Abstract
Gallbladder cancer (GBC) is a highly malignant cancer with poor prognosis. Although long noncoding RNA (lncRNA) H19 has been reported to play vital role in many human cancers, whether it is involved in GBC proliferation is still unknown. This study was designed to explore the effect of H19 in GBC cell proliferation. The expression of H19 and AKT2 were significantly elevated in GBC tissues, and the level of miR-194-5p is markedly decreased. Moreover, the RNA levels of H19 and AKT2 were positively correlated, and H19 elevation was significantly associated with tumor size. Cell proliferation decreased significantly after knockdown of H19 in GBC-SD and NOZ cells and after knockdown of AKT2 in NOZ cells. Results from cell cycle studies indicated that the S phase were significantly decreased after knockdown of H19 in NOZ cells but significantly elevated after overexpression of H19 in GBC-SD cells. Furthermore, knockdown of H19 upregulated miR-194-5p levels, yet significantly decreased miR-194-5p targeting AKT2 gene expression in NOZ cells. Inhibitor against miR-194-5p reversed these effects. In addition, overexpression of H19 in GBC-SD cells downregulated miR-194-5p and markedly increased AKT2 expression, and miR-194-5p mimic reversed these effects. Eventually, GBC cells were arrested in G0/G1-phase after H19 knockdown, inhibition of miR-194-5p markedly promoted cells into S-phase and co-transfection of siH19, and miR-194-5p inhibitor exerted mutually counter-regulated effects on cell cycle. These results suggested that H19/miR-194-5p/AKT2 axis regulatory network might modulate cell proliferation in GBC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
GBC is the most common malignant tumor among extrahepatic bile duct cancers, with over 10,000 new cases each year in the USA [1]. Due to an aggressive biological behavior, early lymph node invasion and distant metastases are commonly seen at the time of diagnosis. Moreover, the onset of GBC is in sheath, where sensitive screening test is lacking, resulting in advanced cancer at diagnosis. These features have made GBC a highly lethal disease with an overall 5-year survival of less than 5 % [2–4]. Therefore, more investigations were needed to uncover the molecular mechanism of gallbladder carcinoma, to develop novel anticancer agents, as well as to discover new biological markers for early diagnosis.
Noncoding RNAs (ncRNAs) are RNA molecules that are not translated into proteins. Those longer than 200 nucleotides were named long noncoding RNAs (lncRNAs). Thus far, only a small part of lncRNAs have been compiled [5]. Even so, a growing body of studies suggested that some lncRNAs were involved in tumorigenesis and may be used as biological marker [6]. Several lncRNAs have been proved to be specifically expressed in specific types of cancer and played important roles in tumor progression. For example, highly upregulated in liver cancer (HULC) was a lncRNA which were identified as striking overexpression in hepatocellular carcinoma (HCC) [7]. Loss of neuroblastoma-associated transcript-1 (NBAT-1) promoted cell proliferation and invasion [8].
LncRNA H19 is a 2.3 kb RNA encoded by H19 gene, which is a maternally expressed imprinted gene that plays a vital role in mammalian development [9]. Breast cancer cells ectopically expressing H19 generated bigger colonies in vitro and xenograft tumors compared to the control [10]. It has also been reported that H19 inactivation was linked to the development of Wilms’ tumor [11]. However, the function of H19in GBC has not been demonstrated in any literature.
Protein kinase B (PKB), also known as Akt, has three isoforms including AKT1, AKT2, and AKT3. AKT2 is a putative oncogene which had been shown to be overexpressed in ovarian cancer [12], breast cancer [13], and pancreatic cancer [14]. Galicia VA and colleagues confirmed that loss of PTEN leads to spontaneous development of liver tumors from progenitor cells. Attenuation of hepatic injury by deletion of Akt2 reduced progenitor cell proliferation and delayed tumor development [15]. To date, the role of AKT2 in GBC is still unclear.
In this study, we found that H19 expression was significantly associated with tumor size and was positively correlated to AKT2 expression. Loss and gain of function studies indicated that H19 and AKT2 both promoted GBC cell proliferation. H19/miR-194-5p/AKT2 axis regulatory network might contribute to GBC progression.
