Abstracts
Using a suite of morphological traits and neutral molecular markers, we have reevaluated the origins of the two cultivated jute species. The nine Corchorus species, which are restricted to both Africa and Asia, were phenotyped for bast fibre yield- and quality-linked traits, morpho-phenetically classified based on those traits’ similarities, genotyped by nuclear microsatellites and haplotyped by chloroplast microsatellites. The two cultivated jute species contain ~70% of the neutral genetic diversity present in their wild relatives. Phylogenetic analyses using 38 highly polymorphic nuclear microsatellites identified C. aestuans as the common ancestor to both the cultivated jute species. C. urticifolius is the nuclear progenitor of C. olitorius (dark jute), but no close nuclear progenitor of C. capsularis (white jute) could be identified. Although either C. aestuans or C. pseudo-olitorius or both appear to be the possible cytoplasmic progenitors of white jute, the matrilineal ancestry of dark jute remains obscure. However, the presence of a single, unique chloroplast haplotype in each of the two cultivated species indicates the involvement of other unknown wild relatives. Taken together with archaeobotanical evidence, our results suggest that dark jute had its origin in equatorial region of east Africa, but was domesticated in India. We could not support an Indo-Myanmar origin of white jute, and possibly it also originated in Africa, but was domesticated in Asia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The origin and phylogeny of the genus Corchorus as a whole or its two domesticated species (C. capsularis L. and C. olitorius L.) has long been controversial (Edmonds 1990; Benor et al. 2010). For much wider diversity of Corchorus species in Africa, an African centre of origin of C. olitorius is nowadays supported, whereas C. capsularis is supposed to have originated in Indo-Myanmar region including south China (Kundu 1951, 1956). These competing views are, however, based on biogeographic diversity and species adaptation (Edmonds 1990).
Implicit in this ongoing debate on the origins of the two domesticated jute species, however, are three apparent caveats. Firstly, recent archaeobotanical evidence supports not only the possibility of an ancient use of jute in the Indus civilization, but also its clear differentiation from silk in that period (Wright 2010). Secondly, as opposed to earlier claims that C. capsularis is not found wild in Africa and Australia, now its presence in both the continents is well documented (Du Puy and Telford 1963; Benor et al. 2010). Thirdly, although both the domesticated jute species are an important source of fibre in Asian countries, they are widely utilized as a mucilaginous leafy vegetable throughout Africa (Grubben and Denton 2004).
Molecular marker-based genetic diversity analyses revealed considerable divergence between the two cultivated jute species, with relatively higher level of allelic diversity in C. olitorius than in C. capsularis despite fairly low genetic diversity at the intraspecific level (Hossain et al. 2002; Basu et al. 2004; Roy et al. 2006; Mir et al. 2008, 2009). Support for a distinct genetic differentiation between the two cultivated jute species was provided by morpho-physiological (Palit et al. 1996), karyomorphological (Maity and Datta 2009) and genomic (Basu et al. 2004; Roy et al. 2006; Haque et al. 2007; Mir et al. 2008) analyses. However, none of these studies could establish a relationship between geographic origin and phenetic classification. This reported lack of correspondence between genetic divergence and geographic diversity is indicative of not only uncertain origins of cultivated jute accessions, but also the possibility of their recent migration or germplasm movement across geographical boundaries (Roy et al. 2006).
Codominant nuclear microsatellites have been shown to be ideal markers for detecting phylogenetically significant diversity within cultivated jute (Mir et al. 2008, 2009). In the present study, we analyzed the nuclear microsatellite diversity in the nine Corchorus species including the two cultivated jute species that encompass wide geographic diversity. Chloroplast microsatellites represent a general marker system for assessing the genetic structure of plant populations. Several mononucleotide repeats resident on intronic and intergenic cpDNA regions are known to be conserved across dicotyledonous angiosperms (Weising and Gardner 1999). We used a set of those consensus microsatellite primers to study mononucleotide repeat-based chloroplast haplotype diversity in Corchorus species. We also sought to phenetically classify the nine Corchorus species based on morphological similarities by using a suite of most distinguishing vegetative growth and bast fibre traits.
