Abstract
Background
The epidermal growth factor receptor (EGFR) is a member of the HER family of growth factors that activates several intracellular signaling pathways promoting proliferation and survival. EGFR over-expression is frequently associated with gene mutation or amplification, thereby constituting a major target for molecular therapies. Recently, a new generation of EGFR inhibitors has been developed with pan-HER properties and irreversible actions. Allitinib® (AST1306) is an orally active, highly selective irreversible inhibitor of the HER family of receptor tyrosine kinases with promising efficacies. In the present study we aimed to investigate the cytotoxicity of allitinib in a large panel of human cancer-derived cell lines and to correlate its efficacy to the mutational status of the EGFR, KRAS, BRAF, PI3KCA and PTEN genes. In addition, we aimed to evaluate the functional role of KRAS mutations in the response to this new inhibitor.
Results
In total 76 different cancer-derived cell lines, representing 11 distinct histological types, were analyzed and classified into three groups: highly sensitive (HS), moderately sensitive (MS) and resistant (R). We found that 28 (36.8 %) cancer-derived cell lines exhibited a HS phenotype, 19 (25.0 %) a MS phenotype and 29 (38.1 %) a R phenotype. Allitinib showed a stronger cytotoxicity in head and neck, esophageal, melanoma and lung cancer-derived cell lines. We found that KRAS mutations were significantly associated with the R phenotype. To substantiate these results, an allitinib-sensitive lung cancer-derived cell line (H292) was transfected with plasmids carrying the two most common activating KRAS mutations (p.G12D and p.G12S). We found that both mutations reverted the allitinib-sensitive phenotype in these cells.
Conclusions
The current study represents the largest in vitro assessment of allitinib cytotoxicity performed to date. Through this study, we identified cancer types that could potentially benefit from this drug. Additionally, our findings suggest that prevalent KRAS mutations constitute potential predictive biomarkers for allitinib response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
EGFR is a receptor tyrosine kinase (RTK) belonging to the HER family of growth factor receptors, which encompasses four representative members: EGFR (ErbB1/HER1), HER2 (ErbB2/neu), HER3 (ErbB3) and HER4 (ErbB4) [1]. These RTKs bind distinct ligands, such as EGF (epidermal growth factor), amphiregulin, betacellulin, epiregulin, neuregulin, heparin-binding EGF and transforming growth factor alpha (TGF-α) [1, 2]. Under normal conditions, when RTKs are stimulated by their ligands, receptor dimerization and autophosphorylation take place, leading to downstream activation of intracellular signaling pathways, mainly the RAS/RAF/MEK/ERK and PI3KCA/AKT pathways, promoting among others cellular proliferation and survival [3, 4].
EGFR over-expression occurs in most solid tumors, including colorectal cancer [5], lung cancer [6], head and neck squamous cell carcinoma [7], pancreatic cancer [8], breast cancer [9] and glioma [10, 11]. In tumor cells, anomalous EGFR activation may occur through several mechanisms, including EGFR gene amplifications [12, 13], the occurrence of activating mutations in the extracellular or tyrosine kinase domains [14–16] or by autocrine/paracrine signaling mechanisms [13]. Anomalous EGFR activation may have an impact on tumor cell behavior and, as such, be implicated in metastatic disease and a poor prognosis [17].
Due to its paramount relevance to cancer, several therapeutic strategies targeting the EGFR have been developed during the past decade [18, 19]. The two main anti-EGFR options currently used in clinical practice are small molecule tyrosine kinase inhibitors (TKIs) and monoclonal antibodies [20]. TKIs such as gefitinib (Iressa®) or erlotinib (Tarceva®) are being used in the treatment of patients with non-small cell lung cancer (NSCLC) and metastatic head and neck cancer [21]. Both inhibitors are employed as first-line therapy in EGFR mutated NSCLCs [14]. Second-generation TKIs consist of small molecules with irreversible actions in target tyrosine kinase domains. In addition, some of these molecules, such as afatinib (Gilotrif®, Boehringer Ingelheim Pharmaceuticals), a recently US FDA approved agent for the treatment of patients with NSCLC refractory to gefitinib or erlotinib, and which can target wild-type EGFR, mutant EGFR (L858R)(L858R/T790M) and HER2, have the ability to inhibit various molecular targets [22, 23]. The US FDA approval of afatinib was based on the observed increase in progression-free survival (PFS) in a multi-center, international clinical trial performed with NSCLC patients, which confirmed the activity of afatinib as first-line therapy in EGFR mutated patients [24]. Afatinib has been included in several clinical trials for other solid tumors, such as head and neck squamous cell carcinomas and breast cancers [25].
Allitinib (Shanghai Allist Pharmaceuticals, China), also known as AST1306, is similar to afatinib a potent irreversible EGFR inhibitor [26]. Allitinib is an anilino-quinazoline compound that has been reported to inhibit EGFR and other members of the HER family, such as HER2 and HER4 [26]. A recent phase I clinic trial has assessed the safety, pharmacologic tolerance and anti-tumor effects of allitinib in patients with breast and lung cancer [27].
Several studies have intended to identify potential predictive biomarkers for anti-EGFR responses [28]. The majority of mutations arising in the tyrosine kinase domain of the EGFR are associated with positive responses to erlotinib- and gefitinib-based therapies [29]. However, it has been reported that secondary EGFR mutations, mainly T790M, may elicit acquired resistance to EGFR tyrosine kinase inhibitors in NSCLC [30]. In other tumors, such as metastatic colorectal carcinoma, KRAS mutations serve as main predictive biomarkers of resistance to anti-EGFR monoclonal antibodies, such as cetuximab and panitumumab [31]. Moreover, mutations in the BRAF, NRAS and PIK3CA (exon 20) genes have been associated with low response rates to these therapies [32]. Mechanisms of de novo resistance were also identified in vitro by secondary activation of the interleukin-6 receptor (IL-6R)/JAK1/STAT3 signaling pathway via autocrine IL-6 secretion [33]. Ongoing trials continue to investigate anti-EGFR therapies in various treatment settings but, due to sensitivity and resistance complexities, the underlying mechanisms have remained unclear.
