Abstract
RNA interference (RNAi)-mediated gene silencing offers a novel treatment and prevention strategy for human immunodeficiency virus (HIV) infection. HIV was found to infect and replicate in human brain cells and can cause neuroinfections and neurological deterioration. We designed dual-antibody-modified chitosan/small interfering RNA (siRNA) nanoparticles to deliver siRNA across the blood-brain barrier (BBB) targeting HIV-infected brain astrocytes as a strategy for inhibiting HIV replication. We hypothesized that transferrin antibody and bradykinin B2 antibody could specifically bind to the transferrin receptor (TfR) and bradykinin B2 receptor (B2R), respectively, and deliver siRNA across the BBB into astrocytes as potential targeting ligands. In this study, chitosan nanoparticles (CS-NPs) were prepared by a complex coacervation method in the presence of siRNA, and antibody was chemically conjugated to the nanoparticles. The antibody-modified chitosan nanoparticles (Ab-CS-NPs) were spherical in shape, with an average particle size of 235.7 ± 10.2 nm and a zeta potential of 22.88 ± 1.78 mV. The therapeutic potential of the nanoparticles was evaluated based on their cellular uptake and gene silencing efficiency. Cellular accumulation and gene silencing efficiency of Ab-CS-NPs in astrocytes were significantly improved compared to non-modified CS-NPs and single-antibody-modified CS-NPs. These results suggest that the combination of anti-Tf antibody and anti-B2 antibody significantly increased the knockdown effect of siRNA-loaded nanoparticles. Thus, antibody-mediated dual-targeting nanoparticles are an efficient and promising delivery strategy for inhibiting HIV replication in astrocytes.

Graphic representation of dual-antibody-conjugated chitosan nanoparticles for the targeted delivery of siRNA across the blood-brain barrier (BBB) for inhibiting HIV replication in astrocytes. a Nanoparticle delivery to the BBB and penetration. b TfR-mediated transcytosis of nanoparticles across the epithelial cells. c B2R-mediated endocytosis of nanoparticles in astrocytes. d The molecular interactions between HIV-1 Tat protein and Cyclin T1 and Tip110 cellular proteins. e A schematic representation of chitosan nanoparticles with its components. RNAPII RNA polymerase II, TAR transactivation response RNA element, LTR long terminal repeat, Ab antibody, CS chitosan, TPP tripolyphosphate
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human immunodeficiency virus (HIV) infection of the central nervous system (CNS) occurs in majority of the patients with AIDS, particularly when the immune system is weak that it can no longer fight off the virus or other threatening infections [1, 2]. HIV invades the CNS within the first few months of infection even when HIV levels are undetectable in the blood. The virus begins to replicate in the CNS independently from viral populations in the blood causing a variety of HIV-associated neurological dysfunctions (HANDs) such as neuropathy, memory loss, neurosyphilis, vacuolar myelopathy, HIV-associated dementia, motor control deficits, and progressive multifocal leukoencephalopathy [3–5]. HIV can also lead to other types of nerve damage outside of the brain, including peripheral neuropathy [4, 6, 7]. The major target cells for HIV-1 infection in the CNS include microglia, macrophages, and astrocytes, but rarely neurons [8, 9]. Astrocytes constitute the most abundant cell type in the brain, greatly outnumbering microglial cells and neurons. The potential of astrocytes to provide HIV sanctuary from anti-viral attacks makes astrocytes important cellular targets for HIV [10]. Moreover, the large number of astrocytes in the brain and their extremely important roles in this organ strongly support the notion that HIV infection of astrocytes contributes to HIV-associated neuropathogenesis. Unlike microglia that express CD4 and chemokine co-receptors CCR5 and CCR3 for HIV infection, astrocytes do not have a detectable level of CD4 receptor expression [11]. HIV infects brain astrocytes using a CD4-independent mechanism, establishing chronic infection with very limited virus production and predominant expression of non-structural HIV components [11, 12]. The virus may then use the astrocytes as a reservoir as it is non-productive once inside. Much progress has been made in terms of the mechanisms of non-productive HIV-1 replication in astrocytes. The unique features of HIV-1 infection of astrocytes have made astrocytes an excellent model for studying molecular mechanisms of CD4-independent HIV-1 entry and regulation of HIV-1 replication.
The success of gene therapy is highly dependent on the delivery vector [13, 14]. Targeted delivery of therapeutic genes such as small interfering RNA (siRNA) to the diseased brain is of great interest [15, 16]. Despite promising features of RNA interference (RNAi) that may be clinically useful, the use of siRNA has several limitations, such as rapid degradation by nucleases, limited stability in the bloodstream, and low delivery efficiency to target cells [17]. Moreover, delivery of drug especially siRNA across the blood-brain barrier (BBB) is intrinsically very limited by the size and biochemical properties of siRNA [18, 19]. The BBB controls the passage of substances from the blood into the CNS, therefore making it difficult for therapeutics to reach the brain [18]. As a result, many researchers have developed delivery systems to overcome these limitations and to enhance targeted delivery and intracellular uptake in the brain [20, 21].
Chitosan (CS) is derived from chitin by partial deacetylation, which is the second most abundant natural polysaccharide and is composed of glucosamine and N-acetyl glucosamine residues [22]. CS has frequently been used in gene delivery applications, including siRNA delivery, due to its ability to form stable complexes with genes via electrostatic interactions between positively charged amino groups in CS and negatively charged nucleic acids [22, 23]. CS is thought to be an ideal drug delivery vehicle due to its biocompatibility, minimal immunogenicity, biodegradability, and its ability to open cellular tight junctions [24].
