Abstract
Molecular mass images of tissues will be biased if differences in the physicochemical properties of the microenvironment affect the intensity of the spectra. To address this issue, we have performed—by means of MALDI-TOF mass spectrometry—imaging on slices and lipidomic analysis in extracts of frontal cortex, both from the same postmortem tissue samples of human brain. An external calibration was used to achieve a mass accuracy of 10 ppm (1σ) in the spectra of the extracts, although the final assignment was based on a comparison with previously reported species. The spectra recorded directly from tissue slices (imaging) show excellent s/n ratios, almost comparable to those obtained from the extracts. In addition, they retain the information about the anatomical distribution of the molecular species present in autopsied frozen tissue. Further comparison between the spectra from lipid extracts devoid of proteins and those recorded directly from the tissue unambiguously show that the differences in lipid composition between gray and white matter observed in the mass images are not an artifact due to microenvironmental influences of each anatomical area on the signal intensity, but real variations in the lipid composition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Mass spectrometry (MS) is an excellent tool for the analysis of the lipidome, due to its intrinsic characteristics [1, 2]: fast acquisition times, robustness, high tolerance impurities, and high dynamic range (from pmol/μl to mmol/μl). Therefore, an increasing number of studies have been published over the last few years that aim at the identification of lipids in biological samples using different mass spectrometry techniques [3–6]. The matrix-assisted laser desorption/ionization (MALDI) imaging mass spectrometry (IMS) technique, introduced by Caprioli et al. [7, 8], permits the scanning of tissue slices directly in the mass spectrometer, and therefore takes the precise anatomical localization of different types of lipids a step further. Such information, which is lost in the extracts, is of singular importance to attempt to understand the physiopathology of the different lipid species, and is even more significant in the case of the central nervous system (CNS), which is composed of a network of very specialized groups of cells and signaling pathways distributed in discrete areas or nuclei of the brain.
Despite the potential applicability of this method and the growing number of papers published recently in the field of lipid IMS [9–11], some questions remain unanswered and numerous concerns have been raised. Most importantly, it is not clear whether the different intensities of lipid signals in different tissues or areas of a tissue are indeed due to real differences in lipid concentrations, or rather to changes in the environment from which the lipids are extracted during the desorption process [12, 13]. Because of the nature of biological tissues, lipid signals might be affected by the presence of and/or interactions with other lipids, proteins, salts, and many other complex biological materials, such as nucleic acids or porphyrins. Given the tissue specificity of both the composition and the regional and subcellular organization of biomolecules, ionization efficiency is likely to be tissue specific.
Improvements in sample preparation protocols for IMS [14] and the employment of new matrices [6] have made it possible to obtain spectra directly from tissue sections with s/n ratios and resolutions that are almost comparable to those obtained from lipid extracts [10]. Such improvements allow us to determine if lipid extraction procedures preserve lipid concentration in the tissues, or if this is modified by differences in the lipids’ physicochemical properties, which result in different extraction ratios.
The present study also involves a proposal for the identification of a number of lipid molecular species in frontal cortex of human brain from postmortem tissue obtained from autopsies, and provides information on the relative abundances and distribution of the proposed lipid species. To carry out a study of this nature, it is necessary to ensure that the original lipid composition is preserved during the extraction protocol, and that mass distribution images reflect the real distribution of the different lipid species in the tissue. In addition, it is necessary to implement a protocol that allows one to obtain a clear correlation between signal distribution and density distribution in IMS experiments. The employment of this technique will be of enormous value in clarifying the physiological role of lipids, not only in the CNS but also in pathological conditions, including neurodegenerative diseases that affect an increasing number of patients worldwide.
The availability of data from both extracts and IMS of sections of postmortem human brain tissue helped us to address some of the questions raised above. We prepared separate lipid extracts from gray and white matter of human frontal cortex that were compared with consecutive slices from the same samples. The anatomical distribution was analyzed with counter-stained slices and MALDI-IMS maps of molecular densities at different resolutions, yielding density maps for several phospholipid species, mainly glycerophosphocholines (GPCho), glycerophosphoethanolamines (GPEtn) and sphingomyelins (SPM). Additionally, the absence of ion suppression between lipids was confirmed by the analysis of lipid classes separated by thin-layer chromatography (TLC-MALDI). The present study offers new and important information on the lipidome of human brain, which helps to increase the limited knowledge of this subject [6, 15–20]. In addition, the procedure used, which involves comparing the results from three different types of samples—extracts of all lipid classes (lipid extracts), individual lipid classes (TLC), and original tissue from consecutive human brain cortex slices—provides a suitable approach to test the ability of IMS to determine the anatomical distribution of lipid species in tissues at a resolution of a few μm (between 50 and 200 μm), a necessary step prior to elucidating their physiological role.