Materials and methods
Patients and samples
Twenty GBC tissue samples and matched adjacent noncancerous gallbladder tissue samples were collected from patients with GBC who had undergone surgery in Eastern Hepatobiliary Surgical Hospital, Shanghai, China (between January 2009 and December 2009). All samples were processed by two professional pathology doctors. No patients recruited into the present study received any treatment prior to surgery. All samples were snap-frozen in liquid nitrogen and stored at −80 °C prior to RNA isolation. GBC patients were staged according to the tumor node metastasis (TNM) staging system (the seventh edition) of the American Joint Committee on Cancer (AJCC) staging system. Informed consent was obtained from all patients. Complete clinicopathological follow-up data of the GBC patients from which the specimens were collected were available. This study was approved by the Human Ethics Committee of Xinhua Hospital at Shanghai Jiao tong University (Shanghai, China).
Cell culture
Three human GBC cell lines (GBC-SD, EH-GB1, and NOZ) were used in this study. NOZ cells were purchased from the Health Science Research Resources Bank (Osaka, Japan). GBC-SD and EH-GB1 cells were purchased from Cell Bank of the Chinese Academy of Science (Shanghai, China). Cells were cultured in Dulbecco’s Modified Eagle’s Medium (Gibco BRL, Grand Island, NY, USA), supplemented with 10 % fetal bovine serum (FBS, HyClone, Invitrogen, Camarillo, CA, USA), and 100 μg/ml penicillin and 100 μg/ml streptomycin (Invitrogen, Carlsbad, CA, USA). Cells were incubated at 37 °C with 5 % CO2.
RNA extraction, reverse transcription, and qRT-PCR
Total RNA was extracted from tissues or cells using TRIzol reagent (TaKaRa, Dalian, China) and then converted into complementary DNAs (cDNAs) using the Primer-Script one step RT-PCR kit (TaKaRa, Dalian, China) following the manufacturer’s instruction. The cDNA templates were amplified by real-time PCR using the SYBR Premix Dimmer Eraser kit (TaKaRa, Dalian, China). Gene expression in each sample was normalized to GADPH. The primer sequences were as follows: GAPDH (forward), 5′-CGGAGTCAACGGATTTGGTCGTAT-3′, GAPDH (reverse), 5′-AGCCTTCTCCATGGTGGTGAAGAC-3′; H19 (forward), 5′-TTCAAAGCCTCCACGACTCT-3′; H19 (reverse), 5′-GCTCACACTCACGCACACTC-3′. AKT2 (forward), 5′-AGCACAGGTTCTTCCTCAGC-3′; AKT2 (reverse), 5′-ACTGGGCGGTAAATTCATCA-3′. Real-time RT-PCR reactions were performed by the ABI7500 system (Applied Biosystems, Carlsbad, CA, USA). Data were presented as mean ± SD from three independent experiments. The relative expression fold change of messenger RNA (mRNA) was calculated by the 2−ΔΔCt method.
Cell transfection
The small interfering RNAs (siRNAs) specifically targeting H19 were synthesized by Shanghai Gene Pharma Co, Ltd. The siRNA sequences for H19 were si-H19-1, 5′-CCAACAUCAAAGACACCAUdTdT-3′, and si-H19-2, 5′-UAAGUCAUUUGCACUGGUUdTdT-3′. The siRNA sequences for AKT2 are AKT2, 5′-GCAGAGATTGTCTCGGCTCTT-3′. The pLV-CMV-Not/BamHI–GFP–puro-H19 (PLv-CMV-H19) was produced for the overexpression for H19 in GBC-SD cells (data not show). Plasmids were transfected into cells using Lipofectamine TM 2000 (Invitrogen, Carlsbad, CA, USA) and were incubated for 48 h according to the manufacturer’s instructions. MiR-194-5p mimics and miR-194-5p inhibitors were transfected into NOZ cells using Lipofectamine TM 2000 (Invitrogen, Carlsbad, CA, USA).
Cell proliferation assay
Cell proliferation was assayed using CCK-8 (Beyotime Biotechnology, China) according to the manufacturer’s instruction. The infected cells were seeded at a density of 1 × 104 cells/well in a 96-well flat-bottom cell culture dish. OD value of cells was detected at 0, 24, 48, 72, and 96 h.
