 2007. The genetic variability of finger millet genotypes was estimated by genotypic variance and phenotypic variance as per Steel (1997). The phenotypic coefficient of variation and genotypic coefficient of variation was calculated according to the formula of Burton and Devane (1953). Broad sense heritability of all traits was calculated according to the formula as described by Falconer et al. (1996) and the genetic advance was determined as described by Johnson et al. (1955). The principal component analysis (PCA) and cluster analysis were carried out by using PAST-3 (Palaeontological Statistics) software.
Results
Variations of morpho-physiological traits
Variations of growth parameters viz. shoot length (SL), root length (RL), fresh weight (Fwt), dry weight (Dwt) and dry matter accumulation (DMA) of different finger millet genotypes were presented in Table 2. The range of SL and RL ranged from 35.55 to 77 cm and 13.75 to 28 cm, respectively. Some indigenous finger millets such as Kurkuti, Sili jhana, Bhadi and Dasara showed significantly (P < 0.05) higher SL and RL compared to other millets. Further, fresh weight (Fwt), dry weight (Dwt) and dry matter accumulation (DMA) significantly varied (P < 0.05) among the studied finger millet genotypes (Table 3). The range of Fwt, Dwt and DMA in different finger millets was varied from 2.63 to 5.36 g Plant−1, 0.68 to 1.75 g Plant−1 and 20.31 to 34.85%, respectively. The DMA was significantly higher in Jhana and Ladu compared to other millets and high yielding genotypes. The leaf area in studied finger millet genotypes ranges from 6.49 to 17.50 cm2. The leaf area was significantly higher in Bhairabi followed by Sili jhana, Taya and Jhana and lowest was observed in Limca genotypes (Table 3). Physiological parameters such as relative water content (RWC) and membrane stability index (MSI) was significantly varied (P < 0.05) among the studied finger millet genotypes. The range of RWC and MSI was varied from 75.94 to 91.81% and 57.94 to 86.61%, respectively among the studied finger millets. Some of the indigenous finger millets viz. Ladu and Limca showed significantly higher RWC compared to other millets, whereas MSI was highest in Sili jhana.
Variations of photosynthetic traits
Variations in the photosynthetic traits in traditional finger millet varieties were studied by gas-exchange measurements and significant (P < 0.05) variation was observed among studied finger millet genotypes (Table 3). There were significant differences between indigenous and modern high-yielding finger millets for photosynthetic rate (PN). The range of PN varied from 10.43 to 19.18 µmol CO2 m−2 s−1 among the studied finger millet genotypes. Some indigenous finger millets such as Jhana, Taya and Dasara showed significantly (P < 0.05) higher PN compared to modern high-yielding finger millets genotypes. Similarly, stomatal conductance (gs) was also significantly varied among the studied finger millet genotypes. Among the finger millet genotypes, the transpiration rate (E) was not significantly (P < 0.05) different. The stomatal conductance (gs) was ranged from 39.69 to 94.35 mMol H2O m−2 s−1 among the finger millet genotypes and some indigenous finger millets such as Jhana, Sili jhana and Dasara showed significantly (P < 0.05) higher gs compared to modern high-yielding finger millets genotypes. The internal CO2 concentration (Ci) was significantly varied from 146.80 to 248.85 mMol m−2 s−1 among the studied finger millet genotypes and highest Ci was observed in Dasara genotypes. Further, water-use efficiency (WUE, PN/E) and carboxylation efficiency (CE, PN/Ci), among tested finger millet genotypes significantly (P < 0.05) varied (Table 3). The range of WUE and CE was varied from 3.35 to 5.86 and 0.06 to 0.14, respectively among the tested finger millet genotypes.
Significant variations of leaf chlorophyll content and SPAD index was observed among the studied finger millet genotypes (Table 3). The range of SPAD index was varied from 19.65 to 24.25 and highest SPAD index was observed in Dasara compared to other genotypes (Table 3). The chlorophyll and carotenoid content significantly varied among the studied finger millet genotypes and the value of chlorophyll and carotenoid ranged from 0.20 to 0.35 mg g−1 fwt and 0.03 to 0.09 mg g−1 fwt, respectively.