In the current study, we investigated the efficacy of allitinib in a large panel of 76 cancer-derived cell lines, and correlated drug response rates with the mutational statuses of the EGFR, KRAS, BRAF, PI3KCA and PTEN genes. In addition we assessed the role of KRAS mutations (G12D and G12S) as putative predictive biomarkers for the response to allitinib.
2 Methods
2.1 Cell culture and reagents
In total 76 different human cancer-derived cell lines were included, representing 11 solid tumor types, i.e., 7 head and neck, 4 esophageal, 9 melanoma, 12 glioma, 8 colon, 4 pancreatic, 6 breast, 3 prostate, 4 bladder, 15 lung and 4 cervical cancer-derived cell lines. The cell lines were maintained in RPMI-1640 or DMEM (GIBCO) medium supplemented with 10 % fetal bovine serum (FBS, GIBCO) and 1 % penicillin and streptomycin (SIGMA) at 37 °C in a humidified 5 % CO2 atmosphere. Further details on the culture conditions and the origins of the cell lines are provided in Supplementary Table 1 [34–41]. A 10 mM stock of the EGFR-specific tyrosine kinase inhibitor (TKI) allitinib (Selleck Chemical, Houston, TX, USA) was prepared and stored at −20 °C. Next, the drug was diluted in dimethyl sulfoxide (DMSO) at various intermediate concentrations and stored at −20 °C until use.
2.2 Cell viability (MTS) assay
The cytotoxic effects of the EGFR inhibitor allitinib were determined using a Cell Titer 96 Aqueous cell proliferation assay (MTS assay, PROMEGA, Madison, WI, USA). To this end, the cells were plated in 96 well plates (maximum 5x103/well) and allowed to adhere overnight. The seeding density of each cell line varied (Supplementary Table 1 and [42–49]) in order to assure that cells did not reach confluence during the drug treatment and to allow a uniform MTS measurement. A representation of the distinct proliferation and survival kinetics of the cell lines tested is depicted in Supplementary Fig. 1. Subsequently, the cells were treated with increasing concentrations of allitinib in culture medium (serum-free) for 72 hours. In order to assess the cytotoxicity of allitinib, the treated cells were incubated with MTS reagent after which the absorbance was measured using an automatic microplate reader (Varioskan, Thermo) at 490 nm. The results were expressed as percentages relative to control (DMSO treated) cells. The IC50 concentrations were calculated through nonlinear regression analyses using GraphPad Prism software version 5. To uniformly classify the cell lines, growth inhibition (GI) values were determined. Mean GI values were established at a fixed dose of 1000 nM (100 % - percentage of viable cells at this dose), and the cell lines were scored as highly sensitive (HS) if GI > 60 %, moderately sensitive (MS) if GI 40–60 % and resistant (R) if GI < 40 %, as previously described using the formula: living cells (%) at 1000 nM - 100 (%) [50]. All the assays were performed in triplicate and repeated at least three times.
2.3 Gene mutation analyses
Mutation analyses of the EGFR hotspots (exons 18, 19, 20 and 21), the KRAS codons 12/13 and 61 and the NRAS codons 12/13 and 61, were performed by direct targeted sequencing as previously described [9, 51, 52]. Briefly, PCR was carried out in a final volume of 15 μl containing 50 ng DNA, 10 μM forward and reverse primers and 7,5 μl HotStar master mix (Qiagen, Hilden, Germany) following the protocol proposed by the manufacturer. The cycling parameters used were: denaturation at 96 °C for 15 minutes, followed by 40 cycles of 96 °C for 45 seconds and 58 °C for 45 seconds for EGFR, 55.5 °C for 45 seconds for KRAS, 72 °C for 45 seconds for NRAS and, in all cases, a final extension at 72 °C for 10 minutes using a Veriti® 96-Well Thermal Cycler (Applied Biosystems, Carlsbad, USA). The resulting PCR products were purified using EXOSAP-IT (Affymetrix, USB) and subjected to direct sequencing using an ABI PRISM BigDye XTerminator in conjunction with a BigDye XTerminator purification kit (Applied Biosystems). The analyses were performed using the Genetic Analyzer ABI PRISM 3500 and SeqScape version 2.7 software packages (Applied Biosystems). To retrieve additional mutation profiles we used information from the Roche Cancer Genome Database (Mutome, DB) [53] for the BRAF, PIK3CA and PTEN genes.
2.4 Authentication of the cell lines
Authentication of all cell lines was performed by short tandem repeat (STR) DNA typing according to the International Reference Standard for Authentication of Human Cell Lines using a panel of 8 STR loci (D5S818, D13S317, D7S820, D16S539, vWA, TH01, TPOX and CSF1P0) plus gender determination (AMEL) using fluorescent labeling primers as reported by Dirks et al. [54]. Briefly, 50 ng DNA was amplified by a multiplex PCR reaction in a total volume of 10 μl using a Qiagen multiplex kit (Qiagen) comprising 0.5 μM of all fluorescent primer pairs plus 1 μM of TH01 primer reinforcement, using a Veriti® 96-Well Thermal Cycler. The cycling parameters used were: denaturation at 95 °C for 15 min, followed by 30 cycles of 95 °C for 30 seconds, 55 °C for 1 minute and 30 seconds, 72 °C for 1 second and a final extension at 72 °C for 30 minutes. The resulting PCR products were diluted 1:100 in nuclease-free water ultrapure (USB, Cleveland, USA) supplemented with 0.3 μl internal size standard 500ROX (Applied Biosystems, Foster City, CA, USA) in 8.7 μl formamide and loaded automatically for capillary electrophoresis using a Genetic Analyzer ABI PRISM 3500 (Applied Biosystems). The analyses were performed using GeneMapper software version 4.1 (Applied Biosystems). The genotyping confirmed the identity of all cell lines, with the exception of U373 and SNB19, which were found to be sub-clones of the U251 cell line.