Transwell invasion assay
Cell invasion assays were performed using 24-well transwell plates (Corning, NY, USA) pre-coated with Matrigel (BD, Franklin Lakes, NJ, USA). Cells were seeded in the upper chambers of inserts at a density of 1 × 105 cells per well with serum free DMEM. DMEM supplement with 10 % serum was added to lower chambers. After 24 h, the inserts were removed and washed; the cells above the Matrigel layer were carefully removed by swabbing, and the cells below the membrane were fixed in 4 % para-formaldehyde for 10 min and stained with 0.1 % crystal violet for 10 min. After being washed in PBS for three times, cells were viewed and photographed under a phase-contrast microscope (Olympus, Tokyo, Japan) and counted from randomly chosen fields.
Cell-cycle analysis
Cells were harvested by trypsinization and 1 × 105 cells were used for analysis. After being washed in PBS and fixed in ice-cold 70 % ethanol overnight at 4 °C, the cells were washed in PBS and incubated in 500-μL staining solution (10 μg/mL propidium iodide and 5 U/mL RNaseA) for 30 min at 37 °C. Cell cycles were examined by a flow cytometer (FACSort; Becton). The cell-cycle populations were determined by ModFit software.
Protein extraction and western blot analysis
Cells were incubated in lysis buffer and centrifuged at 14,000g for 15 min. The BCA protein assay reagent kit was used to measure protein concentration. Protein samples were separated by 10 % SDS-polyacrylamide gels and then transferred to polyvinylidenedifluoride (PVDF) membranes (Millipore, Bedford, MA) and sequentially incubated with indicated primary antibodies against AKT2 and GAPDH (1:1000, Cell Signaling Technology) at 4 °C overnight on a shaker. Bolts were incubated in Horseradish peroxidase (HRP)-conjugated secondary antibody (1:1000, Abcam) at room temperature for 1 h. The blots were developed using enhanced chemiluminescence detection reagents and scanned with a Molecular Imager system (Bio-Rad).
Statistical analysis
Data were analyzed with SPSS 13.0 software. All experiments were repeated at least three times. Data were presented as the mean ± SD. Data from multiple groups were analyzed with one-way analysis of variance. Data from two groups were compared by t tests. Pearson correlation was used to analyze the correlation between H19 and AKT2. P values less than 0.05 were considered statistically significant.
Results
Elevated expression of H19 in GBC
H19 expression in GBC tissues was examined by real-time RT-PCR. Significantly higher H19 expression was observed in GBC samples compared to that in normal samples (Fig. 1a), and higher H19 was associated with bigger tumor size (P = 0.015) (Fig. 1b, Table 1). Moreover, other clinical variables including age, gender, histological grade, clinical stage, and lymphatic metastasis were not associated with H19 expression (Table 1).
Expression levels of H19 in cancer tissues and cells. a H19 expression was significantly upregulated in GBC tissues. H19 expression was measured by qRT-PCR and normalized to GAPDH. b H19 expression level was significantly associated with tumor sizes (tumor size > 5 cm vs tumor size <5 cm, P < 0.05). c Expression levels of H19 in three GBC cell lines (GBC-SD, EH-GB1, and NOZ). Data were presented as fold change over GAPDH. d qRT-PCR analysis of the efficacy of H19 knockdown in NOZ cells. e The efficacy of overexpression of H19 and GBC-SD cells was transfected with plv-CMV-GFP-H19 lentivirus. All data were represented as the mean ± S.D. from three independent experiments. **P < 0.05
To explore the effect of H19 on cell proliferation and invasion, three GBC cell lines (GBC-SD, EH-GB1, NOZ) expressing various levels of H19 were used in this study (Fig. 1c). Quantitative real-time polymerase chain reaction (qRT-PCR) analyses indicated that the expression of H19 in NOZ cells was reduced by 50 % or 80 % after si-H19-1 or si-H19-2 transfection, respectively (Fig. 1d). Cells transfected with si-H19-2 were used as the representative in the following experiments due to more efficient suppression of H19. Overexpression of H19 was also successful in GBC-SD cells transfected PLv-CMV-H19, with a 10.5-fold induction of H19 compared to control cells (Fig. 1e).
