Abstract
Leaf and shoot blight, often accompanied by die-back symptoms, on Eucalyptus species, hybrids and clones have been reported from a number of countries. More than one bacterial species has been found to cause these symptoms. In this study, a leaf disease of E. pellita in Indonesia was investigated. The disease was found primarily on nursery plants and young trees that recovered within the first year of growth. Leaf samples were collected from symptomatic trees, and isolations consistently yielded a Xanthomonas sp. Sequencing of the 16S rRNA gene region and multilocus sequence analysis (MLSA) was performed on 19 of the 61 Xanthomonas isolates obtained. In the MLSA, four genes, namely, dnaK, fyuA, gyrB and rpoD, were sequenced and the isolates were identified as X. perforans. Four representative isolates, at a concentration of 106 CFU/ml, were leaf-infiltrated and spray-inoculated on to E. pellita, tomato and pepper seedlings. The type isolate of X. perforans was included in the pathogenicity trials as a positive control. All four isolates of X. perforans, inclusive of the type isolate, induced bacterial spot symptoms on tomato and pepper seedlings. They also caused water-soaked lesions on the leaves of E. pellita seedlings, characteristic of the symptoms observed in the field. This is the first report of X. perforans infecting leaves of a woody host.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Blight and dieback symptoms on Eucalyptus can be caused by a number of bacterial species in different parts of the world. In Australia, Xanthomonas dyei pv. eucalypti (X. campestris pv. eucalypti) caused leaf blight in eucalypts (Corymbia citriodora) in the 1970s (Truman 1974). In Brazil and Uruguay, the disease, accompanied by dieback, is caused by X. axonopodis pv. eucalyptorum (Ferraz et al. 2018; Gonçalves et al. 2008). Pantoea ananatis (Coutinho et al. 2002) and X. vasicola (Coutinho et al. 2015) are responsible for the disease on Eucalyptus in South Africa. Although not specifically a leaf pathogen, Erwinia psidii infects the vascular tissues of young shoots and, leaves that can result in severe dieback in young Eucalyptus plantations (Arriel et al. 2014; Coutinho et al. 2011).
In earlier studies, Xanthomonas species have been identified based on phenotypic and biochemical properties (Truman 1974; Van den Mooter and Swings 1990). This led to considerable taxonomic confusion and misidentification of bacterial isolates. Other methods used included repetitive sequence PCR (rep-PCR), amplified fragment length polymorphism (AFLP) and restriction fragment length polymorphism (RFLP), which showed high similarities to the genomic clusters described by Rademaker et al. (2000); and Schaad et al. (2005); Vauterin et al. (1995). Observations from single gene-based phylogenies also produced corresponding group homologies (Hauben et al. 1997; Parkinson et al. 2007). Although these techniques were consistent in producing homologous groupings, certain limitations made them unsuitable for routine diagnosis of Xanthomonas. One of these limitations was the apparent difficulty in comparing and standardizing gel profiles between laboratories. Additionally, a lack of resolution of single gene phylogenies was also observed within species. Consequently, a multilocus sequence analysis (MLSA) (Young et al. 2008, 2010) has been used to produce a classification framework using two or more protein-coding genes located on the core genome (Gevers et al. 2005).
The aims of this study were i) to identify the causal agent of a leaf disease that is common on young Eucalyptus pellita trees in Indonesian plantations and ii) to test the pathogenicity of the representative isolates of the putative pathogen on E. pellita seedlings.
Materials and methods
Bacterial isolates
Leaves of young E. pellita trees between 1 and 6-months-old in plantations in Riau, South Sumatra, Indonesia, commonly displayed symptoms of a leaf disease reminiscent of bacterial infection. Symptoms included necrotic tissue typically surrounded by distinct chlorotic zones (Fig. 1). The symptoms were common on nursery plants prior to outplanting and persisted during the first 6 months of growth. Subsequently, the infected leaves typically abscised from the trees that recovered and grew in the absence of symptoms.