2 Experimental Procedures
2.1 Brain Sectioning, Lipid Extraction, and Separation
Samples were obtained from the Brain Bank of the Institute of Neuropathology (University Hospital Bellvitge, Barcelona). Human brains were obtained at autopsy after prior informed consent and with the institutional approval of the ethics committees of the University Hospital Bellvitge and the University of the Basque Country. The samples were obtained from two male subjects (54 and 64 years old) who died suddenly after cardiac infarction and who had shown no evidence of neurological or metabolic disease. The neuropathological study disclosed no abnormalities in the brain. The intervals between death and autopsy were 3 and 11 h, respectively. Following autopsy, the brain samples were immediately frozen at −80 °C. The frozen tissue was brought to −25 °C, and 20 μm thick slices were obtained in a cryostat (Microm, HM550). Two consecutive sections were mounted on conducting slides (standard size slides for a microscope, Bruker Daltonics) and kept at −25 °C until they were introduced into a MALDI metal holder (Bruker Daltonics). A section consecutive to those on the slides was counter-stained with thionine, and a human brain atlas was used for the identification of the different cortical layers analyzed by MS.
Approximately 100 mg of gray and white matter samples from the same tissue were dissected and homogenized at 4 °C in 5 vol ice-cold phosphate buffer solution pH 7.4 using a Polytron (Kinematica) homogenizer. The protein content of the samples was comparable and was estimated in 5 mg protein/100 mg tissue using Pierce reagent [21]. Lipids were exhaustively extracted from homogenates by a modified Bligh and Dyer procedure described earlier [22], with the extraction efficiency for the lipid classes averaging 95–99% according to radiometric analysis performed in parallel. The solvent—chloroform—was removed in a Savant SpeedVac concentrator–evaporator, the lipids were dissolved in toluene, and the extracts were stored under an N2 atmosphere at −80 °C until analyzed. Due to the considerable diversity of lipid polarity, we used six polar and apolar radiolabeled lipid species to evaluate lipid recovery, which were obtained from PerkinElmer NEN Radiochemicals (PerkinElmer, Waltham, MA, USA) and were used without further purification: [1-14C]dipalmitoyl phosphatidylcholine, sphingomyelin [choline, methyl 14C], glycerol tri[1-14C]oleate, [1-14C]dioleyl phosphatidylserine, cholesteryl [1-14C]oleate, and [1-14C] cholesterol. Typically, 10,000 dpm were added to each homogenate, and aliquots of the chloroformic phase were counted for radioactivity in a Packard Tri-Carb liquid scintillation counter.
TLC separation of the lipid extracts from gray and white matter was carried out on silica gel plates (aluminum sheets, 5 × 7.5 cm, Merck 60 RP-18 F254S) as described by Murata et al. [23] using the following mobile phases: chloroform/methanol/water (65:40:5, v/v/v), ethyl acetate/isopropanol/methanol/ethanol/chloroform/0.25% KCl (35:5:15:20:22:9, v/v/v/v/v/v), n-heptane/diisopropyl ether/acetic acid (7:3:0.2, v/v/v), and n-heptane. The correct separation of the different classes of lipids was confirmed by visualization with UV light (254 nm), using primuline. Two microliters of sample were used on each analysis (an overall lipid mass of about 20 μg).
2.2 Sample Preparation for MALDI-TOF, IMS, and TLC-MALDI
A saturated solution of 2-mercaptobenzothiazole (MBT) (Sigma-Aldrich, St. Louis, MO, USA) in methanol was employed as matrix solution for both extracts and tissues, as described previously [9]. Lipid extracts with a total lipid concentration of ∼10 mg/ml were mixed with the matrix solution in a 1:10 (v/v) ratio in order to adjust the total lipid concentration to ca. 1 mg/ml. One microliter of the mixture was spotted into each well of the stainless steel target plate and co-crystallized by evaporation. To obtain the constants of the external calibration polynomial, 36 peaks in the spectrum of the polyethylene glycol–sodium adduct [PEG-Na]+ in the range 400–2,000 Da, obtained with the target plate at various positions around the sample, were used and fitted to 15–25° polynomials, with the 19° polynomial providing the best fit. In this way, an ∼10 ppm accuracy in mass determination was achieved [9].
Spectra were acquired in positive linear and reflectron modes with the aid of a Bruker Reflex IV time-of-flight (TOF) mass spectrometer (Bremen, Germany) equipped with an N2 laser (337 nm, 9 Hz, 100 μJ max energy). Typical settings were: laser energy at 45%, repulsion plate 20.0 kV, extraction plate 16.35 kV, lens voltage 9.9 kV, reflectron field 23.0 kV, detector gain ×5, and extraction pulse delay 200 ns. Nine hundred shots were accumulated for each spectrum of the extracts, and at least ten replicates of each extract were analyzed to assess the reproducibility of the spectra.