Materials and methods
Plant material
Nine diploid (2n = 2× = 14) Corchorus species, viz., C. aestuans L. (WCIJ 037), C. fascicularis Lam. (WCIJ 004), C. pseudo-capsularis Schweinf. (WCIJ 007), C. pseudo-olitorius Islam & Zaid (WCIJ 034), C. tridens L. (WCIJ 046), C. trilocularis L. (WCIJ 072) and C. urticifolius Wight & Arnold (WCIJ 006) including the two cultivated jute species C. capsularis L. (cv. JRC-321 and mutant CMU-010) and C. olitorius L. (cv. JRO-524 and mutant PPO-4) were used in the present study. The selected accessions of wild species have been used in other published studies (Chakraborty and Palit 2009), whereas the selected genotypes of both the cultivated jute species represent leading cultivars in an agricultural setting (global area sown and bast fibre production). Except for C. trilocularis, which is a diffuse undershrub, they are all annual herbs, with distinct growth habits, morphological features and sexual compatibilities (Supplementary Table 1).
Field study
The experiment was laid out in a randomized complete block design with three replications over the two growing seasons. Each genotype was represented by three rows of 10 plants each at a spacing of 10 cm in rows 30 cm apart per replication. At 90 d after sowing, observations on vegetative characters, such as plant height (cm), basal stem diameter (cm), leaf length (cm), leaf breadth (cm), leaf area (cm2) and leaf ratio (average leaf length divided by average leaf breadth) were recorded for 10 randomly selected healthy plants per replication. The matured plants were harvested at 120 d after sowing. After drying, wood and bast fibre yields (g plant−1) were determined. Bundle strength and airflow fineness testers were used to measure tensile strength (g tex−1) and fibre fineness (tex), respectively. Fibre fineness was scored on a 0–5 scale of decreasing fineness, with 5 representing the coarsest fibre.
Morpho-phenetic analysis
Prior to univariate analysis of quantitative data for each character, normality (Shapiro-Wilk test) and equal variance (Bartlett’s test) assumptions were tested (GenStat Version 7.2.0.220; VSN International Ltd., Oxford, UK). Accordingly, the data were suitably transformed and analyzed by one-way analyses of variance followed by multiple comparisons of mean values by Tukey-Kramer honestly significant difference test. Associations between morphological and bast fibre traits were estimated by nonparametric method of Kendall’s coefficient (tau-b) of rank correlation. The dendogram was constructed using programs in NTSYSpc Version 2.20v (Applied Biostatistics, Inc., New York, USA), with resampling of means for each character (1000 bootstrap replicates) and clustering based on UPGMA. A majority-rule consensus tree was compiled from trees that resulted from bootstrap sampling.
Nuclear microsatellite genotyping
DNA was isolated from 2-week-old seedlings, according to Kundu et al. (2011). Altogether 66 nSSRs that exhibited ± polymorphism due to null alleles (Mir et al. 2008, 2009) were initially selected, of which 38 single-locus polymorphic nSSRs were finally selected based on their preliminary screening in all the nine Corchorus species (Supplementary Table 2). The nSSR sequences were amplified according to Mir et al. (2008), and the amplified products were denatured and separated in 10 % denaturing polyacrylamide (19:1) gels in 1.0 × TBE buffer at 80 V cm−1. The amplified products were stained with a silver staining protocol (Sanguinetti et al. 1994), and microsatellite alleles were scored as present (1) or absent (0).