Drug delivery vehicles modified with ligands such as antibodies have been widely used for targeted therapy. Transferrin receptor (TfR), a type II transmembrane protein highly expressed on brain endothelial cells [25, 26], has potential for mediating entry of therapeutics into the brain [27]. Studies have shown that anti-TfR antibodies (TfR-Ab) conjugated to nanoparticles can improve targeted delivery across the BBB [28, 29]. hCMEC/D3 cell line is a widely used model of the human BBB. Transferrin receptors are highly expressed in hCMEC/D3 cells and glioblastomas, making it a popular target for drug delivery to the brain [30–33]. The bradykinin B2 receptor (B2R) is a G protein-coupled receptor, which is a surface antigen present on human brain cells and has been exploited for antibody-based targeted delivery in both preclinical and clinical studies due to rapid internalization after antibody binding. B2R is expressed in many different cells including astrocytes [34, 35] and human astrocytic tumors [36]. It has been reported that B2R is more highly expressed in glioma cells compared to normal astrocytes [37], and the level of expression of B2R may be correlated with the grade of human glioma [38].
In this study, we hypothesized that dual-antibody-conjugated CS nanoparticles could be useful for siRNA delivery, specifically for penetrating the BBB and targeting to astrocytes. Two different sequences of siRNA were loaded in the designed CS nanoparticles, which could silence the following cellular target genes: SART3 and hCycT1 [39, 40]. SART3 encodes Tip110, which regulates Tat transactivation by binding to unphosphorylated RNA polymerase II. Tat is a HIV-1 regulatory protein that is required for efficient viral transcription. hCycT1 encodes Cyclin T1, which besides CDK9, is a subunit of the human positive transcription elongation factor P-TEFb. Tat interacts with hCycT1 to activate the elongation of RNA polymerase II at the HIV-1 promoter. Therefore, the combination delivery of both siRNAs is proposed to enhance the inhibition of HIV replication by different mechanisms. Nanoparticles were prepared from CS and siRNA by a coacervation method. The various physicochemical characteristics such as size, zeta potential, and morphology were investigated. The cytotoxicity, cellular uptake, and gene silencing efficiency of antibody-conjugated CS nanoparticles were evaluated. Overall, our antibody-mediated dual-targeting nanoparticle system can efficiently penetrate across a co-culture model of the BBB in vitro, enhance siRNA uptake and messenger RNA (mRNA) knockdown in astrocytes, demonstrating its utility as a promising strategy for inhibiting HIV replication in the brain.
Materials and methods
Materials
Chitosan (CS, MW = 500 kDa, 86% deacetylation), sodium tripolyphosphate (TPP), glacial acetic acid of analytical grade, and human basic fibroblast growth factor (hbFGF) were obtained from Sigma-Aldrich (Canada). CellTiter 96® AQueous One Solution Cell Proliferation Assay Kit was obtained from Promega (USA). 1-Ethyl-3-(dimethylaminopropyl) carbodiimide (EDC) and N-hydroxysulfosuccinimide (sulfo-NHS) were purchased from G-Biosciences (USA). Eagle’s minimum essential medium (EMEM) was obtained from the American Type Culture Collection (ATCC, USA). EndoGRO-MV complete culture media kit was purchased from EMD Millipore (Canada). SYBR® green supermix reaction mixes (Quanta Biosciences), VWR-qScript™ cDNA supermix (Quanta Biosciences), and diethylpyrocarbonate (DEPC)-treated nuclease free water were purchased from VWR (Canada). Phosphate-buffered saline (PBS), trypsin-EDTA, penicillin-streptomycin solution, and fetal bovine serum (FBS) were purchased from Gibco Life Technologies (Canada). All siRNA were synthesized by Life Technologies. siRNA targeting human SART3 contains the sequence sense 5′-GGAGACAGGAAAUGCCUUATT-3′ and anti-sense 5′-UAAGGCAUUUCCUGUCUCCTT-3′. siRNA against human CycT1 consisted of the following sense 5′-UCCCUUCCUGAUACUAGAATT-3′ and anti-sense 5′-UUCUAGUAUCAGGAAGGGATT-3′. Non-targeting scrambled siRNA was used as a negative control with a sense strand consisting of 5′-UUCUCCGAACGUGUCACGUTT-3′ and an anti-sense strand of 5′-ACGUGACACGUUCGGAGAATT-3′. Fluorescent siRNA used in this study were labeled at the 5′-end of the sense strand with cyanine dye (Cy3) (Cy3-siRNA) for cellular uptake studies. Human B2R antibody was obtained from Abnova (China). Anti-TfR antibody was obtained from AbD Serotec (Canada). Anti-SART3 antibody, anti-CycT1 antibody, and anti-rabbit IgG H&L (DyLight® 488) secondary antibody were purchased from Abcam (USA). FITC-IgG antibody was used as an isotype control (Santa Cruz Biotechnology, USA).
The human cell line U138-MG was purchased from ATCC (USA) and cultured in EMEM supplemented with 10% FBS, 100 u/mg penicillin, and 100 mg/mL streptomycin at 37 °C and 5% CO2. U138-MG is a glioblastoma (astrocytoma) cell line that exhibits properties similar to astrocytes and has been widely used for studying HIV infection and replication of the brain [41, 42]. Immortalized human cerebral microvascular endothelial cell line (hCMEC/D3) was purchased from CELLutions Biosystems and cultured in EndoGRO™-MV complete media kit supplemented with 1 ng/mL hbFGF, 100 u/mg penicillin, and 100 mg/mL streptomycin at 37 °C and 5% CO2.