In the IMS experiments, a uniform coat of matrix was applied using a sprayer (DESAGA, model SG1B) loaded with an MBT/methanol saturated solution. The matrix was sprayed from a distance of 50 cm, with the sample being exposed only when the sprayer was in the stationary regimen. Up to 20 spraying/drying cycles were employed in order to obtain a uniform layer. With this technique, it was demonstrated that resolutions of at least 50 μm can be achieved [9].
The acquisition area was defined with the CreateTarget computer software [24], which generates files in a format compatible with the Bruker Reflex IV spectrometer. Thirty shots at each location were accumulated to construct each individual spectrum, with a total acquisition time of 2–9 h, depending on the size of the scanned area. The spatial resolution—taken as the distance between adjacent points where the spectra were recorded—of the mass images shown in the present study varies from 50 to 200 μm. As there may be potential displacement in the masses along the tissue due to irregularities of the surface and other factors, acquired spectra were aligned by maximizing the correlation with the overall averaged spectrum, normalized using the total ion current, and exported to the computer program Histomass™ in a compatible format using a script written in Mathematica™ 6.0. The data analysis and visualization were carried out with Histomass™ (http://www.noraybio.com/en/histomass.asp).
In the TLC-MALDI experiments, the plates were covered with a saturated solution of MBT that prevented lipid migration, and then placed in the spectrometer using a TLC-MALDI adaptor (Bruker). A much larger amount of matrix than used in the experiments with extracts or tissues was required in order to obtain an intense signal. The separated lipid bands were scanned in the same way as the tissue sections; we defined a rectangular geometry containing the species of interest and scanned it, recording spectra every 100 μm, with the laser energy set at 70% and using reflectron positive detection. The spectra of three separate phospholipid classes, GPCho, GPEtn and SPM, were identified and averaged using the software described for IMS and the probabilistic latent semantic analysis (PLSA) algorithm described below.
2.3 Probabilistic Latent Semantic Analysis (PLSA)
The data from the IMS experiments were analyzed using a modified version of the PLSA algorithm [25], programmed using Mathematica™ 6.0. The program allows us to automatically cluster the spectra obtained from the tissue according to their similarity. Using this program, three groups were obtained without the intervention of the user: one clustering all the spectra recorded in white matter, another one containing all the spectra recorded in gray matter, and a third one with the spectra recorded outside the tissue. The spectra selected in this way were employed to obtain representative spectra of gray and white matter. A detailed description of the algorithm employed can be found in [26, 27]. Briefly, the PLSA algorithm was initially developed for automated text analysis. When applied to the analysis of IMS data, the documents are replaced by spectra, words by m/z channels, and topics by tissues. Thus, each tissue is characterized by a specific distribution [25]. The advantage with respect to other algorithms for the automated analysis of IMS data, like the commonly employed principal component analysis, is that the resulting values are normalized and non-negative. The interested reader is redirected to [25] for more information on its advantages and application.
2.4 Peak Assignment
The human brain samples analyzed in this study are quite complex, containing a large number of lipids that share similar masses. Without using MS/MS it is not possible to discriminate among chemical variants of lipids with identical numbers of acylic carbons and double bonds; that is, with identical masses. Therefore, the identities of the acyl chains and the positions of their double bonds could not be specified in this study. The assignment of lipid species was facilitated by the use of a database, which we created based on the specific lipid composition of the tissues. The experimental values of the peaks’ positions were compared with this database and with the data in Lipid MAPS (http://www.lipidmaps.org/) and Madison Metabolomics Consortium (http://mmcd.nmrfam.wisc.edu/) databases, using a 50 ppm mass accuracy as the tolerance window. Such a large window was chosen so as not to miss any likely candidate, as we rely on the m/z value to perform the assignment. Nevertheless, since the accuracy achieved with our spectrometer (∼10 ppm, 1σ) does not allow us to unequivocally identify many of the peaks present in the spectrum, only the assignments that are in accordance with those reported by previous works are accepted and presented in this study. All of the lipids identified in this way are collected in Table S1 of the Electronic Supplementary Material (ESM). The glycerolipid species numbers (x:y) denote the total lengths and the number of double bonds of the acyl chains, while the sphingolipid and sulfatide species numbers correspond to the length and number of double bonds of the acyl chain added to those of the attached sphing-4-enine (d18:1) or sphinganine (d18:0) base.
3 Results and Discussion
Serial sections from human brain samples of the frontal cortex (area 8) were obtained and two of the sections were covered with MBT, following the procedure described above, and subjected to IMS. A third section was stained to distinguish the different layers of the frontal cortex and the white matter. Simultaneously, two different portions (∼5 mg protein/100 mg tissue) of the sample were processed (following the methodology described) to extract the lipids from gray and white matter separately and subjected to lipidomic analysis.