Symptomatic leaves were randomly collected from different trees widely distributed in plantations and nurseries in South Sumatra. Although these did not form part of a systematic survey, the collections represented a wide distribution of the symptoms in the sampled area. Isolations for bacteria were made from the affected tissues. Diseased tissue was submerged into a 10% (v/v) sodium hypochlorite solution for 1 min and then into 70% (v/v) ethanol for a further minute before being crushed in a sterile mortar containing 1 mL sterile 10 mM phosphate buffer with a pestle. The resulting suspension was streaked on to nutrient agar (NA) (Biolab Diagnostics, Merck, South Africa) and incubated at 28 °C for 48 h. Single colonies were purified. Care was taken to analyse samples from different areas and trees separately. Sixty-one pure cultures of suspected Xanthomonas strains were deposited in the Bacterial Culture Collection (BCC) in the Forestry and Agricultural Biotechnology Institute (FABI), University of Pretoria.
DNA extraction
Single colonies of each isolate were suspended in 1 M phosphate buffer solution [4.0 g NaCl, 0.1 g KCl, 0.72 g Na2HPO4 and 0.12 g KH2PO4 dissolved in 500 mL distilled water; pH 8] and genomic DNA was extracted using the Zymo Research Genomic DNA™ Tissue Miniprep Kit (Inqaba Biotech, South Africa) following the manufacturer’s instructions. The quantity and quality of the DNA were measured using a spectrophotometer, NanoDrop™ 2000 (Thermo Scientific, USA), and gel electrophoresis, respectively.
16S rRNA sequencing
The 16S rRNA region for each DNA template was amplified using universal primers described previously by Coenye et al. (1999). The partial sequences were manually edited using Chromas LITE Sequence Alignment Editor v2.1.1 (Technelysium Pty Ltd., Australia). The sequences were submitted through the BLAST search engine and compared against reference sequences in the GenBank database (National Center for Biotechnology Information, USA). Thereafter, the sequences of the isolates and reference/type isolates of Xanthomonas species obtained using GenBank accession numbers from Hauben et al. (1997) were uploaded on to the MAFFT online sequence alignment program v7.0 (Katoh et al. 2002; Katoh and Standley 2013) to verify sequence orientation and also to perform a multiple sequence alignment. The aligned sequences were exported to BioEdit Sequence Alignment Editor v7.0.9.0 (Hall 1999) and the overhangs trimmed. A best fit evolutionary model was determined using JModeltest v2.0 (Posada 2008; Posada and Crandall 1998) and a maximum likelihood analysis performed using PhyML v3.0 (Guindon et al. 2010; Guindon and Gascuel 2003). The Hasegawa-Kishino-Yano (HKY) model was chosen for the maximum likelihood analysis with rate variation among sites (+G) (Hasegawa et al. 1985). A bootstrap analysis of a 1000 replicates was performed in order to determine confidence in branching points.
Multilocus sequence analysis (MLSA)
Nineteen Xanthomonas isolates were selected for further identification (Table 1) and subjected to a multilocus sequence analysis (MLSA) using primer sets of four house-keeping genes, namely, dnaK, fyuA, gyrB and rpoD, and cycle conditions described by Young et al. (2008). The cycle conditions for the sequencing reactions were as previously described by Coenye et al. (1999). The sequencing products were precipitated using the sodium acetate-ethanol protocol, and the samples were submitted to the DNA sequencing facility at the University of Pretoria, South Africa and sequenced using the ABI3130XL sequencer (Life Technologies, USA).
The partial sequences of the four protein-coding genes were analysed as mentioned previously. The sequences of the isolates were aligned with the reference/type isolates of Xanthomonas spp., obtained from Professor CT Bull, Pennsylvania State University, USA. The partial sequences of the four protein-coding genes dnaK, fyuA, gyrB and rpoD for the Xanthomonas isolates and reference sequences of type species were trimmed to nucleotide lengths of 769, 524, 516 and 701 nt, respectively. Best fit evolutionary models were determined, and maximum likelihood analyses performed for all four protein-coding genes (data not shown) and concatenated dataset (Fig. 2). Stenotrophomonas maltophila ICMP17033T was chosen as the outgroup taxon for these analyses with the exception of the fyuA phylogeny (data not shown). The chosen models for the concatenated dataset were GTR + I + G, TIM2 + I + G for dnaK and fyuA, TIM1 + G for gyrB, and TIM3 + I + G for rpoD. Bootstrap analyses of 1000 replicates were performed for each dataset to define confidence in branching points.