2.5 Generation of stable KRAS mutant cell lines
The lung cancer-derived cell line H292 was used to generate KRAS (p.G12D and p.G12S) mutant cell lines through transfection of EGFP-KRAS fusion constructs. Briefly, plasmids pEGFP.C1.KRAS_wt, pEGFP.C1.KRAS_G12D (GAT) and pEGFP.C1.KRAS_G12S (AGT) (Reniguard Life Sciences Inc.) were transfected in sub-confluent 6-well plates (2.5x105 cells) using Fugene HD (PROMEGA, Madison, WI, USA) according to the manufacturer‘s recommendations. Twenty-four hours after transfection, positive cell lines were selected in culture medium containing neomycin (250 μg/ml) during 2 weeks. After this selection, transfected cells (EGFP-positive) were enriched by flow cytometry using BD FACSAria II (BD Biosciences, Bedford, MA, USA). Next, the cells were expanded in culture flasks. KRAS mutations were confirmed by direct sequencing using the primers hCMV_F: AGCAGAGCTGGTTTAGTGAAC and KRAS_R: CCAAGAGACAGGTTTCTCCATCA. This approach was chosen in order to directly compare the effect of allitinib treatment on wild-type KRAS and p.G12D and p.G12S mutant KRAS cells. For this reason, and as previously reported [55], empty vector controls were not included in these assays.
2.6 Viability and cytotoxicity analyses of KRAS mutant cell lines
To evaluate the effect of KRAS mutations on the sensitivity of H292 cells to allitinib, MTS assays were performed on the wild-type and p.G12D and p.G12S mutant cell lines (see above). To this end, the respective cell lines were seeded in 96-well plates (maximum 5x103/well) and allowed to adhere overnight. Subsequently, the cells were treated with increasing concentrations of allitinib diluted in DMEM (serum-free) for 72 hours to allow IC50 determination. Finally, the viability and cytotoxicity changes were assessed using an ApoTox-Glo assay (PROMEGA, Madison, WI, USA) at fixed doses of allitinib (1 μM and 6 μM) after 72 hours.
2.7 Statistical analyses
Statistical analyses were performed using SPSS software (Statistical Package for Social Sciences; SPSS, Chicago, Ill., USA), version 19. Associations between response rates to allitinib and mutation status of the cell lines were performed using a χ2 test. When the χ2 test assumptions were not met Fisher’s exact test was used. Single comparisons between mutant KRAS H292 cell lines were performed using Student’s t test, and differences between groups were tested using two-way analysis of variance (ANOVA). The analyses were performed using GraphPad Prism version 5 and significance levels were set at p < 0.05.
3 Results and discussion
In order to evaluate the therapeutic efficacy of the tyrosine kinase inhibitor (TKI) allitinib on different solid tumors, we determined its cytotoxicity by MTS assay after exposing a panel of 76 human cancer-derived cell lines to increasing concentrations of this drug (maximum dose 1000 nM) (Table 1). By doing so, we observed IC50 values in the nanomolar (nM) range, varying from 24.01 ± 2.41 nM to 980.21 ± 0.16 nM (Table 1). In 32 cell lines the IC50 values could not be calculated since they did not reach a 50 % cell viability reduction at the maximum dose applied (IC50 > 1000 nM). We observed a wide variation between the IC50 values for each individual tumor type, hampering clear-cut comparisons among them. In order to uniformly classify the cell lines according to their responses to allitinib, we determined growth inhibition (GI) scores, as depicted in Table 1. According to this classification we found that 38.1 % (29/76) of the cell lines was resistant, 25.0 % (19/76) was moderately sensitive and 36.8 % (28/76) was highly sensitive (Table 2 and Fig. 1). The tumor types that showed high percentages of highly sensitive cell lines were esophageal cancer (100 %), head and neck cancer (57.1 %), melanoma (55.6 %), bladder cancer (50 %) and lung cancer (46.6 %). When considering resistant versus sensitive (highly + moderate) cell lines, we found that the lung, breast, melanoma and glioma-derived cell lines were the most sensitive ones, with lower percentages of cell lines considered as being resistant. In contrast, we found that pancreatic and prostate cancer-derived cell lines showed the least effective responses to the drug (Table 2 and Fig. 1). A representation of the distinct proliferation and survival curves of the cell lines is depicted in Supplementary Fig. 1.
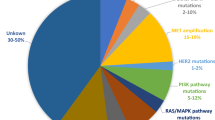
To identify potential biomarkers for allitinib response, we assessed the mutation status of 3 major genes involved in the primary responses to anti-EGFR therapy in solid tumors, i.e., EGFR, KRAS and NRAS (Table 1). We found that only 4 of the cell lines tested (HN13, H1975, PC9 and NCI-H827) harbored mutations in the EGFR gene (H773Y, L858R + T790M and del19). Targeted KRAS sequencing revealed 20 mutations in codon 12/13, across distinct tumor types, with high frequencies in colorectal (6/8 cases), lung (5/10 cases) and pancreatic (3/4 cases) cancers (Table 1). We also identified 2 melanoma cell lines (Colo858 and WM852) with NRAS mutations. The mutation statuses of other important genes (i.e., BRAF, PTEN and PIK3CA), as deduced from literature data, are summarized in Table 1.
Our statistical analysis revealed that the mutation status of the KRAS gene was significantly (p = 0.001) associated with allitinib resistance (Table 3), i.e., the majority of the KRAS mutant cell lines was classified as allitinib resistant (70 %), whereas 70.9 % (39/76) of the KRAS wild-type cell lines was classified as allitinib sensitive (p = 0.001; Table 3). No additional significant associations were found for the other remaining genes (EGFR, NRAS, BRAF, PTEN and PIK3CA) (Table 3).