H19 promotes cell proliferation and invasion in GBC cells
In both NOZ and GBC-SD cell lines, proliferation was inhibited by si-H19 compared to control group in CCK8 assay (Fig. 2a). Flow cytometry analyses suggested that si-H19 transfection decreased the number of NOZ cells in S-phase, whereas overexpression of H19 increased the number of GBC-SD cells in S-phase (Fig. 2b, c). Transwell invasion assay suggested that knockdown of H19 inhibited the invasion of NOZ cells, and overexpression of H19 promoted the invasion of GBC-SD cells (Fig. 2d, e).
Effect of H19 on GBC cell proliferation and cell cycle and invasion. a After H19 knockdown, cell proliferation was evaluated by CCK8 assay in NOZ and GBC-SD cells, respectively. b Flow cytometric analysis was performed following H19 knockdown. c Statistical analysis of NOZ cells with H19 knockdown and that of GBS-SD cells with H19 overexpression. d, e Cell invasion was evaluated by transwell invasion assay after H19 knockdown or ectopic expression in indicated cells. All data were represented as the mean ± S.D. from three independent experiments. **P < 0.05
AKT2 levels are elevated and positively correlate with H19 in GBC
The RNA level of AKT2 is significantly elevated in GBC tissues compared to adjacent normal tissues (Fig. 3a). H19 and AKT2 levels were positively correlated in GBC patients (R2 = 554, P = 0.012) (Fig. 3b). AKT2 knockdown inhibited NOZ cell proliferation (Fig. 3c) and decreased the number of NOZ cells in S-phase (Fig. 3d, e). Transwell invasion assay suggested that knockdown of AKT2 inhibited the invasion of NOZ cell (Fig. 3f, g).
AKT2 expression level positively correlated with H19. a AKT2 is markedly elevated in gallbladder carcinoma. b The RNA level of AKT2 is positively correlated with H19, R2 = 0.554, P = 0.012. c Cell proliferation was examined by CCK8 assay in NOZ cells after knockdown of AKT2. d, e Cell cycle of NOZ cells after knockdown of AKT2. f, g Cell invasion was evaluated by transwell invasion assay after knockdown of AKT2. All data were represented as the mean ± S.D. from three independent experiments. **P < 0.05
H19 negatively regulated miR194-5p in GBC cells
To investigate the intrinsic mechanism of H19 in GBC, starbase 2.0 (http://starbase.sysu.edu.cn/), a computer algorithm, was used to identify microRNA (miRNA) target genes including lncRNAs. MiR-194-5p was predicted as a competing miRNA, and further prediction found that AKT2 might be the target of mir-194-5p. According to previous reports, miR-194 functioned as a tumor suppressor gene in cancer [16]. In this study, miR-194-5p was downregulated in GBC tissues compared to adjacent normal tissues (Fig. 4a). The interaction between H19 and miR-194-5p was validated by transfecting H19-siRNA into human GBC cell line NOZ. qRT-PCR analysis revealed that the levels of miR-194-5p were increased in the NOZ cells transfected with si-H19 (Fig. 4b), and ectopic expression of H19 decreased the miR-194-5p levels in GBC-SD cells (Fig.4c). However, neither miRNA-194-5p mimic nor miRNA-194-5p inhibitor emerged obvious influence in the level of H19 (Fig. 4d). These results suggested a potential negative regulatory mechanism between H19 and miR194-5p.
H19 negatively regulated miR-194-5p expression in GBC. a The expression of miR-194-5p in GBC tissues was significantly downregulated compared to corresponding normal tissues. b The expression of miRNA-194-5p was downregulated by overexpression of H19 in GBC-SD cells. c The expression of miRNA-194-5p was upregulated by knockdown of H19 in NOZ cells. d NOZ cells were transfected with miRNA-194-5p mimics or inhibitor, and H19 was measured by qRT-PCR respectively. All data were represented as mean ± S.D. from three independent experiments. **P < 0.05. n.s. not significant
H19 modulated expression of endogenous miR-194-5p targeting AKT2
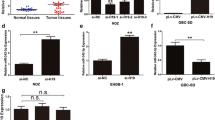
AKT2 is one of the validated targets of miR-194 [16]. Dual luciferase reporter assay was carried out to elucidate whether H19 modulated expression of miR-194-5p by targeting AKT2. The binding sites predicted by starbase 2.0 are shown in Fig. 5a. PmirGLo-H19-wt significantly decreased luciferase activities in HEK293 cells but PmirGLo-H19-mut rescued this effect (Fig. 5b). To further confirm direct binding of H19/miR194-5p/AKT2 in GBC cells, inhibitors and mimics of miR-194-5p were used. qRT-PCR and Western blot analyses revealed that both mRNA and protein levels of AKT2 were decreased significantly by H19 knockdown in NOZ cells. Moreover, co-transfection of miR-194-5p inhibitor and si-H19 attenuated the inhibition of the mRNA and protein levels of AKT2 (Fig. 5c, d). In GBC-SD cells, ectopic expression of H19 upregulated the mRNA and protein levels of AKT2, which was reversed by miRNA-194-5p mimics (Fig. 5e, f). These results indicated that H19 potentially influenced AKT2 by inhibiting the expression of miR-194-5p, suggesting a potential H19/miR-194-5p/AKT2 axis in GBC.
