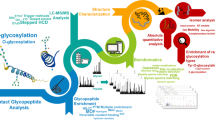
Abstract
Glycomics is the comprehensive study of glycan expression in an organism, cell, or tissue that relies on effective analytical technologies to understand glycan structure–function relationships. Owing to the macro- and micro-heterogeneity of oligosaccharides, detailed structure characterization has required an orthogonal approach, such as a combination of specific exoglycosidase digestions, LC-MS/MS, and the development of bioinformatic resources to comprehensively profile a complex biological sample. Liquid chromatography-electrospray ionization-mass spectrometry (LC-ESI-MS/MS) has emerged as a key tool in the structural analysis of oligosaccharides because of its high sensitivity, resolution, and robustness. Here, we present a strategy that uses LC-ESI-MS/MS to characterize over 200 N- and O-glycans from human saliva glycoproteins, complemented by sequential exoglycosidase treatment, to further verify the annotated glycan structures. Fragment-specific substructure diagnostic ions were collated from an extensive screen of the literature available on the detailed structural characterization of oligosaccharides and, together with other specific glycan structure feature ions derived from cross-ring and glycosidic-linkage fragmentation, were used to characterize the glycans and differentiate isomers. The availability of such annotated mass spectrometric fragmentation spectral libraries of glycan structures, together with such substructure diagnostic ions, will be key inputs for the future development of the automated elucidation of oligosaccharide structures from MS/MS data.

ᅟ
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Glycosylation is a universal post-translational modification of proteins that is widely distributed in nature and enhances the functional diversity of the relatively limited number of proteins translated from the genome. Between 20 % and 50 % of all secreted and cellular proteins have been estimated to be glycosylated [1], thus providing great structural protein diversity. A wide range of functions for glycans has been described, ranging from structural roles, participation in molecular regulatory pathways, and association with various disease conditions. Glycomics (the study of the entire complement of oligosaccharides of an organism) has proven to be challenging compared with other ‘omics’ disciplines. As in these disciplines, integrating bioinformatic platforms with glycomics data acquisition technologies is essential to improve our understanding of the structure–function roles of oligosaccharides.
Unlike proteomics and genomics, the field of glycomics has lagged behind to a large extent because of technical challenges associated with the analysis of oligosaccharides. Proteins and nucleic acids are linear biopolymers that can be more easily sequenced. Glycan structures are not template derived and are substantially more complex, comprising elongated branches and isomeric configurations. Over the last decade, there have been considerable advancements in analytical mass spectrometric techniques to characterize glycan structures.
Mass spectrometry provides the potential capability to fully define the composition, sequence, and topology of complex glycans. In addition to glycosidic linkages, cross ring fragmentation provides valuable structural information [2–5], whereby monosaccharide compositional analysis from MS data can be complemented with multistage/tandem MS to determine the linkage and position of the residues [6]. A wide range of MS instruments are available for glycan analysis; matrix-assisted laser desorption-ionization (MALDI) and electrospray ionization (ESI) are the two most commonly used ionization techniques. MALDI-MS, usually with a time of flight analyser (TOF), is a relatively simple technique for rapid glycan profiling, with a high mass range and tolerance of low concentrations of contaminants. A limitation of the MALDI-TOF-MS is in the analysis of native sialylated glycan species in which sialic acids are often lost during the ionization process. This problem can be circumvented by esterification of the acidic groups [7, 8]. Although this type of analysis provides a rapid snapshot of the potential glycan structures present, it does not provide their structural assignment in terms of sequence and linkage and, therefore, does not provide identification of isobaric glycan isomers. In ESI-MS, the glycan sample can be directly infused through either an off-line ESI-source or by an on-line ESI-source coupled to a separation medium such as high-performance liquid chromatography (LC) or capillary electrophoresis (CE). The direct infusion ESI-MS approach is characterized by its simplicity and high-throughput potential [9, 10]; however, again, isobaric glycan species are not resolved in the analysis of complex samples [11]. Furthermore, ion suppression occurs and low abundance glycans may not be detected. To overcome these issues, ESI-MS is commonly combined with LC to enable the detailed characterization of complex glycan mixtures.
Mass spectrometry coupled to porous graphitized carbon liquid chromatographic (PGC-LC) separation brings an additional dimension to glycan structural analysis. The capability of PGC to separate isomeric glycan structures can be used to create a structure-retention time library of known glycans [12] that can discriminate glycan structural features, such as, for example, the resolution of 2–3/6 sialic acid linkage isomers, and the identification of bisecting GlcNAc on N-linked oligosaccharides [3, 4].
To further assist the elucidation and structural characterization of glycans, exoglycosidases, either sequentially or in array form, can be used to reduce sample complexity and to validate linkage information. Such strategies have been extensively reported for fluorescently labelled N-linked oligosaccharides [13] in which LC-MS is used to validate and monitor individual N-glycans by observing shifts in retention time, the occurrence of glycan mass fragments, and the known specificities of exoglycosidases for particular monosaccharide linkages. Although powerful, this technique has certain drawbacks, such as the lack of enzymes for specific linkages [3]. The difficulties of release, derivatization, and heterogeneity of O-linked structures have also impacted the use of this approach to characterize this class of glycan.
The interpretation of glycan analytical data is time consuming and is a limiting aspect for new entrant laboratories. Recent advances in MS analyses of glycans and the development of some database resources have started to aid in the emergence of glycomics as a key player in biomarker discovery [14] and in the monitoring of the glycosylation of therapeutic proteins [15]. However, most glycan structure determination is still based on experimentally determined manual annotation of the tandem MS spectra using the masses generated from glycosidic as well as cross-ring internal cleavages. Commonly used glycan MS/MS annotation programs are based on algorithms that theoretically generate glycan mass fragments and assign them to experimentally obtained MS peaks. GlycosidIQ [16] and GlycoSearchMS [17] implement a SEQUEST-like [18] database searching algorithm that compares an experimental MS/MS spectrum against a spectral database of theoretically fragmented glycan structures. Similarly, Glyco-Peakfinder [19] and GlycoWorkBench [20] utilize theoretical glycan fragmentation masses generated from the EUROCarbDB database of glycan structures [14]. GLYCH uses a scoring algorithm based on patterns of cross-ring cleavage ions [21], whereas Cartoonist [22, 23] uses biosynthetically plausible structures from a theoretical glycan pool to annotate MS spectra.