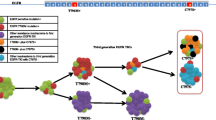
In order to substantiate the abovementioned association, we stably transfected a KRAS wild-type cell line, H292, with two plasmids carrying the most prevalent KRAS mutations (p.G12D and p.G12S) and a control wild-type plasmid. The mutation status of the resulting cell lines was confirmed by sequence analysis (Fig. 2a). As expected, we observed IC50 values of 0.95 ± 0.17 μM for H292 wild-type (WT), a six times increase of IC50 (6.56 ± 0.23 μM) in the H292-KRAS-G12D (GAT) and an eight times increase of IC50 (8.47 ± 0.15 μM) in the H292-KRAS-G12S (AGT) cells (Fig. 2b). Using a fluorescent assay we found that after 72 hours H292-WT cells showed a significant decrease in viability at both allitinib concentrations used (1 μM and 6 μM) compared to the H292-KRAS-G12D and H292-KRAS-G12S cells (p = 0.03 and p = 0.003) (Fig. 2c). In addition, we observed by cytotoxicity fluorescent analyses inverse proportional relations with the viability profiles, i.e., both mutant KRAS cell lines exhibited a significantly lower cytotoxicity compared to the H292-WT cells when exposed to 1 μM and 6 μM allitinib (p = 0.005 and p = 0.006, respectively) (Fig. 2d).
Viability and cytotoxicity analyses of KRAS transfected cell lines (H292-KRAS-wt; H292-KRAS-G12D and H292-KRAS-G12S). (a). Electropherogram of KRAS mutant H292 cell lines; (b). Sensitive response to incremental allitinib (AST) concentrations (MTS assay); Viability (c) and cytotoxicity (d) changes in KRAS mutant H292 cell lines exposed to 1 μM and 6 μM allitinib for 72 hours (ApoTox-Glo assay)
Initial preclinical trials have shown that allitinib preferentially inhibits EGFR and HER2, including the erlotinib/gefitinib resistant EGFR T790M mutant form. The IC50 values obtained in these trials were equivalent to those of the recently US FDA approved cancer drug afatinib [26, 56]. Yet, these promising results were obtained in a limited number of cancer-derived cell lines [26]. Here, we extended these initial studies to 76 cancer-derived cell lines corresponding to 11 distinct tumor types. We found that 29 of the cancer-derived cell lines exhibited a high rate of resistance to allitinib, independent of the tumor type. These results thus indicate that the allitinib response is not tumor type dependent, but rather driven by (anomalous) cellular signaling and genetic mutation profiles. Notably, we observed that the KRAS mutation status was significantly associated with the allitinib response. This observation was substantiated through stable transfection of the KRAS and EGFR wild-type lung cancer-derived cell line H292 with plasmids carrying two common KRAS mutations (p.G12D and p.G12S), i.e., we found a significant increased resistance to allitinib in both mutant KRAS cell lines.
Interestingly, KRAS mutations are currently the better surrogate markers for resistance to anti-EGFR monoclonal antibody therapies, i.e., cetuximab and panitumumab [28, 57]. KRAS mutation leads to a constitutive active form of the protein resulting in constant stimulation of downstream signaling pathways, such as the MAPK and AKT pathways, which contribute to cellular proliferation, survival, neoplastic transformation, migration and metastasis [58]. The role of mutant KRAS in a cell’s response to tyrosine kinase inhibitors, such as gefitinib and erlotinib, is less clear, and its predictive value has been questioned [59]. For these drugs, the most important biomarkers for response are mutations in the tyrosine kinase domain of EGFR, specifically in exons 18, 19, 20 and 21, which frequently occur in primary lung adenocarcinomas [60]. Among the cell lines studied, we found that four lines harbored EGFR mutations. The HN13 head and neck cancer-derived cell line was found to carry a p.H773Y mutation. This mutation is not known as a hotspot mutation associated with the response of lung cancer cells to gefitinib/erlotinib. We found that the cell line showed resistance to allitinib. The lung cancer-derived cell lines PC9 and NCI-H827 both harbor a hotspot in-frame deletion of exon 19 and, accordingly, they showed a sensitive response to allitinib. In Asian and Northern American populations deletions in exon 19 account for 45-50 % of the patients with non-small cell lung cancer (NSCLC), and they have been found to be associated with clinical responsiveness to gefitinib [61]. A recent clinical trial (LUX-Lung 3,6) revealed an advantage of afatinib over chemotherapy in NSCLC patients with EGFR exon 19 deletions [62]. Finally, we found that the lung cancer-derived cell line NCI-H1975 carries a double p.L858R/p.T790M EGFR mutation. The p.T790M EGFR mutation is a well-know acquired EGFR mutation associated with erlotinib/gefitinib resistance [63]. We found that allitinib exhibited a high efficacy in this cell line. Similar findings were reported by Xie et al. [26] using NIH3T3 cells engineered to express the double p.L858R/p.T790M EGFR mutation [26]. These authors also suggested that cell lines exhibiting higher HER2 levels were more sensitive to allitinib.
Allitinib has been entered in a clinical phase I trial for solid tumors and the results of this trial have recently been published [27]. A partial response of patients with breast, lung and gastric cancer was observed, raising high expectations [27]. Based on the present work we believe that other cancers, such as head and neck cancer, melanoma and esophageal cancer, may serve as potential candidates for new allitinib-based clinical trials [64, 65].
In conclusion, the current study represents the largest in vitro assessment of allitinib cytotoxicity to date. We identified tumor types that could potentially benefit from this drug and, importantly, suggest that KRAS mutations constitute a potential predictive biomarker for allitinib response, thereby constituting a step forward in the selection of patients that will benefit most from allitinib therapy.