Preparation and characterization of CS nanoparticles
CS was dissolved in acetic acid solution (0.1 M) while stirring overnight at room temperature to obtain a 1 mg/mL stock solution and then adjusted to pH 5.0. CS/TPP nanoparticles (NPs) were prepared according to the ionotropic gelation process. In brief, 2 mL of TPP aqueous solution (0.125 mg/mL) was added dropwise to 750 μL of CS solution and stirred (800 rpm) for 30 min at room temperature to obtain blank NPs. For preparation of siRNA-loaded CS/TPP NPs (CS-NPs), 5 μL of siRNA solution (5 μg/μL) was added slowly to CS solution with stirring (800 rpm) for 20 min at room temperature, and then, TPP solution was added dropwise to the mixture with stirring (800 rpm) for another 30 min. The CS/TPP weight ratio used throughout this study was 3:1, which was obtained from the results of several trials.
Antibody conjugation to NPs was performed using the EDC/sulfo-NHS coupling reaction method. EDC (0.5 mM) and sulfo-NHS (0.25 mM) were added to a reaction mixture containing antibody (0.5 mg/mL) and NPs and stirred at room temperature. The antibody-conjugated NPs (Ab-CS-NPs) were recovered by centrifugation, washed with water (three or four times), and resuspended in distilled water.
The morphology of the NPs was observed by transmission electron microscopy (TEM) following negative staining with uranyl acetate. The hydrodynamic mean particle size and zeta potential of the NPs were measured by dynamic light scattering (DLS) using the ZetaPALS analyzer (Brookhaven). DLS was performed in water at 25 °C, pH 7.4, and at a viscosity of 0.8872 mPa•s.
In vitro cytotoxicity
Cytotoxicity of CS-NPs and Ab-CS-NPs in U138-MG and hCMEC/D3 cells were determined using the MTS assay (CellTiter 96® AQueous One Solution Cell Proliferation Assay, Promega). U138-MG and hCMEC/D3 cells were seeded in a 96-well plate at 1 × 105 cells per 100 μL medium per well for 24 h, respectively. After 24 h of incubation, the cell culture medium was replaced with 100 μL positive and negative control medium and various treatment groups. Various treatment groups including blank CS-NPs, blank Ab-CS-NPs, CS-NPs, and Ab-CS-NPs (0.1 μg siRNA per well) were diluted in cell culture medium to achieve an appropriate concentration (∼0.07 mg/mL). Cells cultured in medium alone were used as negative control. Cells treated with 1% (v/v) Triton X-100 were used as positive control for the MTS assay. Treatment groups and controls were incubated for 24 h at 37 °C.
Co-culture model
U138-MG and hCMEC/D3 cells were co-cultured to model the BBB in vitro for evaluating transport of NPs across the BBB, cellular uptake, and gene knockdown studies. hCMEC/D3 cells were seeded onto a cell culture insert consisting of a polycarbonate membrane (mean pore size 3.0 μm, effective cell growth area of membrane 0.33 cm2; Corning Inc., Life Sci., MA, USA) at a density of 5 × 104 cells/200 μL medium per well and were cultured until the formation of tight junctions was achieved. This was monitored by measuring trans-epithelial electrical resistance (TEER) daily using a volt-ohm meter (Millicell-ERS-2, Millipore, USA) attached to Endohm-12 chamber electrodes. Cell monolayers with TEER values over 150 Ω were used for the following experiments. The medium was changed in both sides every 2 days. hCMEC/D3 cell monolayers grown in the transwell chambers were transferred to a 24-well plate containing U138-MG cells, which were pre-seeded in 500 μL of complete culture medium at a density of 3 × 104 cells and then co-cultured for another day. The co-culture model was then used in the following cellular uptake and transfection studies.
Cellular uptake and targeting of nanoparticles in vitro
U138-MG or hCMEC/D3 cells were seeded in 24-well plates at a seeding density of 5 × 104 cells/well. The cells were treated with Cy3-siRNA alone, CS-NPs, anti-TfR antibody-CS-NPs [(Ab0 + Ab1)-CS-NPs], anti-B2 antibody-CS-NPs [(Ab0 + Ab2)-CS-NPs], or anti-TfR/anti-B2 dual antibody-CS-NPs [(Ab1 + Ab2)-CS-NPs] loaded with Cy3-siRNA (0.5 μg siRNA/well, ∼0.35 mg/mL NPs) for 4, 12, and 24 h. Ab0 refers to non-targeting scrambled antibody, Ab1 refers to anti-TfR antibody, and Ab2 refers to anti-B2 antibody. Trypan blue (0.4%) was used to quench extracellular fluorescence. The treated cells were then washed with PBS twice and imaged using a fluorescence microscope (Nikon TE2000). The cellular uptake of NPs in the same corresponding cell population was also analyzed by a flow cytometer (FACSCalibur, BD, USA).
The penetration and dual-targeting effect of Ab-CS-NPs were evaluated using the BBB co-culture model. The hCMEC/D3 monolayer (apical side) was treated with 300 μL of fresh medium containing CS-NPs or Ab-CS-NPs (2 μg siRNA/well). After incubation for 24 h, U138-MG cells were harvested and quantified using flow cytometry (FACS Calibur, BD, USA).