The range of Fo, Fm and Fv/Fm was varied from 49.0 to 72.5, 143.5 to 228.0 and 0.630 to 0.712, respectively among the studied finger millets (Table 4). The value of Fo, Fm and Fv/Fm were not significantly (P < 0.05) different among traditional and modern high yielding finger millet genotypes. However, significant (P < 0.05) variation of ETR, qP and NPQ was found among traditional and modern high yielding finger millet genotypes. Some indigenous finger millets such as Jhana, Bhadi, Chilli and Dasara showed higher ETR compared to modern high-yielding finger millets genotypes. The value of qP and NPQ ranged from 0.351 to 0.571 and 0.567 to 1.641, respectively among the studied genotypes. However, the value of NPQ was significantly higher in high-yielding varieties as compared to indigenous finger millets (Table 4).
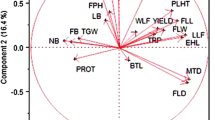
There were significant (P < 0.05) variations of stomatal density (SD), stomata number (SLA), stomatal size (SS) and stomatal index (SI) was found among the studied finger millet genotypes (Fig. S2). The value of SD, SLA, SS and SI was varied from 25.00 to 49.99 no. mm−2, 162.11to 620.36 million LA−1, 1830.65 to 19,234.91 µm2 and 21.83 to 30.30, respectively among the studied finger millet genotypes (Table 5). Some indigenous finger millets such as Jhana, Lala, Kurkuti, Bhadi and Dasara showed higher SD, SLA and SI compared to modern high-yielding finger millets genotypes, whereas SS was significantly higher in high-yielding finger millets genotypes except Limca (Fig. 1).
Relationship between leaf photosynthetic parameters with stomatal traits and dry matter accumulation
Relationship between photosynthetic and stomatal traits in finger millet genotypes was studied by multiple correlation analysis (Table 6). The results showed that a strong positive correlation between PN with gs, WUE, CE and DMA (r = 0.635**, 0.833**, 0.525** and 0.544**, respectively, P < 0.01) was observed whereas, leaf PN was negatively correlated with NPQ (r = − 0.456**, respectively, P < 0.01). The stomatal traits such as SD and SI was positively correlated with leaf PN (r = 0.451*, 0.416*, respectively, P < 0.05), whereas SS was negatively correlated with leaf PN (r = − 0.425*, P < 0.05). The leaf PN in different finger millets was not significantly correlated with leaf area and leaf chlorophyll content.
Genetic variability of morpho-physiological traits
The extent of variability for various morpho-physiological traits in studied finger millet genotypes was evaluated in terms of phenotypic variances (\(\upsigma^{2}_{\text{P}}\)), genotypic variances (\(\upsigma^{2}_{\text{G}}\)), phenotypic coefficient of variation (PCV) and genotypic coefficient of variation (GCV) (Table 7). The GCV was less than that of PCV and low differences were observed between the two for all the studied traits (Table 7). The highest values of GCV and PCV were recorded in traits such as gs, CE, LA, YII, and SLA in different finger millets (Table 7). In the present study, the broad-sense heritability was varied from 1.74 to 99.9 among the studied morpho-physiological traits (Table 7). The genetic advance as a percentage of means (GAM) among the studied traits ranged from 0.01 to 1266.3%. High GAM, along with high heritability, was observed in gs, CI, Chlorophyll, DMA, SL and SLA (Table 7).
Cluster analysis by Bray–Curtis paired linkage revealed the percent of similarity in different morpho-physiological traits among different finger millet genotypes (Fig. 2). The dendrogram is showing the similarity forming three major clusters. The cluster I included Arjuna and Dasara genotypes having more than 91% similarity. The cluster II included 7 genotypes such as Bati, Limca, Biri and Sili jhana in one sub-cluster and Chilika, Jhana, and Chilli in another sub-cluster having 92% similarity. The cluster III included Bhadi Taya, Bhairabi, Kurkuti, Ladu and Lala with similarity 89%.