H19 modulated expression of endogenous miR-194 targeting AKT2. a Predicted binding site of miR194-5p and 3′UTR of H19. b Dual luciferase reporter assay of miR194-5p and 3′UTR of H19. c, d The mRNA and protein levels of AKT2 in NOZ cells transfected with si-NC, si-H19, miRNA-194-5p inhibitor, and si-H19 + miRNA-194-5p inhibitor. e, f The mRNA and protein levels of AKT2 in GBC-SD cells transfected with pLv-CMV, pLv-CMV-H19, miRNA-194-5p mimic, and pLv-CMV-H19 + miRNA-194-5p mimic. All data were represented as mean ± S.D. from three independent experiments. **P < 0.05
The role of H19/miR-194-5p/AKT2 axis in cell cycle modulation of GBC
To further explore the biological role of H19/miR-194-5p/AKT2 axis. We analyzed cell cycle alteration after H19 knockdown and miR-194-5p inhibition. GBC cells were arrested in G0/G1 phase after H19 knockdown. Inhibition of miR-194-5p markedly promoted cells into S-phase and co-transfection of siH19 and miR-194-5p inhibitor exerted mutually counter-regulatory effects on cell cycle (Fig.6a, b).
The role of H19/miR-194-5p/AKT2 axis in cell cycle modulation of GBC. a, b Cell cycle of NOZ cells after knockdown of H19, inhibition of miR-194-5p, co-transfection of siH19 and miR-194-5p inhibitors and si-NC transfection group. All data were represented as mean ± S.D. from three independent experiments. **P < 0.05
Discussion
H19 is an oncofetal gene that was discovered more than 30 years ago [17]. The biological role of H19 in cancer has been commanding attention for many years. The present study showed that H19 was involved in GBC cell proliferation.
The present work first proposed that H19 was significantly upregulated in gallbladder carcinoma tissues. In most recently published literatures, H19 was reported to be markedly elevated in gastric cancer [18], breast cancer [19], and ovarian cancer [20], which function as oncogene based on tissue culture system. On the contrary, H19 was reported to be downregulated in Wilm’s tumor [21, 22], prostate cancer [23], and hepatocellular carcinoma [24]. The role of H19 in colorectal cancer was controverted greatly. Evidence from human samples supported its oncogenic roles [25, 26], but in vivo models suggested that lack of H19 might play a role in the initial step of tumorigenesis [27]. According to statistical analyses, we found that H19 was upregulated in GBC tissues and was significantly associated with tumor size, though a larger sample pool is still needed. Combined in vitro cell cycle and growth curve results, we concluded that H19 may act as oncogene in GBC and promote GBC cell proliferation.
Mechanically, we found that H19 functioned as a competing endogenous RNA (ceRNA) for miR-194-5p, antagonized its function, and suppressed downstream effector AKT2. CeRNA hypothesis was proposed by Seitz in 2009 [28], challenging the classic theory that protein-coding gene needed to be translated into protein to exert its function. Pier Paolo Pandolfi put forward that all types of RNAs may compete with each other for miRNAs, generating trans-regulatory in transcriptome-scale [29]. The present work suggested that H19 acted as a sponge for miR-194-5p, supporting the opinion that H19 functioned as a natural sponge for miRNAs, including miRNA-let7 [30], miR-141 [31], miR-138, miR-200a [26], miR-106a, and miR-17-5p family [32]. Others researchers found that H19 could enhance miR-675 expression to increase bladder cancer growth by regulating p53 activation [33], which indicated a different posttranslational regulatory pathway. Thus, due to the space constraints, we focused on miR-194-5p without further exploring other miRNAs. MiR-194-5p is one of the serum exosomal multi-miRNAs that could be used for detection of esophageal adenocarcinoma [34]. However, dual luciferase reporter assay (Fig.5a, b) indicated that miR-194-5p could bind to 3′UTR of H19 and confirmed a direct interaction between H19 and miR-194-5p.