Provision of glycoinformatic resources, in the form of high-quality experimental databases and data analysis tools, will significantly aid in dealing with the challenges of analytical and structural glycomics, such as it did for genomics and proteomics analysis. One such recent initiative is UniCarb-DB [24], a web-based LC-MS/MS database (www.unicarb-db.org). The current version of UniCarb-DB contains annotated experimental fragmentation spectra of N- and O-glycans predominantly from negative mode electrospray ionization and graphitized carbon LC retention times from previous publications [25–28]. The annotated peak list and tandem mass spectra of interest can be downloaded. The database also contains metadata on the glycan analysis method used, including LC details such as column type, solvents, solvent gradient, and flow rate. The MS metadata includes glycan derivatization, type of instrument used, and mode of data acquisition. UniCarb-DB allows users to query a parent ion mass and obtain its annotated tandem mass spectra, thus enabling the identification or annotation of experimental data by manual comparison at this stage. Automated spectral matching and scoring, using such information as the structure feature ions described in this manuscript, will soon be implemented. The interpretation of tandem MS spectra of oligosaccharides will be greatly simplified by access to this type of glycan LC-MS/MS library if sufficient quality annotated data is available.
Salivary glycans have been previously shown to have a wide variety of structures with various biologically important glycan motifs [5, 26–29] and potential biomarkers [30, 31]. We present a strategy for using a combination of capillary carbon LC retention times, negative ion ESI-MS fragmentation spectra, and the application of a comprehensive set of structural feature fragment ions, to characterize over 200 N- and O-glycans released from salivary glycoproteins. The glycan structures were further validated with exoglycosidase treatments. This study demonstrates that this workflow can result in the comprehensive analysis of a complex biological mixture of protein-derived glycans. This annotated data has been added to the publically available UniCarb-DB database to contribute to an evolving informatics framework that will form the basis of future automated assignment of glycan structures from MS/MS fragmentation spectra.
2 Materials and Methods
2.1 Collection of Saliva
Whole saliva (2 mL) was collected from 10 healthy donors in 2 mL of PBS supplemented with protease inhibitor cocktail (3 mg of antipain-dihydrochloride, 0.5 mg of aprotinin, 0.5 mg of bestatin, 1 mg of chymostatin, 3 mg of E-64, 10 mg of EDTA-Na2, 0.5 mg of leupeptin, 20 mg of Pefabloc SC, 0.5 mg of pepstatin, 3 mg of phosphoramidon), one tablet per 100 mL of buffer (Roche Diagnostics, Mannheim, Germany) and 1 % (vol/vol) Gibco antibiotic-antimycotic, containing 100 units/mL penicillin, 100 μg/mL streptomycin, and 0.25 μg/mL amphotericin-B. The saliva samples were clarified by centrifugation at 3000 × g at 4 °C for 20 min. Proteins were precipitated from the supernatant by addition of ice-cold acetone in the ratio 9:1 and incubated at –20 °C overnight. The precipitated proteins were resolubilized in 4 M urea and protein quantification was performed by Bradford Assay (Sigma-Aldrich, St. Louis, MO).
2.2 N- and O-Linked Glycan Release for Mass Spectrometry Analysis
N- and O-linked glycans were released from saliva according to Jensen et al. [32]. A sample aliquot equivalent to 10 μg of salivary proteins were immobilised by dot blotting onto immobilon-P PVDF membranes for glycan release. The N-linked glycans were released by incubation with PNGase F (3 U) overnight at 37 °C. The released N-linked glycans were incubated with 100 mM NH4COOH pH 5 (final concentration 15 mM) for 60 min at room temperature, and dried in a vacuum centrifuge. The samples were reduced with 20 μL of 1 M NaBH4 in 50 mM KOH at 50 °C for 3 h. The reduction was quenched with 1 μL glacial acetic acid and N-linked glycans were purified using cation exchange columns comprising of 30 μL AG50W-X8 cation-exchange resin (BioRad, Hercules, CA) packed in μC18 ZipTips [32]. The residual borate from the glycan samples were removed by addition of methanol (200 μL) and dried under vacuum.
O-linked glycans were then released by reductive β-elimination from the PNGase F treated proteins by incubating the PVDF spots overnight in 20 μL of 0.5 M NaBH4 in 50 mM KOH at 50 °C for 16 h. The reduction was quenched with 1 μL glacial acetic acid and O-linked glycans were purified using cation exchange columns as described for the N-glycans.
The purified glycans were resuspended in 10 μL of water and analyzed by porous graphitized carbon LC-ESI MS/MS.
2.3 Mass Spectrometry
Both the N- and O-glycan alditols were separated using a commercial 5 μm Hypercarb porous graphitized carbon column, 180 my i.d. × 100 mm, (Thermo Hypersil, Runcorn, UK) maintained at 27 °C. The samples were injected onto the Hypercarb column in 10 mM NH4HCO3, pH 8. Separations of N-glycans were performed over an 85 min gradient of 0 %–45 % (vol/vol) CH3CN in 10 mM NH4HCO3. Similarly the elution of O-glycans was performed over a 45 min gradient of 0 %–90 % (vol/vol) CH3CN in 10 mM NH4HCO3. The flow rate for both the N- and O-glycan separation was 2 μL/min, using a HPLC system (Agilent 1100, Agilient Technologies Inc., CA, USA) and the eluate was introduced directly into the ESI source (Agilent 6330, Agilient Technologies Inc., CA, USA). In the ion-trap MS, the voltage of the capillary outlet was set at 3 kV, and the temperature of the transfer capillary was maintained at 300 °C. The MS spectra were obtained in the negative ion mode over mass range between m/z 200 and 2200. Ions were detected in ion charge control (ICC) (target: 80,000 ions) with an accumulation time of 200 ms. Induced collision was performed at 35 % normalized collision energy and an isolation window of 4 m/z.
2.4 Exoglycosidase Treatment of the Released Glycans
Aliquots of reduced N- and O-glycans released from the salivary proteins of a blood group O individual were digested with an array of exoglycosidase enzyme combinations, following the manufacturer’s recommended concentrations in 50 mM sodium acetate incubation buffer pH 5 at 37 °C for 16 h [13]. Enzymes were removed by carbon-cleanup as previously described [32] before being analyzed by PGC- LC-ESI MS/MS as above.
The enzymes used were as follows: Streptococcus pneumoniae sialidase (α2–3); Arthrobacter ureafaciens sialidase (α2–3, 6, 8); almond meal α-fucosidase (α1–3, 4); bovine testes β-galactosidase (β1–3, 4, 6), purchased from Prozyme (Carlsbad, CA) and Xanthomonas manihotis α-fucosidase (α1-2) from Calbiochem (San Diego, CA).
3 Results and Discussion
3.1 Characterizing Released N- and O-Glycans
The strategy employed to characterize the glycosylation of a complex mixture of salivary glycoproteins is outlined in Supplemental Figure S1. Proteins from whole saliva were first dot-blotted onto PVDF membrane, followed by enzymatic release of N-glycans with PNGase F before the O-glycans were cleaved by subsequent reductive β-elimination. The reduced N- and O-glycans were analyzed by capillary carbon LC-ESI-IT-MS/MS. The resulting glycan tandem mass spectra were manually annotated based on substructure fragment ions that were collated (summarized in Table 1) from an extensive screen of the literature available on the detailed structural characterization of oligosaccharides [2, 4, 5, 27, 33–37]. The structures were further orthogonally confirmed by exoglycosidase treatments.