Molecular genotyping of finger millet genotypes based on SCoT markers
Molecular profiling of the studied finger millet genotypes was carried out by taking 36 SCoT markers. The details of primer and their sequence information were presented in Table S1. Different banding pattern, in the forms of variation in molecular weight of each amplified products for each marker against studied finger millet genotypes, are given in Fig. S3. A representative SCoT profile using primer SCoT7 was shown in Fig. 3. The present investigation used a total of 36 SCoT primers to examine genetic polymorphism, out of total used primers, 33 SCoT primers produced unambiguous and reproducible banding profile with product size ranged from 250 to 2000 bp but 3 SCoT primers such as SCoT-31, SCoT-32 and SCoT-34 failed to amplify the studied millet genotypes. A total of 184 bands were generated by using 33 SCoT primers with an average of 5.57 bands per primer. The number of locus amplified by each primer ranged from 3 to 9 (Table 8). The highest number of fragment (9) was amplified by two primers (SCoT-3 and SCoT-14), whereas lowest number of bands (3) was reported in three primers (SCoT-22, SCoT-28, SCoT-35). A total of 48 polymorphic bands were generated with an average of 1.45 per marker among the studied finger millet genotypes and out of 36 primers 21 primers showed polymorphic bands. The polymorphism percentage with SCoT markers ranged from 11.11 to 100% among the studied primer and the highest polymorphism was obtained in SCoT-18 marker. The level of polymorphism among the 15 studied millet genotypes was obtained by evaluating the polymorphism information content (PIC) values for each primer. The PIC value ranged from 0.02 to 0.29, with an average of 0.071 per primer. Among all the SCoT primers, SCoT-14, SCoT-18 SCoT-20 and SCoT-23 showed the higher PIC value, which implies the potentiality for exploring the genetic diversity of studied millet genotypes. The marker index (MI) was varied from 0.02 to 1.44 among the studied markers and the highest MI was obtained in SCoT-18 markers. Resolving power (RP) ranges from 0.13 (SCoT 3) to 3.07 (SCoT 14), as shown in Table 8. Jaccard’s similarity coefficient calculated the pair-wise genetic similarity and it ranged from 0.872 to 0.981 among the 15 finger millet genotypes (Table 9). The maximum genetic similarity (Jc = 0.981) was between variety Bhadi and Taya; both, are local landraces from Koraput, Odisha. On the other hand, the minimum genetic similarity (Jc = 0.872) was found between variety Arjuna (pure-line selection from OUAT, Bhubaneswar Odisha) and Jhana (local landraces from Koraput).
Representative SCoT profile using primer SCoT7 of 15 Finger millet genotypes. Lane numbers are the same as the serial numbers of the genotypes mentioned in Table 1. Lanes M is the 1 kb ladder used as size markers
The dendrogram was constructed among 15 genotypes of finger millet using binary data that was based on neighbour-joining. Cluster analysis grouped all genotypes in two major clusters I and II. Cluster I included four genotypes all were local landraces of Koraput while cluster II was further subdivided into two sub-clusters IIA and IIB. Cluster IIA included three genotypes namely Bati, Chilli and Sili Jhana were from local landraces of Koraput while cluster IIB included seven genotypes from genotype Chilika is a hybrid from OUAT, Berhampur and Bhairabi is a mutant Bhudha Mandia from OUAT while remaining five genotypes were from local landrace from Koraput. The most distinct genotype namely, Arjuna, which is pure line selection from OUAT, Bhubaneswar, was not grouped in any of the two clusters (Fig. 4).
Discussion
The Koraput district of Odisha is one of the centres of diversity for many food crops and forest species and harbouring rich genetic diversity of millets (Mishra et al. 2012). These millet genotypes may serve as a valuable genetic resource for future crop improvement to meet the demands of food security. Although, indigenous finger millet genotypes have less productivity it provides a great opportunity that ensures food security and livelihood of poor farming communities due to their tolerance capacity to biotic and abiotic stresses. This is the first study on genetic diversity and population structure of different indigenous finger millet genotypes originated from various regions of Koraput.
There was remarkable variation of shoot length (SL), root length (RL), fresh weight (Fwt), dry weight (Dwt) and dry matter accumulation (DMA) was observed among the studied finger millet genotypes. Some indigenous finger millets such as Kurkuti, Sili jhana and Dasara showed remarkably superior growth parameters compared to modern high yielding millet genotypes. Such diversity among the millet genotypes might be related to their genetic origin, genetics of the species and geographical sources where they are grown. These results of morphological variations of millet genotypes were also consistent with the previous report of global collections of millets (Babu et al. 2014) and millets of India (Das et al. 2007). The leaf relative water content (RWC) is important parameters for study of water status of the plant (Lafitte et al. 2002). Some of the indigenous finger millets such as Ladu and Limca showed significantly higher RWC and Sili jhana showed maximum membrane stability index (MSI) compared to other millets, Which suggested that these genotypes may have better shielding mechanism to protect membrane damage and sustain better leaf turgor under prevalent environmental conditions (Chakraborty et al. 2002; Swapna and Shyalaraj 2017).
