Summary
Ongoing investigations into causes and cures for human movement disorders are important toward the elucidation of diseases such as Parkinson’s disease (PD) and dystonia. The use of animal model systems can provide links to susceptibility factors, as well as therapeutic interventions. In this regard, the nematode roundworm, Caenorhabditis elegans, is ideal for examining age-dependent neurodegenerative disease studies. It is genetically tractable, has a short lifespan, and a well-defined nervous system. Green fluorescent protein is readily visualized in C. elegans because it is a transparent organism, thus the nervous system and factors that alter the viability of neurons can be directly examined in vivo. Through expression of the human PD-associated protein (α-synuclein in the worm dopamine neurons), neurodegeneration is observed in an age-dependent manner. Furthermore, expression of the early-onset dystonia-related protein torsinA increases vulnerability to endoplasmic reticulum (ER) stress in C. elegans, because torsinA is located in the ER. Here we provide an overview of collaborative studies we have conducted that collectively demonstrate the usefulness of the nematode model to discern functional effectors of dopaminergic neurodegeneration and ER stress that translate to mammalian data in the fields of PD and dystonia. Taken together, the application of C. elegans toward the evaluation of genetic modifiers for movement disorders research has predictive value and serves to accelerate the path forward for therapeutic interventions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Attributes of Caenorhabditis elegans as a Model System
Many characteristics and physical attributes of the nematode Caenorhabditis elegans are advantageous for use in biomedical research. Reaching a length of only 1.5 mm in adulthood, this microscopic, free-living roundworm is a convenient model organism that is inexpensive and easy to maintain in the laboratory. Unlike larger, more complex mammalian or cell culture models of disease, worms do not require expensive or bulky facilities for the propagation and upkeep of large numbers of animals, as thousands of worms may be cultivated on a single Petri dish of agar seeded with Escherichia coli. C. elegans has an average lifespan of 2 weeks and undergoes a rapid life cycle with hermaphrodites producing eggs continuously from day 3, resulting in the production of approximately 300 progeny per hermaphrodite. This constant replenishment represents a practically infinite source of animals for experimentation and enables researchers to rapidly conduct large genetic and pharmacological screens, processes that would require significantly more time, labor, and resources in a mammalian system. In addition, hundreds of animals may be analyzed for each data point, a clear statistical advantage in comparison with other models in which such sample sizes are impossible.
The complete cell lineage of C. elegans has been known for more than 2 decades. The adult hermaphrodite possesses 959 somatic cells, precisely 302 of which are neurons. The relatively small number of neurons in the worm represents a conveniently quantifiable population of cells that may be used to discern the effects of genetic and environmental factors on neuron health and function. The fate and synaptic connections of the neurons of C. elegans are well characterized [1]. Readily available references for the lineage and connectivity of each of the neurons provide a valuable resource for investigating the phenotypic effects associated with genes of interest [2]. In addition, the transparent anatomy of this nematode body facilitates the use of fluorescent markers, such as green fluorescent protein (GFP) to visualize the localization and function of select gene products, as well as the consequences of genetic and environmental manipulation in vivo [3]. The use of such markers enables researchers to quickly determine the effects of genetic modifiers and pharmacological targets of interest on cell death as a result of neurodegeneration.
The genetic tractability of the nematode system is one of its most useful traits. C. elegans researchers use a broad and growing tool kit for genetic manipulation of the organism. Worms are particularly amenable to genetic crosses, as well as RNA interference (RNAi) and microinjection, techniques used in gene knockdown and overexpression, respectively. Due to the relative ease with which this organism may be genetically altered, many genes can be screened using the organism in a relatively short amount of time to identify key genes of interest. Similar experiments in mammalian models present logistical challenges pertaining to the use of far more costly, time-consuming, and labor-intensive techniques, hindering the progress of genetic screens and unavoidably narrowing the scope of such investigations. Likewise, complementary experiments performed in neuronal cell culture lack some of the advantages displayed by the whole animal approach, such as behavioral phenotypes and an intact nervous system, whereby the physiological responses will reflect normal neuronal connectivity. If primary cell culture is used in an approach, added time and technical difficulty must also be considered. Taken together, C. elegans can serve either as an intermediate level of validation for genetic or chemical analyses or as an initial discovery model.
At least 50% of worm genes have human homologs, and C. elegans orthologs exist for ~70% of known human disease-related genes [4]. The genetic overlap between C. elegans and Homo sapiens , although not as significant as the overlap between humans and more complex model organisms, enables researchers to use this simple model system to study the functions and interactions of genes contributing to basic cellular pathways relevant to human disease. Due to its long history as a model system, a wealth of bioinformatic and functional genomic data from nearly 4 decades of research are available for C. elegans, providing a substantive base from which future studies may be launched.
Although the nervous system of C. elegans is relatively simple, worms share cellular and molecular pathways with higher organisms. In the context of the nervous system, worms use many of the same neurotransmitters as humans, including dopamine (DA), serotonin, and gamma-aminobutyric acid, as well as ion channels, receptors, and vesicular transporters [5, 6]. Moreover, C. elegans is susceptible to many of the toxins used to model neurodegeneration in mammals, including 6-hydroxydopamine and 1-methyl 4-phenyl 1,2,3,6-tetrahydropyridine [7, 8]. This susceptibility enables researchers to use the worm model in evaluating genetic and chemical factors that modulate response to such toxins.
Taken together, these characteristics of C. elegans lend themselves well to rapidly and efficiently conducting large-scale screens of multiple genes that may be involved in the health of the nervous system, the results of which may be further pursued in higher organisms. As a model organism, C. elegans is already being used as a valuable tool for the initial identification of genes linked to evolutionarily conserved cellular pathways, such as cell death and synaptic transmission [9, 10]. Directed investigation into genetic components of such pathways in worm models of human disease may aid in the effort to understand the complex molecular underpinnings of diseases of neurodegeneration.
Parkinson’s Disease
Application of C. elegans toward Investigation of Parkinson’s Disease
Parkinson’s disease (PD) is the second most common neurodegenerative disease affecting approximately 1 million Americans, with an estimated 7 to 10 million people affected worldwide. Symptomatically, PD is characterized by the presence of tremors, impaired balance, and bradykinesia with severity that progresses in time. Although there are rare cases of early onset PD, it is primarily a disease of aging, typically affecting people age 60 and older. It is estimated that 1 to 2% of the population age 65 years and older is affected by this disease. Pathologically, PD results from the progressive loss of dopamine neurons selectively from the substantia nigra region of the midbrain. The cellular hallmarks of PD are intracellular proteinaceous inclusions termed Lewy bodies, which can be detected (postmortem) in the brains of PD patients.