The systematic nomenclature introduced by Domon and Costello [38] is used with minor modifications for N-glycans [34] and O-glycans [5]. The annotation and interpretation of fragment ion spectra was further characterized with the assistance of software tools, GlycoMod [39] and GlycoWorkBench [20].
3.2 N-Glycans
The structures of the salivary N-glycans were structurally diverse encompassing the high mannose, hybrid and complex N-glycan subclasses.
3.2.1 Neutral N-Glycan Fragmentation
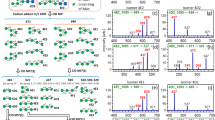
Although it is acknowledged that the use of mass spectrometry to differentiate glycan isomers and closely related structures is challenging, various studies have shown the value of mass spectrometry in characterizing isomers through approaches such as ion-mobility mass spectrometry, multistage fragmentation, and LC-MS [12, 34, 40–42]. Here, we demonstrate the ability of porous graphitized carbon LC in combination with negative ion tandem mass spectrometry to resolve isobaric N-glycan isomers from a complex biological sample. For example, in Figure 1f, three isomers of composition Hex4HexNAc5dHex1 and two isomers of Hex3HexNAc5dHex1 are separated over 13 min in a 45 min gradient run. In addition to the retention time differences, ions specific to structural features in the negative mode fragmentation were important in differentiating these closely related structures (Figure 1a-d, Table 1). Fragmentation of the chitobiose core of hybrid and complex neutral N-glycans produced similar ions as described by Harvey [2] and allowed the confirmation of the presence of the chitobiose core (Y2 m/z 5711– and Z2 m/z 5531– ions, with core fucose attached; and m/z 3891– and m/z 4071– when core fucose is absent, Figure 1). The chemical reduction of the reducing end GlcNAc suppresses the formation of the 2,4A cross ring cleavage described for unreduced N-glycans without the core fucose [33, 34, 43], although the 2,4A4 cleavage of the chitobiose GlcNAc linked to the branching mannose (m/z 11541– in Figure 1a and b) and 2,4A5 cleavage of m/z 13161– (in Figure 1c, d, and e) was present in high abundance. The presence of a fucose at the 6-position of the reducing-terminal GlcNAc residue produced signature ions (Z1 ion at m/z 3501– and a Y1 ion at m/z 3681–) corresponding to the composition of the reduced GlcNAc plus the 6-linked fucose (Figure 1), whereas outer arm fucosylation of GlcNAc was differentiated by the non-reducing end B and C cleavage ions of m/z 3481– and 3661– respectively (as seen on O-glycans in Figure 2).
Characterisation of neutral N-glycan structural features by negative ion PGC-LC ESI-IT-MS/MS. MS/MS spectra shown in panels (a)-(e) are derived from the doubly charged negative ions of m/z 832.62– at 37 min (a) and at 48 min (b), m/z 913.82– at 38.5 min (c), at 39.6 min (d) and at 50 min (e). The extracted ion chromatogram of m/z 832.62– (Hex3HexNAc5dHex1) and m/z 913.82– (Hex4HexNAc5dHex1) exemplifying the PGC-LC separation of the structural isomers is shown in (f). Monosaccharide symbols are as defined by Consortium for Functional Glycomics (CFG) [54] with linkages shown by the Oxford Glycobiology Institute (UOXF) system [55]
Harvey reported the D and [D -18] ions as diagnostic of the composition of the N-glycan 6-antenna [34], and these were found to give prominent signals in the spectra of glycans without a bisecting GlcNAc residue (Figure 1). The D ion arises from the loss of from the chitobiose core (C ion cleavage between Manβ1-4GlcNAc), and the substituents forming the 3-antennae (Z ion cleavage between Manα1-3Man); thus the D-ion mass corresponds to the composition of the 6-arm antenna plus the two remaining branching core mannoses. For example, in Figure 1d the N-glycan 6-antenna monosaccharide composition was identified from ions m/z 5261– [D ion] together with m/z 5081– [D ion-18], which corresponds to a GlcNAc-Man-Man chain. The D-221 ion [34] in negative ion fragment spectra of N-glycans is formed from the additional loss of the bisecting β1-4 linked GlcNAc and has been found to be diagnostic for bisecting N-linked structures. In Figure 1a and c, the tandem MS of the bisecting structures of m/z 832.62– (37 min) and m/z 913.82– (38.5 min) had the same 6-antenna composition giving rise to an abundant D-221 ion at m/z 508–1. In contrast, another structure with the same parent ion mass m/z 913.82– (Figure 1d), that elutes 0.7 min later (39.6 min), contained the D-221 ion of m/z 6701–, which corresponds to the additional presence of a terminal galactose on its 6-arm antenna. The D and [D-18] ions (m/z 5081– and 526–1) of the even later eluting m/z 913.82– isomer (Figure 1e, 50 min) clearly indicated that this structure is not bisected with the 6-antenna containing a non-galactosylated GlcNAc. The fragmentation (D and [D-18], O,4A3) and retention time pattern were similar to the related later eluting m/z 832.62– (48 min) isomers (Figure 1a, b, and f). The composition of the 3-antenna was further determined by the presence of a cross-ring cleavage O,4A fragment ion of the α1-3 linked mannose (also referred to as the E ion [34]). The occurrence of this ion at m/z 6691– [GlcNAc (Gal-GlcNAc)-C4H5O3] in Figure 1e and the absence of the B or C ions corresponding to the linear composition of HexNAc-Gal-GlcNAc classifies this isomer as a triantennary structure with two substitutions on the 3-antenna. Similarly, in Figure 1b, this O,4A2 fragment ion of m/z 5071– indicated that the glycan (m/z 832.62– at 48 min) has two GlcNAc substitutions on the 3-antennae.
The 1,3A cross-ring cleavages (also referred to as the F ion [34]) of the 3- or 6-linked mannose antenna residues are useful in determining the non-reducing end compositions. The spectra of complex glycans containing Gal-GlcNAc antennae exhibited a prominent fragment at m/z 4241– (Figures 1c, d, and e), while m/z 2621– corresponded to the terminal GlcNAc residue. Sagi et al. reported the formation of this cross link cleavage to be more favorable for the 6-mannose antennae [44].
3.2.2 Sialylated N-Glycan Fragmentation
In general, sialylated structures produce low intensity tandem mass spectra in the negative mode, as most charge is retained on the acidic groups rather than on the hydroxyl groups. Although most of the fragment ions of the mono-sialylated glycan isomers predominantly contained the acidic groups, their fragmentation still provided information. The tandem mass spectra of mono-sialylated N-glycans mainly showed the B ion cleavages of Neu5Ac and the Neu5Ac-Gal-GlcNAc chain as signals at m/z 2901– and 6551–, respectively, irrespective of the linkages and attachment to the N-glycan core as shown by the isomers in Supplemental Figure S2a–d. The m/z 9791– corresponded to a D ion formed by both the α2-3 and α2-6 linked sialic acid, and indicated the presence of sialylated lactosamine on the 6-antenna (Supplemental Figure S2a and c). However, the fragmentation of the doubly and triply sialylated glycan masses observed in the MS spectra was dominated by the loss of the sialic acids. The fragmentation spectra contained sufficient information to confirm the composition, but other structural feature ions were suppressed. Permethylation of sialylated glycans has been shown to provide more information on multi-sialylated species in the positive mode [45, 46]; further MSn fragmentation of the daughter ions is also an alternate approach [41].