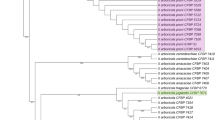
A Maximum Likelihood tree representing the concatenated phylogeny of the Xanthomonas isolates and reference isolates. Bootstrap values after 1000 replicates are indicated as percentages (only values 50 and above are shown). Stenotrophomonas maltophila ICMP17033T was included as an outgroup. The scale bar represents the number of substitutions per site. Isolates marked with an asterisk (*) represent sequences originally from the study by Ferraz et al. 2018. Postscript ‘T’ refers to type isolate of the species.
Pathogenicity trial
Inoculum preparation
Four isolates (EPK8, EPK15, EPK28 and EPK43) were selected and grown on NA and incubated for 48 h at 28 °C. Single colonies obtained from the pure cultures were then suspended in 25 mL nutrient broth (NB) (Biolab Diagnostics, Merck, South Africa) and the samples were placed in a shaking incubator at 120 rpm for 18 h at 28 °C. The bacterial suspensions were adjusted to a concentration of 106 cfu/mL (OD600 = 0.01) using 10 mM phosphate buffer solution.
Seedling inoculations
Eucalyptus pellita seeds were cultivated in a 12.5 cm pot with potting soil 60 dm3 (Garden Master, South Africa (SA)), vermiculite 15 dm3 (Bark Unlimited, SA) and river sand mixture (2:1:1), and kept in a phytotron at approx. 25 °C with 16 h day and 8 h night conditions. The emerging E. pellita seedlings were transplanted into small plastic bags using the same soil mixture described above. The nutrients were supplemented using organic fertiliser containing 80 g/kg N, 20 g/kg P, 58 g/kg K (4:1:3) (Nitrosol, Efekto, SA) once each week before seedling emergence, and subsequently once bi-weekly. The transplanted seedlings were maintained under the same greenhouse conditions mentioned above. The Solanum lycopersicum (tomato) cv. ‘Moneymaker’ and Capsicum annum (pepper) cv. ‘Jupiter’ seedlings were sourced from a nursery (Sunshine seedlings, Pietermaritzburg, SA) and maintained under the same greenhouse conditions as for the E. pellita seedlings.
Fifteen seedlings each of 1-year-old Eucalyptus pellita, tomato cv. Moneymaker and pepper cv. Jupiter, both 6 weeks old, were used in the inoculation trial with three replications per isolate. Prior to the inoculations, the seedlings were placed in plastic square containers, respective to the isolate used for inoculation, and arranged inside the phytotron following a randomised design. Plants were inoculated with one of the four isolates of Xanthomonas perforans. Pantoea ananatis LMG20103 was used as the positive control for the E. pellita inoculations and sterile phosphate buffer as the negative control. Xanthomonas perforans ICMP16690T was included as a positive control for the inoculation of tomato and pepper seedlings.
For each seedling, 2–3 leaves were infiltrated with the bacterial suspension at multiple sites along the main vein, flooding the leaf panels, using a 1 mL blunt-end insulin syringe in the case of E. pellita. Tomato and pepper leaves were spray-inoculated on both the abaxial and adaxial sides until run-off. The plants were covered with plastic bags to induce a humid environment for 48 h and maintained in an enclosed phytotron at approximately 26 °C with 16 h day and 8 h night cycles. After 48 h, the plastic bags were removed and the plants were monitored for development of disease symptoms. The presence or absence of typical water-soaked lesions on inoculated plants was recorded every 24 h for 1 month. The inoculation trial was repeated once, but in the second trial, the plants of E. pellita were spray-inoculated as opposed to infiltration with the bacteria.
Re-isolation procedure
A subset of inoculated leaves showing leaf spot symptoms, and leaves inoculated with 10 mM phosphate buffer, were collected and surface sterilized by submersion into a 10% (v/v) sodium hypochlorite solution for 1 min and then into 70% (v/v) ethanol for a further minute. The leaves were rinsed with sterile distilled water three times, and 1 mm2 leaf pieces from the leading edge of lesions were macerated in 10 mM phosphate buffer solution and 30 μL of the resulting suspension streaked on to NA. The plates were incubated at 28 °C for 48 h.