Gene knockdown in vitro
The gene knockdown efficiency of CS-NPs and (Ab1 + Ab2)-CS-NPs was evaluated using the BBB co-culture model. Culture medium (300 μL) containing NPs was added into the upper apical chamber of each transwell (2 μg siRNA/well). After incubation for 24 h, medium containing NPs was removed and replaced with equal volume of fresh complete medium, and the cells were further incubated for an additional 12 h. The cells were collected and total RNA was isolated. Real-time PCR was performed to determine the expressions of SART3 and hCycT1 mRNA level. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the housekeeping gene for normalization of results. The comparative CT (ΔΔCT) method was used to quantitate relative SART3 and hCycT1 mRNA expression levels, comparing treated samples to non-treated controls.
Inhibition of HIV-associated protein expression in U138-MG cells
HIV-associated SART3 and CycT1 protein expressions were also analyzed to confirm gene silencing in U138-MG cells. After transfection, the U138-MG cells were fixed with 4% paraformaldehyde for 20 min, permeabilized with 0.1% Triton X-100 for 10 min, and blocked with 3% BSA for 1 h. The cells were then incubated overnight with primary monoclonal anti-SART3 and anti-CycT1 antibody at 4 °C, followed by incubation with FITC-conjugated secondary antibody (1:500) for 1 h at room temperature. The cells treated with medium alone were used as negative control. The cells stained with FITC-labeled non-specific antibody were used as isotype stain control. After the final washes with PBS, the cells were mounted using an anti-fade mounting fluid and analyzed by flow cytometry (FACSCalibur, BD, USA). Cells were gated to evaluate 10,000 viable cells per experiment. The mean fluorescence intensity of fluorescence-positive cell population was determined. The extent of SART3 and CycT1 silencing with different formulations was expressed relative to the cells treated with the scrambled siRNA of the corresponding formulations.
Statistical analysis
All data are presented as means ±SD calculated from at least three independent experiments. The statistical analysis of significant differences among experimental groups was determined by a two-way ANOVA test. p < 0.05 was considered statistically significant.
Results and discussion
Preparation and characterization of nanoparticles
In this study, the CS-NPs were formulated via the ionic gelation method with TPP and siRNA. The formulation was optimized to the mass ratio of 750:250:25:5 (CS:TPP:siRNA:Ab). The particle size, polydispersity index, and zeta potential of NPs were characterized by DLS. Measurements were done in triplicate and calculated using the refractive index and viscosity of water (Table 1). The results showed that all of the NPs ranged between 150 and 250 nm in size with a positive surface charge, which is suitable for cellular uptake. Antibody conjugation efficiency was ∼70% and the density of antibody was ∼3.5 μg Ab/mg NP. The morphology of the NPs was observed and imaged by TEM (Fig. 1). The representative TEM images of NPs exhibited spherical structure with moderate and uniform particle size.
Cytotoxicity of chitosan nanoparticles
Cytotoxicity of blank CS-NPs, blank Ab-CS-NPs, CS-NPs, and Ab-CS-NPs were evaluated in U138-MG and hCMEC/D3 cells by MTS assay. The cells treated with 1% Triton X-100 were used as positive control. As shown in Fig. 2, the viabilities of both U138-MG and hCMEC/D3 cells treated with NPs were higher than 90%. There was no significant difference in cell viability between treatment groups and negative control group. There was some slight reduction in cell viability for the Ab-CS-NPs in U138-MG cells. In comparison to hCMEC/D3 cells, the U138-MG cells appeared to be more sensitive to the Ab-CS-NPs. The enhanced cytotoxicity of U138-MG cells following treatment with Ab-CS-NPs could be attributed to enhanced cellular uptake.
Cytotoxicity evaluation of various NPs in U138-MG and hCMEC/D3 cells by MTS assay. The cells were incubated with various NPs for 24 h. The concentration of siRNA was 0.1 μg siRNA per well per 100 μL medium. The cells treated with 1% Triton X-100 were used as positive control. The cells without treatment were used as negative control. Blank Ab-CS-NPs and Ab-CS-NPs were conjugated with anti-TfR and anti-B2 antibodies (w/w = 1:1). The data were presented as mean ± SD (n = 5). *p < 0.05, ***p < 0.001 vs. negative control
Cellular uptake and targeting
Receptor-mediated endocytosis is an effective way for enhancing drug uptake rate. Therefore, receptor-mediated uptake can be achieved by decorating corresponding ligands on the surface of nanocarriers. Targeting delivery of siRNA via internalizing cell surface receptors is an appealing strategy to enhance target-specific uptake. We explored the use of a monoclonal antibody directed against TfR and B2R to induce rapid receptor-mediated endocytosis, making the receptor an attractive gateway for intracellular delivery of siRNA. In this study, the antibody conjugation is expected to reduce non-specific cellular uptake and increase the selectivity of receptor-mediated delivery. Thus, the ability of CS-NPs, anti-TfR antibody-CS-NPs [(Ab0 + Ab1)-CS-NPs], anti-B2 antibody-CS-NPs [(Ab0 + Ab2)-CS-NPs], and anti-TfR/anti-B2 dual antibody-CS-NPs [(Ab1 + Ab2)-CS-NPs] to enhance Cy3-siRNA internalization was evaluated in hCMEC/D3 and U138-MG cells. Ab0 refers to non-targeting scrambled