Genotypic data analyses
For each nSSR locus, the number of observed alleles (n a ), the effective number of alleles (n e ) (Kimura and Crow 1964), the Levene’s observed heterozygosity (H o ) (Levene 1949) and the Nei’s unbiased genetic diversity (H e ) (Nei 1973) were calculated by using the software POPGENE Version 1.32 (Yeh et al. 1999). PIC for each nSSR locus was calculated as \( \mathrm{PIC}=1\text{-}\sum\nolimits_{{\mathrm{i}=1}}^{\mathrm{n}} {{{{\left( {\mathrm{Pi}} \right)}}^2}} \), where P i is the frequency of the ith allele and n is the number of alleles (Botstein et al. 1980). The allelic richness (R s ) that estimates the number of alleles independent of the sample size (El Mousadik and Petit 1996) was calculated by using the software FSTAT (Goudet 1995). The Shannon’s information index (Shannon and Weaver 1949) was calculated by using POPGENE. The number of private alleles (Mohammad et al. 2004) and their average frequencies were estimated from allele distributions by using MS Office Excel. For genetic diversity estimates (n a , n e , H e and R s ), the differences between the cultivated and wild Corchorus groups were tested by non-parametric Wilcoxon signed-paired rank test after Bonferroni correction.
Genetic analysis was performed by using the software DARwin5 Version 5.0.158 (Perrier and Jacquemoud-Collet 2006). A distance matrix was computed by using Jaccard’s coefficients and 1,000 replicate bootstraps. The dendogram was constructed by using the NJ method employing the weighted version of Saitou and Nei (1987) based on a principle of parsimony that finds a tree close to the true phylogenetic tree (Rohlf 2005). Since a radial tree is the better option if it is not known as to where the root lies (Hall 2001), the data were presented as a radial tree. The NJ tree resolved C. aestuans L. as an outgroup. It was grafted a posteriori on a tree built on the unit set without this outgroup by using the least-squares grafting method of DARwin5, and the grafting point was regarded as the ancestral root (Perrier et al. 2003). Concordance of the nSSR data to the morphological data was tested by two-way Mantel test.
Chloroplast microsatellite haplotyping
Ten pairs of consensus chloroplast microsatellite primers (Weising and Gardner 1999) were used to characterize mononucleotide SSR variation in chloroplast DNA (Table 1). The PCR was performed employing the same protocol as above, except for initial denaturation (94 °C for 5 min) and final extension (72 °C for 8 min), with an annealing temperature of 50 °C. Amplified products were separated using a non-denaturing polyacrylamide gel (8 %, 29:1) in 0.5 × Tris-glycine buffer and stained with a modified silver staining protocol (Sanguinetti et al. 1994).
For cpSSR allele detection and scoring, Gene Profiler Version 4.05 (Scanalytics, Fairfax, USA) was used. Alleles were generated using linkage analysis database module of the software. Chloroplast haplotypes were resolved, and haplotype statistics, such as the effective number of haplotypes (N e ), the number of private haplotypes (P), haplotypic richness (R h ) (Petit et al. 1998), the Nei’s unbiased genetic diversity (H e ) (Nei 1973) and average genetic distance between individuals (D 2 sh ) (Morgante et al. 1998) were calculated by using the software HAPLOTYPE ANALYSIS Version 1.05 (Eliades and Eliades 2009). Since length polymorphisms caused by variable mononucleotide repeats are not suitable for phylogenetic analysis, chloroplast haplotypes were mapped onto the nSSR NJ tree.
Results and discussion
Morphological and morpho-phenetic analyses
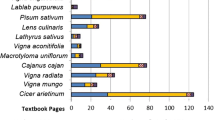
Morphological traits were highly polymorphic/diverse and significantly different among the Corchorus species (Fig. 1). Our results showed that all the wild jute species produced finer fibre with lower tensile strength, while both the cultivated jute species produced relatively coarser fibre with higher tensile strength. Non-parametric Kendall’s coefficients of rank correlations showed varying degrees of positive associations between the seven out of eight quantitative traits (Table 2). Except for leaf ratio, all traits were significantly associated with fibre yield plant−1.