Identification of recovered bacterial isolates
Genomic DNA from single colonies of bacterial cultures was extracted using the ZR Genomic DNA™ Tissue Miniprep Kit following the manufacturer’s instructions. The small ribosomal sub-unit gene region of the recovered isolates was sequenced using PCR based techniques mentioned previously. In the case of a positive 16S rRNA sequence identity for Xanthomonas, the rpoD gene region was sequenced.
Results
16S rRNA sequencing and multilocus sequence analysis (MLSA) classification
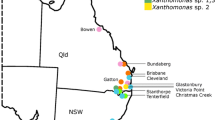
The results of the BLAST search across the GenBank database revealed a 99% sequence similarity with Xanthomonas (61), Pantoea stewartii (1) and Pseudomonas oryzihabitans (1). Only the Xanthomonas strains were further analysed. These represented different sampled areas in Indonesia and spanning different periods of sample collection.
The four protein-coding gene datasets, namely, dnaK, fyuA, gyrB, and rpoD, were used to generate a concatenated dataset. Congruency was tested using the four sequence datasets and based on the results of the partition-homogeneity test (p > 0.05) the datasets were concatenated. The nucleotide sequence for the combined dataset was 2510 nt in length. The concatenated phylogeny (Fig. 2) revealed a clustering of the E. pellita isolates from Indonesia with Xanthomonas perforans. The topology of this clade was consistent with the observations of the individual gene phylogenies (data not shown).
Sub-clustering was observed between members of X. perforans which may be indicative of the presence of potential sub-populations within this species. The Indonesian isolates formed a discrete sub-group separate from the reference isolate, X. perforans ICMP109, the type isolate, X. perforans ICMP16690 and isolates LPF581, LPF583, LPF596, LPF597 and LPF598 obtained from symptomatic leaves of Eucalyptus urophylla x E. globulus clones in Brazil (Fig. 2).
Pathogenicity trial
Symptom development was slow in the early stages of infection on E. pellita seedlings (Table 2). However, at 5 days post-inoculation (dpi), water-soaking was observed on leaves of seedlings infiltrated with EPK28 and EPK43. A mild to weak reaction was observed on E. pellita leaves infiltrated with EPK8, EPK15, X. perforans ICMP16690T and P. ananatis LMG20103. No reaction was observed for the negative control. By 18 dpi, the water-soaked lesions had expanded into reddish-brown necrotic lesions surrounded by chlorosis resembling the symptoms observed in the field (Fig. 3). Eucalyptus seedlings inoculated with X. perforans ICMP16690T became chlorotic at the site of inoculation, indicative of a typical HR reaction caused by a non-host pathogen (Fig. 3e). These observations were consistent throughout the period of the pathogenicity trial. However, the lesions on spray-inoculated E. pellita seedlings were extremely small, less than 2 mm in size (data not shown).
Symptoms caused by Xanthomonas perforans infiltrated into 1 year old Eucalyptus pellita leaves.a) Brown necrotic lesions caused by EPK43; b) faint red-brown chlorotic lesions caused by EPK15; c) dark red-brown lesions surrounded by chlorosis caused by EPK8 and d) EPK28; e) chlorotic lesions caused by Xanthomonas perforans type isolate, ICMP16690T. f) A mild hypersensitive reaction was observed on E. pellita leaves infiltrated with Pantoea ananatis LMG20103. Pictures were taken at 18 days post-inoculation. Postscript ‘T’ refers to type isolate of the species
Characteristic leaf spot symptoms were observed on tomato seedlings from 3 dpi onwards, inoculated with the three representative Xanthomonas perforans isolates, including X. perforans ICMP16690T (Fig. 4). In contrast, only one of the representative isolates (EPK43), and the type strain (ICMP16690T) had caused leaf spots on pepper seedlings, albeit weak, by the time of termination of the trial (Fig. 4g, h). Similar results were observed for the tomato and pepper seedlings spray-inoculated with the three representative X. perforans isolates in the second pathogenicity trial (data not shown). Leaf spots were absent from the negative controls of all three plant species.