Multiple genes have been linked to PD, including PARK1 or α-synuclein (α-syn), which were discovered as a cause for familial PD in 1997 [11]. The α-syn aggregates, normally found primarily in the presynaptic termini, form the majority of the insoluble, intracellular protein fibrils that comprise Lewy bodies. Both overexpression of wild-type (WT) α-syn and mutations in α-syn have been linked to rare, early onset forms of PD, but inclusions containing α-syn are also observed in the more common sporadic cases of PD. Although the precise cellular function of α-syn is not yet known, recent work has suggested that it normally serves as a molecular chaperone in the formation of SNARE complexes at the synapse, suggesting a role in synaptic transmission [12].
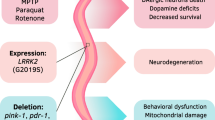
In C. elegans, genetic analyses of PD-related genes are facilitated by the presence of worm orthologs for 6 known familial PD genes, such as PARK8/LRRK2, PARK6/PINK1, and a homolog for the Gaucher disease-linked glucocerebrosidase gene product [13]. These 7 gene products are listed in Table 1. C. elegans does not possess an ortholog for α-syn, thus overexpression of both wild-type and mutant human α-syn may be carried out in the worm without concern for functional interference by endogenous α-syn. Expression of human α-syn under the DA transporter (dat-1) promoter results in age- and dose-dependent neurodegeneration of DA neurons (Fig. 1); α-syn-induced degeneration of neurons in C. elegans has been reported by several laboratories [14–17]. For example, when α-syn mRNA levels are expressed at lower levels in 1 transgenic line (Fig. 2a), DA neurodegeneration is reduced in comparison to another line in which α-syn is expressed at higher levels (Fig. 2b, c).
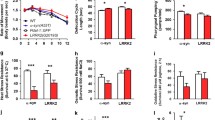
α-synuclein (α-syn) expression causes age- and dose-dependent neurodegeneration. (a) Semi-quantitative RT-PCR of 2 integrated transgenic lines containing both P dat-1 ::α-syn and P dat-1 ::green fluorescent protein (GFP). “α-syn low” exhibits lower α-syn expression than “α-syn high.” (b) The increased α-syn expression in the “α-syn high” transgenic line correlates with greater neurodegeneration compared to that of the “α-syn low” transgenic line, indicating a dose-dependent effect of α-syn on dopamine (DA) neuron survival. (c) Age-dependent DA neurodegeneration resulting from high α-syn dose. Parts (a) and (b) reprinted with permission (Hamamichi S, Rivas RN, Knight AL, Caldwell KA, Caldwell GA. Hypothesis-based RNAi screening identifies neuroprotective genes in a Parkinson’s disease model. Proc Natl Acad Sci USA 2008;105:728-733). (Copyright 2008, National Academy of Sciences, USA). GFP = green fluorescent protein
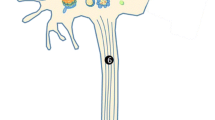
Dopamine (DA) neurons of Caenorhabditis elegans hermaphrodites. (a) The cell bodies and processes are highlighted using green fluorescent protein (GFP) driven from the DA transporter promoter (Pdat-1::GFP). The 6 anterior DA neurons include 2 pairs of cephalic neurons (CEPs) (arrows) and 1 pair of anterior deirid neurons (ADEs) (large arrowheads). One pair of posterior deirid (PDE) neurons (small arrowheads) is also shown. (b) Magnification of the anterior region of C. elegans shows only the 6 anterior-most DA neurons highlighted by GFP (P dat-1 ::GFP) with the 4 CEPs (arrows) and 2 ADE neurons (arrowheads) highlighted. (c) A 7-day-old worm co-expressing both GFP and α-syn in DA neurons display neurodegeneration. Most worms within a population expressing α-syn in the DA neurons are missing anterior DA neurons by day 7. In this example, 3 CEP neurons (arrows) are intact while there are no ADE neurons. (d) Schematic representation of C. elegans anterior DA neurons. The 6 anterior DA neurons of the worm are subclassified in pairs as 2 dorsal CEPs, 2 ventral CEPs, and 2 ADEs. Note the connectivity between dorsal CEPs and ADEs. The dorsal CEPs are postsynaptic to the ADEs, whereas there is no connection between the ventral CEPs and the ADEs
Dopamine Neurons in C. elegans
There are precisely 8 dopaminergic neurons in C. elegans hermaphrodites, identified by their lineage names; they can be visualized using a GFP reporter driven from a DA neuron-specific promoter [18] (Fig. 2a, b, d). Among the 6 DA neurons located in the anterior region of the worm, there are 2 anterior deirid (ADE) neurons and 4 cephalic (CEP) neurons, whereas in the posterior region of the worm there are 2 posterior deirid neurons. Although the 2 ventral CEPs do not have synaptic connections to the ADEs, the 2 dorsal CEPs form synapses with the ADEs (Fig. 2d).
The biosynthesis, processing, packaging, and transport of DA in the worm involve evolutionarily conserved pathways (Table 1). In addition, DA signaling has been linked to several behavioral phenotypes in C. elegans, including a decrease in locomotion, defecation, and egg laying with exposure to exogenous DA. Another DA signaling related phenotype is the basal slowing response, a decrease in locomotion that occurs when a worm enters a bacterial lawn. The basal slowing response is inhibited by disrupting the function of cat-2, the gene encoding worm tyrosine hydroxylase [19]. The small number of DA neurons in C. elegans, their functional similarity to those of humans, and the presence of quantifiable behavioral phenotypes related to DA signaling facilitate the use of the worm as a model system for studying DA neuron biology [20].
Usefulness of C. elegans α-Synuclein Model for Analysis of Neuroprotective Gene Products
Our laboratory has used the reproducible and readily quantifiable DA neurodegeneration following co-expression of α-syn and GFP in nematode DA neurons to examine the effect of modifiers of α-syn toxicity in an in vivo model of α-syn neurodegeneration (Fig. 2c). In a screen for gene products contributing to α-syn toxicity, altered levels of 14-3-3 proteins were identified in the substantia nigra of mice overexpressing human WT α-syn [21]. 14-3-3 s are highly conserved proteins that negatively regulate diverse cell death pathways [22] and have been found to co-localize with α-syn in PD patient Lewy bodies. This time it has been demonstrated that transfection of some 14-3-3 isoforms reduces the formation of α-syn inclusions and protects against rotenone and MPP + toxicity in human neuronal cell culture [23]. To evaluate the protective effects of 14-3-3 s against α-syn neurodegeneration in vivo, our laboratory created double transgenic worms expressing 3 specific 14-3-3 isoforms (ε, γ, and θ) and α-syn under the dat-1 promoter. We determined that expression of both 14-3-3θ and ftt-2, the worm homologue to 14-3-3θ, significantly protected worm DA neurons from α-syn neurodegeneration in this model, thus validating the effect of 14-3-3 s on α-syn toxicity in a more biologically relevant context. Moreover, by crossing α-syn expressing worms into an ftt-2 mutant background, we observed that resulting progeny no longer exhibited neuroprotection in the absence of 14-3-3θ [23] (Fig. 3a). Such traditional genetic analysis augments overexpression data and is easily performed within C. elegans.