Figure 1 shows a comparison between the stained section and the images obtained by using the IMS method; each image shows the distribution along the tissue of a different species (= mass channel), and it is constructed to represent the integral of a single mass channel against the spatial coordinates at which the spectra were acquired. In this figure, some representative lipid species are shown, together with the proposed assignment. Another additional image represents the three main components obtained from the LSA-PLSA analysis, and this is shown in three different colors. The algorithm groups the spectra into three differentiated regions: one corresponding to the areas where there is only matrix (in white), another area corresponding to the gray matter (in blue), and a third region corresponding to the white matter (in red). Accordingly, the IMS images, in black-green-yellow scale, show that the diverse lipid molecular species found in human brain cortex exhibit a different and specific distribution; while some lipid species are evenly distributed along the whole section, like 760.59 (assigned as GPCho 34:1 + H+, see Table S1 of the ESM) or 753.58 (SPM 36:1 + Na+), others show a strong propensity to appear either in white matter, like 786.60 (GPCho 36:2 + H+), 788.61 (GPCho 36:1 + H+) and 746.59 (GPEtn 36:1 + H+), or in gray matter, like 734.56 (GPCho 32:0 + H+) and 828.55 (GPCho 38:6 + Na+). Two different areas of the section in Figure 1 were scanned at a resolution of 50 μm, which should be enough to distinguish between the six different layers of the cortex (Figure S1 of the ESM); slight differences in lipid density along the cortex are apparent. The cortical layers differ mainly in the type of neurons and fiber projections (dendrites and axons). The differences in lipid composition between these layers (I–VI) are not as clear as they are for white and gray matter.
Distribution of some representative lipid species in a tissue section of frontal cortex from human brain, together with a consecutive stained section and the representation of the principal components identified by LSA-PLSA. The section was 20 × 15 mm and the spectra were recorded with 200 μm spacing, resulting in 7500 spectra. Thirty shots were accumulated at each point with the spectrometer working in reflectron mode. Laser power was set at 55%. Thionine staining is shown with a bar of 0.5 cm to illustrate the dimensions of the section. The average spectra recorded by accumulating 900 shots along each type of matter are shown in figure 2, while the average of each of the components is depicted in figure 3. See Table S1 of the ESM for the assignment of each m/z to a lipid molecule
The same section was employed to record averaged spectra over white and gray matter separately (900 shots per spectrum, Figure 2), prior to recording the results that are shown in Figure 1. This means that during the ca. 48 h that the section was inside the spectrometer (which was the time taken to record the three scans and the averaged spectra) there was no degradation of the lipids or variation in the signal intensity due to matrix sublimation. This result is demonstrated by comparing the IMS images in Figures 1 and S1 with those in Figure S2 of the ESM, which were obtained from the sequential section scanned at 100 μm resolution, and by comparing the spectra obtained from the tissue in each IMS experiment that were almost identical (data not shown).
Comparison between mass spectra obtained from extracts of lipids from human gray and white brain matter (upper traces), and the average spectra recorded directly from the tissue slice, selecting only either white or gray matter (lower traces). Each spectrum is the average of 900 shots along the tissue, with the laser power at 55% and using MBT matrix. Detection is performed in positive reflectron mode
The assignment of the mass channels represented in Figure 1 was performed by comparing the spectra recorded from the tissue with the assigned spectra of the extract, which were recalibrated with a polymer following the external calibration protocol described in the “Experimental Procedures” section. The spectra resulting from gray matter are very similar in terms of the relative abundances of the different lipid species, and this was also the case for the white matter. The intensity was higher in the extracts, but they correspond to the accumulation of 900 shots instead of the 30 used in each slice coordinate.
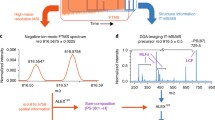
Figure 2 shows a comparison between the 710–840 Da regions (where the strongest signals appear) of the spectra obtained from the extracts of gray and white matter (upper traces), and the averaged spectra of 900 shots (in groups of 30 shots per location) recorded directly from gray and white matter from tissue slices (lower traces). The s/n ratio of the spectra recorded directly from the tissue is almost as good as that of the spectra from the extract, although the signal intensity is approximately one order or magnitude lower: 6500 counts compared with over 40,000 counts.
Additionally, the average of the spectra classified using the LSA-PLSA probabilistic analysis shows an improved s/n ratio, equivalent to that obtained from the extracts (Figure 3). The comparison between the gray and white matter spectra from extracts shows, for example, changes in the relative intensities of GPCho 32:0, GPCho 38:6, and GPEt 38:6. Meanwhile, other species such as GPCho 36:2 are found predominantly in white matter. Nevertheless, we did not find that any of the identified species were exclusively located in one of these main regions or types of matter of the human brain cortex. The differences correlate with different density distributions along the tissue. For example, the peak at 734.560 Da corresponding to GPCho 32:0 + H+ is more intense in the spectrum from the gray matter extract than in the white matter extract, in close agreement with the IMS presented in Figure 1, and with the differences observed between the averaged spectra recorded directly from the tissue. Also, the peak at 766.57 Da, assigned as GPEtn 38:5 + H+, is more intense in the white matter spectra (both in extracts and tissue) than in the corresponding gray matter spectrum. A detailed view of that region of the mass spectrum is offered in Figure S3 of the ESM. From the above, one can conclude that the distribution maps obtained using IMS correspond to real variations in the lipid concentrations, and that they are not due to changes in the environmental features of the anatomical areas that are analyzed.