We considered here the majority-rule (>50 % bootstrap support) consensus tree produced by average taxonomic distance because of its universal use in numerical taxonomy (Rohlf 2005). The nine Corchorus species were grouped in the two major clusters, each with a >75 % bootstrap support-cluster I with both the cultivated jute species together with a mutant of C. olitorius, and cluster II with all the wild jute species together with a mutant of C. capsularis (Fig. 2). In the wild species cluster, C. aestuans, C. pseudo-capsularis and C. urticifolius were grouped in a distinct sub-cluster (IIb), while the remaining species formed a separate sub-cluster (IIc). Based on morpho-phenetic data, C. aestuans was found to be closely related to C. pseudo-capsularis (sub-cluster IIa). The suite of bast fibre yield- and quality-linked traits provided a high degree of phenetic support to both the cultivated jute species in relation to their wild relatives. Chakraborty and Palit (2009) have also found them to be closely related based on a large number of morphological characters.
A majority-rule consensus unweighted pair-group method arithmetic average tree constructed from the eight bast fibre yield- and quality-linked traits’ similarities. Photographs from left to right show seed, leaf and capsule of individual species, and the numbers at nodes represent the occurrence in 1,000 bootstrap replicates
Nuclear microsatellite polymorphisms
Summary statistics were computed for each nSSR locus (Supplementary Table 3) and for the whole sample and each of the two groups (cultivated and wild) over the 38 nSSR loci (Table 3). A total of 222 alleles was detected, with 95 and 183 alleles distributed in cultivated and wild jute, respectively. Maximum numbers of alleles were observed in C. fascicularis (59) followed by C. aestuans (54), while the least in C. capsularis (28). The number of observed alleles per locus (n a ) varied from 4 to 8, with an average (n o ) of 5.8 alleles per locus, whereas the effective number of alleles per locus (n e ) varied from 2.3 to 7.6, with an average of 4.6 alleles per locus. For n o , n e and the allelic richness (R s ), there was significant variation between the two Corchorus groups. The average genetic diversity (H e ) was 0.81 for the whole sample, and it was significantly higher in the wild (0.78) than in the cultivated jute (0.56). Overall, the average genetic diversity (H e ) in wild Corchorus species was ~40 % higher than that in the cultivated jute species. However, the average level of the Levene’s observed heterozygosity (H o ) was relatively low (0.31) for the whole sample. The average PIC was significantly higher in the wild (0.72) than in the cultivated (0.43) jute. Similarly, Shannon’s information index (I) was significantly higher in the wild (1.43) than in the cultivated (0.77) jute, with an average of 1.60 for the whole sample (Table 3).
With increasing allele frequency, the number of alleles decreased (Fig. 3). The proportion of low (< 0.1 ≤ 0.3) frequency alleles was markedly higher in the wild (156) than in the cultivated (42) jute. Alleles with a frequency of <0.1 were not detected in the cultivated jute. In contrast, the proportions of medium (>0.3 ≤ 0.6) and high (>0.6) frequency alleles were higher in the cultivated (41 and 12, respectively) than in the wild (26 and 1, respectively) jute. Out of the 222 alleles, there were 166 private alleles; 39 in the cultivated and 127 in the wild jute. The cultivated and wild jute shared a total of 56 alleles, with an avergae frequency of 0.26.