The Caenorhabditis elegans α-synuclein model is predictive for gene analysis in dopamine (DA) neuroprotection assays (a). Gene products identified in mammalian systems can be further evaluated for efficacy in protecting DA neurons from α-synuclein-induced DA neurodegeneration by overexpression in C. elegans DA neurons [23, 31]. (b) Yeast screens for suppressors of α-synuclein toxicity have yielded gene candidates that were subsequently examined in C. elegans and drosophila for the ability to reduce the neurodegeneration associated with α-synuclein overexpression. Candidates that reduce toxicity in both yeast and invertebrate neuronal systems also protected rat primary neurons from degeneration following transfection with A53T α-synuclein [24, 28]. (c) A C. elegans hypothesis-based RNA interference (RNAi) screen for effectors of α-synuclein protein misfolding [14] identified candidates that were reported to cause neurodegeneration in a mouse knockout model (ATG7 [37]), as a risk factor for PD (ULK2 [36]), or neuroprotective in human cells when overexpressed (VPS41 [33]). MPP = 1-methyl-4-phenylpyridine
The ease of producing multiple transgenic lines of C. elegans has also allowed us to examine the impact of an array of genes on α-syn-induced dopaminergic neurodegeneration. For example, 2 Rab gene products involved in regulating vesicular trafficking in an α-syn-dependent manner were implicated as modifiers of α-syn toxicity in a yeast model for PD. RAB1, a Rab guanosine triphosphatase involved in endoplasmic reticulum (ER)-to-Golgi vesicle trafficking, and RAB8A, a close paralog to Rab1 that is localized to post-Golgi vesicles and functions in targeting secretory vesicles to the plasma membrane, both co-localized with α-syn clusters in an in vitro model of α-syn-induced trafficking defects. Though expression of RAB1 in yeast suppressed α-syn cytotoxicity, expression of RAB8A did not. The combined use of multiple animal models supported the roles of these proteins in α-syn toxicity. Suppression of cytotoxicity was identified in Drosophila (Rab1), rat neuronal cell cultures (Rab1 and Rab8A), and in C. elegans (Rab1 and Rab8A), where we showed that elevated expression of these specific Rab GTPases rescues DA degeneration induced by human α-syn overexpression [24, 25] (Fig. 3b). New molecules that impact this cellular mechanism have been discovered in yeast and validated across multiple organisms, including C. elegans [26].
Likewise, multiple α-syn toxicity modifiers have been identified through yeast screens using the propensity of this single-celled eukaryote toward ER stress and cell death with the expression of α-syn [27]. Yeast YPK9 (a lysosomal ATPase) was identified in such a screen as a suppressor of α-syn toxicity. Our laboratory has investigated the role of this protein in the DA neurons of the multicellular worm. We have shown that expression of the worm YPK9 ortholog, W08D2.5 (catp-6), suppressed α-syn-induced DA neurodegeneration in the worm. The human YPK9 ortholog (ATP13A2) has been shown to suppress α-syn-induced cell death in rat primary neuronal culture [28] (Fig. 3b). Notably, mutations in ATP13A2 (PARK9) have been shown to cause Kufor-Rakeb syndrome, an autosomal recessive form of early onset parkinsonism and dementia [29]. Mutations in this gene cause a loss of function of the gene product, whereas expression of the normal gene is elevated in the substantia nigra of sporadic PD patients [30].
Lysosomal dysfunction is also associated with α-syn aggregation via a defect in another gene product. In a mouse model of Cathepsin D (CD) deficiency, α-syn accumulates intracellularly without increased α-syn mRNA, suggesting that deficiencies in α-syn protein clearance and not increased expression of α-syn may play a role in the pathology of synucleinopathies. We used the genetic tractability of C. elegans to investigate the effect of human CD on α-syn-induced neurodegeneration. First, we co-expressed human CD within the DA neurons of α-syn-expressing worms and found significant neuroprotection against α-syn-induced neurodegeneration, demonstrating that human CD has a protective effect against α-syn toxicity in vivo. In addition, multiple transgenic lines were generated to further characterize the role of other cathepsin family members in neuroprotection. We found that overexpression of Cathepsin B or Cathepsin L, as well as 2 enzymatic CD mutants, failed to protect worm DA neurons against α-syn neurodegenedration, demonstrating that this effect is both specific to CD and contingent based on its enzymatic function [31] (Fig. 3a).
C. elegans α-Synuclein Model Provides Both a Rapid and Predictive Means for Screening Modifiers
An advantage of the nematode system is the rapidity by which bioassay development can be expedited. Using transgenic C. elegans, we established a strategy for investigating genetic and environmental susceptibility factors contributing to the α-syn-induced DA neurodegeneration. We first overexpressed an α-syn::GFP fusion in the body wall muscles of the worm, whereby visible cytoplasmic aggregation of α-syn was observed. Co-expression of a worm chaperone-like protein (TOR-2) attenuated this effect, leading to a diffuse presence of α-syn in the cytoplasm [14]. Using this genetic background, our laboratory performed a hypothesis-based RNAi screen of ~900 potential α-syn interactors, ultimately identifying 20 genes that enhance misfolding of α-syn in the body wall muscles of the worm. Seven positive hits were further characterized through functional analyses, whereby 5 genes were found to protect against α-syn neurodegeneration when co-expressed with α-syn in the worm DA neurons. Candidates identified through this RNAi screen included DJ-1 and PINK-1, worm orthologs of known recessive PD genes, as well as C. elegans homologs of the human genes encoding VPS41, ULK2, and ATG7 [14] (Fig. 3c).
Although the isolation of DJ-1 and PINK1 served as internal controls for our screening strategy, other research has bolstered the identification of VPS41, ULK2, and ATG7 as modifiers of neurotoxicity. For example, VPS41 has been previously implicated in Golgi to lysosome trafficking in yeast [32] and is strongly expressed in human DA neurons in the substantia nigra. Because of its identification in our screen, the protective capacity of VPS41 has been further analyzed in both the worm and human cell culture. We found that overexpression of human VPS41 protects worm DA neurons from α-syn neurodegeneration, demonstrating that the protective functions of this protein are evolutionarily conserved. VPS41 has also been shown to protect SH-SY5Y cells against both 6-hydroxydopamine (6-OHDA) and rotenone treatment, and to reduce the aggregation of insoluble α-syn in SH-SY5Y cells following rotenone treatment [33] (Fig. 3c).
