Abstract
Purpose
Cholangiocarcinoma (CCA) is a highly invasive malignant tumor originating from the bile duct epithelium. Tweety homolog 3 (TTYH3) is a member of the family of calcium-activated chloride channels, which have several biological functions. Here, we aimed to investigate the expression and biological function of TTYH3 in CCA.
Methods
The mRNA and protein expression levels of TTYH3 were investigated in primary human CCA tissues and normal tissues. The DNA methylation levels of three CpG sites in the TTYH3 promoter region were evaluated using pyrosequencing. The effect of TTYH3 expression on proliferation, apoptosis, migration and invasion were assessed in HUCCT1 and QBC939 cells. Xenograft models were developed to substantiate its role in the development of CCA. Western blot analysis was used to investigate the mechanistic role of TTYH3 in regulating CCA progression.
Results
We found that TTYH3 was highly expressed both at the mRNA and protein levels in CCA (p = 0.0001) and that the expression levels were significantly related to a poor overall survival of the patients (p = 0.0019). The DNA methylation levels of three CpG sites in the TTYH3 promoter region were significantly lower in CCA tissues compared to normal tissues (p < 0.05). In vitro studies indicated that TTYH3 can promote the proliferation, migration and invasion of the CCA cells. TTYH3 overexpression significantly promoted tumor progression and cellular proliferation in vivo as indicated by Ki‐67 expression. In addition, we found that exogenous TTYH3 overexpression induced epithelial-mesenchymal transition (EMT) in CCA as indicated by expression changes in E-cadherin, N-cadherin and vimentin. The EMT process was found to occur through the Wnt/β-catenin signaling pathway, with simultaneous changes in P-GSK3β and β-catenin levels.
Conclusions
Our data indicate that DNA hypomethylation-induced overexpression of TTYH3 regulates CCA development and metastasis through the Wnt/β-catenin pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cholangiocarcinoma (CCA) is a highly invasive malignant tumor originating from the epithelium of the bile duct, and its incidence is increasing worldwide. It accounts for approximately 2 % of global cancer-related deaths each year [1]. Although surgical resection is the best treatment for CCA, a low surgical resection rate and post-operative early recurrence and metastasis are the main reasons for the low overall survival. Additionally, resistance to chemotherapy and the nonavailability of targeted drugs limit the survival of CCA patients. Therefore, investigation of the molecular mechanisms underlying the development and metastasis of CCA is necessary to identify new therapeutic targets.
Ca2+-activated chloride channels (CaCCs) comprise a group of membrane proteins that function as ligand-gated ion channels for chloride [2]. CaCCs have been identified in different epithelial cells, endothelial cells and smooth muscle cells [3]. These proteins regulate different biological functions in different cells, including neuronal excitability, epithelial mucus secretion and proliferation [4, 5]. In addition, CaCCs play important roles in cancer development and progression. TMEM16A, the best studied CaCC, has been related to the progression of several types of cancer [6, 7]. It has e.g. been reported that overexpression of TMEM16A promotes the growth, migration and invasion, and inhibits the apoptosis of hepatocellular carcinoma cells [8]. CLCA, another member of the CaCC family, has been reported to be involved in tumor metastasis and regulation of the cell cycle and apoptosis [9,10,11].
Tweety homolog 3 (TTYH3) is a member of the tweety family of proteins that act as Ca2+-activated chloride channels. The members of this family play a role in several cellular processes, including cell adhesion, division and tumorigenesis [12]. TTYH1 has been found to be associated with brain cancer [13,14,15,16] and to regulate embryonic neural stem cell characteristics by enhancing the activity of the Notch signaling pathway [17]. TTYH2 has been found to be upregulated in colon and renal cell carcinomas [18, 19], and downregulation of TTYH2 has been found to limit epithelial-mesenchymal transition (EMT) in osteosarcoma [20]. According to a bioinformatic study, TTYH3, the third member of the family, is upregulated in gastric cancer and associated with a poor prognosis [21], but its biological function has not been validated. Also, the biological function of TTYH3 in CCA is elusive.
Here, we investigated the expression of TTYH3 in CCA tissues and cells and the methylation status of CpG sites within its promoter region. We studied the biological function of TTYH3 in CCA cell proliferation, migration, invasion and apoptosis in vitro and in CCA tumor formation in vivo. We also investigated the molecular mechanism underlying TTYH3-mediated regulation of CCA progression.