Our results supported that H19 might partly modulate AKT2 by competing with miR-194-5p. AKT2 (v-AKT murine thymoma viral oncogene homologue 2), a pro-survival protein, is activated by the phosphatidylinositol 3′ kinase (PI3K) pathway [35]. In hepatocellular carcinoma cells, inhibition of lncRNAH19 and miR-675 expression can promote cell migration and invasion via AKT/GSK-3β/Cdc25A signaling pathway [36]. Akt/mTOR signaling pathway was associated with cell proliferation of gallbladder carcinoma [37]. Moreover, AKT2 has been found to be associated with breast cancer [38] and hepatocellular carcinoma [39]. However, all above research studies pointed out that AKT2 play a role in cancer biology, but the upstream regulatory pathway of AKT2 in GBC remains largely unknown. Our results verified that the expression levels AKT2 in GBC samples were higher than those in normal samples. It is interesting that H19 expression is positively correlated with AKT2 gene expression. Further studies indicated that H19 modulated AKT2 expression through inhibiting miR-194-5p level, eventually regulating cell cycle in GBC cells.
In our cohort of GBC patients, sample capacity was limited due to low incidence of GBC in our community; most of the GBC patients were admitted at advanced stages that were contraindication for surgery, and samples could not be obtained from these patients. The small sample size in our study was also the main reason for the insignificance between high H19 group and control of several clinical variables in the Table 1.
In conclusion, our data showed that both of H19 and AKT2 promoted cell proliferation in gallbladder carcinoma. H19 was markedly upregulated in GBC tissues and was significantly associated with tumor size. Specially, H19 promoted GBC cell proliferation by inhibiting miR-194-5p targeting of AKT2. Our findings suggested that 19 could serve as a novel target for treatment or prognostic marker of GBC.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA: Cancer J Clin. 2015;65(1):5–29. doi:10.3322/caac.21254.
Kanthan R, Senger JL, Ahmed S, Kanthan SC. Gallbladder Cancer in the 21st Century. J Oncol. 2015;2015:967472. doi:10.1155/2015/967472.
Zhu AX, Hong TS, Hezel AF, Kooby DA. Current management of gallbladder carcinoma. Oncologist. 2010;15(2):168–81. doi:10.1634/theoncologist.2009-0302.
Rakic M, Patrlj L, Kopljar M, Klicek R, Kolovrat M, Loncar B, et al. Gallbladder cancer. Hepatobiliary Surg Nutr. 2014;3(5):221–6. doi:10.3978/j.issn.2304-3881.2014.09.03.
Managadze D, Lobkovsky AE, Wolf YI, Shabalina SA, Rogozin IB, Koonin EV. The vast, conserved mammalian lincRNome. PLoS Comput Biol. 2013;9(2):e1002917. doi:10.1371/journal.pcbi.1002917.
Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014;5(4):e944014. doi:10.4161/21541272.2014.944014.
Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM, et al. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007;132(1):330–42. doi:10.1053/j.gastro.2006.08.026.
Pandey GK, Mitra S, Subhash S, Hertwig F, Kanduri M, Mishra K, et al. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell. 2014;26(5):722–37. doi:10.1016/j.ccell.2014.09.014.
Brunkow ME, Tilghman SM. Ectopic expression of the H19 gene in mice causes prenatal lethality. Genes Dev. 1991;5(6):1092–101.
Lottin S, Adriaenssens E, Dupressoir T, Berteaux N, Montpellier C, Coll J, et al. Overexpression of an ectopic H19 gene enhances the tumorigenic properties of breast cancer cells. Carcinogenesis. 2002;23(11):1885–95.
Zhang Y, Shields T, Crenshaw T, Hao Y, Moulton T, Tycko B. Imprinting of human H19: allele-specific CpG methylation, loss of the active allele in Wilms tumor, and potential for somatic allele switching. Am J Hum Genet. 1993;53(1):113–24.