Furthermore, a novel susceptibility locus for PD (called PARK16) has been identified by a genome-wide association study (GWAS) of multiple PD populations. Included in this locus is the gene encoding RAB7L1, another Ras-related GTP-binding protein. RAB7L1 contains an single nucleotide polymorphism (SNP) within a noncoding region that has recently been associated with the disease through PD patient GWAS [34]. Notably, Rab7 interacts with VPS41 and VPS39, components of the homotypic fusion and vacuole protein sorting (HOPS) complex that promote the formation and fusion of the t-SNARE complex [35]. This connection between VPS41, which was identified from our screen as a modifier of α-syn toxicity and RAB7, a novel PD-related locus identified via GWAS, which opens new directions for future experimental design.
A single nucleotide polymorphism in ULK2, a conserved serine-threonine kinase, identified in our RNAi screen, was also identified as 1 of only 6 significant genes in a GWAS of PD patients that were linked to the disease [36] (Fig. 3c). Another gene product that was positive in our screen (ATG7), which is a lysosomal protein essential for autophagy, displayed correlative results in mammals. Specifically, knockout of ATG7 results in a strong neurodegenerative phenotype in mice [37] (Fig. 3c). Taken together, these collective data demonstrate the predictive power of C. elegans as a model for studying the dopamine neurodegeneration associated with a complex mammalian disease. Furthermore, these data provide evidence for the usefulness of screening in nematodes for rapid identification of gene products potentially involved in disease pathology.
Chemical Modifiers are Readily Analyzed in C. elegans α-Synuclein Dopamine Neurodegeneration Models
In addition to its value for screening for genetic contributors to disease, C. elegans is a useful model for investigating the effects of pharmacological compounds on cellular aspects of disease. For example, we have used our worm model for DA neurodegeneration to evaluate the protective effects of the plecomacrolide antibiotic bafilomycin against α-syn toxicity. Bafilomycin A1 and B1 have been previously shown to protect cerebellar granule neurons against the disruption of lysosome function and subsequent cell death resulting from treatment with the anti-malarial drug chloroquine [38]. Further work has shown that treatment with bafilomycin A1 attenuates chloroquine-induced cell death in both naïve and undifferentiated SH-SY5Y human neuroblastoma cells. To determine the effect of this antibiotic on the toxicity of α-syn aggregation, our laboratory exposed worms expressing human WT α-syn in the DA neurons to bafilomycin during larval development. We found significant protection against α-syn DA neurodegeneration, confirming the functional effect of bafilomycin in a cellular context and strengthening the case for lysosomal dysfunction as a mechanism involved in neurodegeneration [39].
Another pharmacological use for C. elegans is in testing medications already on the market for potential repurposing. As an example, we exposed C. elegans DA neurodegeneration models to the common analgesic acetaminophen to determine if protective effects could result from treatment. Exposure to acetaminophen did not protect DA neurons from degeneration in our worm model for α-syn-induced neurodegeneration. Notably, however, exposure to acetaminophen did protect C. elegans DA neurons from degeneration induced by 2 other mechanisms.
DA neuron death can be induced in C. elegans via acute exposure to 6-OHDA, which results in the production of reactive oxygen species (ROS) and DA neuron death following short-term exposure [7]. Only low concentrations of acetaminophen protected C. elegans DA neurons. However, 6-OHDA induces DA neurodegeneration, which is both rapid and acute, and this effect may be too strong to accurately evaluate the neuroprotective influence of acetaminophen. Therefore, as a supplemental strategy to examine the efficacy of acetaminophen in an ROS model, we also evaluated animals that overexpress tyrosine hydroxylase (TH), the rate-limiting enzyme in the production of DA. Overexpression of the C. elegans CAT-2 gene product (the worm homolog for TH) leads to neurodegeneration that progresses with time as animals age [40], presumably leading to slower ROS formation. Using the TH overexpression model, we treated worms with varying concentrations of acetaminophen and found that all concentrations tested significantly suppressed TH-induced DA neurodegeneration [41].
Although a majority of studies conducted in C. elegans have focused on α-syn models, recently LRRK2 neurodegenerative worm models have also emerged [42, 43]. Significantly, data on small molecule inhibitors of LRRK2 kinase activity and DA neuron loss in C. elegans were confirmed as attenuating neurodegeneration and PD-associated phenotypes in drosophila, as well as both in vitro and in vivo mammalian models of LRRK2-induced neurodegeneration [43, 44].
C. elegans is also being used to investigate several potential environmental toxicants for their contribution to DA neurodegeneration. Included in these studies is the evaluation of exposures to manganese, uranium, methylmercury (MeHgCl), select botanicals, and bacterially produced secondary metabolites [45–49]. Given the unmet need in understanding environmental contributors to neurodegenerative diseases, the mechanistic capacity, experimental accessibility, and limited regulatory requirements surrounding invertebrate research render C. elegans an attractive system for continued expansion of molecular toxicological analyses.
These combined studies exemplify the manner by which C. elegans can be exploited for mechanistic evaluation and validation of chemical modifiers of neurodegeneration. It is believed that chemical uptake occurs primarily through the intestine, but absorption via the cuticle is speculated to occur as well. In this regard, we examined the data gathered from a few researchers who have calculated external versus internal chemical concentrations in worms following chemical exposures. Their data revealed vastly differing results for the fold difference of chemical concentration between the external environment and uptake in animals, whereby the external concentration of different molecules ranged from 8× to 66× more concentrated (Table 2) [50–52]. A more in-depth analysis was performed following MeHgCl exposure in C. elegans, where it was demonstrated that the internal accumulation of this chemical was dose-, developmental stage-, and time-dependent [47]. Thus, the initial exposure parameters for a given substance should be determined empirically. Often, if a phenotypic effect is elicited, than it is assumed that the chemical has been internalized. There are 2 common procedures for exposing worms to chemicals. These include the addition of compounds to the surface of solid agar media on a Petri dish, whereby the animals are exposed by crawling on the surface of the agar, or soaking worms directly in liquid media with chemicals. Furthermore, the dosage of the chemical and the time of administration will vary according to phenotypic response. Importantly, C. elegans are also viable in low concentrations of several different solvents (ethyl acetate, ethanol, methanol, and dimethyl sulfoxide), thus providing experimental flexibility. Therefore, variables, such as mode and time of exposure, as well as solvent, make it difficult to form generalized guidelines for determining the amount of chemical required to be absorbed into nematodes and/or yield a response.
Although drug screening in C. elegans is an efficient starting point for identification of drug targets, it does not supplant mammalian translational studies. Chemicals must be validated in cell culture and animal models, whereby these assays recapitulate some aspects of the disease state under investigation. As previously described, the external/internal concentration of chemical exposure in worms is variable. A recent C. elegans study, in which thousands of chemicals were screened, derived a model for identifying chemical substructures with both increased and decreased likelihood for bioaccumulation and bioactivity in worms [53]. These guidelines are an important resource for designing chemical libraries specifically suited for screening in C. elegans.