2 Materials and methods
2.1 Analysis of TTYH3 expression and methylation using the TCGA database
GEPIA (http://gepia2.cancer-pku.cn) is a tool for the visualization of data deposited in the Cancer Genome Atlas (TCGA). The relative expression of TTYH3 in the TCGA database was reflected by setting the following screening criteria of the box plots “cancer name: CHOL (CHOL = CCA), |Log2FC| Cutoff: 1 and P-value Cutoff: 0.01”. UALCAN (http://ualcan.path.uab.edu/index.html) is an online website to acquire the methylation status of genes in certain cancerous and their normal tissues in the TCGA database.
2.2 Cell culture and transfection
The human biliary epithelial cell line HIBEpic and the CCA cell lines HUCCT1, QBC939, RBE and FRH0201 were purchased from the Shanghai Institute for Biological Sciences, Chinese Academy of Sciences. Dulbecco’s Modified Eagle Medium (DMEM; HyClone, USA) supplemented with 10 % fetal bovine serum (FBS, BI, Israel) and 1 % penicillin and streptomycin was used for culturing the cells in a humidified incubator (Thermo Scientific, USA) at 37 °C and 5 % CO2. The TTYH3 and negative control siRNAs used were obtained from GenePharma Co. Ltd. (Shanghai, China). The plasmids FLAG-NC and FLAG-TTYH3 were obtained from GeneChem Co. Ltd. (Shanghai, China). Cells were transiently transfected by incubating them with siRNAs and Xfect Transfection Reagent (Takara, Japan) for 48 h or with the plasmids and Lipofectamine 2000 (Invitrogen, USA) for 24 h. To obtain stable HUCCT1 cells with a high TTYH3 expression, a lentiviral vector harboring TTYH3 (Hanbio Biotechnology, Shanghai, China) was transfected into the cells after which they were selected using puromycin.
2.3 CCA tissue samples and ethics statement
We collected thirty pairs of tumor and para-tumor tissues from patients who underwent hepatectomy from 2016 to 2019 at the affiliated hospital of Qingdao University. All the patients were confirmed as CCA by post-operative pathology and their clinical and follow-up data were integrated. This study complies with the declaration of Helsinki. The sample collection was approved by the ethics committee of the affiliated hospital of Qingdao University. No preoperative radiotherapy or chemotherapy was given to any of the patients. The tissue samples were promptly preserved in RNA protective additive-filled microtubes and stored at -80 °C.
2.4 Quantitative real-time PCR analysis
Total RNA was extracted from the cells and the tissue samples using a RNAiso Plus kit (total RNA extraction reagent, Takara Inc. Japan) following the manufacturer’s instructions. Total RNA was quantified using the A260/A280 ratio, and the corresponding complementary DNA was obtained using a PrimeScript™ RT reagent Kit (Takara Inc. Japan). Quantitative real-time PCR (qRT-PCR) was performed on a LightCycler 480 (Roche, Basel, Switzerland) using Takara TB Green™ Premix Ex Taq™ II (Tli RNaseH Plus, Takara Inc. Japan). The thermocycling conditions were: pre-incubation at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 20 s, annealing at 60 °C for 30 s and extension at 72 °C for 20 s. The experiments were performed in triplicate. A melting curve was analyzed for each sample to verify the specificity of the amplification. GAPDH was used as internal control.
2.5 Methylation detection by pyrophosphate sequencing
DNA was isolated from CCA and normal tissues of 16 patients using a genomic DNA extraction kit (QIAGEN DNeasy kit, Germany). A QiagenEpiTect Bisulfite Kit (Qiagen, Germany) was used for methylation. The primer design software used was PyroMark Assay Design 2.0 (Qiagen, Germany). PCR amplification was carried out for 40 cycles and pyrosequencing was carried out on a pyrosequencing detector (PyroMark Q96 ID, QIAGEN, Germany) according to the manufacturer’s instructions.
2.6 Cell viability and colony assays
Cells were inoculated in a 96-well cell culture plate (2000 cells/well) and incubated overnight at 37 °C and 5 % CO2. A Cell Counting Kit-8 (CCK-8) assay was used to assess cell viability at different time points. The cells were cultured in 100 µl DMEM supplemented with 10 µl CCK-8 reagent per well for 1 h at 37 °C. Next, optical density (OD) was measured at 450 nm absorbance. For the colony assay, 1000 transfected cells per well were seeded in a six-well culture plate and cultured for 14 days at 37 °C and 5 % CO2. The cells were fixed with 10 % formalin and stained with crystal violet, after which colonies were counted.
2.7 Cell migration and invasion assays
Following transfection and culture, cells of the control group and the experimental group were counted and placed in trans-well chambers for the migration assay (Transwell assay). CCA cells (105 cells) were mixed with serum-free DMEM and added to the chambers and 600 µl DMEM supplemented with 10 % FBS was added to the bottom of the 24-well cell culture plate, after which the chambers were placed in the wells of the plate. For the invasion assay, the chambers were pre-treated with Matrigel (BD Biosciences, USA) according to the manufacturer’s instructions. After culturing in the incubator for 10–24 h, methanol was added to the chambers for fixing the cells that had migrated or invaded. The cells were stained with crystal violet, observed and counted under a microscope.