Together, the two cultivated jute species contained ~70% of the nuclear microsatellite diversity present in their wild relatives, reflecting the proportion of wild progenitors’ neutral genetic diversity that is typically retained by today’s modern crops (Gepts 2004; Allaby et al. 2008). The nSSR allele frequency indicates that the wild jute contains many alleles that are missing from the cultivated jute. This is reflected not only in the proportions of the total allelic diversity represented by the wild (82 %) versus cultivated (43 %) jute, but also in the distributions and patterns of private alleles. This reduction in neutral genetic diversity may be due to genetic drift in the form of domestication bottlenecks (Gepts 2004). It is well known that C. capsularis has a much lower level of neutral genetic diversity than C. olitorius (Basu et al. 2004; Roy et al. 2006; Haque et al. 2007; Akter et al. 2008; Mir et al. 2008, 2009). We have also recorded the least nSSR polymorphism in the former. The reduced diversity in C. capsularis compared to C. olitorius indicates its relatively recent origin as a species. Based on Nei’s measure of neutral gene diversity, Allender and King (2010) have also drawn the same conclusion for Brassica napus, in relation to B. oleracea and B. rapa.
nSSR neighbor-joining tree
Jaccard’s NJ tree separated two broad groups, with the tree rooted on C. aestuans: i) C. olitorius including its mutant together with C. urticifolius, and ii) C. capsularis including its mutant together with C. fascicularis, C. pseudo-capsularis, C. pseudo-olitorius, C. tridens and C. trilocularis (Fig. 4). The two-way Mantel test showed lower concordance of the nSSR results to the morphological data, as revealed by a low correlation value between the two data sets (r = 0.35). Notwithstanding, the nSSR data still supported a close relationship between C. fascicularis, C. pseudo-olitorius, C. tridens and C. trilocularis. However, the two cultivated jute species were not closely related, in agreement with earlier results of AFLP (Basu et al. 2004), ISSR (Roy et al. 2006), SSR (Basu et al. 2004; Akter et al. 2008; Mir et al. 2008), RAPD (Hossain et al. 2002; Haque et al. 2007; Roy et al. 2006) and STMS (Roy et al. 2006) analyses. C. urticifolius was found to be closely related to C. olitorius, and therefore it could be the closest extant relative of dark jute. In contrast, the nSSR results failed to identify a close relative of C. capsularis.
Phylogenetic tree based on a distance matrix of nine Corchorus species examined with 38 single-locus nuclear microsatellites. The tree was constructed by using Jaccard’s weighted version of the Neighbor-Joining method, with its topology shown as an inset. Unique chloroplast haplotype of each species as resolved by characterizing mononucleotide repeat-based chloroplast microsatellite variation over the eight intronic and intergenic cpDNA regions are indicated. Numbers at nodes represent % bootstrap support out of 1,000 bootstrap replicates
Our results suggest that C. aestuans could be a common ancestor to both the cultivated jute species. It is sexually compatible with both C. capsularis and C. olitorius (Arangzeb 1994). Roy et al. (2006) have earlier advocated that it could be a progenitor of C. olitorius, with a possibility that both of them might have evolved from a common ancestor. Recent morpho-molecular grouping (Chakraborty and Palit 2009) and karyomorphological analysis (Maity and Datta 2009) have also shown them to be primary relatives. In addition, a close relationship between C. aestuans and C. urticifolius may further allude to the former as an ancestor to C. olitorius, possibly introgressed via natural hybridization with C. urticifolius. The possibility of natural hybridization between the ancestral species of C. olitorius has been suggested earlier (Mir et al. 2008). Of the Corchorus species, C. aestuans is the most cosmopolitan and is not only capable of growing in extremely diverse habitats (Edmonds 1990; Benor et al. 2010), but is also present throughout the world across continents. It is thus logical to assume that both the cultivated jute species might have evolved from C. aestuans or one of its very similar ancestral species. It has been shown that they are allopatric with certain common alleles and may have originated from a closely related common ancestor (Basu et al. 2004).