antibody, Ab1 refers to anti-TfR antibody, and Ab2 refers to anti-B2 antibody. Non-targeting scrambled antibody was also evaluated in (Ab0 + Ab1)-CS-NPs and (Ab0 + Ab2)-CS-NPs to ensure that each NP group contained the same amount of antibody conjugation. Cellular uptake was expressed as the fluorescence intensity of cells that internalized Cy3-siRNA-labeled NPs. As shown in Fig. 3a–d, the process of cellular uptake was time dependent. Ab-CS-NPs increased the cellular uptake in hCMEC/D3 and U138-MG cells when compared to the CS-NPs. The cellular uptake of (Ab0 + Ab1)-CS-NPs was 68.9 ± 38.7, 63.4 ± 7.90, and 57.7 ± 13.3% higher than that of CS-NPs in hCMEC/D3 cells at 4, 12, and 24 h, respectively. The cellular uptake of (Ab1 + Ab2)-CS-NPs was 63.0 ± 52.6, 72.9 ± 23.5, and 64.1 ± 9.93% higher than that of CS-NPs in hCMEC/D3 cells at 4, 12, and 24 h, respectively. The cellular uptake of (Ab0 + Ab2)-CS-NPs was 26.9 ± 19.9, 36.2 ± 12.4, and 35.7 ± 1.75% higher than that of CS-NPs in U138-MG cells at 4, 12, and 24 h, respectively. The cellular uptake of (Ab1 + Ab2)-CS-NPs was 30.9 ± 21.1, 43.2 ± 26.5, and 44.1 ± 10.1% higher than that of CS-NPs in U138-MG cells at 4, 12, and 24 h, respectively. Furthermore, there is no significant difference in cell uptake between (Ab0 + Ab1)-CS-NP- and (Ab1 + Ab2)-CS-NP-treated groups in hCMEC/D3 cells or between (Ab0 + Ab2)-CS-NP- and (Ab1 + Ab2)-CS-NP-treated groups in U138-MG cells. The main reason for this is the fact that receptor-mediated endocytosis promoted the internalization of NPs.
Cellular uptake and targeting of NPs in hCMEC/D3 and U138-MG cells. a Representative fluorescence images of hCMEC/D3 cells treated with Cy3-siRNA alone, Cy3-siRNA loaded CS-NPs, (Ab0 + Ab1)-CS-NPs, and (Ab1 + Ab2)-CS-NPs for 4, 12, and 24 h. Cy3-siRNA is represented in red. b Quantitative analysis of the cellular uptake of Cy3-siRNA alone, CS-NPs, (Ab0 + Ab1)-CS-NPs, and (Ab1 + Ab2)-CS-NPs in hCMEC/D3 cells using flow cytometry. c Representative fluorescence images of U138-MG cells treated with Cy3-siRNA alone, Cy3-siRNA-loaded CS-NPs, (Ab0 + Ab2)-CS-NPs, and (Ab1 + Ab2)-CS-NPs for 4, 12, and 24 h. d Quantitative analysis of the cellular uptake of Cy3-siRNA alone, CS-NPs, (Ab0 + Ab2)-CS-NPs, and (Ab1 + Ab2)-CS-NPs in U138-MG cells using flow cytometry. e Dual targeting and transport across BBB co-culture model in vitro. The cellular uptake of Cy3-siRNA alone, CS-NPs, (Ab0 + Ab1)-CS-NPs, (Ab0 + Ab2)-CS-NPs, and (Ab1 + Ab2)-CS-NPs in hCMEC/D3 and U138-MG cells were quantitated using flow cytometry. Ab0 refers to non-targeting scrambled antibody, Ab1 refers to anti-TfR antibody, and Ab2 refers to anti-B2 antibody. At different time intervals after incubation, cells were stained with DAPI and imaged by fluorescence microscopy. Magnification ×20. The data were presented as mean ± SD (n = 3). *p < 0.05, ***p < 0.001
In order to confirm the role of antibody-mediated targeting and enhanced penetration ability of the NPs, cellular uptake of NPs was also evaluated using the hCMEC/D3 and U138-MG BBB co-culture model. The in vitro co-culture model better represents the interaction between endothelial cells and astrocytes in comparison to monolayer cell experiments. The co-culture model was also designed to evaluate NP penetration across the BBB. Penetration through cells, rather than uptake into cells, was more physiologically relevant compared to studying the effects of NPs on monolayer of cells. As indicated in Fig. 3e, all treatment groups containing Cy3-siRNA alone, CS-NPs, (Ab0 + Ab1)-CS-NPs, (Ab0 + Ab2)-CS-NPs, and (Ab1 + Ab2)-CS-NPs were able to penetrate across the hCMEC/D3 cell monolayer and to be taken up by U138-MG cells within 24 h. (Ab0 + Ab1)-CS-NPs and (Ab1 + Ab2)-CS-NPs possessed significantly higher uptake than that of CS-NPs in hCMEC/D3 cells, which may be due to the expression of TfR. Quantitatively, the cellular uptake of (Ab0 + Ab1)-CS-NPs and (Ab1 + Ab2)-CS-NPs was 28.6 ± 12.9 and 40.6 ± 2.51% higher than that of CS-NPs in fluorescence intensity at 24 h. Although the fluorescence intensity of hCMEC/D3 cells treated with (Ab1 + Ab2)-CS-NPs showed a little higher than (Ab0 + Ab1)-CS-NPs, there was no statistically significant difference between them. Moreover, CS-NPs and (Ab0 + Ab2)-CS-NPs showed almost the same uptake in hCMEC/D3 cells, which indicated that anti-B2 antibody conjugation had no impact on the uptake of NP in the hCMEC/D3 cell monolayer. Interestingly, all the antibody conjugation groups containing (Ab0 + Ab1)-CS-NPs, (Ab0 + Ab2)-CS-NPs, and (Ab1 + Ab2)-CS-NPs enhanced the uptake of NPs in U138-MG cells in comparison to CS-NPs, suggesting that anti-TfR antibody and anti-B2 antibody play specific roles in mediating cellular uptake. Anti-TfR antibody may have improved the cell uptake and penetration of (Ab0 + Ab1)-CS-NPs in hCMEM/D3 cells and also recognized the TfR on the surface of U138-MG cells. CS-NPs and (Ab0 + Ab2)-CS-NPs showed almost the same cellular uptake in hCMEC/D3 cells, but the cellular uptake of (Ab0 + Ab2)-CS-NPs in U138-MG cells was 2.54 times higher than that of CS-NPs. This may be attributed to the anti-B2 antibody that can recognize B2R on the surface of U138-MG cells. Dual-targeting (Ab1 + Ab2)-CS-NPs showed the highest uptake in the U138-MG cells, which was approximately six times higher than that of the non-targeting CS-NPs. This result may be due to the additive effects of the dual-antibody conjugation.