Comparison between the average of the 710–840 Da regions of the spectra corresponding to the two components found in the LSA-PLSA analysis of the section in Fig. 1. The inset shows a detail of the 803–840 Da region, in which the s/n ratio can be appreciated
The IMS experiments carried out directly in the tissue slices offer a second advantage: the absence of “hot spots” that can influence the relative intensities of the peaks. Furthermore, if an algorithm like the LSA-PLSA used here is employed, it is possible to identify not only the spectra of each region and to average them, but also to identify different regions, based on the most common molecules that are present, which is especially interesting in studies related to pathological processes. The PLSA results are spectra representative of the relative concentrations of the molecules in the tissue and a greatly improved s/n ratio. Thus, the average spectra in Figure 3 show a s/n ratio comparable to, or even better than, those obtained from extracts (Figure 2). The inset in Figure 3 shows a detailed view of the 803–840 Da region of the average spectrum over the white matter to emphasize this point. The comparison between the spectra in Figures 2 and 3 makes it clear that the differences in the distributions of the different lipid species found in the IMS experiments are not due to artifacts inherent to the technique. For example, the peaks in the region around 830 Da in the average spectrum of the gray matter in Figure 3 are weaker than in white matter. The predominant peak in that region is at 826.565 Da, which is assigned to GPCho 36:1 + K+, and this is accompanied in the spectra from the tissue presented in Figure 2 by some other peaks for which no satisfactory assignment was found. In summary, most of the differences between the spectra from extracts and those from the LSA-PLSA average are probably due to differences in the abundance of adducts and to the existence of non-lipid species in the tissue slices. Consequently, a representative spectrum of each region of the tissue was obtained by applying a mathematical procedure to get the averaged spectrum.
An additional experiment was carried out to test the existence of ion suppression among lipids. It is known that the classes with quaternary amines (for example GPCho) yield a stronger signal, even at lower concentrations, than other lipid species. The different abundances of such charged species in the two types of tissues (white or gray matter) in our samples could affect the detection of the rest of the lipids, even if they are present in similar abundances. Thus, we applied a separation protocol (TLC) to samples of lipid extracts from white and gray matter and then recorded the MALDI spectra of three main classes of lipids—GPCho, GPEtn, and SPM—on the TLC plate. The resulting spectra (Figures S3, S4 and S5 of the ESM) are in agreement with the IMS results. For example, a clear increase in the intensity of the GPCho 32:0 is observed from white to gray matter, while the GPCho 36:1 exhibits the opposite pattern (Figure S3 of the ESM). Also, it is clear from Figure S4 of the ESM that the overall amount of GPEtn is considerably greater in white matter than in gray matter. Finally, while the SPM 36:1 concentration does not vary greatly from white to gray matter, there is a subtle decrease in SPM 34:1, in agreement with that observed in the IMS experiments.
The question that still remains unclear is if the conclusions reached about the validity of the IMS images obtained from lipids in the present work hold for other type of molecules and tissues. Brain tissue is very rich in lipids (which account for about 60% of its dry weight), so variations in the concentrations of other components, like proteins, may not result in significant changes in the intensity of the peaks, as demonstrated throughout the present paper. However, it cannot be ruled out that in the particular case of protein-rich tissues, variations in the concentrations of certain proteins or other major components may result in changes in the relative intensities of the lipid peaks. Interactions between proteins and lipids are known to be tissue specific, so studies that are specially designed for this purpose are necessary.
Despite the fact that little is known about human lipid species distribution and composition, our results are in accordance with others previously reported. In this context, it has been demonstrated [19, 28, 29] that human brain is mainly composed of cholesterol and phospholipids. Among the latter, the most abundant classes are GPCho, GPEtn and SPM, which are precisely the species detected most prominently in the present work. Using LC/ESI, Isaac et al. determined the presence of GPCho 38:6, 36:4, 34:1 and 36:1 and SPM 34:1, 36:1 and 42:2 species in human mesencephalon [29]. All of them were detected in the present study, and the GPCho species exhibit a strong signal. Moreover, O’Brien et al. determined the relative abundance of some phospholipid species between gray and white matter in human brain [19], concluding that the SPM species are more abundant in white matter. We also observe a tendency of some of the SPM species to be located in white matter (Figure 1 and Figures S1 and S2 of the ESM). Söderberg et al. reached the same conclusion, but in addition, they estimated a phosphatidylethanolamine/phosphatidilcholine (GPEtn/GPCho) ratio of ∼1.7 for white matter and of 0.8 for gray matter [30], in agreement with the present work, in which GPEtn is more abundant in white matter.