Variations in the photosynthetic traits in indigenous finger millets genotypes were studied by gas-exchange measurements, chlorophyll fluorescence and stomatal traits and the results were compared with the modern high-yielding genotypes. Some indigenous finger millets such as Jhana, Taya and Dasara showed superior photosynthetic rate, stomatal conductance, water-use efficiency (WUE, PN/E) and carboxylation efficiency (CE, PN/Ci) compared to modern high-yielding finger millets genotypes. The leaf photosynthesis is regulated by various internal and environmental parameters (Shi et al. 2005). The variation observed in photosynthetic traits among the genotypes might be related to their origin and genetics of the varieties. In the present study, leaf photosynthetic rate of millet genotypes was lower than the previous reports (Shankar et al. 1990; Subrahmanyam 2000). The lower values of photosynthetic rate may be due to high temperatures during measurements. In an earlier study of Xie et al. (2011) reported that there is a decline in photosynthesis and grain yield, with atmospheric air temperature higher than 35 °C. Results of the present study indicated that indigenous genotypes showed better photosynthetic efficiency but unable to increase grain yield, because the increase in leaf photosynthesis may not necessarily lead to increase in grain yield as reported in different millets (Subrahmanyam 2000) and rice (Gu et al. 2013). The results suggested that the indigenous finger millets associated with high leaf photosynthetic rate, which can contribute this trait for crop improvement despite their other disadvantageous phenotypes (Subrahmanyam 2000; Vikram et al. 2016). Leaf chlorophyll content is an important plant pigment regulates the photosynthetic process by absorbing solar energy from the environment, as they indicate the light-harvesting capacity of plants (Panda et al. 2018). Some indigenous finger millets such as Jhana, Taya and Dasara showed better chlorophyll and carotenoid status in comparison to modern high-yielding finger millets genotypes.
In addition, Chl fluorescence measurement helps to assess the intrinsic photosynthetic performance of plants, which provides essential information regarding the function of PSII under prevalent environmental conditions (Batra et al. 2016). The F0, Fm, Fv/Fm, YII, NPQ, and qP is widely used Chl fluorescence parameters in plant physiology studies (Murchie and Lawson 2013). There were no significant differences in values of Fo, Fm, Fv/Fm and Y (II) observed among studied finger millet varieties (Table 4). However, some indigenous finger millets such as Jhana, Lala, Bhadi, Chilli and Dasara showed maximum ETR and showed better photochemical activity in the prevalent environmental condition as compared to the other genotypes.
Stomata are the key player in plants which regulates the exchange of CO2 and water vapour between atmosphere and surrounding (Buckley 2005; Kondamudi et al. 2016). The stomatal traits such as stomatal size, number and density controls the leaf gas exchange in plants (Panda et al. 2018). However, very little information is available on the relationship between leaf gas exchange and stomatal traits in finger millet (Subrahmanyam 2000). Some indigenous finger millets such as Jhana, Lala, Kurkuti, Bhadi and Dasara showed better stomatal traits and showed adaptive mechanism to cope with changing environment compared to modern high-yielding finger millet genotypes (Subrahmanyam 2000).
Based on the result of multiple correlations, the rate of photosynthesis (PN) was not significantly influenced by leaf pigments (Chl and carotenoid contents) and leaf area. A strong positive correlation between PN with gs, CE and WUE, were observed. This result indicated that the observed variations in PN in different millets genotypes were not based on pigment content or leaf area, but related to the leaf CE and WUE as has been reported earlier in rice and other crops (Yeo et al. 1994; Kiran et al. 2013; Haritha et al. 2017). Further, leaf PN showed a significant positive correlation with DMA (P < 0.05), which supports the previous reports in other crops (Evans 2013; Puteh et al. 2014) for increased biomass in plants is associated with higher photosynthetic rate. The findings also suggested that the number and density of stomata in the leaf are the major regulating factor for leaf photosynthesis than the shape and size in millets. This results consistent with the earlier finding in different plant species (Chandra and Das 2000; Giuliani et al. 2011).
Based on genetic variability study, high PCV and GCV values were recorded in the traits such as gs, CE, LA, YII, and SLA in different finger millets. This indicates the existence of substantial variability for such characters based on which selection of finger millet genotype may be useful. These characters showing ample variation are efficient for selection in a breeding program (Mohapatra et al. 2017). The GCV was less than that of PCV and low differences were observed between them for all the morphological traits in the studied finger millets. This indicates that environment least influence high contribution of genotypic effect for phenotypic expression of these traits and above characters. In the present study, high GAM along with high heritability was observed in gs, CI, Chlorophyll, DMA, SL and SLA parameters. It indicates that these characters would be beneficial as a base for selection in finger millet improvement.