Gene knockdown in vitro
For siRNA-mediated gene silencing, it is difficult to achieve 100% knockdown efficiency. As a result, in order to maximize the inhibition of HIV, a combination of siRNA-targeting SART3 and hCycT1 were loaded into the NPs and evaluated with hopes that this will enhance the inhibition of HIV replication by different mechanisms. Downregulation of SART3 and hCycT1 expression mediated by CS-NPs and (Ab1 + Ab2)-CS-NP-loaded SART3 siRNA and/or hCycT1 siRNA in U138-MG cells was evaluated in vitro using the hCMEC/D3 and U138-MG BBB co-culture model at both mRNA and protein level (Fig. 4). As expected, CS-NPs and (Ab1 + Ab2)-CS-NPs significantly reduced SART3 and hCycT1 mRNA expressions when compared to non-treated control. As shown in Fig. 4a, greater SART3 and hCycT1 gene knockdown efficiencies were observed in (Ab1 + Ab2)-CS-NP-treated U138-MG cells in comparison to CS-NPs. For example, qPCR data showed a significant gene knockdown efficiency by 80.9 ± 5.01% for SART3 and 66.6 ± 5.88% for hCycT1 in U138-MG cells treated by (Ab1 + Ab2)-CS-NPs loaded with SART3 siRNA and hCycT1 siRNA, which was approximately 1.57- and 1.56-fold higher than that of CS-NPs, respectively. Furthermore, the knockdown efficiencies of SART3 and hCycT1 mRNA in U138-MG cells treated by NPs loaded with a combination of SART3 siRNA and hCycT1 siRNA were significantly higher than cells treated with NPs loaded with SART3 siRNA or hCycT1 siRNA alone, which may be due to the synergistic silencing effects of the combination siRNA. Changes to Tip110 and Cyclin T1 protein expressions were assessed using flow cytometry after immunofluorescence staining. As shown in Fig. 4b, Tip110 and Cyclin T1 proteins were expressed in great abundance in the untreated U138-MG cells and Lipofectamine 2000-treated U138-MG cells, whereas treatment of U138-MG cells with (Ab1 + Ab2)-CS-NPs significantly reduced Tip110 and/or cyclin T1 protein expressions compared to CS-NPs. The results supported our qPCR gene knockdown studies. These studies demonstrated the potential for additive silencing effects of SART3 siRNA and hCycT1 siRNA and dual-antibody-mediated specific targeted delivery of siRNA for inhibition of HIV replication in astrocytes.
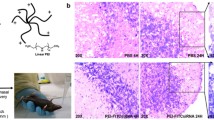
Evaluation of SART3 and hCycT1 mRNA and protein levels in U138-MG cells using the hCMEC/D3 and U138-MG BBB co-culture model. a Knockdown of SART3 and hCycT1 of mRNA expressions in U138-MG cells after treated with SART3 siRNA and/or hCycT1 siRNA-loaded CS-NPs and (Ab1 + Ab2)-CS-NPs (anti-TfR and anti-B2 conjugation, dual targeting to TfR and B2R), respectively. Lipofectamine 2000 was used as a positive control. The expressions of SART3 and hCycT1 mRNA were determined by qPCR. SART3 and hCycT1 mRNA expressions were normalized to GAPDH mRNA. The relative percentage of SART3 or hCycT1 mRNA expression levels from treated was normalized to that of the non-treated cells (negative control), which was defined as 100%. b Quantification of Tip110 and Cyclin T1 protein expressions in U138-MG cells by flow cytometry after treatment with SART3 siRNA and/or hCycT1 siRNA-loaded CS-NPs and (Ab1 + Ab2)-CS-NPs (anti-TfR and anti-B2 conjugation, dual targeting to TfR and B2R), respectively. Data represents the mean ± SD; n = 3. *p < 0.05, **p < 0.01, ***p < 0.001
Conclusions
In the present study, we demonstrated the potential of dual-antibody-mediated receptor targeting for siRNA delivery across the BBB to inhibit HIV replication in astrocytes. Anti-Tf antibody and anti-B2 antibody was successfully conjugated to CS-NPs by carboxyl-to-amine cross-linking using carbodiimide EDC and sulfo-NHS. The cellular uptake of (Ab1 + Ab2)-CS-NPs was significantly increased compared with CS-NPs, (Ab1 + Ab0)-CS-NPs, and (Ab2 + Ab0)-CS-NPs. The expression levels of SART3 and hCycT1 in U138-MG cells were significantly reduced by the delivery of SART3 and hCycT1 siRNA using dual-antibody-conjugated CS-NPs. The designed (Ab1 + Ab2)-CS-NPs may provide a new strategy for developing siRNA-targeted delivery to potential HIV-1 reservoirs in the brain. Movement from single-targeted NPs to dual-targeted NPs is the direction that drug delivery system development appears to be heading. Dual-ligand modified NPs are a potential strategy to improve the efficiency of drug delivery, especially in brain drug delivery. Based on our promising results, the dual-targeting NPs will be used to inhibit HIV replication in HIV-infected astrocytes. In the future, we may construct a novel 3D co-culture model, in which three kinds of HIV-1 infected cell lines including microglia, macrophages, and astrocytes can be surrounded by BBB cells. The targeting ability and penetration efficiency of the dual-targeting NPs will be evaluated using this 3D co-culture model.