Polytarchou C, Iliopoulos D, Hatziapostolou M, Kottakis F, Maroulakou I, Struhl K, et al. Akt2 regulates all Akt isoforms and promotes resistance to hypoxia through induction of miR-21 upon oxygen deprivation. Cancer Res. 2011;71(13):4720–31. doi:10.1158/0008-5472.CAN-11-0365.
Fohlin H, Perez-Tenorio G, Fornander T, Skoog L, Nordenskjold B, Carstensen J, et al. Akt2 expression is associated with good long-term prognosis in oestrogen receptor positive breast cancer. Eur J Cancer. 2013;49(6):1196–204. doi:10.1016/j.ejca.2012.12.006.
Nitsche C, Edderkaoui M, Moore RM, Eibl G, Kasahara N, Treger J, et al. The phosphatase PHLPP1 regulates Akt2, promotes pancreatic cancer cell death, and inhibits tumor formation. Gastroenterology. 2012;142(2):377–87 e1-5. doi:10.1053/j.gastro.2011.10.026.
Galicia VA, He L, Dang H, Kanel G, Vendryes C, French BA, et al. Expansion of hepatic tumor progenitor cells in Pten-null mice requires liver injury and is reversed by loss of AKT2. Gastroenterology. 2010;139(6):2170–82. doi:10.1053/j.gastro.2010.09.002.
Zhao HJ, Ren LL, Wang ZH, Sun TT, Yu YN, Wang YC, et al. MiR-194 deregulation contributes to colorectal carcinogenesis via targeting AKT2 pathway. Theranostics. 2014;4(12):1193–208. doi:10.7150/thno.8712.
Pachnis V, Belayew A, Tilghman SM. Locus unlinked to alpha-fetoprotein under the control of the murine raf and Rif genes. Proc Natl Acad Sci U S A. 1984;81(17):5523–7.
Zhou X, Yin C, Dang Y, Ye F, Zhang G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci Report. 2015;5:11516. doi:10.1038/srep11516.
Vennin C, Spruyt N, Dahmani F, Julien S, Bertucci F, Finetti P, et al. H19 non coding RNA-derived miR-675 enhances tumorigenesis and metastasis of breast cancer cells by downregulating c-Cbl and Cbl-b. Oncotarget. 2015;6(30):29209–23. doi:10.18632/oncotarget.4976.
Medrzycki M, Zhang Y, Zhang W, Cao K, Pan C, Lailler N, et al. Histone h1.3 suppresses h19 noncoding RNA expression and cell growth of ovarian cancer cells. Cancer Res. 2014;74(22):6463–73. doi:10.1158/0008-5472.CAN-13-2922.
Steenman MJ, Rainier S, Dobry CJ, Grundy P, Horon IL, Feinberg AP. Loss of imprinting of IGF2 is linked to reduced expression and abnormal methylation of H19 in Wilms’ tumour. Nat Genet. 1994;7(3):433–9. doi:10.1038/ng0794-433.
Honda S, Arai Y, Haruta M, Sasaki F, Ohira M, Yamaoka H, et al. Loss of imprinting of IGF2 correlates with hypermethylation of the H19 differentially methylated region in hepatoblastoma. Br J Cancer. 2008;99(11):1891–9. doi:10.1038/sj.bjc.6604754.
Zhu M, Chen Q, Liu X, Sun Q, Zhao X, Deng R, et al. lncRNA H19/miR-675 axis represses prostate cancer metastasis by targeting TGFBI. FEBS J. 2014;281(16):3766–75. doi:10.1111/febs.12902.
Zhang L, Yang F, Yuan JH, Yuan SX, Zhou WP, Huo XS, et al. Epigenetic activation of the MiR-200 family contributes to H19-mediated metastasis suppression in hepatocellular carcinoma. Carcinogenesis. 2013;34(3):577–86. doi:10.1093/carcin/bgs381.
Tsang WP, Ng EK, Ng SS, Jin H, Yu J, Sung JJ, et al. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis. 2010;31(3):350–8. doi:10.1093/carcin/bgp181.
Liang WC, Fu WM, Wong CW, Wang Y, Wang WM, Hu GX, et al. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget. 2015;6(26):22513–25. doi:10.18632/oncotarget.4154.