Symptoms caused by Xanthomonas perforans spray-inoculated on leaves of six-week-old Solana lycopersicum (tomato) cv. ‘Moneymaker’ and Capsicum annum (pepper) cv. ‘Jupiter’ seedlings. On tomato seedlings, a) necrotic lesions surrounded by chlorosis caused by EPK8; b) necrotic lesions resulting in curling at the leaf tip and chlorosis caused by EPK28; c) chlorotic and marginal necrotic lesions caused by EPK43 and d) the type isolate, ICMP16690T. e) No symptoms were observed on pepper leaves spray-inoculated with EPK8 and f) EPK28; g) a single necrotic lesion on the leaf margin caused by EPK43. And h) necrotic lesions surrounding the leaf margin and on the leaf lamina caused by the type isolate of Xanthomonas perforans, ICMP16690T. Pictures were taken 4 days post-inoculation. Postscript ‘T’ refers to type isolate of the species
Xanthomonas perforans was recovered from all of the E. pellita, tomato and peppers samples selected for re-isolation.
Discussion
This study identified the causal agent of a commonly occurring leaf disease of Eucalyptus pellita in nurseries and plantations in Indonesia as X. perforans. To the best of our knowledge, this is the first report of X. perforans causing a leaf disease on a woody host. Xanthomonas perforans is a well-known pathogen causing bacterial leaf spot of tomato and pepper in many parts of the world (Jones et al. 1995; Stall et al. 1994; Vicente et al. 2006). Its ability to cause disease symptoms in Eucalyptus pellita was surprising and suggests a host shift (Coutinho et al. 2015; Wingfield et al. 2008) possibly from tomato or pepper plants grown in the region.
The disease of E. pellita considered in this study appears to be of relatively minor importance. The symptoms can first appear in young nursery plants and persist when trees are established in plantations. This can result in the loss of young leaves that appears to slow establishment, but the infected leaves are lost during the first few months of tree growth, which is typically very rapid reaching up to 3 m in the first 6 months. By that time, most infected leaves have been lost, and trees grow free of symptoms.
The origin of infection of E. pellita plants by X. perforans is unknown, but the fact that seedlings first display typical symptoms as they mature in the nursery suggests that infection originates at that time. Infection possibly occurs via contaminated irrigation water and/or from seed, which could be internally infected with the bacterium. Xanthomonas perforans is known to be internally seedborne in both tomato and pepper (Jones et al. 1986). These questions require further study and their understanding should help to manage the problem. The fact that tomato plants were shown to be susceptible to infection by X. perforans in this study might suggest that the bacterium originated on tomato but that question requires further investigation.
The phylogenies of the Xanthomonas isolates from E. pellita based on the four protein-coding genes, and concatenated dataset were congruent, in most cases, for the main clusters investigated in this study. Overall, although highly supported by all phylogenies, the taxonomic position of the Indonesian isolates relative to the type isolate, X. perforans ICMP16690, remained ambiguous, with these isolates residing in a sub-lineage within the X. perforans cluster (Fig. 2). A MLSA phylogeny constructed with a nucleotide dataset contains a high level of substitution saturation, homoplasy and codon bias (Palmer et al. 2017). However, protein-coding genes are still widely used as phylogenetic markers in numerous research studies for routine diagnosis and taxonomic purposes (Coutinho et al. 2015; Parkinson et al. 2007; Rodriguez-R et al. 2012; Young et al. 2008, 2010). Taken collectively, in order to determine if there is indeed a sub-population within this cluster, a true phylogeny will need to be reconstructed using core-genomic data. The taxonomic position of X. alfalfae ssp. alfalfae ICMP5718, shown in the concatenated phylogeny presented in this study needs clarification. The concatenated phylogeny suggests that this species was incorrectly identified as an X. alfalfae strain while it is, in fact, X. perforans, therefore, making MLSA approach a robust classification tool for identifying known and unknown isolates up to but not limited to species level.