The spectra of sialylated glycans lack most of the antenna specific ions observed for neutral glycans (see above). Previous studies have reported various diagnostic ions that could differentiate between the α2-6 and the α2-3 linkages in the negative ionization mode, such as the m/z 3061– ion corresponding to 0,2A2–CO2 cross link cleavage of the sialic acid attached Gal of unreduced small oligosaccharides [4, 36, 47]. We could not observe these diagnostic ions in our analyses of α2-6 and α2-3 sialylated glycan isomers released from glycoproteins. Deguchi et al. [37] suggested that the formation of the 0,2A2–CO2 diagnostic ion could be dependent on the collision gas. Other studies [4, 36, 47], using argon gas for CID observed the presence of this ion, whereas it was absent when the collision gas was helium, as used here. Nevertheless, our data (Supplemental Figure S2 and 4) show that the intensity ratio of the B cleavage products [corresponding to the sialic acid (m/z 2901–) and sialylated lactosamine loss (m/z 6551–)] can be used as an indicator of the linkage. Sialic acids α2-3 linked to the Gal are more labile (higher ion intensity of m/z 2901) than α2-6 linked sialic acids (Supplemental Figure S2a-d). This observation is consistent with previous studies by Seymour et al. on sialylated milk free oligosaccharides and Deguchi et al. on monosialylated N-glycans [37, 43]. Additionally α2-6 linked sialylated N-glycans contained fragment ions originating from the sequential loss of the Gal and GlcNAc monosaccharides, respectively, (m/z 7022– and 6002–) from the abundant 2,4A6 ion (m/z 782.82–) (Supplemental Figure S2a and b). These specific cleavages were consistently present in other α2-6 linked sialylated N-glycans, while absent in the α2-3 linked variants. The longer retention of α2-3 linked sialylated N-glycans over the α2-6 variants by PGC-LC (Supplemental Figure S2e) provides further confirmation of the type of sialylated linkage. This selective order of elution by PGC-LC conferred by sialic acid linkage position has been reported by other studies [12, 48]. Additionally, the sequential enzymatic desialylation by Streptococcus pneumoniae sialidase (α2-3) and Arthrobacter ureafaciens sialidase (α2-3, 6, 8; ABS) orthogonally confirmed the sialic acid linkage (data not shown).
3.3 O-Glycans
The mucins in saliva are a rich source of O-glycan structures [5, 27, 28], that comprise various O-glycan core types, different fucose linkages, as well as O-glycan sulphation and sialylation. The assignment of O-glycan structural complexity and diversity requires sequence-specific tandem mass spectral fragment ions as well as additional diagnostic ions that correspond to their particular linkages and configurations.
3.3.1 Neutral O-Glycan Fragmentation: Differentiating Core Type Fragmentation
There have been up to 8 O-GalNAc linked core structures described in mammalian mucins [49]. Cores 1 and 8 contain a Gal linked by a β1-3 or α1-3 linkage, respectively, to the reducing end GalNAc, while cores 3, 5, 6, and 7 contain a HexNAc (GalNAc or GlcNAc) linked to the reducing GalNAc. Core 2 (Galβ1-3(GlcNAcβ1-6)GalNAcol) and core 4 (GlcNAcβ1-3(GlcNAcβ1-6)GalNAcol) are substituted with Gal and GlcNAc linked β1-6 to the reducing end GalNAc of core 1 and core 3, respectively. Differentiating between mucin oligosaccharide cores differing only in linkage (such as between cores 1 and 8) is particularly difficult in mass spectrometric glycan analysis, and in these cases the origin of the sample is used to differentiate such isomers. For example, core 1 and core 2 are ubiquitously found in human mucins, whereas core 6 has been reported only in seminal fluid, meconium, and ovarian cyst [50].
The singly C-3 substituted reducing end terminus GalNAcol (core 1: Galβ1-3GalNAcol and core 3: GlcNAcβ1-3GalNAcol) can be easily identified by the CID loss of this moiety (loss of 223 Da) from the pseudomolecular ion as previously described [5, 35, 50]. Figure 2a and b illustrate that core 1 type structural isomers with the corresponding B type ion (m/z 8181–) formed with the loss of the reducing end GalNAcol from the parent ion mass of 1041.31–. Core 1 type structures were further confirmed by the reported loss of 108 Da from the [M – H]1– precursor ion (resulting in m/z 9331– in Figure 2a and b) through the partial loss of side groups from the reducing end GalNAcol [5].
The reported archetypal Y and Z ions (m/z 3841– and 3661– [5, 35, 50]) representing the cleavage of the core 1 type Galβ1-3GalNAcol were also observed (Figure 2a and b; Table 1). In core 2 type glycans, in addition to these ions (m/z 3841– and 3661–) the GlcNAc β1-6 linked to the GalNAcol generated m/z 4251–, 4071-, and 3891– (Figure 2b, c, and d; Table 1).
The fragmentation of core 2 and core 4 structures also yielded a diagnostic cross-ring cleavage 0,4A previously reported by Schulz et al. [30]. This cleavage is between C-4 and C-5 of the reducing end GalNAcol and can aid in determining the composition of the substituents of the C-6 linkage. For example, in Figure 2c, d, and e, for the three core 2 isomers, the 0,4A cleavage of the GalNAcol resulted in the m/z 716–1, 5701–, and 5701–, respectively, corresponding to the C-6 fragment compositions of (Fuca1-2Galβ1-3(Fucβ1-4)GlcNAc1β-), (Galβ1-4(Fucα1-3)GlcNAc1β-), and (Fucα1-2Galβ1-4GlcNAc1β-), respectively.
3.3.2 Type 1 and Type 2 Chain Elongations and Terminal Fucose Linkages
Terminal N-acetylglucosamine residues can be substituted with β1-4 galactose to generate the type 2 lactosamine structure (Gal β1-4GlcNAc), whereas the β1-3 linkage of galactose gives rise to the type 1 lactosamine (Gal β1-3GlcNAc). Characteristic structural feature ions of type 1 structures such as those reported by Chai et al. [36] on milk oligosaccharide standards are of low intensity or absent in mucin type glycans [5, 35]. However the 0,2A cross ring cleavage, with a concurrent loss of water from the 4 linked GlcNAc (as seen on Figure 2e [m/z 4091–]) is characteristic of the type 2 lactosamine chain, provided the GlcNAc is not fucosylated. Karlsson et al. [5] proposed from MS3 experiments on this fragment that the GlcNAc moiety is converted to an enone type structure.