Chloroplast haplotype diversity
Except for the loci rpl20-rps12 intergenic and ORF 74b-psbB intergenic, all cpSSR loci were amplified in Corchorus species. All of them, excepting trnK intron, were polymorphic and informative (Fig. 5). A total of 30 alleles was detected over the eight loci (Table 1). In total, nine unique chloroplast haplotypes (haplo-1 to haplo-9) were resolved (Table 4; Fig. 4). The haplotype of white jute was more similar to those of C. aestuans and C. pseudo-olitorius, while the haplotype of dark jute was relatively more similar to those of C. aestuans, C. pseudo-olitorius and C. urticifolius. Intra-group haplotype statistics are shown in Table 5. The wild jute had higher haplotypic richness (R h ), genetic diversity (H e ) and mean genetic distance between individuals (D 2 sh ) than the cultivated jute. The two cultivated species contained ~70% of the chloroplast haplotype diversity present in their wild relatives.
Non-denaturing polyacrylamide gels showing monomorphism at the trnK intron and polymorphism at the trnG intron, ORF 77-ORF 82 intergenic and rpl2-rps19 intergenic chloroplast microsatellite loci. M, 100-bp ladder; 1, C. capsularis cv. JRC-321; 2, C. capsularis mt. CMU-010; 3, C. olitorius cv. JRO-524; 4, C. olitorius mt. PPO-4; 5, C. aestuans; 6, C. fascicularis; 7, C. pseudo-capsularis; 8, C. pseudo-olitorius; 9, C. tridens; 10, C. trilocularis; 11, C. urticifolius
What is intriguing in our study is that none of the wild species haplotypes are shared by either of the two domesticated jute species. We further haplotyped 19 and 20 indigenous plus exotic cultivars of white and dark jute, respectively (Supplementary Table 4). However, each cultivated species was characterized by a single chloroplast haplotype. C. olitorius cvs JRO-632 (indigenous) and Sudan Green (exotic from Sudan) were found to represent the same haplotype (haplo-2). Also, the haplotype of another exotic cv. Tanganyika-1 (exotic from Tanzania) was found to be similar to that of the indigenous cultivars of C. olitorius. Similarly, C. capsularis accessions from Africa (CEX-033), Australia (Cap Australia Green) and South America (Solimos) shared the same haplotype (haplo-4) of those from South Asia.
Considering the magnitude of the allele size variation, it seems most likely that the haplotypes of both the cultivated jute species are the result of introgression/chloroplast capture (Wills and Burke 2006). The allele size variation due to variable mononucleotide repeats in noncoding cpDNA is known to increase with phylogenetic distance between the taxa (Weising and Gardner 1999). Thus, on the maternal lineage, both the cultivated jute species are not phylogenetically close in agreement with earlier report that suggested their matrilineal origins to be different (Basu et al. 2004). Since the haplotypes of C. aestuans and C. pseudo-olitorius are more similar to that of C. capsularis than C. olitorius, either of the wild species, the haplotypes of which are very similar to each other, is a much more likely matrilineal progenitor of C. capsularis. But it is not possible at this stage to identify a putative matrilineal progenitor of C. olitorius.
Did both the cultivated jute species originate in Africa, but were independently domesticated in Asia?
Our study supports an African origin of dark jute, possibly in equatorial region of east Africa because its closest extant relative C. urticifolius is only restricted to this region. Archaeobotanical results also suggest its origin in Africa, but introduction to India together with many African crops that reached India in prehistory (Blench 2003). This finds support from the recent human Y-chromosome evidence that indicates an African origin of Dravidian agriculture in southern India (Winters 2010). Interestingly, however, archaeobotanical evidence suggests that C. urticifolius is of south Asian origin and reached Africa at an early period (Blench 2003). Taken together, it is thus most likely that dark jute had its origin in Africa from C. aestuans and C. urticifolius, but was domesticated in India through the development of an ennobled type from its African wild type. Benor et al. (2012) have hypothesized that dispersal of dark jute might have occurred via the Mediterranean-Indian trade route. However, there is no archaeobotanical evidence for C. capsularis, except for that it was used in China and India in early history in clear distinction with silk (Gopal 1961). The possibility of the dispersal of white jute from Africa to Asia (south China) via the Nile valley along the ancient silk route cannot be ruled out. However, in the absence of corresponding African, Chinese or Indian archaeobotanical evidence for white jute, this view would continue to be broadly speculative.