This is the first report of SCoT marker-based genetic diversity and population structure in finger millet genotypes of Koraput. SCoT marker technique used in the current study is simple, low cost, fast, effective and highly reproducible. In the present investigation, a set of 36 SCoT primers were used to examine genetic polymorphism, out of the total, 33 SCoT primers produced unambiguous and reproducible banding profile with 250–2000 bp product size but 3 SCoT primers such as SCoT-31, SCoT-32 and SCoT-34 failed to amplify the studied millet genotypes. Significant level of polymorphism were detected as reported in the present study complies with earlier investigations using some of these SCoT primers in rose (Henuka et al. 2015) and Grape (Guo et al. 2012). It was observed that all the markers used in the present study showed polymorphism at various loci which may be useful in breeding for improving the genotypes of finger millets and as well as for the identification of the appropriate genotypes suitable for specific environmental conditions. Among all the SCoT primers, SCoT-14, SCoT-18 SCoT-20 and SCoT-23 showed the higher PIC value and marker index, which implies the potentiality for exploring the genetic diversity of studied millet genotypes. Based on the genetic similarity analysis it is revealed that some of the indigenous finger millet genotypes such as Jhana, Lala, Kurkuti, Ladu, Bhadi and Taya showed highest genetic disimilarity with modern high yielding genotype Arjuna. The mean genetic distance among the studied varieties was 0.934, which indicates that a very low level of genetic diversity among the studied finger millet genotypes. The low level of genetic diversity might be due to the similar origin, ecotype and speciation as millets were collected only from, Koraput.
Conclusion
Significant variability was observed for leaf photosynthetic traits in studied finger millets of Koraput, which need conservation and sustainable utilization for future crop improvement programs. The major morpho-physiological traits such as stomatal conductance, dry matter accumulation, shoot length and stomata per leaf area are played a pivotal role and are the significant determinants of phenotypic diversity among studied finger millet genotypes. The positive association of photosynthesis with dry matter accumulation indicates that some of the genotypes remarkably have more photosynthetic rate along with better plant biomass accumulation. These parameters might be instrumental for superior phenotypic selection of finger millet improvement program. Further, SCoT markers were polymorphic and revealed a moderate level of genetic diversity and provided information on population structure among the finger millet genotypes. Based on the genetic similarity analysis it is revealed that some of the indigenous finger millet genotypes such as Jhana, Lala, Kurkuti, Ladu, Bhadi and Taya showed highest genetic dissimilarity with modern high yielding genotype Arjuna. These genotypes can be considered as the potential genetic resources for a breeding program. The information generated in this study will be valuable for breeding and conservation of finger millet genotypes. Further research is aim to elucidate the marker-trait association mapping and yield evaluation in field condition in these genotypes and could be used in finger millet breeding program.
Abbreviations
- CE:
-
Carboxylation efficiency
- Ci:
-
Internal CO2 concentration
- CV:
-
Coefficient of variance
- DMA:
-
Dry matter accumulation
- Dwt:
-
Dry weight
- ETR:
-
Electron transfer rate
- Fo:
-
Minimum fluorescence yield obtained with dark-adapted leaf
- Fm:
-
Maximum fluorescence yield obtained with dark-adapted leaf
- Fv/Fm:
-
Maximal photochemical efficiency of PS II
- Fwt:
-
Fresh weight
- gs :
-
Stomatal conductance
- LA:
-
Leaf area
- LSD:
-
Least significance difference
- MSI:
-
Membrane stability index
- NPQ:
-
Non-photochemical quenching
- PN :
-
Photosynthetic rate
- RL:
-
Root length
- RWC:
-
Relative water content
- SL:
-
Shoot length
- qP:
-
Photochemical quenching
- WUE:
-
Water use efficiency
- Y(II):
-
Yield of PSII photochemistry
References
Arnon DI (1949) Copper enzymes in isolated chloroplasts, polyphenol oxidase in Beta vulgaris. Plant Physiol 24:1–15
Assefa A, Fetene M, Tesfaye K (2013) Agro-morphological, physiological and yield related performances of Finger millet [Eleusine coracana (L.) Gaertn.] accessions evaluated for drought resistance under field condition. Asian J Agric Rural Dev 3(10):709–720
Babu BK, Dinesh P, Agrawal PK, Sood S, Chandrasekhar C (2014) Comparative genomics and association mapping approaches for blast resistant genes in finger millets using SSRs. PLoS One 9(6):e99182
Babu BK, Sood S, Agrawal PK, Chandrasekhar C, Kumar A, Kumar A (2017) Molecular and phenotypic characterization of 149 finger millet accessions using microsatellite and agro-morphological markers. Proc Natl Acad Sci India Sect B Biol Sci 87(4):1217–1228
Bashir EMA, Ali AM, Ali AM, Mohamed ETI, Melchinger AE, Parzies HK, Haussmann BIG (2015) Genetic diversity of Sudanese pearl millet (Pennisetum glaucum (L.) R. Br.) landraces as revealed by SSR Markers, and relationship between genetic and agro morphological diversity. Genet Resour Crop Evol 62:579–591
Batra NG, Kumari N, Sharma V (2016) Photosynthetic performance of Ocimum sanctum morphotypes in a semiarid region. J Herbs Spices Med Plants 22(3):211–224
Buckley TN (2005) The control of stomata by water balance. New Phytol 168:275–292
Burton GW, Devane EH (1953) Estimating heritability in tall fescue (Festuca arundinacea) from replicated clonal material. Agron J 45(10):478–481
Chakraborty U, Dutta S, Chakraborty BN (2002) Response of tea plants to water stress. Biol Plant 45:557–562
Chandra K, Das AK (2000) Correlation and interaction of physiological parameters in rice under rainfed transplanted condition. J Crop Res 19:251–254
Collard BCY, Mackill DJ (2009) Start codon targated (SCoT) polymorphism: a simple, novel DNA marker techniques for generating gene-targeted markers in plants. Plant Mol Biol Rep 27:86–93
Das S, Mishra RC, Rout GR, Aparajita S (2007) Genetic variability and relationships among thirty genotypes of finger millet (Eleusine coracana L. Gaertn.) using RAPD markers. Z Naturforschung 62:116–122
Dida MM, Srinivasachary R, Ramakrishnan S, Bennetzen JL, Gale MD, Devos KM (2007) The genetic map of finger millet, Eleusine coracana. Theor Appl Genet 114:321–332
Evans JR (2013) Improving photosynthesis. Plant Physiol 162:1780–1793
Falconer DS, Mackay TF, Frankham R (1996) Introduction to quantitative genetics (4th edn). Trends Genet 12:280
FAO (2007) FAOSTAT database. http://www.faostat.fao.org. Accessed 18 Mar 2019
Giuliani R, Koteyeva N, Voznesenskaya E, Evans MA, Cousins AB, Edwards GE (2011) Coordination of leaf photosynthesis, transpiration and structural traits in rice and wild relatives (genus oryza). Plant Physiol 162:1632–1651
González L, González-Vilar M (2001) Determination of relative water content. In: Reigosa Roger MJ (ed) Handbook of plant ecophysiology techniques. Springer, Dordrecht, pp 207–212
Gu J, Yin X, Stomph TJ, Struick PC (2013) Can exploiting natural genetic variation in leaf photosynthesis contributes to increasing rice productivity? A simulation analysis. Plant Cell Environ 37(1):22–34
Guo DL, Zhang JY, Liu CH (2012) Genetic diversity in some grape varieties revealed by SCoT analyses. Mol Biol Rep 39:5307–5313
Gupta NK, Meena SK, Gupta S, Khandelwal SK (2002) Gas exchange, membrane permeability, and ion uptake in two species of Indian jujube differing in salt tolerance. Photosynthetica 40:535–539
Gupta SM, Arora S, Mirza N, Pande A, Lata C, Puranik S, Kumar J, Kumar A (2017) Finger millet: a “certain” crop for an “uncertain” future and a solution to food insecurity and hidden hunger under stressful environments. Front Plant Sci 8:643
Haritha G, Vishnukiran T, Yugandhar P, Sarla N, Subrahmanyam D (2017) Introgressions from Oryza rufipogon increase photosynthetic efficiency of KMR3 rice lines. Rice Sci 24(2):85–96