2.8 Scratch wound-healing assay
Cells were cultured in a six-well cell culture plate and incubated overnight at 37 °C and 5 % CO2. On the subsequent day, the cells were transiently transfected. Next, the cells were washed twice with phosphate-buffered saline (PBS), followed by making cross scratches at the bottom of the wells using a 200 µl pipette tip. Images of the ‘wounds’ were captured at 0 and 24 h using microscopy (Nikon, Japan).
2.9 Flow cytometric apoptosis assay
Transfected cells were evaluated using an Annexin V-PI apoptosis assay kit, by staining with Annexin V and Propidium Iodide (PI) (Basin, Shanghai, China), as per the manufacturer’s instructions. Cell apoptosis was measured using a BD FACSCalibur flow cytometer (BD Biosciences, USA).
2.10 Western blotting
Total proteins from transfected cells and tissues were extracted using a lysis solution comprising radioimmunoprecipitation assay (RIPA) buffer, phenylmethylsulfonyl fluoride (PMSF) and protease inhibitor cocktail (PIC) in a ratio of 100:1:1. The protein concentrations were measured after which electrophoresis was performed and the proteins were transferred to polyvinylidene fluoride (PVDF) membranes (0.45 μm; Millipore, USA), activated by treating with methyl alcohol for 2 min. The proteins were stained using labelled antibodies and the protein bands were visualized using a chemiluminescent kit (Affinity, USA).
2.11 Immunohistochemistry (IHC) and survival analysis
Paraformaldehyde-fixed (4 %) and paraffin-embedded tissue sections were deparaffinized in xylene and sequentially hydrated with absolute ethyl alcohol, 95 % ethyl alcohol, 90 % ethyl alcohol and 85 % ethyl alcohol. Antigen was retrieved using a citrate buffer (pH 6). Next, the sections were incubated with a primary anti-TTYH3 antibody (1:100; Thermo Fisher, ab2649109, USA) at 4 °C overnight, followed by incubation with a polymer-HRP secondary antibody (ZSGB-Bio, MA2522, Beijing, China) for 2 h at 25 °C. The sections were counterstained with hematoxylin. The results were evaluated based on the intensity and degree of staining. TTYH3 staining intensity was marked as 0 (none), 1 (weak), 2 (medium) and 3 (strong). The staining degree was scored according to the percentage of positive cells as follows: 0 (< 5 %), 1 (5–25 %), 2 (26–50 %), 3 (51–75 %) and 4 (> 75 %). The score of each section was calculated according to the following formula: staining intensity score x staining degree score. A total score < 6 is regarded as low expression, and > 6 as high expression. Images of each section were acquired using a Pannoramic SCAN (3DHISTECH, Hungary). Survival analysis was performed based on the TTYH3 expression level.
2.12 Xenograft model
All animal experiments were carried out at the specific-pathogen-free medical animal laboratory of the Affiliated Hospital of Qingdao University and approved by the Animal Ethics Committee of the Affiliated Hospital of Qingdao University. Ten male BALB/c athymic nude mice (4-week-old) were purchased from Charles River Laboratory Animal Technology Co., Ltd. (https://www.vitalriver.com/) and reared in an environment free of pathogens and suitable for the growth of mice, for 5 days according to the guidelines of animal care institutions. A stable HUCCT1 cell line overexpressing TTYH3 was successfully developed and subcutaneously injected into the mice. The 10 mice were divided into two groups (5 mice for lentivirus-TTYH3 and 5 for lentivirus-nc). Each mouse received an injection of 5 × 106 cells suspended in PBS. The sizes of the tumors were recorded every 6 days and they were excised after one month. The sizes of the tumors were measured using a Vernier caliper, after which the volumes of the tumors were calculated as follows: Volume = Length × Width2. After removal, the final volumes of the tumors were calculated and the tumors were preserved for subsequent experiments.
2.13 Statistical analysis
Data analysis was performed using GraphPad Prism 7.0 software (San Diego, CA, USA). All experimental data are expressed as mean ± SD. Statistical significance of the results was analyzed using chi-square test for the patients in Table 1 and Student’s t-test for two-group comparisons. p < 0.05 was considered statistically significant. All experiments were performed in triplicate.