Abbreviations
- cpSSR:
-
Chloroplast simple sequence repeat
- MJM:
-
Meerut jute marker
- NJ:
-
Neighbor joining
- nSSR:
-
Nuclear simple sequence repeat
- PIC:
-
Polymorphism information content
References
Akter J, Islam MS, Sajib AA, Ashraf N, Haque S, Khan H (2008) Microsatellite markers for determining genetic identities and genetic diversity among jute cultivars. Austral J Crop Sci 1:97–107
Allaby GA, Fuller DQ, Brown TA (2008) The genetic expectations of a protracted model for the origins of domesticated crops. Proc Nat Acad Sci USA 105:13982–13986
Allender CJ, King GJ (2010) Origins of the amphiploid species Brassica napus L. investigated by chloroplast and nuclear molecular markers. BMC Plant Biol 10:54
Arangzeb S (1994) Cross compatibility of eight wild species of jute with cultivars and among themselves. Ph. D. Dissertation, Dhaka University, Dhaka
Basu A, Ghosh M, Meyer R, Powell W, Basak SL, Sen SK (2004) Analysis of genetic diversity in cultivated jute determined by means of SSR markers and AFLP profiling. Crop Sci 44:678–685
Benor S, Blattner FR, Demissew S, Hammer K (2010) Collection and ethnobotanical investigation of Corchorus species in Ethiopia: potential leafy vegetables for dry regions. Genet Resour Crop Evol 57:293–306
Benor S, Demissew S, Hammer K, Blattner FR (2012) Genetic diversity and relationships in Corchorus olitorius (Malvaceae s. l.) inferred from molecular and morphological data. Genet Resour Crop Evol 59:1125–1146
Blench RM (2003) The movement of cultivated plants between Africa and India in prehistory. In: Neumann K, Butler A, Kahlheber S (eds) Food, fuel and feeds: Progress in African archaeobotany. Heinrich-Barth-Institut, Köln, pp 273–292
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Human Genet 32:314–331
Chakraborty A, Palit P (2009) Morphological and molecular characterization of nine species of Corchorus (L.) grown in India with special reference to their interrelationship. J Bot Soc Bengal 63:89–99
Du Puy DJ, Telford IRH (1963) Flora of Australia, volume 50. Oceanic Islands 2. Australian Government Publishing Service, Melbourne
Edmonds JM (1990) Herbarium survey of African Corchorus L. species. International Board for Plant Genetic Resources, Rome
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa L. Skeels) endemic to Morocco. Theor Appl Genet 92:832–839
Eliades N-G, Eliades DG (2009) Haplotype analysis: Software for analysis of haplotypes data. Forest genetics and forest tree breeding. Georg-August University Göttingen, Göttingen
Gepts P (2004) Crop domestication as a long-term selection experiments. Plant Breed Rev 24:1–43
Gopal L (1961) Textiles in ancient India. J Econ Social Hist Orient 4:53–69
Goudet J (1995) FSTAT (Version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Grubben GJH, Denton OA (2004) Plant resources of tropical Africa. 2. Vegetables. PROTA Foundation, Wageningen
Hall BG (2001) Phylogenetic trees made easy: A how-to manual for molecular biologists. Sinauer Associates, Sunderland
Haque S, Begum S, Sarker RH, Khan H (2007) Determining genetic diversity of some jute varieties and accessions using RAPD markers. Plant Tissue Cult Biotechnol 17:183–191
Hossain MB, Haque S, Khan H (2002) DNA fingerprinting of jute germplasm by RAPD. J Biochem Mol Biol 35:414–419
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Kundu BC (1951) Origin of jute. Indian J Genet Plant Breed 11:95–99
Kundu BC (1956) Jute: world’s foremost bast fibre. I. Botany, agronomy, diseases and pests. Econ Bot 10:103–133