Henuka R, Raju D, Janakiram N (2015) Characterization and analysis of genetic diversity among different species of rose (Rosa species) using morphological and molecular marker. Indian J Agric Sci 85(2):240–245
Johnson HW, Robinson HF, Comstock RE (1955) Estimates of genetic and environmental variability in soybeans. Agron J 47(7):314–318
Kiran TV, Rao YV, Subrahmanyam D, Rani NS, Bhadana VP, Rao PR, Voleti SR (2013) Physiological trait associated with high yield potential in rice. In: Mew TW, Brar DS, Peng S, Dawe D, Hardy B (eds) Rice science: innovations and impact for livelihood. IRRI, Los Banos, pp 117–145
Kondamudi R, Swamy KN, Rao YV, Kiran TV, Suman K, Rao DS, Rao PR, Subrahmanyam D, Sarla N, Ramana BK, Voleti SR (2016) Gas exchange, carbon balance and stomatal traits in wild and cultivated rice (Oryza sativa L.) genotypes. Acta Physiol Plant 38(16):1–9
Kumar A, Sharma D, Tiwari A, Jaiswal JP, Singh NK, Sood S (2016) Genotyping-by-sequencing analysis for determining population structure of finger millet germplasm of diverse origins. Plant Genome 9(2):3–15
Lafitte HR, Courtois B, Arraudeau M (2002) Genetic improvement of rice in aerobic systems: progress from yield to genes. Field Crops Res 75(2–3):171–190
Lichtenthaler HK, Wellburn AR (1983) Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem Soc Trans 11:591–592
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence practical guide. J Exp Bot 51:659–668
Mishra S, Chaudhury SS (2012) Ethnobotanical flora used by four major tribes of Koraput, Odisha, India. Genet Resour Crop Evol 59(5):793–804
Mishra S, Chaudhury SS, Nambi VA (2012) Strengthening of traditional seed selection practices with improved knowledge and skills of tribal farm families in Koraput District. Indian J Tradit Knowl 11(3):461–470
Mishra SS, Behera PK, Panda D (2019) Genotypic variability for drought tolerance-related morpho-physiological traits among indigenous rice landraces of Jeypore tract of Odisha, India. J Crop Improv 33:254–278
Mohapatra PP, Bhoi S, Maity TK, Majhi A, Tarafdar J (2017) Genetic variability, heritability and genetic advance studies in onion. J Crop Weed 13(3):32–34
Mukami A, Ngetich A, Mweu C, Oduor OR, Muthangya M, Mbinda WM (2019) Differential characterization of physiological and biochemical responses during drought stress in finger millet varieties. Physiol Mol Biol Plants 25(4):837–846
Murchie EH, Lawson T (2013) Chlorophyll fluorescence analysis: a guide to good practice and understanding some new applications. J Exp Bot 64(13):3983–3998
Panda D, Mahakud A, Mohanty B, Mishra SS, Barik J (2018) Genotypic variation of photosynthetic gas exchange and stomatal traits in some traditional rice (Oryza sativa L.) landraces from Koraput, India for crop improvement. Physiol Mol Biol Plants 12(5):973–983
Panda D, Sailaja NH, Padhan B, Lenka K (2019) Sprouting-associated changes in nutritional and physic functional properties of indigenous millets from Koraput, India. In: Proceedings of the national academy of sciences, India section B: biological sciences, pp 1–8
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Puteh AB, Mondal MMA, Ismail MR, Latif MA (2014) Grain sterility in relation to dry mass production and distribution in rice (Oryza sativa L.). Biol Med Res Int. https://doi.org/10.1155/2014/302179
Radoglou KM, Jarvis PG (1990) Effects of CO2 enrichment on four poplar clones, I. growth and leaf anatomy. Ann Bot 65(1):617–626
Rajendran HAD, Muthusamy R, Stanislaus AC, Krishnaraj T, Kuppusamy S, Ignacimuthu S, Al-Dhabi NA (2016) Analysis of molecular variance and structure in Southern Indian finger millet genotypes using three different molecular markers. J Crop Sci Biotechnol 19(4):275–283
Ramakrishnan M, Ceasar SA, Duraipandiyan V, Al-Dhabi NA, Ignacimuthu S (2016) Using molecular markers to assess the genetic diversity and population structure of finger millet (Eleusine coracana (L.) Gaertn.) from various geographical regions. Genet Resour Crop Evol 63:361–376
Roldan-Ruiz I, Calsyn E, Gilliland TJ, Coll R, van Eijk MJT, de Loose M (2000) Estimating genetic conformity between related ryegrass (Lolium) varieties. 2. AFLP characterization. Mol Breed 6:593–602