References
Su ZZ, Kang DC, Chen Y, Pekarskaya O, Chao W, Volsky DJ, et al. Identification and cloning of human astrocyte genes displaying elevated expression after infection with HIV-1 or exposure to HIV-1 envelope glycoprotein by rapid subtraction hybridization, RaSH. Oncogene. 2002;21:3592–602.
Hurwitz AA, Berman JW, Lyman WD. The role of the blood-brain barrier in HIV infection of the central nervous system. Adv Neuroimmunol. 1994;4:249–56.
Gray F, Lescs MC, Keohane C, Paraire F, Marc B, Durigon M, et al. Early brain changes in HIV infection: neuropathological study of 11 HIV seropositive, non-AIDS cases. J Neuropathol Exp Neurol. 1992;51:177–85.
Skolasky RL, Dal Pan GJ, Olivi A, Lenz FA, Abrams RA, McArthur JC. HIV-associated primary CNS lymorbidity and utility of brain biopsy. J Neurol Sci. 1999;163:32–8.
Davies J, Everall IP, Weich S, Glass J, Sharer LR, Cho ES, et al. HIV-associated brain pathology: a comparative international study. Neuropathol Appl Neurobiol. 1998;24:118–24.
Gelman BB, Chen T, Lisinicchia JG, Soukup VM, Carmical JR, Starkey JM, et al. The national neuroAIDS tissue consortium brain gene array: two types of HIV-associated neurocognitive impairment. PLoS One. 2012;7:e46178.
Gray F, Adle-Biassette H, Chretien F, Lorin de la Grandmaison G, Force G, Keohane C. Neuropathology and neurodegeneration in human immunodeficiency virus infection. Pathogenesis of HIV-induced lesions of the brain, correlations with HIV-associated disorders and modifications according to treatments. Clin Neuropathol. 2001;20:146–55.
Zhang YL, Ouyang YB, Liu LG, Chen DX. Blood-brain barrier and neuro-AIDS. Eur Rev Med Pharmacol Sci. 2015;19:4927–39.
Strelow LI, Janigro D, Nelson JA. The blood-brain barrier and AIDS. Adv Virus Res. 2001;56:355–88.
Gorry P, Purcell D, Howard J, McPhee D. Restricted HIV-1 infection of human astrocytes: potential role of nef in the regulation of virus replication. J Neurovirol. 1998;4:377–86.
Reeves JD, Hibbitts S, Simmons G, McKnight A, Azevedo-Pereira JM, Moniz-Pereira J, et al. Primary human immunodeficiency virus type 2 (HIV-2) isolates infect CD4-negative cells via CCR5 and CXCR4: comparison with HIV-1 and simian immunodeficiency virus and relevance to cell tropism in vivo. J Virol. 1999;73:7795–804.
Klein RS, Williams KC, Alvarez-Hernandez X, Westmoreland S, Force T, Lackner AA, et al. Chemokine receptor expression and signaling in macaque and human fetal neurons and astrocytes: implications for the neuropathogenesis of AIDS. J Immunol. 1999;163:1636–46.
Guo X, Huang L. Recent advances in nonviral vectors for gene delivery. Acc Chem Res. 2012;45:971–9.
Mintzer MA, Simanek EE. Nonviral vectors for gene delivery. Chem Rev. 2009;109:259–302.
Chen Y, Liu L. Modern methods for delivery of drugs across the blood-brain barrier. Adv Drug Deliv Rev. 2012;64:640–65.
Lockman PR, Mumper RJ, Khan MA, Allen DD. Nanoparticle technology for drug delivery across the blood-brain barrier. Drug Dev Ind Pharm. 2002;28:1–13.
Whitehead KA, Langer R, Anderson DG. Knocking down barriers: advances in siRNA delivery. Nat Rev Drug Discov. 2009;8:129–38.
Mathupala SP. Delivery of small-interfering RNA (siRNA) to the brain. Expert Opin Ther Pat. 2009;19:137–40.
Pardridge WM. shRNA and siRNA delivery to the brain. Adv Drug Deliv Rev. 2007;59:141–52.
Wong HL, Chattopadhyay N, Wu XY, Bendayan R. Nanotechnology applications for improved delivery of antiretroviral drugs to the brain. Adv Drug Deliv Rev. 2010;62:503–17.