References
Y. Yarden, G. Pines, The ERBB network: at last, cancer therapy meets systems biology. Nat. Rev. Cancer 12, 553–563 (2012)
S.L. Jeppe Knudsen, A.S. Wai Mac, L. Henriksen, B. Deurs, L.M. Grovdal, EGFR signaling patterns are regulated by its different ligands. Growth Factors 32, 155–163 (2014)
G. Tarcic, Y. Yarden, MAP Kinase activation by receptor tyrosine kinases: in control of cell migration. Methods Mol. Biol. 661, 125–135 (2010)
S.B. Prasad, S.S. Yadav, M. Das, A. Modi, S. Kumari, L.K. Pandey, S. Singh, S. Pradhan, G. Narayan, PI3K/AKT pathway-mediated regulation of p27(Kip1) is associated with cell cycle arrest and apoptosis in cervical cancer. Cell. Oncol. 38, 215–225 (2015)
F. Ciardiello, N. Kim, T. Saeki, R. Dono, M.G. Persico, G.D. Plowman, J. Garrigues, S. Radke, G.J. Todaro, D.S. Salomon, Differential expression of epidermal growth factor-related proteins in human colorectal tumors. Proc. Natl. Acad. Sci. U. S. A. 88, 7792–7796 (1991)
F.R. Hirsch, M. Varella-Garcia, P.A. Bunn Jr., M.V. Di Maria, R. Veve, R.M. Bremmes, A.E. Baron, C. Zeng, W.A. Franklin, Epidermal growth factor receptor in non-small-cell lung carcinomas: correlation between gene copy number and protein expression and impact on prognosis. J. Clin. Oncol. 21, 3798–3807 (2003)
L. Sweeny, N.R. Dean, J.S. Magnuson, W.R. Carroll, E.E. Helman, S.O. Hyde, R.L. Desmond, E.L. Rosenthal, EGFR expression in advanced head and neck cutaneous squamous cell carcinoma. Head Neck 34, 681–686 (2012)
M.L. Fjallskog, M.H. Lejonklou, K.E. Oberg, B.K. Eriksson, E.T. Janson, Expression of molecular targets for tyrosine kinase receptor antagonists in malignant endocrine pancreatic tumors. Clin. Cancer Res. 9, 1469–1473 (2003)
J.S. Reis-Filho, C. Pinheiro, M.B. Lambros, F. Milanezi, S. Carvalho, K. Savage, P.T. Simpson, C. Jones, S. Swift, A. Mackay, R.M. Reis, J.L. Hornick, E.M. Pereira, F. Baltazar, C.D. Fletcher, A. Ashworth, S.R. Lakhani, F.C. Schmitt, EGFR amplification and lack of activating mutations in metaplastic breast carcinomas. J. Pathol. 209, 445–453 (2006)
D.A. Bax, N. Gaspar, S.E. Little, L. Marshall, L. Perryman, M. Regairaz, M. Viana-Pereira, R. Vuononvirta, S.Y. Sharp, J.S. Reis-Filho, J.N. Stavale, S. Al-Sarraj, R.M. Reis, G. Vassal, A.D. Pearson, D. Hargrave, D.W. Ellison, P. Workman, C. Jones, EGFRvIII deletion mutations in pediatric high-grade glioma and response to targeted therapy in pediatric glioma cell lines. Clin. Cancer Res. 15, 5753–5761 (2009)
M. Viana-Pereira, J.M. Lopes, S. Little, F. Milanezi, D. Basto, F. Pardal, C. Jones, R.M. Reis, Analysis of EGFR overexpression, EGFR gene amplification and the EGFRvIII mutation in Portuguese high-grade gliomas. Anticancer Res. 28, 913–920 (2008)
K. Iida, K. Nakayama, M.T. Rahman, M. Rahman, M. Ishikawa, A. Katagiri, S. Yeasmin, Y. Otsuki, H. Kobayashi, S. Nakayama, K. Miyazaki, EGFR gene amplification is related to adverse clinical outcomes in cervical squamous cell carcinoma, making the EGFR pathway a novel therapeutic target. Br. J. Cancer 105, 420–427 (2011)
R. Zandi, A.B. Larsen, P. Andersen, M.T. Stockhausen, H.S. Poulsen, Mechanisms for oncogenic activation of the epidermal growth factor receptor. Cell. Signal. 19, 2013–2023 (2007)
K. Kobayashi, K. Hagiwara, Epidermal growth factor receptor (EGFR) mutation and personalized therapy in advanced nonsmall cell lung cancer (NSCLC). Target. Oncol. 8, 27–33 (2013)
H.K. Gan, A.H. Kaye, R.B. Luwor, The EGFRvIII variant in glioblastoma multiforme. J. Clin. Neurosci. 16, 748–754 (2009)
B.S. Paugh, X. Zhu, C. Qu, R. Endersby, A.K. Diaz, J. Zhang, D.A. Bax, D. Carvalho, R.M. Reis, A. Onar-Thomas, A. Broniscer, C. Wetmore, J. Zhang, C. Jones, D.W. Ellison, S.J. Baker, Novel oncogenic PDGFRA mutations in pediatric high-grade gliomas. Cancer Res. 73, 6219–6229 (2013)
K. Laimer, G. Spizzo, G. Gastl, P. Obrist, T. Brunhuber, D. Fong, V. Barbieri, S. Jank, W. Doppler, M. Rasse, B. Norer, High EGFR expression predicts poor prognosis in patients with squamous cell carcinoma of the oral cavity and oropharynx: a TMA-based immunohistochemical analysis. Oral Oncol. 43, 193–198 (2007)
A.W. Burgess, Y.I. Henis, N.E. Hynes, T. Jovin, A. Levitzki, R. Pinkas-Kramarski, Y. Yarden, EGF receptor family: twisting targets for improved cancer therapies. Growth Factors 32, 74–81 (2014)
S.H. Ou, Second-generation irreversible epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (TKIs): a better mousetrap? a review of the clinical evidence. Crit. Rev. Oncol. Hematol. 83, 407–421 (2012)