Tandem mass spectra of glycans carrying the H-antigen (α1-2 fucose linkage to Gal) showed the presence of B and C ion cleavages of these fragments with m/z 3071– and m/z 3251– (Figure 2c, d, and e). In smaller glycans these ions were accompanied by the B-C2H4O2 ion of m/z 2471– and 1,3A cross ring cleavage signals of the H-antigen, which included the fucose and part of the C2 linked galactose at m/z 2051– (Table 1).
The unusual Z/Z cleavage of the di-substituted GlcNAc is a fragmentation specific to Lewis type epitopes [5]. In Figure 2e, the Z/Z cleavage of the GlcNAc corresponded to the signal observed at m/z 5511–. Karlsson et al. suggested that this fragmentation results from the formation of a conjugated diene by dual elimination of the Gal and Fuc residues from the GlcNAc [5]. The extension of the Lewis a into Lewis b epitopes and Lewis x into Lewis y epitopes does not inhibit the formation of this diene as seen on Figure 2a, c, and d, where the Z/Z of both these isomers resulted in a fragment ion of m/z 5511–, 5511–, and 6971–, respectively. These fragments were further accompanied by the loss of acetyl and acetate groups from the parent ion; for example, m/z 5211– (Figure 2a and c) corresponds to ZZ-CH2O and, similarly, m/z 6251– and m/z 6731– (Figure 2d) represent ZZ-C3H4O2 and ZZ-C2H2O, respectively (Figure 2d and e).
Mass spectra of negative ion fragmentation of Lewis a and x antigen standards have been described previously [36, 43]. Both structures produce an abundant Z ion cleavage of the 3-linked residue. For Lewis x, the abundant Z fragment ion was m/z 3641– (Figure 2b, d, and e) but for Lewis a, the Z ion was m/z 3481– (Figure 2a and c), suggesting that the C3 linkage to the GlcNAc is labile relative to the C4. The fragment spectra also showed a 2,4X signal corresponding to the 4-linked residue from the abundant Z ion that is present for both Lewis type epitopes. This ion corresponds to a 60 u loss from the corresponding Z ion fragments resulting in m/z 3041– for Lewis x (Figure 2a and c) and m/z 2881– for Lewis a (Figure 2d). These Z and 2,4X ion cleavages were successfully used to resolve the isomers in Figure 2 in which the occurrence of ions m/z 3481– and m/z 2881– (Figure 2a and c) indicated that the Fuc was α1-3 linked to GlcNAc. The structures in Figure 2b and d have the analogous Z and 2,4X cleavage ions with m/z 3641– and m/z 3041– indicating that the Gal is β1-4 linked to the GlcNAc forming a Lewis a epitope.
3.3.3 Sialylated O-Glycan Fragmentation
Interpretation of the fragmentation of sialylated O-glycans poses a similar problem to that of acidic N-glycans, with losses of Neu5Ac in the form of B ions that diminish the abundances of the A, and X-type cleavage ions that can provide additional information on the structure [43] (Supplemental Figure S3 compared with Figure 2). In human mucin O-glycans, sialic acid can be either α2-3 linked to a Gal residue or α2-6 linked to the GalNAcol [35]. The sialylated O-glycan isomers of composition Neu1Hex1HexNAc2 of m/z of 8781–, which differ in sialic acid linkage and core type, eluted at 25 and 27 min as shown in Supplemental Figure S3c.
In Supplemental Figure S3a, the Z1β and Y1β ions (m/z 4951– and m/z 5131–) are diagnostic of the sialic acid linkage to the core GalNAc (GalNAcol + Neu5Ac). A fragment ion generated via a 0,2X-type cleavage of the α2-6 sialic acid corresponding to a mass difference of 221 Da relative to the precursor ion, was present at m/z 6571–. In contrast, the low intensity m/z 6571– as shown in Supplemental Figure S3b corresponded to Z1α cleavage representing the composition (Neu5Acα2-3Galβ1-3 GalNAcol). The observation that the α2-6 sialylated isomer elutes earlier than the α2-3 isomer by graphitized carbon LC is consistent with the retention difference observed for the sialylated N-glycans.
3.3.4 Sulphated O-Glycan Fragmentation
Sulphation is a modification found on various O-glycans; however, determining the position of this monosaccharide substitution remains a technical challenge. In Supplemental Figure S4 the isomers of m/z 8131– (Sulph1NeuAc1Hex1HexNAc2) differing only in the sulphate substitution to Gal and GlcNAc are separated using PGC-LC. The presence of the sulphate is indicated by fragment ions at m/z 4441– (HSO3-Gal-GlcNAc) and 5901– (HSO3-Gal-Fuc-GlcNAc) in both sulphated O-glycan spectra (Supplemental Figure S4). As shown by Robbe et al. [35, 50] previously, the fragment ion at m/z 2411– and 2821– are diagnostic for locating the sulphate group on Gal and GalNAc residues, respectively. The sulphate linkage to the monosaccharide residues can be further investigated through subsequent MSn analysis of the sulphated fragment ions, including m/z 4441– (HSO3-Gal-GlcNAc) and 5901– (HSO3-Gal-Fuc-GlcNAc). The tandem mass spectra of these sulphated isomers also provided additional information, such as the location of the fucose on the GlcNAc (m/z 4281– in Supplemental Figure S4a and m/z 3661– in Supplemental Figure S4b).
3.4 Exoglycosidase Treatment and Confirmation of Glycan Structures
Sequential exoglycosidase treatment of the global glycan profiles showed dramatic differences, and these changes were instrumental in validating isomeric structures annotated by the MS/MS and LC retention time analysis. For example, Supplemental Figure S5 illustrates the sequential exoglycosidase digestions used to confirm the structural annotation of nine isomeric glycans of m/z 1039.42– (Hex2HexNAc2dHex3 + Man3GlcNAc2), which had been initially assigned by diagnostic and structural feature fragment ions and chromatographic behavior. The glycans were desialylated with Arthrobacter ureafaciens sialidase (α2-3, 6, 8; ABS) to reduce the complexity that could arise from interference by biantennary disialylated glycans of the similar mass of m/z 1038.82– that the ion trap mass accuracy could not distinguish. The bovine testes β-galactosidase (β1-3, 4, 6; BTG) digests isomers with a free terminal galactose residue but its activity is inhibited [51] by the presence of Lewis epitopes (Supplemental Figure S5b). This enabled the confirmation of unmodified LacNAc (Gal-GlcNAc) structures in this isomer. The Lewis x/a epitopes were further confirmed by the digestion of the glycan isomers with BTG in combination with almond meal α-fucosidase (α1-3, 4; AMF). The tandem mass spectra of the undigested isomers after treatment with the BTG + AMF exoglycosidases contained the two signature ions (previously discussed) for α1, 2 fucose linked to type 1 and type 2 galactose residues, with the third fucose linked to the N-glycan core. These three isomers (Supplemental Figure S5c) were further confirmed by the use of bovine kidney α-fucosidase which removed all the remaining α2, 6-linked core fucose residues (Supplemental Figure S5d).