In a recent paper, Goto-Inoue et al. [31] applied a TLC-Blot-MALDI-IMS procedure to lipid extracts from human frontal gyrus white and gray matter and hippocampus white and gray matter, and analyzed the differences in lipid composition for GPCho and SPM species. They found that the main GPCho peak corresponds to the 34:1 species, and also identified GPCho 32:0 and 36:1 in the mixture. They also report the identification of SPM 34:1, 36:1 38:1 and 42:2 species in the extracts. The analysis of the relative concentrations of SPM in white and gray matter led them to conclude that SPM 36:1 is present at similar abundances in both types of matter, while SPM 38:1 is more abundant in gray matter and SPM 42:2 is more abundant in white matter. If we compare the results from Goto-Inoue et al. with those reported in this paper (Figures S3, S4 and S5 of the ESM), they are similar except for the SPM 38:1 white/gray relative abundance. In our samples, we find the opposite trend to that reported by Goto-Inoue. A possible reason for this difference may be that while they study the hippocampus, our samples are from the frontal cortex.
Some recent studies using MALDI-MS and MALDI-IMS have determined the distributions of several lipid species in rat brain [9, 32–35]. In all these studies, a difference in lipid abundance between white and gray matter has been observed. For example, Woods et al., using DHA matrix, obtained spectra from cerebellar cortex and peduncule (i.e., gray and white matter) of rats, and among the species identified, those yielding the strongest signals were GPCho 30:0, 34:1 and 36:1 and SPM 36:1, which are also the species that give the strongest signal in the spectra shown in the present study. Furthermore, their study points to a higher abundance of GPCho 32:0 in gray matter, while GPCho 36:1 is more abundant in white matter. We observed the same relative abundance in the human samples, as depicted in Figure 1 and Figures S1 and S2 of the ESM. Jackson et al. also used the ion mobility method to study some additional cerebral areas [34]; [36], observing a different distribution of diverse phospholipid species. For example, while GPCho 34:1 is distributed uniformly along the tissue, GPCho 36:1 is more abundant in white matter and GPCho 38:6 is more abundant in gray matter, as has also been observed in the present work. The same tendency was observed by Cha and Yeung using colloidal graphite as matrix [35]. In summary, all of the above cited studies report that while some lipid species are evenly distributed, others are more abundant in one type of matter, and although different techniques and matrices were used, their results are in agreement with the results we have obtained.
The present study proposes a distribution of several lipid species within white and/or gray matter of the human frontal cortex, area 8. The differences in the relative abundances of each specific lipid molecule must be related to different physiological roles. In this sense, there are virtually no neurons in white matter, which is mainly formed by glial cells and the “wiring” of the brain, the axons, that are isolated by the myelin envelopment enriched in saturated fatty acids. Moreover, it is known that the lipid/protein ratio is higher for white matter, and that glycerophosphoinositols or GPCho are not only essential for the integrity of the membranes, but also for intracellular signaling processes [37]. However, the anatomical localization of precise molecules by IMS in the brain is already raising new questions related to what this specific localization means in relation to the functions of specific lipids in the brain and different diseases which affect the CNS.
4 Conclusions
In this study we analyzed the lipid composition of the frontal cortex (area 8) from postmortem human brain samples, using both lipid extracts and IMS on frozen sections. The comparison between spectra from extracts and tissue slices points to a very efficient extraction protocol that does not modify the original lipid composition. There is a clear difference in lipid distribution between gray and white matter, both in extracts and in the tissue, demonstrating that the density maps obtained from IMS experiments and by probabilistic analysis correspond to real differences in the lipid compositions of the different areas of the tissue. This is further confirmed by the spectra recorded from some lipid classes that were separated by using TLC. Such a conclusion opens the field to applying this IMS approach in the study of lipid distribution in different kinds of tissue samples, including frozen samples from human autopsies.