However, if the chemical of interest does not possess the optimal molecular scaffold for promoting bioavailability in C. elegans, it still might result in a phenotypic response, albeit at high concentrations. In these cases, the chemical concentrations used in worm assays may not necessarily reflect an accurate measure of efficacy in translational studies and should be used as loose guidelines to initiate mammalian studies. Additional problems related to translational dose response analyses include differences in cell permeability, the experimental time course (important for diseases of aging), and sensitivity to solvents. A survey of a few worm-to-cell culture translational studies reveals wide ranging fold differences between these experimental systems, wherein 0.5× to 160× chemical was used in the C. elegans studies (Table 3). It should be noted, however, that in most of these studies, only a single concentration of chemical was examined in either the worm or cell culture experiments [26, 45, 54]. Thus, it is likely that the minimum fold difference required for a biological response is not accurately represented in these studies, nor was it the goal for the cited literature. However, in the case of 1 such study [39], comparative dose response curves were performed for both C. elegans and SH-SY5Y cells, and the study revealed a 160-fold difference in the drug required for a similar phenotypic response (Table 3).
Nevertheless, the limitations and caveats of the C. elegans assays previously described are countered by savings in both time and expense. Furthermore, conserved metabolic pathways and mechanisms can be further examined quickly using a host of mutant strains (or RNAi) in combination with chemical exposures, and these can be combined with microarray or proteomic studies as well. In this regard, inclusion of C. elegans models as in vivo assay systems in the drug discovery process can provide an intermediate level of screening for lead identification or optimization and functional analysis.
Dystonia
Dystonia is an inclusive term used to define a series of clinically related disorders characterized by involuntary muscle contractions affecting different areas, often twisting parts of the body into abnormal movements or postures [55]. Mechanistic in-roads toward understanding and treatment of dystonia have been limited, but significant progress has been made on specific forms of the disorder. Most research in animal models has focused on DYT1 or early onset generalized torsion dystonia (EOTD), the most common and severe heritable form of the disease, which is linked to a single codon deletion within the gene encoding torsinA [56]. The pathology of EOTD is poorly understood. There is a lack of apparent cellular degeneration associated with the neurological dysfunction of EOTD that complicates the effort to search for molecular mechanisms contributing to this disease. To circumvent this limitation, efforts have focused on uncovering functions for torsinA and its potential implications in disease.
The absence of the dystonia-associated glutamic acid (ΔE302) residue in torsinA results in the formation of aberrant membranous inclusions and redistribution of the protein to the nuclear envelope in neurons [57–59], thereby resulting in a net loss of native torsinA function at the ER. Other studies have implicated interactions between torsinA and cytoskeletal proteins termed nesprins that mediate connections between the nuclear envelope and cytoskeleton [60].
Human torsinA is a member of the large, structurally and functionally diverse ATPases Associated with cellular Activity (AAA+) family of ATPase proteins [61]. These proteins use energy released by ATP hydrolysis to remodel their target substrates and are involved in processes that can be broadly subclassified as having roles in microtubule binding/severing, nondestructive recycling of proteins, or protein quality control [62]. Torsins are an evolutionarily conserved subfamily of the AAA + proteins, with orthologs across all sequenced invertebrate and mammalian genomes, which appear to have arisen uniquely in metazoans. Unlike any other AAA + protein, however, the subcellular location of all characterized torsin homologs, to date, is the lumen of the ER [63–65]. Significantly, human fibroblasts from DYT1 patients also display a deficit for the processing of proteins through the secretory pathway [66].
Application of C. elegans to Dystonia Research
The gap in functional understanding of torsinA led to the development of C. elegans models to evaluate torsin activity in vivo. Initial studies in C. elegans on torsin-related proteins were focused on an embryonically expressed, ER-localized ortholog termed, OOC-5. C. elegans ooc-5 mutants are defective for nuclear rotation and spindle positioning in early worm embryogenesis [63, 67]. Evidence from in vitro biochemical studies and transgenic nematodes expressing variants of OOC-5 revealed that redox changes associated with intrinsic disulfide bond formation appear to affect the binding of ATP and ADP, and that mutants exhibit a low embryo hatch rate compared with control animals [68]. Studies on another torsinA ortholog that is neuronally expressed (TOR-2), as well as ectopically expressed human torsinA, showed that torsin proteins function in a neuroprotective capacity to combat cellular stressors that induce oxidative damage (6-OHDA, overproduction of DA), as well protein misfolding [40]. Overexpression of human torsinA, TOR-2, or OOC-5 in C. elegans also reduced the aggregation of polyglutamine-repeat containing proteins [69]. Interestingly, a physical association between torsinA and the α-syn aggregates of Lewy bodies in PD patients further suggests a role for the gene in misfolded protein management [70]. This is further supported by co-expression of worm TOR-2 ameliorating α-syn::GFP misfolding in the body wall muscles of C. elegans [14].
The absence of neurodegeneration in dystonia suggests that more subtle causes of cellular dysfunction underlie the disease state. Moreover, the fact that only 30 to 40% of DYT1 carriers display symptoms is further indication that disease penetrance is potentially an outcome of unknown environmental influences and/or cellular processes that are more stochastic in nature [71]. Taken together, functional studies in C. elegans imply a cellular role for torsinA as a potential redox-regulated protein with chaperone-like activity. These data correlate with a hypothesis whereby dystonia is a manifestation of a failure in a torsinA-mediated mechanism that normally acts to maintain homeostatic conditions of protein function in neurons.
To address this hypothesis, Chen et al. [72] generated several transgenic nematode lines that expressed different variants of human torsinA to evaluate their differential impact on ER homeostasis and the unfolded protein response. These experiments demonstrated that unaltered WT torsinA dramatically attenuated ER stress induced by tunicamycin, whereas mutant torsinA (ΔE) lost this cytoprotective property (Fig. 4a-d). Co-expression of both WT and ΔE together resulted in a level of ER stress equal to that of mutant expressed in the absence of WT, in congruence with dysfunction exhibited by a dominant negative loss-of-function. A structure function analysis of torsinA in C. elegans further delineated that the stress alleviating activity of torsinA was limited to isoforms predicted to localize at the ER. Importantly, through the use of mouse embryonic fibroblasts (MEFs) isolated from torsinA knockout mice, it was further demonstrated that the loss of endogenous torsinA leads to increased sensitivity to ER stress (Fig. 4d). These data were extended by a subsequent report showing that torsinA associates with components of the ER-associated degradation pathway, facilitating processing and selective degradation of improperly folded proteins, including the mutant cystic fibrosis transmembrane conductance regulator (CFTR) protein (Fig. 4d) [73].