3 Results
3.1 TTYH3 is upregulated in CCA tissues and correlates with a poor prognosis
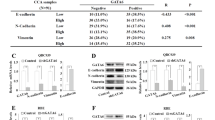
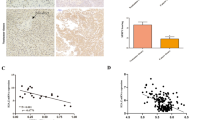
We analyzed TTYH3 expression in nine normal and 36 CCA tissues in the TCGA database (Fig. 1a). We found that the expression of TTYH3 was significantly higher in the CCA tissues than in the normal tissues (p < 0.05). We next evaluated the expression of TTYH3 in primary CCA tissues and their corresponding para-carcinoma tissues from 30 patients admitted to our hospital using qRT-PCR. We found that the expression of TTYH3 mRNA was markedly higher in the CCA tissues compared to the para-carcinoma tissues (p < 0.05, Fig. 1b). To evaluate its relation to CCA prognosis, we collected tissue sections of 71 patients who underwent radical CCA resection between 2005 and 2018. The clinicopathological information of the 71 patients is listed in Table 1. Using IHC we again found that the expression level of TTYH3 in the CCA tissues was markedly higher than that in normal tissues and that TTYH3 was localized at the cell membrane (Fig. 1c). The patients were divided into two groups, according to IHC staining: a low expression group with 41 patients and a high expression group with 30 patients. Subsequent Kaplan-Meier analysis indicated that a high TTYH3 expression was associated with a shorter overall survival (p = 0.0019, Fig. 1d).
TTYH3 is highly expressed in CCA and regulated by DNA methylation. (a) In the TCGA database, TTYH3 expression in tumor tissues is significantly higher than that in normal tissues. (b) qRT-PCR analysis of TTYH3 mRNA expression in normal and tumor tissues of 30 patients. (c) IHC analysis of TTYH3 protein expression in para-cancerous normal tissues and CCA tissues. (d) Kaplan-Meier analysis of patients with different TTYH3 expression levels. (e) TTYH3 DNA methylation level in the TCGA database. (f) DNA methylation levels of three CpG sites within the TTYH3 promoter region evaluated using pyrophosphate sequencing. (g) Representative sequencing result of CpG site CG23195008
3.2 TTYH3 methylation in CCA tissues is lower than that in normal tissues
Using the TCGA database, we found that the methylation level of TTYH3 in CCA tissues was significantly lower than that in normal tissues (p < 0.05, Fig. 1e). Subsequent pyrophosphate sequencing indicated that the methylation levels of the three TTYH3 methylation sites (CG00798876, CG23195008 and CG00177218) in CCA tissues were lower than those in the corresponding normal tissues (Fig. 1f, p < 0.05). The CpG methylation level of CG23195008 in a CCA tissue and its paired normal tissue is shown in Fig. 1g.
3.3 TTYH3 promotes the proliferation, migration and invasion of CCA cells in vitro
TTYH3 expression was evaluated using qRT-PCR in four CCA cell lines (Fig. 2a). We found that the expression was lowest in HUCCT1 cells and highest in QBC939 cells (p < 0.05). Therefore, QBC939 was selected for TTYH3 knockdown and HUCCT1 for TTYH3 overexpression analyses. After transfection and culturing of the cells, the transfection efficiencies were confirmed using qRT-PCR and found to be effective (Fig. 2b and c). We selected the cells with the highest knockdown efficiency for subsequent functional analyses.
TTYH3 promotes CCA cell proliferation, migration and invasion, and inhibits CCA cell apoptosis. (a) Relative TTYH3 expression levels in HIBEpiC and four CCA cell lines. (b) TTYH3 overexpression and knockdown efficiencies in HUCCT1 and QBC939 cells. (c) CCK-8 proliferation assay. (d) Effect of TTYH3 upregulation and downregulation on CCA colony growth. (e) Effect of TTYH3 upregulation and downregulation on the migration and invasion of CCA cells using Transwell analyses. (f) Scratch wound-healing analysis of CCA cell migration induced by TTYH3. (g) Flow cytometry showing a decreased apoptosis rate in HUCCT1 cells with upregulated TTYH3 expression and an elevated apoptosis rate in QBC939 cells with downregulated TTYH3 expression
Using a CCK-8 proliferation assay we found that TTYH3 overexpression promoted HUCCT1 cell proliferation, while TTYH3 knockdown inhibited the proliferation of QBC939 cells (Fig. 2d). The colony assay showed that TTYH3 overexpression increased the number of HUCCT1 colonies, compared to those in the negative control group. TTYH3 knockdown led to fewer QBC939 colonies (Fig. 2e). The Transwell assays indicated that TTYH3 upregulation promoted HUCCT1 cell migration and invasion, while its knockdown markedly reduced the migration and invasion of QBC939 cells (Fig. 2f). Moreover, 24 h after scratching, the wounded area of HUCCT1 cells transfected with the vector plasmid was significantly larger than that of those transfected with the TTYH3 overexpression plasmid. Together these results indicate that TTYH3 overexpression induces the migration and invasion of CCA cells, whereas its downregulation inhibits the migration and invasion of CCA cells (Fig. 2g).