Abbreviations
- CNS:
-
Central nervous system
- GPCho:
-
Glycerophosphocholines
- GPEtn:
-
Glycerophosphoethanolamines
- GPG:
-
Glycerophosphoglycerols
- GPIns:
-
Glycerophosphoinositols
- GPSer:
-
Glycerophosphoserines
- IMS:
-
Imaging mass spectrometry
- MALDI:
-
Matrix-assisted laser desorption/ionization
- MBT:
-
2-Mercaptobenzothiazole
- MS:
-
Mass spectrometry
- PLSA:
-
Probabilistic latent semantic analysis
- SPM:
-
Sphingomyelins
- TLC:
-
Thin-layer chromatography
- TOF:
-
Time of flight
References
Schiller, J., Suss, R., Fuchs, B., Muller, M., Zschornig, O., Arnold, K.: MALDI-TOF MS in lipidomics. Front. Biosci. 12, 2568–2579 (2007)
Carrasco-Pancorbo, A., Navas-Iglesias, N., Cuadros-Rodriguez, L.: From lipid analysis towards lipidomics, a new challenge for the analytical chemistry of the 21st century. Part 1: Modern lipid analysis. Trends Anal. Chem. 28, 263–278 (2009)
Dennis, E.A.: Lipidomics joins the omics evolution. Proc. Natl. Acad. Sci. U.S.A. 106, 2089–2090 (2009)
Bisogno, T., Piscitelli, F., Di Marzo, V.: Lipidomic methodologies applicable to the study of endocannabinoids and related compounds: Endocannabinoidomics. Eur. J. Lipid Sci. Technol. 111, 53–63 (2009)
Han, X.L., Jiang, X.T.: A review of lipidomic technologies applicable to sphingolipidomics and their relevant applications. Eur. J. Lipid Sci. Technol. 111, 39–52 (2009)
Fuchs, B., Schiller, J.: Application of MALDI-TOF mass spectrometry in lipidomics. Eur. J. Lipid Sci. Technol. 111, 83–98 (2009)
Caprioli, R.M., Farmer, T.B., Gile, J.: Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal. Chem. 69, 4751–4760 (1997)
Caprioli, R.M., Farmer, T.B., Zhang, H.Y., Stoeckli, M.: Molecular imaging of biological samples by MALDI MS. Abstr. Pap. Am. Chem. Soc. 214, 113 (1997). ANYL
Astigarraga, E., Barreda-Gomez, G., Lombardero, L., Fresnedo, O., Castano, F., Giralt, M.T., Ochoa, B., Rodriguez-Puertas, R., Fernandez, J.A.: Profiling and imaging of lipids on brain and liver tissue by matrix-assisted laser desorption/ionization mass spectrometry using 2-mercaptobenzothiazole as a matrix. Anal. Chem. 80, 9105–9114 (2008)
Chen, R.B., Hui, L.M., Sturm, R.M., Li, L.J.: Three dimensional mapping of neuropeptides and lipids in crustacean brain by mass spectral imaging. J. Am. Chem. Soc. Mass Spectrom. 20, 1068–1077 (2009)
Trim, P.J., Atkinson, S.J., Princivalle, A.P., Marshall, P.S., West, A., Clench, M.R.: Matrix-assisted laser desorption/ionisation mass spectrometry imaging of lipids in rat brain tissue with integrated unsupervised and supervised multivariant statistical analysis. Rapid Commun. Mass Spectrom. 22, 1503–1509 (2008)
Murphy, R.C., Hankin, J.A., Barkley, R.M.: Imaging of lipid species by MALDI mass spectrometry. J. Lipid Res. 50, S317–S322 (2009)
Heeren, R.M.A., Smith, D.F., Stauber, J., Kukrer-Kaletas, B., MacAleese, L.: Imaging mass spectrometry: hype or hope? J. Am. Soc. Mass Spectrom. 20, 1006–1014 (2009)
Hankin, J.A., Barkley, R.M., Murphy, R.C.: Sublimation as a method of matrix application for mass spectrometric imaging. J. Am. Soc. Mass Spectrom. 18, 1646–1652 (2007)
Kohler, M., Machill, S., Salzer, R., Krafft, C.: Characterization of lipid extracts from brain tissue and tumors using Raman spectroscopy and mass spectrometry. Anal. Bioanal. Chem. 393, 1513–1520 (2009)
Baek, R.C., Martin, D.R., Cox, N.R., Seyfried, T.N.: Comparative analysis of brain lipids in mice, cats, and humans with Sandhoff disease. Lipids 44, 197–205 (2009)
Martinez, M., Mougan, I.: Fatty acid composition of human brain phospholipids during normal development. J. Neurochem. 71, 2528–2533 (1998)
Rouser, G., Galli, G., Kritchev, G.: Lipid composition of normal human brain and variations in various diseases. Pathol. Biol. 15, 195–200 (1967)
O’Brien, J.S., Sampson, E.L.: Lipid composition of normal human brain: gray matter white matter and myelin. J. Lipid Res. 6, 537–544 (1965)
Rouser, G., Lieber, E., Galli, C., Kritchevsky, G., Heller, D.: Lipid composition of normal human brain. Fed. Proc. 23, 228 (1964)
Palacios, L., Ochoa, B., Gomez Lechón, M.J., Castell, J.V., Fresnedo, O.: Overexpression of SND p102, a rat homologue of p100 coactivator, promotes the secretion of lipoprotein phospholipids in primary hepatocytes. Biochim. Biophys. Acta. 1761, 698–708 (2006)