TorsinA-mediated response to endoplasmic reticulum (ER) stress. (a-c) Fluorescent micrographs depicting Caenorhabditis elegans strains expressing the ER stress reporter hsp-4::green fluorescent protein (GFP) alone (a), with wild-type (WT) torsinA (b), or with mutant torsinA (ΔE) (c). (d) Transgenic C. elegans expressing no torsinA (hsp-4::GFP alone), or overexpressing torsinA (ΔE), or WT/ΔE torsinA have a statistically enhanced ER stress response (based on GFP pixel intensities) compared to C. elegans overexpressing WT torsinA, in which the ER stress response is significantly reduced. These data were validated in mouse embryonic fibroblasts (MEFs) isolated from WT (+/+) or torsinA knockout animals (-/-) whereby the absence of endogenous torsinA increases ER stress response ([from dithiothreitol (DTT) or tunicamycin]) in torsinA -/- MEFs [72]. (e) WT torsinA also reduces the ER stress response resulting from co-expression of mutant cystic fibrosis transmembrane conductance regulator (CFTR Δ508). Likewise, in a comparison of the ER stress response between DYT1 patient fibroblasts (WT/ΔE) and control fibroblasts (WT), the patient cells are more sensitive to ER stress [73]. (f) Human genetic studies have shown that individuals who carry the torsinA (ΔE) allele are far more likely to be nonmanifesting carriers if they also have a polymorphism at position 216 that is protective in trans (216H) and individuals with the D216 allele are far more likely to show the symptoms of DYT1 dystonia [74]. In C. elegans, co-overexpression of the protective 216H and the mutant ΔE form of torsinA result in significantly reduced ER stress response compared to ΔE alone is similar to WT torsinA [72]. Conversely, significantly enhanced ER stress is observed following co-expression of D216 and ΔE torsinA [72]
C. elegans Bioassays for Genetic and Chemical Modifiers of TorsinA Activity are Predictive
Human genetic analyses by Risch et al. [74] showed that that a “protective” polymorphism in torsinA (D216H) was found at a higher frequency in nonmanifesting DYT1 carriers, and was decreased in affected DYT1 dystonia patients, when carried in trans to the ΔE allele [74]. In this context, combinations of transgenic C. elegans engineered to simulate these clinically relevant genetic modifiers of disease susceptibility for EOTD exhibited a direct functional concordance to changes in ER stress response [72]. As the value of any animal model is the means by which it recapitulates the human disease state at either a phenotypic and/or functional level, these collective results indicate that ER-associated C. elegans assays functionally mirror the disease modifying impact of dystonia-related mutations (Fig. 4d).
Key to such studies is the ability to generate multiple lines of isogeneic nematodes carrying human transgene variants that facilitate a comprehensive functional and quantitative evaluation of torsinA activity in significant numbers of animals. This advantage of C. elegans research can also facilitate systematic drug discovery efforts in distinct genetic backgrounds. Cao et al. [54] reported results of a C. elegans-based small molecule screen to identify compounds that modulate the chaperone-like activity of torsinA. Using multiple transgenic lines and thousands of animals, molecules that selectively altered the activity of WT versus ΔE torsinA with statistically substantial effect could be delineated; this facilitated prioritization for downstream experimentation in mammalian models. Strikingly, one compound, ampicillin, was shown to restore functional capacity of torsinA to human DYT1 dystonia patient fibroblasts defective in protein processing. Administration of this drug also reversed behavioral abnormalities in a DYT1 knock-in mouse model of EOTD [54, 75]. Although an antibiotic is not ideal for chronic dosing regimens in dystonia, the identification of a compound that selectively modifies torsinA activity across multiple cellular and organismal models represents a significant molecular lead toward novel treatment alternatives. Beyond this example of the predictive nature of C. elegans, the worm model represents both an expeditious and cost-effective bottleneck in the translational path.
Conclusion
Here we have outlined distinct facets of C. elegans models that have provided significant insights into how specific disease-related proteins contribute to the dysfunction associated with Parkinson’s disease and dystonia. Pioneering technical advances, such as RNAi and GFP, generated as an outcome of basic science using C. elegans, have already enhanced translational studies in various systems and represent the powerful promise of the nematode system. Screens for genetic and chemical modifiers of disease-associated phenotypic readouts in C. elegans may provide further insights or therapeutic leads for movement disorders. Finally, as genomic studies continue to expand the informational database of human genetic variation, the integrated application of worm-based assays to discern the functional significance of specific genetic changes has the promise of substantially accelerating our understanding of disease mechanism and susceptibility.
References
White JG, Southgate E, Thomson JN, Brenner S. The structure of the nervous system of Caenorhabditis elegans. Philos Trans R Soc Lond 1986;275:327-348.
Altun ZF, Hall DH. In : WormAtlas available at http://www.wormatlas.org. Accessed September 16, 2011.
Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC. Green fluorescent protein as a marker for gene expression. Science 1994;263:802-805.
Schwarz EM. Genomic classification of protein-coding gene families. In: The C. elegans Research Community, ed. WormBook 2005, doi/10.1895/wormbook.1.29.1, http://www.wormbook.org.
Chalfie M, White J. The nervous system. In: Wood WB, ed. The Nematode Caenorhabditis elegans, Cold Spring Harbor Laboratory Press, Cold Spring Harbor: NY 1988.
Bargmann CI. Neurobiology of the C. elegans Genome. Science 1998;282:2028-2033.
Nass R, Hall DH, Miller DM 3 rd, Blakely RD. Neurotoxin-induced degeneration of converted dopamine neurons in Caenorhabditis elegans. Proc Natl Acad Sci USA 2002;99:3264-3269.
Braungart E, Gerlach M, Riederer P, Baumeister R, Hoener MC. Caenorhabditis elegans MPP + model of Parkinson’s disease for high-throughput drug screenings. Neurodegener Dis 2004;1:175-183.
Potts MB, Cameron S. Cell lineage and cell death: Caenorhabditis elegans and cancer research. Nat Rev Cancer 2011;11:50-58.
Richmond J. Synaptic function. In: The C. elegans Research Community, ed., WormBook 2007; doi:10.1895/wormbook.1.69.1, http://www.wormbook.org.
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M. Alpha-synuclein in Lewy bodies. Nature 1997;388:839-840.
Burré J, Sharma M, Tsetsenis T, Buchman V, Etherton MR, Südhof TC. α-synuclein promotes SNARE-complex assembly in vivo and in vitro. Science 2010;329:1663-1667.
Mazzulli JR, Xu YH, Sun Y, et al. Gaucher disease glucocerebrosidase and α-synuclein form a bidirectional pathogenic loop in synucleinopathies. Cell 2011;146:37-52.
Hamamichi S, Rivas RN, Knight AL, Cao S, Caldwell KA, Caldwell GA. Hypothesis-based RNAi screening identifies neuroprotective genes in a Parkinson’s disease model. Proc Natl Acad Sci USA 2008;105:728-733.