3.4 TTYH3 affects apoptosis and EMT of CCA cells via the Wnt/β-catenin pathway
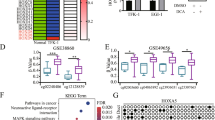
Using flow cytometry, we found that TTYH3 upregulation inhibited apoptosis in HUCCT1 cells, while TTYH3 downregulation promoted apoptosis in QBC939 cells (Fig. 2h). The expression of BAX and BCL2 corroborated the flow cytometry results (Fig. 3a and b).
TTYH3 regulates EMT through the Wnt/ β-catenin signaling pathway in CCA cells. (a, b) Western blot analysis of the effect of TTYH3 overexpression in HUCCT1 cells and TTYH3 downregulation in QBC939 cells on the expression of apoptosis-related proteins. (c, d) Western blot analysis of the effect of TTYH3 overexpression in HUCCT1 cells and TTYH3 downregulation in QBC939 cells on the expression of EMT-related proteins. (e, f, g and h) Participation of TTYH3 in the EMT process of CCA cells via the Wnt/β-catenin pathway
To evaluate whether TTYH3 is involved in the process of EMT, the expression of the EMT-related proteins N-cadherin, E-cadherin and vimentin was evaluated using Western blotting. We found that TTYH3 overexpression in HUCCT1 cells led to downregulation of the expression of E-cadherin and upregulation of that of N-cadherin and vimentin (Fig. 3c). TTYH3 knockdown in QBC939 cells significantly upregulated the expression of E-cadherin and downregulated that of N-cadherin and vimentin (Fig. 3d). These results indicate that TTYH3 may play a key role in the EMT of CCA cells. Based on the expression of β-catenin, GSK3β (ser 9) and P-GSK3β (ser 9) (Fig. 3e and f), we conclude that TTYH3 regulates CCA cell EMT through the Wnt/β-catenin pathway. In addition, we examined expression levels of Wnt/β-catenin pathway downstream targets and found that TTYH3 overexpression in HUCCT1 cells upregulated the expression levels of Met, c-Myc and cyclin D1 (Fig. 3g). TTYH3 knockdown in QBC939 markedly downregulated the expression levels of Met, c-Myc and cyclin D1 (Fig. 3h).
3.5 TTYH3 upregulation enhances tumorigenesis in vivo
We established a subcutaneous xenograft model to investigate the role of TTYH3 in vivo. One month after inoculation, the tumors were removed, weighed, and photographed. We found that the volumes of the tumors in the negative group were significantly lower than those in the TTYH3 overexpression group (Fig, 4a, b). The expression of TTYH3 in the overexpression group was significantly higher than that in the negative group (p < 0.05, Fig. 4c). Immunohistochemistry showed that Ki-67 was significantly overexpressed in the TTYH3-high group (p < 0.05, Fig. 4d).
4 Discussion
CCA is a highly invasive malignant tumor with few treatment options and an overall poor survival. We found that TTYH3 is upregulated in CCA and that this upregulation is associated with the poor survival. We also found that TTYH3 enhanced the viability, migration and invasion, and reduced the apoptosis of CCA cells. TTYH3 overexpression in CCA cells dramatically promoted tumor growth in vivo. TTYH3 overexpression may promote EMT in CCA cells through the Wnt/ β-catenin signaling pathway.
CaCCs comprise a group of ion channel proteins for the transport of chloride, which have different biological functions in epithelial and smooth muscle cells. The structurally different members of the family, TMEM16A and CLCA1, have been found to be involved in the development of various cancers, including liver and colorectal cancer [6, 10, 11, 22,23,24,25]. The TTYH3 gene is located on chromosome 7p22.3 and is also a member of the CaCC family. Others have reported that TTYH3 upregulation is associated with a poor prognosis in patients with gastric cancer [21], but the underlying molecular mechanism was not studied.
Both genetic and epigenetic modifications influence the occurrence and development of tumors and the most common epigenetic modification is DNA methylation [26]. Mutation in the TMEM16A gene has been reported to be related to the occurrence and development of various types of cancer [3]. Therefore, we reviewed the CBioPortal (http://www.cbioportal.org/) database and found that the mutation frequency of TMEM16A in CCA was 2.9 %, while that of TTYH3 was negative. Therefore, we hypothesized that TTYH3 may be regulated through methylation, which was confirmed using data on the UALCAN website. The methylation of TTYH3 in primary samples was analyzed using pyrophosphate sequencing to show that hypomethylation of TTYH3 may promote the development of CCA.