Gupta U, Jain NK. Non-polymeric nano-carriers in HIV/AIDS drug delivery and targeting. Adv Drug Deliv Rev. 2010;62:478–90.
Saranya N, Moorthi A, Saravanan S, Devi MP, Selvamurugan N. Chitosan and its derivatives for gene delivery. Int J Biol Macromol. 2011;48:234–8.
Aktas Y, Yemisci M, Andrieux K, Gursoy RN, Alonso MJ, Fernandez-Megia E, et al. Development and brain delivery of chitosan-PEG nanoparticles functionalized with the monoclonal antibody OX26. Bioconjug Chem. 2005;16:1503–11.
Trapani A, De Giglio E, Cafagna D, Denora N, Agrimi G, Cassano T, et al. Characterization and evaluation of chitosan nanoparticles for dopamine brain delivery. Int J Pharm. 2011;419:296–307.
Jefferies WA, Brandon MR, Hunt SV, Williams AF, Gatter KC, Mason DY. Transferrin receptor on endothelium of brain capillaries. Nature. 1984;312:162–3.
Kissel K, Hamm S, Schulz M, Vecchi A, Garlanda C, Engelhardt B. Immunohistochemical localization of the murine transferrin receptor (TfR) on blood-tissue barriers using a novel anti-TfR monoclonal antibody. Histochem Cell Biol. 1998;110:63–72.
Jones AR, Shusta EV. Blood-brain barrier transport of therapeutics via receptor-mediation. Pharm Res. 2007;24:1759–71.
Qian ZM, Li H, Sun H, Ho K. Targeted drug delivery via the transferrin receptor-mediated endocytosis pathway. Pharmacol Rev. 2002;54:561–87.
Lee HJ, Engelhardt B, Lesley J, Bickel U, Pardridge WM. Targeting rat anti-mouse transferrin receptor monoclonal antibodies through blood-brain barrier in mouse. J Pharmacol Exp Ther. 2000;292:1048–52.
Weksler B, Romero IA, Couraud PO. The hCMEC/D3 cell line as a model of the human blood brain barrier. Fluids Barriers CNS. 2013;10:16.
Ohtsuki S, Ikeda C, Uchida Y, Sakamoto Y, Miller F, Glacial F, et al. Quantitative targeted absolute proteomic analysis of transporters, receptors and junction proteins for validation of human cerebral microvascular endothelial cell line hCMEC/D3 as a human blood-brain barrier model. Mol Pharm. 2013;10:289–96.
Calzolari A, Larocca LM, Deaglio S, Finisguerra V, Boe A, Raggi C, et al. Transferrin receptor 2 is frequently and highly expressed in glioblastomas. Transl Oncol. 2010;3:123–34.
Voth B, Nagasawa DT, Pelargos PE, Chung LK, Ung N, Gopen Q, et al. Transferrin receptors and glioblastoma multiforme: current findings and potential for treatment. J Clin Neurosci. 2015;22:1071–6.
Stephens GJ, Cholewinski AJ, Wilkin GP, Djamgoz MB. Calcium-mobilizing and electrophysiological effects of bradykinin on cortical astrocyte subtypes in culture. Glia. 1993;9:269–79.
Cholewinski AJ, Stevens G, McDermott AM, Wilkin GP. Identification of B2 bradykinin binding sites on cultured cortical astrocytes. J Neurochem. 1991;57:1456–8.
Raidoo DM, Sawant S, Mahabeer R, Bhoola KD. Kinin receptors are expressed in human astrocytic tumour cells. Immunopharmacology. 1999;43:255–63.
Wang YB, Peng C, Liu YH. Low dose of bradykinin selectively increases intracellular calcium in glioma cells. J Neurol Sci. 2007;258:44–51.
Zhao Y, Xue Y, Liu Y, Fu W, Jiang N, An P, et al. Study of correlation between expression of bradykinin B2 receptor and pathological grade in human gliomas. Br J Neurosurg. 2005;19:322–6.
Long L, Thelen JP, Furgason M, Haj-Yahya M, Brik A, Cheng D, et al. The U4/U6 recycling factor SART3 has histone chaperone activity and associates with USP15 to regulate H2B deubiquitination. J Biol Chem. 2014;289:8916–30.
Chiu YL, Cao H, Jacque JM, Stevenson M, Rana TM. Inhibition of human immunodeficiency virus type 1 replication by RNA interference directed against human transcription elongation factor P-TEFb (CDK9/cyclinT1). J Virol. 2004;78:2517–29.
Kozlowski MR, Sandler P, Lin PF, Watson A. Brain-derived cells contain a specific binding site for Gp120 which is not the CD4 antigen. Brain Res. 1991;553:300–4.
Weber J, Clapham P, McKeating J, Stratton M, Robey E, Weiss R. Infection of brain cells by diverse human immunodeficiency virus isolates: role of CD4 as receptor. J Gen Virol. 1989;70(Pt 10):2653–60.
Acknowledgments
This study was funded by a Manitoba Health Research Council (MHRC) Establishment Grant awarded to Dr. Emmanuel A. Ho. Dr. Jijin Gu was supported by a MHRC Post-doctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Gu, J., Al-Bayati, K. & Ho, E.A. Development of antibody-modified chitosan nanoparticles for the targeted delivery of siRNA across the blood-brain barrier as a strategy for inhibiting HIV replication in astrocytes. Drug Deliv. and Transl. Res. 7, 497–506 (2017). https://doi.org/10.1007/s13346-017-0368-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13346-017-0368-5