Lakso M, Vartiainen S, Moilanen AM, et al. Dopaminergic neuronal loss and motor defecits in Caenorhabditis elegans overexpressing human alpha-synuclein. J Neurochem 2003;86:165-172.
Kuwahara T, Koyama A, Gengyo-Ando K, et al. Familial Parkinson mutant alpha-synuclein causes dopamine neuron dysfunction in transgenic Caenorhabditis elegans. J Biol Chem 2006;281:334-340.
Karpinar DP, Balija MB, Kügler S, et al. Pre-fibrillar alpha-synuclein variants with impaired beta-structure increase neurotoxicity in Parkinson's disease models. Mol Syst Biol 2009;28:3256-3268.
Nass R, Miller DM, Blakely RD. C. elegans: a novel pharmacogenetic model to study Parkinson’s disease. Parkinsonism Related Disord 2001;7:185-191.
Sawin ER, Ranganathan R, Horvitz HR. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron 2000;26:619-631
Vidal-Gadea A, Topper S, Young L, et al. Caenorhabditis elegans selects distinct crawling and swimming gaits via dopamine and serotonin. Proc Natl Acad Sci USA 2011;108:17504-17509.
Yacoubian TA, Cantuti-Castelvetri I, Bouzou B, et al. Transcriptional dysregulation in a transgenic model of Parkinson disease. Neurobiol Dis 2008;29:515-528.
Porter GW, Khuri FR, Fu H. Dynamic 14-3-3/client protein interactions integrate survival and apoptotic pathways. Semin Cancer Biol 2006;16:193-202.
Yacoubian TA, Slone SR, Harrington AJ, et al. Differential neuroprotective effects of 14-3-3 proteins in models of Parkinson's disease. Cell Death Dis 1;e2; doi:10.1038/cddis.2009.4
Cooper AA, Gitler AD, Cashikar A, et al. Alpha-synuclein blocks ER-Golgi traffic and Rab1 rescues neuron loss in Parkinson's models. Science 2006;313:324-328.
Gitler AD, Bevis BJ, Shorter J, et al. The Parkinson's disease protein alpha-synuclein disrupts cellular Rab homeostasis. Proc Natl Acad Sci USA 2008;105:145-150.
Su LJ, Auluck PK, Outeiro TF, et al. Compounds from an unbiased chemical screen reverse both ER-to-Golgi trafficking defects and mitochondrial dysfunction in Parkinson's disease models. Dis Model Mech 2010;3:194-208.
Yeger-Lotem E, Riva L, Su LJ, et al. Bridging high-throughput genetic and transcriptional data reveals cellular responses to α-synuclein toxicity. Nat Genet 2009;41:316-323.
Gitler AD, Chesi A, Geddie ML, et al. Alpha-synuclein is part of a diverse and highly conserved interaction network that includes PARK9 and manganese toxicity. Nat Genet 2009;41:308-315.
Ramirez A, Heimbach A, Gründemann J, et al. Hereditary parkinsonism with dementia is caused by mutations in ATP13A2, encoding a lysosomal type 5 P-type ATPase. Nat Genet 2006;38:1184-1191.
Klein C, Lohmann-Hedrich K. Impact on recent genetic findings in Parkinson’s disease. Curr Opin Neurol 2007;20:453-464.
Qiao L, Hamamichi S, Caldwell KA, et al. Lysosomal enzyme cathepsin D protects against alpha-synuclein aggregation and toxicity. Mol Brain 2008;1:17.
Bowers K, Stevens TH. Protein transport from the late Golgi to the vacuole in the yeast Saccharomyces cerevisiae. Biochim Biophys Acta 2005;1744:438-454.
Ruan Q, Harrington AJ, Caldwell KA, Caldwell GA, Standaert DG. VPS41, a protein involved in lysosomal trafficking, is protective in Caenorhabditis elegans and mammalian cellular models of Parkinson’s disease. Neurobiol Dis 2010;37:330-338.
Tucci A, Nalls MA, Houlden H, et al. Genetic variability at the PARK16 locus. Eur J Hum Genet 2010;18:1356-1359.
Brett CL, Plemel RL, Lobinger BT, Vignali M, Fields S, Merz AJ. Efficient termination of vacuolar Rab GTPase signaling requires coordinated action by a GAP and a protein kinase. J Cell Biol 2008;182:1141-1151.
Fung HC, Scholz S, Matarin M, et al. Genome-wide genotyping in Parkinson’s disease and neurologically normal controls: first stage analysis and public release of data. Lancet Neurol 2006;5:911-916.
Komatsu M, Waguri S, Chiba T, et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 2006;441:880-884.
Shacka JJ, Klocke BJ, Shibata M, et al. Bafilomycin A1 inhibits chloroquine-induced death of cerebellar granule neurons. Mol Pharmacol 2006;69:1125-1136.
Pivtoraiko VN, Harrington AJ, Mader BJ, et al. Low-dose bafilomycin attenuates neuronal cell death associated with autophagy-lysosome pathway dysfunction. J Neurochem 2010;114:1193-1204.
Cao S, Gelwix CC, Caldwell KA, Caldwell GA. Torsin-mediated protection from cellular stress in the dopaminergic neurons of Caenorhabditis elegans. J Neurosci 2005;25:3801-3812.
Locke CJ, Fox SA, Caldwell GA, Caldwell KA. Acetaminophen attenuates dopamine neuron degeneration in animal models of Parkinson's disease. Neurosci Lett 2008;439:129-133.
Yao C, El Khoury R, Wang W, et al. LRRK2-mediated neurodegeneration and dysfunction of dopaminergic neurons in a Caenorhabditis elegans model of Parkinson's disease. Neurobiol Dis 2010;40:73-81.
Liu Z, Hamamichi S, Lee BD, et al. Inhibitors of LRRK2 kinase attenuate neurodegeneration and Parkinson-like phenotypes in Caenorhabditis elegans and Drosophila Parkinson's disease models. Hum Mol Genet 2011;20:3933-3942.
Lee S, Liu H, Lin W, Guo H, Lu B. LRRK2 Kinase regulates synaptic morphology through distinct substrates at the presynaptic and postsynaptic compartments of the drosophila neuromuscular junction. J Neurosci 2010;30:16959-16969.
Caldwell KA, Tucci ML, Armagost J, et al. Investigating bacterial sources of toxicity as an environmental contributor to dopaminergic neurodegeneration. PLoS One 2009;4:e7277.
Helmcke KJ, Syversen T, Miller DM 3 rd, Aschner M. Characterization of the effects of methylmercury on Caenorhabditis elegans. Toxicol Appl Pharmacol 2009;240:265-272.
Jiang GC, Hughes S, Stürzenbaum SR, Evje L, Syversen T, Aschner M. Caenorhabditis elegans metallothioneins protect against toxicity induced by depleted uranium. Toxicol Sci 2009;111:345-354.