Tumor invasion and metastasis are main factors that influence the prognosis of CCA. EMT is an important biological process underlying the migration and invasion of tumor cells derived from epithelial cells [27,28,29,30]. Calcium (2+)-activated large-conductance chloride (-) channel proteins are associated with the invasion and metastasis of cancer, including the process of EMT. miRNA-381 has been reported to inhibit EMT in gastric cancer by targeting TMEM16A expression [31], and CLCA1 has been reported to inhibit EMT in colorectal cancer by downregulating the Wnt/β-catenin signaling pathway [25]. Following activation of the canonical Wnt/β-catenin pathway, β-catenin induces the transcription and translation of downstream target genes accelerating the EMT process [32]. Here, we found that upregulation of TTYH3 in CCA cells enhanced their migration and invasiveness. TTYH3 overexpression markedly enhanced the expression of mesenchymal markers, such as N-cadherin and vimentin, and reduced the expression of the epithelial marker E-cadherin. TTYH3 downregulation exhibited the opposite effect. TTYH3 overexpression downregulated Bax and upregulated Bcl2 expression. TTYH3 knockdown had the opposite effect. Our data, therefore, substantiate the notion that TTYH3 upregulation promotes CCA progression via EMT.
There are some limitations to this study. TTYH3 is an ion channel and, therefore, its function may have been influenced by chloride transportation. There is also a possibility that TTYH3 activates the GSK3β/β-catenin pathway through kinase activity. These options need to be addressed in future studies.
In conclusion, we found that TTYH3 is markedly overexpressed in CCA cells and tissues, and that it plays an important role in promoting the progression of CCA. TTYH3 may be involved in CCA initiation through DNA hypomethylation and in CCA progression by regulating EMT through the Wnt/ β-catenin pathway. TTYH3 may serve as a potent therapeutic target for CCA and/or as a biomarker for CCA prognosis (Fig. 5).
Data availability
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- CCA:
-
cholangiocarcinoma
- CaCC:
-
Ca2+-activated chloride channel
- TTYH3:
-
Tweety homolog 3
- qRT-PCR:
-
quantitative real-time PCR
- EMT:
-
epithelial-mesenchymal transition
References
J.M. Banales, J.J.G. Marin, A. Lamarca, P.M. Rodrigues, S.A. Khan, L.R. Roberts, V. Cardinale, G. Carpino, J.B. Andersen, C. Braconi, D.F. Calvisi, M.J. Perugorria, L. Fabris, L. Boulter, R.I.R. Macias, E. Gaudio, D. Alvaro, S.A. Gradilone, M. Strazzabosco, M. Marzioni, C. Coulouarn, L. Fouassier, C. Raggi, P. Invernizzi, J.C. Mertens, A. Moncsek, S. Rizvi, J. Heimbach, B.G. Koerkamp, J. Bruix, A. Forner, J. Bridgewater, J.W. Valle, G.J. Gores, Cholangiocarcinoma 2020: the next horizon in mechanisms and management. Nat. Rev. Gastroenterol. Hepatol. 17, 557–588 (2020)
L. Ferrera, O. Zegarra-Moran, L.J. Galietta, Ca2+-activated Cl- channels. Compr. Physiol. 1, 2155–2174 (2011)
H. Wang, L. Zou, K. Ma, J. Yu, H. Wu, M. Wei, Q. Xiao, Cell-specific mechanisms of TMEM16A Ca(2+)-activated chloride channel in cancer. Mol. Cancer 16, 152 (2017)
Y.D. Yang, H. Cho, J.Y. Koo, M.H. Tak, Y. Cho, W.S. Shim, S.P. Park, J. Lee, B. Lee, B.M. Kim, R. Raouf, Y.K. Shin, U. Oh, TMEM16A confers receptor-activated calcium-dependent chloride conductance. Nature 455, 1210–1215 (2008)
J. Eggermont, Calcium-activated chloride channels: (un)known, (un)loved? Proc. Am. Thorac. Soc. 1, 22–27 (2004)
U. Duvvuri, D.J. Shiwarski, D. Xiao, C. Bertrand, X. Huang, R.S. Edinger, J.R. Rock, B.D. Harfe, B.J. Henson, K. Kunzelmann, R. Schreiber, R.S. Seethala, A.M. Egloff, X. Chen, V.W. Lui, J.R. Grandis, S.M. Gollin, TMEM16A induces MAPK and contributes directly to tumorigenesis and cancer progression. Cancer Res. 72, 3270–3281 (2012)