Bacterial diseases of Eucalyptus spp. have become important as these trees are increasingly grown in plantations globally. The leaf disease on E. pellita caused by X. perforans is one of a number of bacterial leaf diseases of Eucalyptus spp. including those caused by X. axonopodis (Gonçalves et al. 2008), P. ananatis (Coutinho et al. 2002), and X. vasicola (Coutinho et al. 2015). These diseases are generally not considered to be serious with damage mostly occurring in seedlings during propagation and in the early stages of plantation development. This is in contrast to more serious problems associated with bacteria such as two Ralstonia spp., Ralstonia solanacearum and R. pseudosolanacearum (Carstensen et al. 2017; Coutinho et al. 2000; Dianese and Dristig 1993) that are associated with wilt of Eucalyptus in various parts of the world and Erwinia psidii (Arriel et al. 2014; Coutinho et al. 2011) that causes a serious wilt disease in young plantations in South America. These diseases add to growing challenges facing Eucalyptus plantation forestry globally (Wingfield et al. 2008).
References
Arriel D et al (2014) Wilt and die-back of Eucalyptus spp. caused by Erwinia psidii in Brazil. For Pathol 44:255–265
Carstensen G, Venter S, Wingfield M, Coutinho T (2017) Two Ralstonia species associated with bacterial wilt of Eucalyptus. Plant Pathol 66:393–403
Coenye T, Falsen E, Vancanneyt M, Hoste B, Govan JR, Kersters K, Vandamme P (1999) Classification of Alcaligenes faecalis-like isolates from the environment and human clinical samples as Ralstonia gilardii sp. nov. Int J Syst Bacteriol 49:405–413
Coutinho T, Roux J, Riedel KH, Terblanche J, Wingfield M (2000) First report of bacterial wilt caused by Ralstonia solanacearum on eucalypts in South Africa. For Pathol 30:205–210
Coutinho T, Preisig O, Mergaert J, Cnockaert M, Riedel K-H, Swings J, Wingfield M (2002) Bacterial blight and dieback of Eucalyptus species, hybrids, and clones in South Africa. Plant Dis 86:20–25
Coutinho TA, Brady CL, van der Vaart M, Venter SN, Telechea N, Rolfo M, Perez C, Wingfield MJ (2011) A new shoot and stem disease of Eucalyptus species caused by Erwinia psidii. Australas Plant Pathol 40:55–60
Coutinho T, Westhuizen L, Roux J, McFarlane S, Venter S (2015) Significant host jump of Xanthomonas vasicola from sugarcane to a Eucalyptus grandis clone in South Africa. Plant Pathol 64:576–581
Dianese J, Dristig M (1993) Screening Eucalyptus selections for resistance to bacterial wilt caused by Pseudomonas solanacearum. In: Hartman G, Hayward A (eds) International bacterial wilt symposium. ACIAR Proceedings, Kaoshiung, p 206
Ferraz HGM, Badel JL, da Silva Guimarães LM, Reis BP, Tótola MR, Gonçalves RC, Alfenas AC (2018) Xanthomonas axonopodis pv. eucalyptorum pv. Nov. causing bacterial leaf blight on eucalypt in Brazil. Plant Pathol J 34:269–285
Gevers D et al (2005) Re-evaluating prokaryotic species. Nat Rev Microbiol 3:733–739
Gonçalves R, Douglas L, Oliveira J, Maffia L, Cascardo J, Alfenas A (2008) Etiology of bacterial leaf blight of Eucalyptus in Brazil. Trop Plant Pathol 33:180–188
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hasegawa M, Kishino H, T-a Y (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Hauben L, Vauterin L, Swings J, Moore E (1997) Comparison of 16S ribosomal DNA sequences of all Xanthomonas species. Int J Syst Bacteriol 47:328–335
Jones J, Pohronezny K, Stall R, Jones J (1986) Survival of Xanthomonas campestris pv. vesicatoria in Florida on tomato crop residue, weeds, seeds, and volunteer tomato plants. Phytopathology 76:430–434
Jones J, Stall R, Scott J, Somodi G, Bouzar H, Hodge N (1995) A third tomato race of Xanthomonas campestris pv. vesicatoria. Plant Dis 79:395–398