Sakamma S, Umesh KB, Girish MR, Ravi SC, Satishkumar M, Bellundagi V (2018) Finger millet (Eleusine coracana (L.) Gaertn.) production system: status, potential, constrains and implications for improving small farmer’s welfare. J Agric Sci 10(1):162–179
Shankar AG, Udayakumar M, Prasad TG (1990) Genotypic variability for net photosynthesis in finger millet (Eleusine coracana G.) genotypes: an approach to identity high CER types. J Agron Crop Sci 165:240–252
Sharma SS, Negi MS, Sinha P, Kumar K, Tripathi SB (2011) Assessment of genetic diversity of biodiesel species Pongamia pinnata accessions using AFLP and three endonuclease-AFLP. Plant Mol Biol Rep 29:12–18
Shi T, Bibby TS, Jiang L, Irwin AJ, Falkowski PG (2005) Protein interactions limit the rate of evolution of photosynthetic genes in cyanobacteria. Mol Biol Evol 22:2179–2189
Shrestha S, Brueck H, Asch F (2012) Chlorophyll index, photochemical reflectance index and chlorophyll fluorescence measurements of rice leaves supplied with different N levels. J Photochem Photobiol B Biol 113:7–13
Singh TS, Sharma HO (2018) Trend and growth of small millets production in Madhya Pradesh as compared to India. Int J Agric Sci 10(1):4983–4986
Singh A, Negi MS, Rajagopal J, Bhatia S, Tomar UK, Srivastava PS, Lakshmikumaran M (1999) Assessment of genetic diversity in Azadirachta indica using AFLP markers. Theor Appl Genet 99:272–279
Steel R (1997) Analysis of variance I: the one-way classification. Princ Proced Stat Biom Approach 974:139–203
Subrahmanyam D (2000) Genotype variability in photosynthetic characteristic in finger millet. Photosynthetica 38(1):105–109
Swapna S, Shyalaraj KS (2017) Screening for osmotic stress responses in rice varieties (Oryza sativa L.) under drought condition. Rice Sci 24(5):253–263
Vikram P, Kadam S, Singh BP, Lee YJ, Pal JK, Singh S, Singh ON, Swamy BPM, Thiyagarajan K, Singh S, Singh NK (2016) Genetic diversity analysis reveals importance of green revolution gene (Sd1 locus) for drought tolerance in rice. Agric Res 5(1):1–12
Wakista PW, Dasanayaka PN, Illeperuma RJ, Perera SACN (2017) Assessment of genetic diversity of a representative sample of finger millet (Eleusine coracana (L.) Gaertn.) collected from four different district of Sri Lanka using SSR markers. Sri Lankan J Biol 2(1):1–8
Xie XJ, Shen SH, Li YX, Zhao XY, Li BB, Xu DF (2011) Effect of photosynthetic characteristic and dry matter accumulation of rice under high temperature at heading stage. Afr J Agric Res 6(7):1931–1940
Yeo ME, Yeo AR, Flower TJ (1994) Photosynthesis and photorespiration in the genus Oryza. J Exp Bot 45(5):553–560
Yoshida S, Forno DA, Cook JH, Gomes KA (1976) Routine procedure for growing rice plants in culture solution. In: Lab manual for physiological studies of rice, 3rd edn, pp 61–65. The International Rice Research Institute, Los Banos
Acknowledgements
The authors are grateful to Head, Department of Biodiversity and Conservation of Natural Resources for providing necessary facilities for the work. The Regional Director, MS Swaminathan Research Foundation (MSSRF), Jeypore, Odisha is highly acknowledged for providing the millet seeds for the experiment. Financial assistance was provided by University Grants Commission, New Delhi, Govt. of India for providing Non-NET Fellowship (Grant No. CUO/MPHIL/2018/NonNET/02).
Author information
Authors and Affiliations
Contributions
NHS and DP designed the experiments, cultivated the plants and performed the measurement of morphological traits. NHS, PKB and KL performed the measurement of physiological and biochemical traits. SSS and SKL performed the molecular profiling. DP analyzed the data and wrote the paper. All authors read and provided helpful discussions for the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Panda, D., Sailaja, N.H., Behera, P.K. et al. Genetic diversity of under-utilized indigenous finger millet genotypes from Koraput, India for crop improvement. J. Plant Biochem. Biotechnol. 30, 99–116 (2021). https://doi.org/10.1007/s13562-020-00557-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-020-00557-w