E. Raymond, S. Faivre, J.P. Armand, Epidermal growth factor receptor tyrosine kinase as a target for anticancer therapy. Drugs 60 (Suppl 1), 15–23 (2000). discussion 41–2
A. Argiris, M. Ghebremichael, J. Gilbert, J.W. Lee, K. Sachidanandam, J.M. Kolesar, B. Burtness, A.A. Forastiere, Phase III randomized, placebo-controlled trial of docetaxel with or without gefitinib in recurrent or metastatic head and neck cancer: an eastern cooperative oncology group trial. J. Clin. Oncol. 31, 1405–1414 (2013)
R.T. Dungo, G.M. Keating, Afatinib: first global approval. Drugs 73, 1503–1515 (2013)
S.M. Kim, M.R. Yun, Y.K. Hong, F. Solca, J.H. Kim, H.J. Kim, B.C. Cho, Glycolysis inhibition sensitizes non-small cell lung cancer with T790M mutation to irreversible EGFR inhibitors via translational suppression of Mcl-1 by AMPK activation. Mol. Cancer Ther. 12, 2145–2156 (2013)
A. Passaro, B. Gori, F. de Marinis, Afatinib as first-line treatment for patients with advanced non-small-cell lung cancer harboring EGFR mutations: focus on LUX-Lung 3 and LUX-Lung 6 phase III trials. J. Thorac. Dis. 5, 383–384 (2013)
N.U. Lin, E.P. Winer, D. Wheatley, L.A. Carey, S. Houston, D. Mendelson, P. Munster, L. Frakes, S. Kelly, A.A. Garcia, S. Cleator, M. Uttenreuther-Fischer, H. Jones, S. Wind, R. Vinisko, T. Hickish, A phase II study of afatinib (BIBW 2992), an irreversible ErbB family blocker, in patients with HER2-positive metastatic breast cancer progressing after trastuzumab. Breast Cancer Res. Treat. 133, 1057–1065 (2012)
H. Xie, L. Lin, L. Tong, Y. Jiang, M. Zheng, Z. Chen, X. Jiang, X. Zhang, X. Ren, W. Qu, Y. Yang, H. Wan, Y. Chen, J. Zuo, H. Jiang, M. Geng, J. Ding, AST1306, a novel irreversible inhibitor of the epidermal growth factor receptor 1 and 2, exhibits antitumor activity both in vitro and in vivo. PLoS One 6, e21487 (2011)
J. Zhang, J. Cao, J. Li, Y. Zhang, Z. Chen, W. Peng, S. Sun, N. Zhao, J. Wang, D. Zhong, X. Zhang, J. Zhang, A phase I study of AST1306, a novel irreversible EGFR and HER2 kinase inhibitor, in patients with advanced solid tumors. J. Hematol. Oncol. 7, 22 (2014)
S. Siena, A. Sartore-Bianchi, F. Di Nicolantonio, J. Balfour, A. Bardelli, Biomarkers predicting clinical outcome of epidermal growth factor receptor-targeted therapy in metastatic colorectal cancer. J. Natl. Cancer Inst. 101, 1308–1324 (2009)
M.S. Tsao, A. Sakurada, K. Ding, S. Aviel-Ronen, O. Ludkovski, N. Liu, A. Le Maitre, D. Gandara, D.H. Johnson, J.R. Rigas, L. Seymour, F.A. Shepherd, Prognostic and predictive value of epidermal growth factor receptor tyrosine kinase domain mutation status and gene copy number for adjuvant chemotherapy in non-small cell lung cancer. J. Thorac. Oncol. 6, 139–147 (2011)
K. Takezawa, I. Okamoto, J. Tanizaki, K. Kuwata, H. Yamaguchi, M. Fukuoka, K. Nishio, K. Nakagawa, Enhanced anticancer effect of the combination of BIBW2992 and thymidylate synthase-targeted agents in non-small cell lung cancer with the T790M mutation of epidermal growth factor receptor. Mol. Cancer Ther. 9, 1647–1656 (2010)
W. De Roock, B. Claes, D. Bernasconi, J. De Schutter, B. Biesmans, G. Fountzilas, K.T. Kalogeras, V. Kotoula, D. Papamichael, P. Laurent-Puig, F. Penault-Llorca, P. Rougier, B. Vincenzi, D. Santini, G. Tonini, F. Cappuzzo, M. Frattini, F. Molinari, P. Saletti, S. De Dosso, M. Martini, A. Bardelli, S. Siena, A. Sartore-Bianchi, J. Tabernero, T. Macarulla, F. Di Fiore, A.O. Gangloff, F. Ciardiello, P. Pfeiffer, C. Qvortrup, T.P. Hansen, E. Van Cutsem, H. Piessevaux, D. Lambrechts, M. Delorenzi, S. Tejpar, Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 11, 753–762 (2010)
Z.X. Yuan, X.Y. Wang, Q.Y. Qin, D.F. Chen, Q.H. Zhong, L. Wang, J.P. Wang, The prognostic role of BRAF mutation in metastatic colorectal cancer receiving anti-EGFR monoclonal antibodies: a meta-analysis. PLoS One 8, e65995 (2013)
S.M. Kim, O.J. Kwon, Y.K. Hong, J.H. Kim, F. Solca, S.J. Ha, R.A. Soo, J.G. Christensen, J.H. Lee, B.C. Cho, Activation of IL-6R/JAK1/STAT3 signaling induces de novo resistance to irreversible EGFR inhibitors in non-small cell lung cancer with T790M resistance mutation. Mol. Cancer Ther. 11, 2254–2264 (2012)
L.B. Cardeal, E. Boccardo, L. Termini, T. Rabachini, M.A. Andreoli, C. di Loreto, A. Longatto Filho, L.L. Villa, S.S. Maria-Engler, HPV16 oncoproteins induce MMPs/RECK-TIMP-2 imbalance in primary keratinocytes: possible implications in cervical carcinogenesis. PLoS One 7, e33585 (2012)