Ruiz, J.I., Ochoa, B.: Quantification in the subnanomolar range of phospholipids and neutral lipids by monodimensional thin-layer chromatography and image analysis. J. Lipid Res. 38, 1482–1489 (1997)
Murata, N., Sato, N., Omata, T., Kuwabara, T.: Separation and characterization of thylakoid and cell-envelope of the blue-green alga (Cyanobacterium) Anacystis nidulans. Plant Cell Physiol. 22, 855–866 (1981)
Clerens, S., Ceuppens, R., Arckens, L.: Create target and analyze this! new software assisting imaging mass spectrometry on Bruker Reflex IV and Ultraflex II instruments. Rapid Commun. Mass Spectrom. 20, 3061–3066 (2006)
Hanselmann, M., Kirchner, M., Renard, B.Y., Amstalden, E.R., Glunde, K., Heeren, R.M.A., Hamprecht, F.A.: Concise representation of mass spectrometry images by probabilistic latent semantic analysis. Anal. Chem. 80, 9649–9658 (2008)
Ding, C.H.Q.: A probabilistic model for Latent Semantic Indexing. J. Am. Soc. Inf. Sci. Tech. 56, 597–608 (2005)
Ding, CHQ. A similarity-based probability model for latent semantic indexing. In: Sigir’99: Proceedings 22nd International Conference on Research and Development in Information Retrieval (1999) pp 58–65
Rouser, G., Fleischer, S., Yamamoto, A.: Two dimensional thin layer chromatographic separation of polar lipids and determination of phospholipids by phosphorus analysis of spots. Lipids 5, 494–496 (1970)
Isaac, G., Bylund, D., Mansson, J.E., Markides, K.E., Bergquist, J.: Analysis of phosphatidylcholine and sphingomyelin molecular species from brain extracts using capillary liquid chromatography electrospray ionization mass spectrometry. J. Neurosci. Meth. 128, 111–119 (2003)
Soderberg, M., Edlund, C., Kristensson, K., Dallner, G.: Lipid compositions of different regions of the human brain during aging. J. Neurochem. 54, 415–423 (1990)
Goto-Inoue, N., Hayasaka, T., Taki, T., Gonzalez, T.V., Setou, M.: A new lipidomics approach by thin-layer chromatography-blot-matrix-assisted laser desorption/ionization imaging mass spectrometry for analyzing detailed patterns of phospholipid molecular species. J. Chromatogr. A 1216, 7096–7101 (2009)
Woods, A.S., Jackson, S.N.: Brain tissue lipidomics: direct probing using matrix-assisted laser desorption/ionization mass spectrometry. AAPS J. 8, E391–E395 (2006)
Jackson, S.N., Wang, H.Y.J., Woods, A.S.: In situ structural characterization of phosphatidylcholines in brain tissue using MALDI-MS/MS. J. Am. Soc. Mass Spectrom. 16, 2052–2056 (2005)
Jackson, S.N., Ugarov, M., Egan, T., Post, J.D., Langlais, D., Schultz, J.A., Woods, A.S.: MALDI-ion mobility-TOFMS imaging of lipids in rat brain tissue. J. Mass Spectrom. 42, 1093–1098 (2007)
Cha, S.W., Yeung, E.S.: Colloidal graphite-assisted laser desorption/ionization mass spectrometry and MSn of small molecules. 1. Imaging of cerebrosides directly from rat brain tissue. Anal. Chem. 79, 2373–2385 (2007)
Jackson, S.N., Wang, H.Y.J., Woods, A.S.: In situ structural characterization of glycerophospholipids and sulfatides in brain tissue using MALDI-MS/MS. J. Am. Soc. Mass Spectrom. 18, 17–26 (2007)
Farooqui, A.A., Horrocks, L.A., Farooqui, T.: Glycerophospholipids in brain: their metabolism, incorporation into membranes, functions, and involvement in neurological disorders. Chem. Phys. Lipids 106, 1–29 (2000)
Acknowledgments
This study was supported by the Spanish Ministry of Education and Science (SAF2007-60211), the Basque Government (IT-325-07, IT-336-10, IT-440-10 and SAI07/46), Biscay County Council (DIPE07/05), Carlos III Health Institute (FIS PI070628), and UPV/EHU GIU07/50. IM and AV are recipients of UPV/EHU graduate fellowships. GB is supported by a grant from the UPV/EHU Researchers Specialization Program.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Additional information as noted in text. This material may be found in the online version at …
ESM 1
(DOC 5478 kb)
Rights and permissions
About this article
Cite this article
Veloso, A., Astigarraga, E., Barreda-Gómez, G. et al. Anatomical Distribution of Lipids in Human Brain Cortex by Imaging Mass Spectrometry. J. Am. Soc. Mass Spectrom. 22, 329–338 (2011). https://doi.org/10.1007/s13361-010-0024-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-010-0024-5