Yoshimizu T, Miroglio A, Ripoche MA, Gabory A, Vernucci M, Riccio A, et al. The H19 locus acts in vivo as a tumor suppressor. Proc Natl Acad Sci U S A. 2008;105(34):12417–22. doi:10.1073/pnas.0801540105.
Seitz H. Redefining microRNA targets. Curr Biol : CB. 2009;19(10):870–3. doi:10.1016/j.cub.2009.03.059.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146(3):353–8. doi:10.1016/j.cell.2011.07.014.
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013;52(1):101–12. doi:10.1016/j.molcel.2013.08.027.
Zhou X, Ye F, Yin C, Zhuang Y, Yue G, Zhang G. The Interaction Between MiR-141 and lncRNA-H19 in Regulating Cell Proliferation and Migration in Gastric Cancer. Cell Physiol Biochem : Int J Exp Cell Physiol Biochem Pharmacol. 2015;36(4):1440–52. doi:10.1159/000430309.
Imig J, Brunschweiger A, Brummer A, Guennewig B, Mittal N, Kishore S, et al. miR-CLIP capture of a miRNA targetome uncovers a lincRNA H19-miR-106a interaction. Nat Chem Biol. 2015;11(2):107–14. doi:10.1038/nchembio.1713.
Liu C, Chen Z, Fang J, Xu A, Zhang W, Wang Z. H19-derived miR-675 contributes to bladder cancer cell proliferation by regulating p53 activation. Tumour Biol : J Int Soc Oncodev Biol Med. 2015. doi:10.1007/s13277-015-3779-2.
Chiam K, Wang T, Watson DI, Mayne GC, Irvine TS, Bright T, et al. Circulating Serum Exosomal miRNAs As Potential Biomarkers for Esophageal Adenocarcinoma. J Gastrointest Surg : Off J Soc Surg Aliment Tract. 2015;19(7):1208–15. doi:10.1007/s11605-015-2829-9.
Franke TF, Hornik CP, Segev L, Shostak GA, Sugimoto C. PI3K/Akt and apoptosis: size matters. Oncogene. 2003;22(56):8983–98. doi:10.1038/sj.onc.1207115.
Lv J, Ma L, Chen XL, Huang XH, Wang Q. Downregulation of LncRNAH19 and MiR-675 promotes migration and invasion of human hepatocellular carcinoma cells through AKT/GSK-3beta/Cdc25A signaling pathway. J Huazhong Univ Sci Technol Med Sci = Hua zhong ke ji da xue xue bao Yi xue Ying De wen ban = Huazhong keji daxue xuebao Yixue Yingdewen ban. 2014;34(3):363–9. doi:10.1007/s11596-014-1284-2.
Zhang P, Guo Z, Wu Y, Hu R, Du J, He X, et al. Histone deacetylase inhibitors inhibit the proliferation of gallbladder carcinoma cells by suppressing AKT/mTOR signaling. PLoS One. 2015;10(8):e0136193. doi:10.1371/journal.pone.0136193.
Chin YR, Yuan X, Balk SP, Toker A. PTEN-deficient tumors depend on AKT2 for maintenance and survival. Cancer Discovery. 2014;4(8):942–55. doi:10.1158/2159-8290.CD-13-0873.
Xu X, Sakon M, Nagano H, Hiraoka N, Yamamoto H, Hayashi N, et al. Akt2 expression correlates with prognosis of human hepatocellular carcinoma. Oncol Rep. 2004;11(1):25–32.
Acknowledgments
We thank the Eastern Hepatobiliary Surgical Hospital and Institute, The Second Military University, Shanghai, for their help. This work was supported by the National Natural Science Foundation of China (Grant numbers 81272747and 81572297) and Doctorial innovation fund of Shanghai Jiao Tong University School of Medicine. We thank Dr. Qiu Lei, who helped us with language editing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Shou-Hua Wang and Xiao-Cai Wu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, SH., Wu, XC., Zhang, MD. et al. Long noncoding RNA H19 contributes to gallbladder cancer cell proliferation by modulated miR-194-5p targeting AKT2. Tumor Biol. 37, 9721–9730 (2016). https://doi.org/10.1007/s13277-016-4852-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-4852-1