V. Kannen, H. Hintzsche, D.L. Zanette, W.A. Silva Jr., S.B. Garcia, A.M. Waaga-Gasser, H. Stopper, Antiproliferative effects of fluoxetine on colon cancer cells and in a colonic carcinogen mouse model. PLoS One 7, e50043 (2012)
B.M. Costa, J.S. Smith, Y. Chen, J. Chen, H.S. Phillips, K.D. Aldape, G. Zardo, J. Nigro, C.D. James, J. Fridlyand, R.M. Reis, J.F. Costello, Reversing HOXA9 oncogene activation by PI3K inhibition: epigenetic mechanism and prognostic significance in human glioblastoma. Cancer Res. 70, 453–462 (2010)
D.A. Bax, S.E. Little, N. Gaspar, L. Perryman, L. Marshall, M. Viana-Pereira, T.A. Jones, R.D. Williams, A. Grigoriadis, G. Vassal, P. Workman, D. Sheer, R.M. Reis, A.D. Pearson, D. Hargrave, C. Jones, Molecular and phenotypic characterisation of paediatric glioma cell lines as models for preclinical drug development. PLoS One 4, e5209 (2009)
M. Cardinali, H. Pietraszkiewicz, J.F. Ensley, K.C. Robbins, Tyrosine phosphorylation as a marker for aberrantly regulated growth-promoting pathways in cell lines derived from head and neck malignancies. Int. J. Cancer 61, 98–103 (1995)
M.F. Calmon, R.V. Rodrigues, C.M. Kaneto, R.P. Moura, S.D. Silva, L.D. Mota, D.G. Pinheiro, C. Torres, A.F. de Carvalho, P.M. Cury, F.D. Nunes, I.N. Nishimoto, F.A. Soares, A.M. da Silva, L.P. Kowalski, H. Brentani, C.F. Zanelli, W.A. Silva Jr., P. Rahal, E.H. Tajara, D.M. Carraro, A.A. Camargo, S.R. Valentini, Epigenetic silencing of CRABP2 and MX1 in head and neck tumors. Neoplasia 11, 1329–1339 (2009)
P.T. Hennessey, M.F. Ochs, W.W. Mydlarz, W. Hsueh, L. Cope, W. Yu, J.A. Califano, Promoter methylation in head and neck squamous cell carcinoma cell lines is significantly different than methylation in primary tumors and xenografts. PLoS One 6, e20584 (2011)
J.F. Sousa, R. Torrieri, R.R. Silva, C.G. Pereira, V. Valente, E. Torrieri, K.C. Peronni, W. Martins, N. Muto, G. Francisco, C.A. Brohem, C.G. Carlotti Jr., S.S. Maria-Engler, R. Chammas, E.M. Espreafico, Novel primate-specific genes, RMEL 1, 2 and 3, with highly restricted expression in melanoma, assessed by new data mining tool. PLoS One 5, e13510 (2010)
J. Afonso, L.L. Santos, V. Miranda-Goncalves, A. Morais, T. Amaro, A. Longatto-Filho, F. Baltazar, CD147 and MCT1-potential partners in bladder cancer aggressiveness and cisplatin resistance. Mol. Carcinog. 54, 1451–1466 (2015)
F. Morais-Santos, V. Miranda-Goncalves, S. Pinheiro, A.F. Vieira, J. Paredes, F.C. Schmitt, F. Baltazar, C. Pinheiro, Differential sensitivities to lactate transport inhibitors of breast cancer cell lines. Endocr. Relat. Cancer. 21, 27–38 (2014)
O. Martinho, F. Pinto, S. Granja, V. Miranda-Goncalves, M.A. Moreira, L.F. Ribeiro, C. di Loreto, M.R. Rosner, A. Longatto-Filho, R.M. Reis, RKIP inhibition in cervical cancer is associated with higher tumor aggressive behavior and resistance to cisplatin therapy. PLoS One 8, e59104 (2013)
V. Silva, M. Rosa, A. Tansini, J. Lima, C. Jones, L. Pianowski, R. Reis, Cytotoxic activity of semi-synthetic ingenol derived from Euphorbia tirucalli on a large panel of human cancer cell lines. J. Clin. Oncol. 31, (2013)
R.M. Reis, V.A.O. Silva, M.N. Rosa, A. Tansini, J.P.D.S.N. Lima, P.L.F.C. Jones, Cytotoxic effect of euphol from Euphorbia tirucalli on a large panel of human. cancer cell lines. J. Clin. Oncol. 31, abstract (2013)
O. Martinho, R. Silva-Oliveira, V. Miranda-Goncalves, C. Clara, J.R. Almeida, A.L. Carvalho, J.T. Barata, R.M. Reis, In vitro and in vivo analysis of RTK inhibitor efficacy and identification of its novel targets in glioblastomas. Transl. Oncol. 6, 187–196 (2013)
R.H. Shoemaker, The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 6, 813–823 (2006)
F. Pinto, N. Pertega-Gomes, M.S. Pereira, J.R. Vizcaino, P. Monteiro, R.M. Henrique, F. Baltazar, R.P. Andrade, R.M. Reis, T-box transcription factor brachyury is associated with prostate cancer progression and aggressiveness. Clin. Cancer Res. 20, 4949–4961 (2014)
G.E. Konecny, R. Glas, J. Dering, K. Manivong, J. Qi, R.S. Finn, G.R. Yang, K.L. Hong, C. Ginther, B. Winterhoff, G. Gao, J. Brugge, D.J. Slamon, Activity of the multikinase inhibitor dasatinib against ovarian cancer cells. Br. J. Cancer 101, 1699–1708 (2009)