3.5 Interpretation of MS/MS Data
Different MS instruments, separation techniques and experimental conditions result in different retention times, fragmentation profiles and structural ion features of glycans. Current glycan structural bioinformatics analysis programs do not incorporate this information, and mostly depend on in silico generated theoretical fragments to annotate experimental tandem mass spectra. This approach rarely assigns isomeric glycan structures since theoretical fragmentation of these isomers often yields exactly the same “possible” fragment ion masses. An example of the in silico fragment assignment by the tool GlycoWorkBench [20] to the MS/MS spectra of the α2-6 monosialylated biantennary N-glycan isomer sialylated on the 6 arm, from Supplemental Figure S2a, is shown in Supplemental Figure S6. The tandem MS spectra of this isomer matched to the α2-6 monosialylated biantennary N-glycan sialylated on both the 6 branching mannose and the 3 branching mannose antenna. The annotation and coverage statistics generated by GlycoWorkBench revealed identical fragments and fragment ion coverage percentage (Supplemental Figure S6), and was unable to distinguish between the two isomers. On the other hand, using the abundant structural feature D ion, m/z 979.32– in Supplemental Figure S2a is able to clearly distinguish between the two isomers. The defining nature of this single feature ion in determining the structure of such closely related isomers emphasises the potential for the inclusion of these types of fragment ions into spectral matching scoring algorithms. This type of software will significantly increase the confidence of the assignment of glycan structures and their isomers.
4 Conclusion
To achieve comprehensive structural assignment and discrimination, isobaric glycoforms need to be well resolved and separated prior to MS/MS analysis. PGC-LC provides this additional dimension of retention time selectivity based on glycan structural characteristics, and has been shown to reliably construct a retention time library of isomeric glycans [12]. Although the chromatographic behavior, in terms of the order of elution of closely related structures, by PGC-LC is highly reproducible [12, 52], absolute LC retention time shifts are commonly observed. To overcome this issue, the retention time shifts could be aligned algorithmically based on matching of glycans common to each spectra and verified by their fragmentation spectra. Alternatively, the use of internal standards to correct the retention time shifts could be utilized as previously demonstrated by Pabst et al. [12]. The latter, however, would be difficult to uniformly establish across different laboratories. Recently, a graphitized carbon nano-LC chip configuration [53] has been released, which shows promise to be able to deliver the reproducible retention times required for inter-laboratory comparisons.
Here, we present a strategy used to characterize over 200 released N- and O-linked glycans, including closely related isomers, using a combination of online PGC-LC coupled to negative ion ESI-IT MS/MS detection; these structures have been orthogonally confirmed by exoglycosidase digestions. Specifically, we demonstrate how diagnostic and signature fragment ions produced by negative ion MS/MS fragmentation, supported by the elution selectivity of PGC retention of isomeric glycans, can be used to characterize the glycosylation of a complex biological sample. Furthermore we present a comprehensive list of these structural feature ions.
This combination of mass, retention order, MS/MS spectra, and specific fragment ions can be used as determinants of heterogeneous glycan structures. The annotated spectra described in this study have been stored in the UniCarb-DB database [24]. We plan to incorporate the information on the specific structural feature ions into algorithms capable of accurately annotating and scoring the spectral matching of experimental MS/MS spectra with the database. The approach to characterize glycan structures through experimentally determined spectral databases together with specific signature fragment ions will increase the confidence level of glycan structural assignment. It would also effectively reduce the required number of specific exoglycosidase digestions currently used to validate these structures. In this way, we can move one step closer to the automated, high confidence assignment of glycan structures in PGC-LC ESI MS/MS glycomics experiments. The availability of such bioinformatic tools would greatly facilitate the interpretation of the complex MS fragmentation of oligosaccharides that is essential if new discoveries are to be made as to the function of these structurally heterogeneous molecules.
References
Khoury, G.A., Baliban, R.C., Floudas, C.A.: Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci. Rep. 1(90), 1–5 (2011)
Harvey, D.J.: Fragmentation of negative ions from carbohydrates. Part 2. Fragmentation of high-mannose N-linked glycans. J. Am. Soc. Mass Spectrom. 16, 631–646 (2005)
Harvey, D.J., Dwek, R.A., Rudd, P.M.: Determining the structure of glycan moieties by mass spectrometry. Curr. Protoc. Protein Sci. 43, Unit 12.17.1–12.17.18 (2006)
Harvey, D.J., Rudd, P.M.: Fragmentation of negative ions from N-linked carbohydrates. Part 5. Anionic N-linked glycans. Int. J. Mass Spectrom 305, 120–130 (2011)
Karlsson, N.G., Schulz, B.L., Packer, N.H.: Structural determination of neutral O-linked oligosaccharide alditols by negative ion LC-electrospray-MSn. J. Am. Soc. Mass Spectrom. 15, 659–672 (2004)
Pabst, M., Altmann, F.: Glycan analysis by modern instrumental methods. Proteomics 11, 631–643 (2011)
Geyer, H., Geyer, R.: Strategies for analysis of glycoprotein glycosylation. Biochim. Biophys. Acta 1764, 1853–1869 (2006)
Bielik, A.M., Zaia, J.: Historical overview of glycoanalysis. Methods Mol. Biol. 600, 9–30 (2010)
Stumpo, K.A., Reinhold, V.N.: The N-glycome of human plasma. J. Proteome Res. 9, 4823–4830 (2010)
Prien, J.M., Prater, B.D., Cockrill, S.L.: A multi-method approach toward de novo glycan characterization: a Man-5 case study. Glycobiology 20, 629–647 (2010)