A. Carles, R. Millon, A. Cromer, G. Ganguli, F. Lemaire, J. Young, C. Wasylyk, D. Muller, I. Schultz, Y. Rabouel, D. Dembele, C. Zhao, P. Marchal, C. Ducray, L. Bracco, J. Abecassis, O. Poch, B. Wasylyk, Head and neck squamous cell carcinoma transcriptome analysis by comprehensive validated differential display. Oncogene 25, 1821–1831 (2006)
C. Zhang, J. Liu, Z. Han, X. Cui, D. Peng, Y. Xing, Inhibition of TMEM16A suppresses growth and induces apoptosis in hepatocellular carcinoma. Int. J. Clin. Oncol. 25, 1145–1154 (2020)
M.E. Loewen, G.W. Forsyth, Structure and function of CLCA proteins. Physiol. Rev. 85, 1061–1092 (2005)
J.A. Kim, Y.S. Kang, Y.S. Lee, Role of Ca2+-activated Cl- channels in the mechanism of apoptosis induced by cyclosporin A in a human hepatoma cell line. Biochem. Biophys. Res. Commun. 309, 291–297 (2003)
D. Wohlrab, F. Markwardt, Influence of ion channel blockers on proliferation and free intracellular Ca2+ concentration of human keratinocytes. Skin Pharmacol. Appl. Skin Physiol. 12, 257–265 (1999)
A.D. Halleran, M. Sehdev, B.A. Rabe, R.W. Huyck, C.C. Williams, M.S. Saha, Characterization of tweety gene (ttyh1-3) expression in Xenopus laevis during embryonic development. Gene Expr. Patterns 17, 38–44 (2015)
E. Jung, M. Osswald, J. Blaes, B. Wiestler, F. Sahm, T. Schmenger, G. Solecki, K. Deumelandt, F.T. Kurz, R. Xie, S. Weil, O. Heil, C. Thome, M. Gommel, M. Syed, P. Haring, P.E. Huber, S. Heiland, M. Platten, A. von Deimling, W. Wick, F. Winkler, Tweety-Homolog 1 drives brain colonization of gliomas. J. Neurosci. 37, 6837–6850 (2017)
C.L. Kleinman, N. Gerges, S. Papillon-Cavanagh, P. Sin-Chan, A. Pramatarova, D.A. Quang, V. Adoue, S. Busche, M. Caron, H. Djambazian, A. Bemmo, A.M. Fontebasso, T. Spence, J. Schwartzentruber, S. Albrecht, P. Hauser, M. Garami, A. Klekner, L. Bognar, J.L. Montes, A. Staffa, A. Montpetit, P. Berube, M. Zakrzewska, K. Zakrzewski, P.P. Liberski, Z. Dong, P.M. Siegel, T. Duchaine, C. Perotti, A. Fleming, D. Faury, M. Remke, M. Gallo, P. Dirks, M.D. Taylor, R. Sladek, T. Pastinen, J.A. Chan, A. Huang, J. Majewski, N. Jabado, Fusion of TTYH1 with the C19MC microRNA cluster drives expression of a brain-specific DNMT3B isoform in the embryonal brain tumor ETMR. Nat. Genet. 46, 39–44 (2014)
M. Stefaniuk, L. Swiech, J. Dzwonek, K. Lukasiuk, Expression of Ttyh1, a member of the Tweety family in neurons in vitro and in vivo and its potential role in brain pathology. J. Neurochem. 115, 1183–1194 (2010)
C.A. Matthews, J.E. Shaw, J.A. Hooper, I.G. Young, M.F. Crouch, H.D. Campbell, Expression and evolution of the mammalian brain gene Ttyh1. J. Neurochem. 100, 693–707 (2007)
J. Kim, D. Han, S.H. Byun, M. Kwon, J.Y. Cho, S.J. Pleasure, K. Yoon, Ttyh1 regulates embryonic neural stem cell properties by enhancing the Notch signaling pathway, EMBO Rep. 19, e45472 (2018)
F.K. Rae, J.D. Hooper, H.J. Eyre, G.R. Sutherland, D.L. Nicol, J.A. Clements, TTYH2, a human homologue of the Drosophila melanogaster gene tweety, is located on 17q24 and upregulated in renal cell carcinoma. Genomics 77, 200–207 (2001)
Y. Toiyama, A. Mizoguchi, K. Kimura, J. Hiro, Y. Inoue, T. Tutumi, C. Miki, M. Kusunoki, TTYH2, a human homologue of the Drosophila melanogaster gene tweety, is up-regulated in colon carcinoma and involved in cell proliferation and cell aggregation. World J. Gastroenterol. 13, 2717–2721 (2007)
D.K. Moon, Y.J. Bae, G.R. Jeong, C.H. Cho, S.C. Hwang, Upregulated TTYH2 expression is critical for the invasion and migration of U2OS human osteosarcoma cell lines. Biochem. Biophys. Res. Commun. 516, 521–525 (2019)