Kundu A, Sarkar D, Bhattacharjee A, Topdar N, Sinha MK, Mahapatra BS (2011) A simple ethanol wash of the tissue homogenates recovers high-quality genomic DNA from Corchorus species characterized by highly acidic and proteinaceous mucilages. Electron J Biotechnol 14:1
Levene H (1949) On a matching problem arising in genetics. Ann Math Stat 20:91–94
Maity S, Datta AK (2009) Karyomorphology in nine species of jute (Corchorus L., Tiliaceae). Cytologia 74:273–279
Mir RR, Rustgi S, Sharma S, Singh R, Goyal A, Kumar J, Gaur A, Tyagi AK, Khan H, Sinha MK, Balyan HS, Gupta PK (2008) A preliminary genetic analysis of fibre traits and the use of new genomic SSRs for genetic diversity in jute. Euphytica 161:413–427
Mir RR, Banerjee S, Das M, Gupta V, Tyagi AK, Sinha MK, Balyan HS, Gupta PK (2009) Development and characterization of large-scale simple sequence repeats in jute. Crop Sci 49:1687–1694
Mohammad A, Al-Rabab’ AH, Williams CG (2004) An ancient bottleneck in the lost pines of central Texas. Mol Ecol 13:1075–1084
Morgante M, Felice N, Vendramin GG (1998) Analysis of hypervariable chloroplast microsatellites in Pinus halepensis reveals a dramatic bottleneck. In: Karp A (ed) Molecular tools for screening biodiversity: Plants and animals. Chapman and Hall, London, pp 402–412
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Nat Acad Sci USA 70:3321–3323
Palit P, Sasmal BC, Bhattacharryya AC (1996) Germplasm diversity and estimate of genetic advance of four morpho-physiological traits in a world collection of jute. Euphytica 90:49–58
Perrier X, Jacquemoud-Collet JP (2006) DARwin software, version 5.0.158. http://darwin.cirad.fr/darwin
Perrier X, Flori A, Bonnot F (2003) Data analysis method. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science Publishers, Enfield, pp 43–76
Petit RJ, El Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Rohlf FJ (2005) NTSYSpc: Numerical taxonomy and multivariate analysis system version 2.2. Applied Biostatistics Inc, New York
Roy A, Bandyopadhyay A, Mahapatra AK, Ghosh SK, Singh NK, Bansal KC, Koundal KR, Mohapatra T (2006) Evaluation of genetic diversity in jute (Corchorus species) using STMS, ISSR and RAPD markers. Plant Breed 125:292–297
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sanguinetti CJ, Dias Neto E, Simpson AJG (1994) Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. Biotechniques 17:915–919
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Weising K, Gardner RC (1999) A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42:9–19
Wills DM, Burke JM (2006) Chloroplast DNA variation confirms a single origin of domesticated sunflower (Helianthus annuus L.). J Hered 97:403–408
Winters CA (2010) Y-chromosome evidence of an African origin of Dravidian agriculture. Int J Genet Mol Biol 2:30–33
Wright RP (2010) The ancient Indus. Urbanism, economy, and society. Cambridge University Press, New York
Yeh FC, Yang R-C, Boyle T (1999) POPGENE, version 1.31: Microsoft Window-based freeware for population genetic analysis. http://www.ualberta.ca/~fyeh/fyeh
Acknowledgments
The research was supported by Department of Biotechnology, Ministry of Science and Technology, Government of India (File No. BT/PR/7143/AGR/16/672/2006). Comments and suggestions on the manuscript from two anonymous reviewers are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kundu, A., Topdar, N., Sarkar, D. et al. Origins of white (Corchorus capsularis L.) and dark (C. olitorius L.) jute: a reevaluation based on nuclear and chloroplast microsatellites. J. Plant Biochem. Biotechnol. 22, 372–381 (2013). https://doi.org/10.1007/s13562-012-0165-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-012-0165-7