L.S. Yamane, C. Scapulatempo-Neto, L. Alvarenga, C.Z. Oliveira, G.N. Berardinelli, E. Almodova, T.R. Cunha, G. Fava, W. Colaiacovo, A. Melani, J.H. Fregnani, R.M. Reis, D.P. Guimaraes, KRAS and BRAF mutations and MSI status in precursor lesions of colorectal cancer detected by colonoscopy. Oncol. Rep. 32, 1419–1426 (2014)
O. Martinho, A. Gouveia, M. Viana-Pereira, P. Silva, A. Pimenta, R.M. Reis, J.M. Lopes, Low frequency of MAP kinase pathway alterations in KIT and PDGFRA wild-type GISTs. Histopathology 55, 53–62 (2009)
The Roche Cancer Genome Database, 2014 [cited 2014; Available from: http://rcgdb.bioinf.uni-sb.de/MutomeWeb/(2014)
W.G. Dirks, S. Faehnrich, I.A. Estella, H.G. Drexler, Short tandem repeat DNA typing provides an international reference standard for authentication of human cell lines. ALTEX 22, 103–109 (2005)
M. Monticone, E. Biollo, M. Maffei, A. Donadini, F. Romeo, C.T. Storlazzi, W. Giaretti, P. Castagnola, Gene expression deregulation by KRAS G12D and G12V in a BRAF V600E context. Mol. Cancer 7, 92 (2008)
D. Li, L. Ambrogio, T. Shimamura, S. Kubo, M. Takahashi, L.R. Chirieac, R.F. Padera, G.I. Shapiro, A. Baum, F. Himmelsbach, W.J. Rettig, M. Meyerson, F. Solca, H. Greulich, K.K. Wong, BIBW2992, an irreversible EGFR/HER2 inhibitor highly effective in preclinical lung cancer models. Oncogene 27, 4702–4711 (2008)
A. Lievre, J.B. Bachet, D. Le Corre, V. Boige, B. Landi, J.F. Emile, J.F. Cote, G. Tomasic, C. Penna, M. Ducreux, P. Rougier, F. Penault-Llorca, P. Laurent-Puig, KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res. 66, 3992–3995 (2006)
Y. Pylayeva-Gupta, E. Grabocka, D. Bar-Sagi, RAS oncogenes: weaving a tumorigenic web. Nat. Rev. Cancer 11, 761–774 (2011)
P.J. Roberts, T.E. Stinchcombe, KRAS mutation: should we test for it, and does it matter? J. Clin. Oncol. 31, 1112–1121 (2013)
A.F. Gazdar, Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 28(Suppl 1), S24–31 (2009)
T.J. Lynch, D.W. Bell, R. Sordella, S. Gurubhagavatula, R.A. Okimoto, B.W. Brannigan, P.L. Harris, S.M. Haserlat, J.G. Supko, F.G. Haluska, D.N. Louis, D.C. Christiani, J. Settleman, D.A. Haber, Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 350, 2129–2139 (2004)
J.C. Yang, Y.L. Wu, M. Schuler, M. Sebastian, S. Popat, N. Yamamoto, C. Zhou, C.P. Hu, K. O'Byrne, J. Feng, S. Lu, Y. Huang, S.L. Geater, K.Y. Lee, C.M. Tsai, V. Gorbunova, V. Hirsh, J. Bennouna, S. Orlov, T. Mok, M. Boyer, W.C. Su, K.H. Lee, T. Kato, D. Massey, M. Shahidi, V. Zazulina, L.V. Sequist, Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): analysis of overall survival data from two randomised, phase 3 trials. Lancet Oncol. 16, 141–151 (2015)
N. Godin-Heymann, I. Bryant, M.N. Rivera, L. Ulkus, D.W. Bell, D.J. Riese 2nd, J. Settleman, D.A. Haber, Oncogenic activity of epidermal growth factor receptor kinase mutant alleles is enhanced by the T790M drug resistance mutation. Cancer Res. 67, 7319–7326 (2007)
S. Derks, B. Diosdado, Personalized cancer medicine: next steps in the genomic era. Cell Oncol. 38, 1–2 (2015)
L. Prudkin, P. Nuciforo, Obstacles to precision oncology: confronting current factors affecting the successful introduction of biomarkers to the clinic. Cell Oncol. 38, 39–48 (2015)
Acknowledgments
This study was partially supported by FINEP (MCTI/FINEP/MS/SCTIE/DECIT-01/2013 - FPXII-BIOPLAT) and the Assistance Program and Incentive Research (PAIP), Barretos Cancer Hospital São Paulo, Brazil. A.L.C and R.M.R are recipients of a National Counsel of Technological and Scientific Development (CNPq) scholarship. M.N.R is recipient of a CNPq scholarship (380434/2015-6) and O.C.M is recipient of a Portuguese Foundation for Science and Technology (FCT) scholarship (SFRH/BPD/108351/2015).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
Proliferation curves of bladder, breast, colon, esophageal, head and neck, glioma, melanoma, lung and pancreatic cancer-derived cell lines, treated with increasing concentrations of allitinib for 72 hours. (GIF 22 kb)
Supplementary Table 1
Cancer-derived cell lines, origins, culture conditions and seeding densities. (DOC 146 kb)
Rights and permissions
About this article
Cite this article
Silva-Oliveira, R.J., Silva, V.A.O., Martinho, O. et al. Cytotoxicity of allitinib, an irreversible anti-EGFR agent, in a large panel of human cancer-derived cell lines: KRAS mutation status as a predictive biomarker. Cell Oncol. 39, 253–263 (2016). https://doi.org/10.1007/s13402-016-0270-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-016-0270-z