Jiao, J., Zhang, H., Reinhold, V.N.: High Performance IT-MS sequencing of glycans (spatial resolution of ovalbumin isomers). Int. J. Mass Spectrom. 303, 109–117 (2011)
Pabst, M., Bondili, J.S., Stadlmann, J., Mach, L., Altmann, F.: Mass + retention time = structure: a strategy for the analysis of N-glycans by carbon LC-ESI-MS and its application to fibrin N-glycans. Anal. Chem. 79, 5051–5057 (2007)
Royle, L., Campbell, M.P., Radcliffe, C.M., White, D.M., Harvey, D.J., Abrahams, J.L., Kim, Y.G., Henry, G.W., Shadick, N.A., Weinblatt, M.E., Lee, D.M., Rudd, P.M., Dwek, R.A.: HPLC-based analysis of serum N-glycans on a 96-well plate platform with dedicated database software. Anal. Biochem. 376, 1–12 (2008)
Rakus, J.F., Mahal, L.K.: New technologies for glycomic analysis: toward a systematic understanding of the glycome. Annu Rev Anal Chem (Palo Alto, CA) 4, 367–392 (2011)
Wacker, C., Berger, C.N., Girard, P., Meier, R.: Glycosylation profiles of therapeutic antibody pharmaceuticals. Eur. J. Pharm. Biopharm. 79, 503–507 (2011)
Joshi, H.J., Harrison, M.J., Schulz, B.L., Cooper, C.A., Packer, N.H., Karlsson, N.G.: Development of a mass fingerprinting tool for automated interpretation of oligosaccharide fragmentation data. Proteomics 4, 1650–1664 (2004)
Lohmann, K.K., von der Lieth, C.W.: GlycoFragment and GlycoSearchMS: web tools to support the interpretation of mass spectra of complex carbohydrates. Nucleic Acids Res. 32, W261–266 (2004)
Eng, J.K., Mccormack, A.L., Yates, J.R.: An approach to correlate tandem mass-spectral data of peptides with amino-acid-sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994)
Maass, K., Ranzinger, R., Geyer, H., von der Lieth, C.W., Geyer, R.: “Glyco-peakfinder”—de novo composition analysis of glycoconjugates. Proteomics 7, 4435–4444 (2007)
Ceroni, A., Maass, K., Geyer, H., Geyer, R., Dell, A., Haslam, S.M.: GlycoWorkbench: a tool for the computer-assisted annotation of mass spectra of glycans. J. Proteome Res. 7, 1650–1659 (2008)
Tang, H., Mechref, Y., Novotny, M.V.: Automated interpretation of MS/MS spectra of oligosaccharides. Bioinformatics 21, 431–439 (2005)
Goldberg, D., Sutton-Smith, M., Paulson, J., Dell, A.: Automatic annotation of matrix-assisted laser desorption/ionization N-glycan spectra. Proteomics 5, 865–875 (2005)
Goldberg, D., Bern, M., Li, B.S., Lebrilla, C.B.: Automatic determination of O-glycan structure from fragmentation spectra. J. Proteome Res. 5, 1429–1434 (2006)
Hayes, C.A., Karlsson, N.G., Struwe, W.B., Lisacek, F., Rudd, P.M., Packer, N.H., Campbell, M.P.: UniCarb-DB: a database resource for glycomic discovery. Bioinformatics 27, 1343–1344 (2011)
Estrella, R.P., Whitelock, J.M., Packer, N.H., Karlsson, N.G.: The glycosylation of human synovial lubricin: implications for its role in inflammation. Biochem. J. 429, 359–367 (2010)
Issa, S., Moran, A.P., Ustinov, S.N., Lin, J.H., Ligtenberg, A.J., Karlsson, N.G.: O-linked oligosaccharides from salivary agglutinin: Helicobacter pylori binding sialyl-Lewis x and Lewis b are terminating moieties on hyperfucosylated oligo-N-acetyllactosamine. Glycobiology 20, 1046–1057 (2010)
Karlsson, N.G., Thomsson, K.A.: Salivary MUC7 is a major carrier of blood group I type O-linked oligosaccharides serving as the scaffold for sialyl Lewis x. Glycobiology 19, 288–300 (2009)
Everest-Dass, A.V., Jin, D., Thaysen-Andersen, M., Nevalainen, H., Kolarich, D., Packer, N.H.: Comparative structural analysis of the glycosylation of salivary and buccal cell proteins: innate protection against infection by C. albicans. Glycobiology 22, 1465–1479 (2012)
Guile, G.R., Harvey, D.J., O’Donnell, N., Powell, A.K., Hunter, A.P., Zamze, S., Fernandes, D.L., Dwek, R.A., Wing, D.R.: Identification of highly fucosylated N-linked oligosaccharides from the human parotid gland. Eur. J. Biochem. 258, 623–656 (1998)
Guttman, A.: Analysis of monosaccharide composition by capillary electrophoresis. J. Chromatogr. A 763, 271–277 (1997)
Drake, P.M., Cho, W., Li, B., Prakobphol, A., Johansen, E., Anderson, N.L., Regnier, F.E., Gibson, B.W., Fisher, S.J.: Sweetening the pot: adding glycosylation to the biomarker discovery equation. Clin. Chem. 56, 223–236 (2010)
Jensen, P.H., Karlsson, N.G., Kolarich, D., Packer, N.H.: Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 7, 1299–1310 (2012)
Harvey, D.J.: Fragmentation of negative ions from carbohydrates. Part 3. Fragmentation of hybrid and complex N-linked glycans. J. Am. Soc. Mass Spectrom. 16, 647–659 (2005)
Harvey, D.J., Royle, L., Radcliffe, C.M., Rudd, P.M., Dwek, R.A.: Structural and quantitative analysis of N-linked glycans by matrix-assisted laser desorption ionization and negative ion nanospray mass spectrometry. Anal. Biochem. 376, 44–60 (2008)
Robbe, C., Michalski, J.C., Capon, C.: Structural determination of O-glycans by tandem mass spectrometry. Methods Mol. Biol. 347, 109–123 (2006)
Chai, W., Piskarev, V.E., Mulloy, B., Liu, Y., Evans, P.G., Osborn, H.M., Lawson, A.M.: Analysis of chain and blood group type and branching pattern of sialylated oligosaccharides by negative ion electrospray tandem mass spectrometry. Anal. Chem. 78, 1581–1592 (2006)
Deguchi, K., Takegawa, Y., Ito, H., Miura, N., Yoshioka, S., Nagai, S., Nakagawa, H., Nishimura, S.: Structural assignment of isomeric 2-aminopyridine-derivatized monosialylated biantennary N-linked oligosaccharides using negative-ion multistage tandem mass spectral matching. Rapid Commun. Mass Spectrom. 20, 412–418 (2006)
Domon, B., Costello, C.E.: A systematic nomenclature for carbohydrate fragmentations in FAB-MS/MS spectra of glycoconjugates. Glycoconj. J. 5, 397–409 (1988)
Cooper, C.A., Gasteiger, E., Packer, N.H.: GlycoMod—a software tool for determining glycosylation compositions from mass spectrometric data. Proteomics 1, 340–349 (2001)
Plasencia, M.D., Isailovic, D., Merenbloom, S.I., Mechref, Y., Novotny, M.V., Clemmer, D.E.: Resolving and assigning N-linked glycan structural isomers from ovalbumin by IMS-MS. J. Am. Soc. Mass Spectrom. 19, 1706–1715 (2008)
Ito, H., Yamada, K., Deguchi, K., Nakagawa, H., Nishimura, S.: Structural assignment of disialylated biantennary N-glycan isomers derivatized with 2-aminopyridine using negative-ion multistage tandem mass spectral matching. Rapid Commun. Mass Spectrom. 21, 212–218 (2007)