S.K. Saha, P.K. Biswas, M. Gil, S.G. Cho, High expression of TTYH3 is related to poor clinical outcomes in human gastric cancer. J. Clin. Med. 8, 1762 (2019)
V. Walia, Y. Yu, D. Cao, M. Sun, J.R. McLean, B.G. Hollier, J. Cheng, S.A. Mani, K. Rao, L. Premkumar, R.C. Elble, Loss of breast epithelial marker hCLCA2 promotes epithelial-to-mesenchymal transition and indicates higher risk of metastasis. Oncogene 31, 2237–2246 (2012)
R.B. West, C.L. Corless, X. Chen, B.P. Rubin, S. Subramanian, K. Montgomery, S. Zhu, C.A. Ball, T.O. Nielsen, R. Patel, J.R. Goldblum, P.O. Brown, M.C. Heinrich, M. van de Rijn, The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am. J. Pathol. 165, 107–113 (2004)
J. Chenevert, U. Duvvuri, S. Chiosea, S. Dacic, K. Cieply, J. Kim, D. Shiwarski, R.R. Seethala, DOG1: a novel marker of salivary acinar and intercalated duct differentiation. Mod. Pathol. 25, 919–929 (2012)
X. Li, W. Hu, J. Zhou, Y. Huang, J. Peng, Y. Yuan, J. Yu, S. Zheng, CLCA1 suppresses colorectal cancer aggressiveness via inhibition of the Wnt/beta-catenin signaling pathway. Cell Commun. Signal 15, 38 (2017)
Y. Yu, M. Zhang, N. Wang, Q. Li, J. Yang, S. Yan, X. He, G. Ji, L. Miao, Epigenetic silencing of tumor suppressor gene CDKN1A by oncogenic long non-coding RNA SNHG1 in cholangiocarcinoma. Cell Death Dis. 9, 746 (2018)
M.A. Nieto, R.Y. Huang, R.A. Jackson, J.P. Thiery, Emt: 2016. Cell 166, 21-45 (2016)
S. Gupta, A. Maitra, Matter of life or death? Cell 164, 840–842 (2016)
C.J. David, Y.H. Huang, M. Chen, J. Su, Y. Zou, N. Bardeesy, C.A. Iacobuzio-Donahue, J. Massague, TGF-beta tumor suppression through a lethal EMT. Cell 164, 1015–1030 (2016)
T. Chen, Y. You, H. Jiang, Z.Z. Wang, Epithelial-mesenchymal transition (EMT): A biological process in the development, stem cell differentiation, and tumorigenesis. J. Cell. Physiol. 232, 3261–3272 (2017)
Q. Cao, F. Liu, K. Ji, N. Liu, Y. He, W. Zhang, L. Wang, MicroRNA-381 inhibits the metastasis of gastric cancer by targeting TMEM16A expression. J. Exp. Clin. Cancer Res. 36, 29 (2017)
P. Zhou, Y. Li, B. Li, M. Zhang, Y. Liu, Y. Yao, D. Li, NMIIA promotes tumor growth and metastasis by activating the Wnt/beta-catenin signaling pathway and EMT in pancreatic cancer. Oncogene 38, 5500–5515 (2019)
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant number 81600490), the Taishan Scholars Program of Shandong Province (grant number 2019010668), the Shandong Higher Education Young Science and Technology Support Program (grant number 2020KJL005), the China Postdoctoral Science Foundation (grant number 2016M602098) and the Qingdao Postdoctoral Science Foundation (grant number 2016046).
Author information
Authors and Affiliations
Contributions
Xue Weijie, Dong Bingzi, Wang Yixiu, Yang Chenyu and Xie Yuwei performed the experiments, analyzed the data and wrote the manuscript. Dong Bingzi revised the manuscript and illustrated all figures. Zhao Yanjie provided experimental guidance. Niu Zhaojian and Zhu Chengzhan designed the study. Zhu Chengzhan provided the funding support, analyzed the data and wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The Ethics Committee of the Affiliated Hospital of Qingdao University approved this study. Written informed consent for participation was obtained from each patient.
Informed consent
Written informed consent for publication was obtained from each patient.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xue, W., Dong, B., Zhao, Y. et al. Upregulation of TTYH3 promotes epithelial-to-mesenchymal transition through Wnt/β-catenin signaling and inhibits apoptosis in cholangiocarcinoma. Cell Oncol. 44, 1351–1361 (2021). https://doi.org/10.1007/s13402-021-00642-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-021-00642-9