Vanduyn N, Settivari R, Wong G, Nass R. SKN-1/Nrf2 inhibits dopamine neuron degeneration in a Caenorhabditis elegans model of methylmercury toxicity. Toxicol Sci 2010;118:613-624.
Jadiya P, Khan A, Sammi SR, Kaur S, Mir SS, Nazir A. Anti-Parkinsonian effects of Bacopa monnieri: insights from transgenic and pharmacological Caenorhabditis elegans models of Parkinson's disease. Biochem Biophys Res Commun 2011;413:605-610.
McColl G, Killilea DW, Hubbard AE, Vantipalli MC, Melov S, Lithgow GJ. Pharmacogenic analysis of lithium-induced delayed aging in Caenorhabditis elegans. J Biol Chem 2007;283:350-357.
Davies AG, Pierce-Shimomura JT, Kim H, et al. A central role of the BK potassium channel in behavioral responses to ethanol in C. elegans. Cell 2003;115:655-666.
Evanson K, Huang C, Yamben I, Covey DF, Kornfeld K. Anticonvulsant medications extend worm life-span. Science 2005;307:258-262.
Burns AR, Wallace IM, Wildenhain J, et al. A predictive model for drug bioaccumulation and bioactivity in Caenorhabditis elegans. Nat Chem Biol 2010;6:549-557.
Cao S, Hewett JW, Yokoi F, et al. Chemical enhancement of torsinA function in cell and animal models of torsion dystonia. Dis Model Mech 2010;3:386-396.
Breakefield XO, Blood AJ, Li Y, Hallett M, Hanson PI, Standaert DG. The pathophysiological basis of dystonias. Nat Rev Neurosci 2008;9:222-234.
Ozelius LJ, Hewett JW, Page CE, et al. The early-onset torsion dystonia gene (DYT1) encodes an ATP-binding protein. Nat Genet 1997;17:40-48.
Goodchild RE, Dauer WT. Mislocalization to the nuclear envelope: An effect of the dystonia-causing torsinA mutation. Proc Natl Acad Sci USA 2004;101:847-852.
Naismith TV, Heuser JE, Breakefield XO, Hanson PI. TorsinA in the nuclear envelope. Proc Natl Acad Sci USA 2004;101:7612-7617.
Torres GE, Sweeney AL, Beaulieu JM, Shashidharan P, Caron MG. Effect of torsinA on membrane proteins reveals a loss of function and a dominant-negative phenotype of the dystonia-associated DeltaE-torsinA mutant. Proc Natl Acad Sci USA 2004;101:15650-15655.
Nery FC, Zeng J, Niland BP, et al. TorsinA binds the KASH domain of nesprins and participates in linkage between nuclear envelope and cytoskeleton. J Cell Sci 2008;121:3476-3486.
Neuwald AF, Aravind L, Spouge JL, Koonin EV. AAA+: a class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genom Res 1999;9:27-43.
Hanson PI, Whiteheart SW. AAA + proteins: have engine, will work. Nat Rev Mol Cell Biol 2005;6:519-529.
Basham SE, Rose LS. The Caenorhabditis elegans polarity gene ooc-5 encodes a Torsin-related protein of the AAA ATPase superfamily. Development 2001;128:4645-4656.
Kustedjo K, Bracey MH, Cravatt BF. TorsinA and its torsion dystonia-associated mutant forms are lumenal glycoproteins that exhibit distinct subcellular localizations. J Biol Chem 2000;275:27933-27939.
Liu Z, Zolkiewska A, Zolkiewski M. Characterization of human torsinA and its dystonia-associated mutant form. Biochem J 2003;374:117-122.
Hewett JW, Tannous B, Niland BP, et al. Mutant torsinA interferes with protein processing through the secretory pathway in DYT1 dystonia cells. Proc Natl Acad Sci USA 2007;104:7271-7276.
Basham SE, Rose LS. Mutations in ooc-5 and ooc-3 disrupt oocyte formation and the reestablishment of asymmetric PAR protein localization in two-cell Caenorhabditis elegans embryos. Dev Biol 1999;215:253-263.
Zhu L, Wrabl JO, Hayashi AP, Rose LS, Thomas PJ. The torsin-family AAA + protein OOC-5 contains a critical disulfide adjacent to Sensor-II that couples redox state to nucleotide binding. Mol Biol Cell 2008;19:3599-3612.
Caldwell GA, Cao S, Sexton EG, Gelwix CC, Bevel JP, Caldwell KA. Suppression of polyglutamine-induced protein aggregation in Caenorhabditis elegans by torsin proteins. Hum Mol Genet 2003;12:307-319.
Sharma N, Hewett J, Ozelius LJ, et al. A close association of torsinA and alpha-synuclein in Lewy bodies: a fluorescence resonance energy transfer study. Am J Pathol 2001;159:339-344.
Bressman SB. Dystonia: phenotypes and genotypes. Rev Neurol (Paris) 2003;159:849-856.
Chen P, Burdette AJ, Porter JC, et al. The early-onset torsion dystonia-associated protein, torsinA, is a homeostatic regulator of endoplasmic reticulum stress response. Hum Mol Genet 2010;19:3502-3515.
Nery FC, Armata IA, Farley JE, et al. TorsinA participates in endoplasmic reticulum-associated degradation. Nat Commun 2:393. doi: 10.1038/ncomms1383.
Risch NJ, Bressman SB, Senthil G, Ozelius LJ. Intragenic Cis and Trans modification of genetic susceptibility in DYT1 torsion dystonia. Am J Hum Genet 2007;80:1188-1193.
Dang MT, Yokoi F, McNaught KS, et al. Generation and characterization of Dyt1 DeltaGAG knock-in mouse as a model for early-onset dystonia. Exp Neurol 2005;196:452-463.
Acknowledgments
We would like to thank all the members of the Caldwell Laboratory for their contributions, collegiality, and teamwork pertaining to this research. Special thanks go to all our collaborators and colleagues for their collective contributions to the scientific findings summarized. We are sincerely grateful to the many organizations and patient groups that have funded movement disorders research for their insightful support in the establishment and continued application of the worm model system in our laboratory and others. Paige Dexter is a Parkinson Association of Alabama and Goldwater Scholar; she was also a University of Alabama-Howard Hughes Medical Institute Undergraduate Researcher from the Howard Hughes Medical Institute Undergraduate Science Program Grant to The University of Alabama.
Required Author Forms
Disclosure forms provided by the authors are available with the online version of this article.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 510 kb)
Rights and permissions
About this article
Cite this article
Dexter, P.M., Caldwell, K.A. & Caldwell, G.A. A Predictable Worm: Application of Caenorhabditis elegans for Mechanistic Investigation of Movement Disorders. Neurotherapeutics 9, 393–404 (2012). https://doi.org/10.1007/s13311-012-0109-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13311-012-0109-x