Prien, J.M., Huysentruyt, L.C., Ashline, D.J., Lapadula, A.J., Seyfried, T.N., Reinhold, V.N.: Differentiating N-linked glycan structural isomers in metastatic and nonmetastatic tumor cells using sequential mass spectrometry. Glycobiology 18, 353–366 (2008)
Seymour, J.L., Costello, C.E., Zaia, J.: The influence of sialylation on glycan negative ion dissociation and energetics. J. Am. Soc. Mass Spectrom. 17, 844–854 (2006)
Sagi, D., Peter-Katalinic, J., Conradt, H.S., Nimtz, M.: Sequencing of tri- and tetraantennary N-glycans containing sialic acid by negative mode ESI QTOF tandem MS. J. Am. Soc. Mass Spectrom. 13, 1138–1148 (2002)
Mechref, Y., Kang, P., Novotny, M.V.: Differentiating structural isomers of sialylated glycans by matrix-assisted laser desorption/ionization time-of-flight/time-of-flight tandem mass spectrometry. Rapid Commun. Mass Spectrom. 20, 1381–1389 (2006)
Zaia, J.: Mass spectrometry of oligosaccharides. Mass Spectrom. Rev. 23, 161–227 (2004)
Wheeler, S.F., Harvey, D.J.: Negative ion mass spectrometry of sialylated carbohydrates: discrimination of N-acetylneuraminic acid linkages by MALDI-TOF and ESI-TOF mass spectrometry. Anal. Chem. 72, 5027–5039 (2000)
Nakano, M., Saldanha, R., Gobel, A., Kavallaris, M., Packer, N.H.: Identification of glycan structure alterations on cell membrane proteins in desoxyepothilone B resistant leukemia cells. Mol. Cell. Proteom. 10(M111), 009001–009012 (2011)
Brockhausen, I., Schachter, H., Stanley, P.: In Essentials of Glycobiology. Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Certozzi, C.R., Hart, G.W., Etzler, M.E., Eds., Cold Spring Harbor, Cold Spring Harbor Laboratory Press, pp. 115–128 (2009)
Robbe, C., Capon, C., Coddeville, B., Michalski, J.C.: Diagnostic ions for the rapid analysis by nano-electrospray ionization quadrupole time-of-flight mass spectrometry of O-glycans from human mucins. Rapid Commun. Mass Spectrom. 18, 412–420 (2004)
Kolarich, D., Weber, A., Turecek, P.L., Schwarz, H.P., Altmann, F.: Comprehensive glyco-proteomic analysis of human α1-antitrypsin and its charge isoforms. Proteomics 6, 3369–3380 (2006)
Ruhaak, L.R., Miyamoto, S., Kelly, K., Lebrilla, C.B.: N-Glycan profiling of dried blood spots. Anal. Chem. 84, 396–402 (2012)
Hua, S., An, H.J., Ozcan, S., Ro, G.S., Soares, S., DeVere-White, R., Lebrilla, C.B.: Comprehensive native glycan profiling with isomer separation and quantitation for the discovery of cancer biomarkers. Analyst 136, 3663–3671 (2011)
Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Marth, J.D., Bertozzi, C.R., Hart, G.W., Etzler, M.E.: Symbol nomenclature for glycan representation. Proteomics 9, 5398–5399 (2009)
Harvey, D.J., Merry, A.H., Royle, L., Campbell, M.P., Rudd, P.M.: Symbol nomenclature for representing glycan structures: extension to cover different carbohydrate types. Proteomics 11, 4291–4295 (2011)
Acknowledgment
The authors acknowledge Dr. Niclas Karlsson and Dr. Catherine Hayes for their ongoing collaboration with the UniCarb-DB database project. This research project was facilitated by access to the Australian Proteomics Analysis Facility (APAF) established under the Australian Government NCRIS program.
This work was funded by Macquarie University Research Excellence Scheme postgraduate scholarship and Cystic Fibrosis Australia postgraduate studentship grant, Australian National Data Service (ANDS) grant DC12A and National eResearch Collaboration Tools and Resources (NeCTAR) grant RT016.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Figure S1
Workflow for glycan analysis including bioinformatic support for the annotation and interpretation of the generated MS/MS spectra. N- and O-linked glycans were released sequentially from salivary proteins dot blotted on to PVDF membranes, by PNGase F and reductive β-elimination and subsequently analyzed by PGC LC-ESI-IT MS/MS (PNG 158 kb)
Figure S2
Identification and confirmation of sialic acid linkage and arm location of monosialyated biantennary N-glycan isomers. MS/MS spectra were obtained from the doubly charged negative ion parent ions of m/z 965.82– (NeuAc1Hex5HexNAc4) eluting at 44.5 min (a), 45.5 min (b), 52.5 min (c), and 53.5 min (d); (e) shows the extracted ion chromatogram of the four m/z 965.82- isomers (JPEG 243 kb)
Figure S3
Identification and characterization of sialylated O-glycan isomers. (a) and (b) The negative ion fragmentation spectra of m/z 878.51– (NeuAc1Hex1HexNAc2) isomers eluting at 25 and 26.7 min, respectively. The extracted ion chromatogram of the m/z 878.52– isomers is shown in (c) (JPEG 86 kb)
Figure S4
Identification and characterization of sulphated O-glycan isomers. Negative ion fragmentation spectra of m/z 8131– (Sulph1Hex1HexNAc2) isomers eluting at 21.8 min (a) and 22.5 min (b) are shown; (c) shows an extracted ion chromatogram of m/z 8131– isomers (JPEG 90 kb)
Figure S5
Extracted ion chromatograms of m/z 1039.42– (Hex5HexNAc4dHex3) N-glycan isomers. The glycan structural and linkage isomers were orthogonally confirmed by exoglycosidase treatments. ABS: Arthrobacter ureafaciens sialidase (α2–3,6,8). BTG: bovine testes β-galactosidase (β1–3,4,6). AMF: Almond meal α-fucosidase (α1–3,4). BKF: bovine kidney α-fucosidase (α1–2, 6) (JPEG 64 kb)
Figure S6
Glycan annotation report generated by GlycoWorkBench tool [20] of monosialyated biantennary N-glycan isomers of m/z 965.82– (NeuAc1Hex5HexNAc4) differing in α2-6 sialylation of the antennae. (a) MS/MS spectra of the doubly charged negative ion eluting at 44.5 min and its GlycoWorkBench generated annotation based on in silico fragmentation (b) MS/MS spectra of the doubly charged negative ion eluting at 45.5 min and its Glycoworkbench generated annotation based on in silico fragmentation. To simplify the figure, only singly charged glycan fragment masses above a 3 % intensity threshold are displayed. (c) Statistical coverage from the two isomers generated by Glycoworkbench (JPEG 207 kb)
Rights and permissions
About this article
Cite this article
Everest-Dass, A.V., Abrahams, J.L., Kolarich, D. et al. Structural Feature Ions for Distinguishing N- and O-Linked Glycan Isomers by LC-ESI-IT MS/MS. J. Am. Soc. Mass Spectrom. 24, 895–906 (2013). https://doi.org/10.1007/s13361-013-0610-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-013-0610-4