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Katoh K, Misawa K, Ki K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Palmer M et al (2017) Phylogenomic resolution of the bacterial genus Pantoea and its relationship with Erwinia and Tatumella. Antonie Van Leeuwenhoek 110:1287–1309
Parkinson N, Aritua V, Heeney J, Cowie C, Bew J, Stead D (2007) Phylogenetic analysis of Xanthomonas species by comparison of partial gyrase B gene sequences. Int J Syst Evol Microbiol 57:2881–2887
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Rademaker J et al (2000) Comparison of AFLP and rep-PCR genomic fingerprinting with DNA-DNA homology studies: Xanthomonas as a model system. Int J Syst Evol Microbiol 50:665–677
Rodriguez-R LM, Grajales A, Arrieta-Ortiz ML, Salazar C, Restrepo S, Bernal A (2012) Genomes-based phylogeny of the genus Xanthomonas. BMC Microbiol 12:43. https://doi.org/10.1186/1471-2180-12-43
Schaad NW et al (2005) Reclassification of Xanthomans campestris pv. citri (ex Hasse 1915) Dye 1978 forms A, B/C/D, and E as X. smithii subsp. citri (ex Hasse) sp., nov. nom. rev. comb. nov., X. fuscans subsp. aurantifolii (ex Gabriel 1989) sp. nov. nom. rev. comb. nov., and X. alfalfae subsp. citrumelo (ex Riker and Jones) Gabriel et al. 1989 sp. nov. nom. rev. comb. nov.; X. campestris pv. malvacearum (ex Smith 1901) Dye 1978 as X. smithii subsp. smithii nov. comb. nov. nom. nov.; X. campestris pv. alfalfae (ex Riker and Jones 1935) sp. nov. nom. rev.; and "var fuscans" of X. campestris pv. phaseoli (ex Smith 1987) Dye 1978 as X. fuscans subsp. fuscans sp. nov. Syst Appl Microbiol 28:494–518
Stall RE et al (1994) Two genetically diverse groups of strains included in Xanthomonas campestris pv. vesicatoria. Int J Syst Bacteriol 44:47–53
Truman R (1974) Die-back of Eucalyptus citriodora caused by Xanthomonas eucalypti sp. n. Phytopathology 64:143–144
Van den Mooter M, Swings J (1990) Numerical analysis of 295 phenotypic features of 266 Xanthomonas strains and related strains and as improved taxonomy of genus. Int J Syst Bacteriol 40:348–369
Vauterin L, Hoste B, Kersters K, Swings J (1995) Reclassification of Xanthomonas. Int J Syst Evol Microbiol 45:472–489
Vicente JG, Everett B, Roberts SJ (2006) Identification of isolates that cause a leaf spot disease of brassicas as Xanthomonas campestris pv. raphani and pathogenic and genetic comparison with related pathovars. Phytopathology 96:735–745
Wingfield M, Slippers B, Hurley B, Coutinho T, Wingfield B, Roux J (2008) Eucalypt pests and diseases: growing threats to plantation productivity. South For 70:139–144
Young J, Park D-C, Shearman H, Fargier E (2008) A multilocus sequence analysis of the genus Xanthomonas. Syst Appl Microbiol 31:366–377
Young J, Wilkie J, Park DC, Watson D (2010) New Zealand strains of plant pathogenic bacteria classified by multi-locus sequence analysis; proposal of Xanthomonas dyei sp. nov. Plant Pathol 59:270–281
Acknowledgements
This study was funded by the Tree Protection Co-operative Programme (TPCP), University of Pretoria and the National Research Foundation (NRF). K.N. Bophela received the NRF/DST Innovation Masters scholarship for financial support during the tenure of the research study. The authors would like to thank the staff at the DNA sequencing facility at the University of Pretoria for assistance with DNA sequencing and staff members of the Forestry and Agricultural Biotechnology Institute (FABI) for the administrative assistance during the study period.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest exists.
Rights and permissions
About this article
Cite this article
Bophela, K.N., Venter, S.N., Wingfield, M.J. et al. Xanthomonas perforans: a tomato and pepper pathogen associated with bacterial blight and dieback of Eucalyptus pellita seedlings in Indonesia. Australasian Plant Pathol. 48, 543–551 (2019). https://doi.org/10.1007/s13313-019-00657-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-019-00657-9