Abstract
MicroRNAs (miRNAs) play a key role in tumor metastasis based on their capacity to regulate the expression of tumor-related genes. Over-expression of key genes such as c-MYC and CTNNB1 (encoding β-catenin) in Wnt/β-catenin-dependent and ROCK1 in Wnt/β-catenin-independent signaling pathways (Rho/Rho-associated kinase (ROCK) signaling pathway) has already been identified as the hallmarks of many tumors, and their role in breast cancer has also been investigated and confirmed. miR-340 characterization as an onco-suppressor miRNA has been previously reported. However, the mechanism by which it inhibits metastasis has not been completely elucidated. Quantitative real-time PCR (qPCR), Western blot, and luciferase assays were used to confirm the effect of miR-340 on the 3′-untranslated region (UTR) of the target genes. Lentiviral particles containing miR-340 were also used to evaluate the effect of miR-340 restoration on cell proliferation, migration, and invasion in vitro in the invasive MDA-MB-231 cell line. By applying bioinformatic approaches for the prediction of miRNAs targeting 3′-UTRs of CTNNB1, c-MYC, and ROCK1, we found out that miR-340 could dramatically down-regulate metastasis by targeting Wnt signaling in breast cancer cells. In the current study, analyzing miR-340 by reverse transcription quantitative PCR (RT-qPCR) in MDA-MB-231 showed that it was remarkably down-regulated in the metastatic breast cancer cell line. We found that restoration of miR-340 in the invasive breast cancer cell line, MDA-MB-231, suppresses the expression of the target genes’ messenger RNA (mRNA) and protein and, as a result, inhibits tumor cell invasion and metastasis. Our findings highlight the ability of bioinformatic approaches to find miRNAs targeting specific genes. By bioinformatic analysis, we confirmed the important role of miR-340 as a pivotal regulator of breast cancer metastasis in targeting previously validated (ROCK1) and potentially novel genes, i.e., (CTNNB1 and c-MYC).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is considered as the most common malignant disorder in women and accounts for more than 458,000 deaths annually worldwide [1]. Metastasis is the most troublesome complication that may occur during cancers and increases tumor-related death in these patients [2]. Although the mechanism of metastasis has not yet been fully elucidated, known pathways such as Wnt signaling pathway (β-catenin-dependent and β-catenin-independent) have been shown to contribute to metastasis.
Aberrations in the Wnt signaling pathway have been frequently reported in the pathogenesis of several tumors, including breast cancer [3, 4]. Wnt signaling can affect cell fate by influencing proliferation, migration, and morphology of normal cells [5]. Over-expression of factors involved in this pathway which contribute to metastasis has been found in malignant breast cancer cells [6]. The Wnt pathway that involves β-catenin as a key signaling protein is known as Wnt/β-catenin (canonical) pathway. β-Catenin has been identified as the hallmark of metastatic breast cancer and is able to trans-activate its target oncogenes such as Cyclin D1 and c-MYC [7]. The canonical Wnt signaling pathway is mainly involved in cell proliferation and survival [8], and an aberrant expression of nuclear β-catenin has been observed in the invasive forms of primary tumors [8–10]. Its accumulation in the nucleus or cytoplasm has been reported in approximately 60–63 % of patients with breast carcinomas [11, 12].
Alternatively, there is a β-catenin-independent Wnt pathway (non-canonical), which does not involve β-catenin but Rho-associated kinase (ROCK) protein as the key element. This pathway regulates cell morphology and migration through Rho and ROCK proteins. In cancers, this pathway is dysregulated and results in migration and metastasis of cancerous cells [13]. The Wnt/β-catenin-dependent pathway is related to Wnt/β-catenin-independent pathway through the effect of ROCK by controlling c-MYC activation [14]. Therefore, it is important to consider both pathways in the metastasis mechanism.
The Wnt signaling pathway is tightly controlled by key metastasis-related genes and non-coding RNAs such as microRNAs (miRNAs) [14, 15]. miRNAs are evolutionarily conserved, naturally abundant, small non-coding RNAs that target 3′-untranslated region (UTR) of messenger RNAs (mRNAs) in a sequence-specific manner and degrade mRNAs or inhibit their translation [16]. miR-340 has been studied as a putative tumor suppressor in several cancers including neurofibromatosis type 1, neuroblastoma, ovarian tumor, and gastric cancer [17–19]. The over-expression of miR-340 suppresses cell migration, invasion, and metastasis in these cancers [20]. However, little is known about its exact role in breast cancer metastasis.
In the present study, we investigated the potential regulatory role of miR-340 in Wnt signaling pathway in MDA-MB-231 cell line, which is triple negative and one of the most metastatic models of breast cancer, by means of luciferase assay, reverse transcription quantitative PCR (RT-qPCR), and Western blot. Migration and invasion assays were also used to more precisely investigate the possible inhibitory effect of miR-340 on breast cancer metastasis.
Experimental procedures
Prediction of miRNAs targeting 3′-UTR of Wnt signaling pathway genes
Three of the most important Wnt signaling pathway downstream target genes (CTNNB1, ROCK1, and c-MYC) were selected based on previously published works [21–23] and KEGG pathway enrichment analysis (http://www.genome.jp/kegg/). Also, miRNA prediction databases, including TargetScan [24], miRanda [25], and miRWalk [26], were applied to predict miRNAs targeting 3′-UTR of CTNNB1, c-MYC, and ROCK1 mRNAs [27]. Among all the predicted miRNAs, miR-340, having at least one target site in the 3′-UTR of each target gene, had the highest score.
Plasmid construction
Pre-miR-340 with the flanking sequences (∼200 bp at each end) was PCR-amplified, purified by QIAamp DNA mini kit (Qiagen, Germany), and then cloned into the NotI and MluI sites of the pLEX-JRed-TurboGFP® (pLJTG) vector (Open Biosystems, USA).
The 3′-UTRs of human CTNNB1, c-MYC, and ROCK1 genes were amplified by PCR. The purified PCR fragments containing XhoI and NotI restriction sites were first cloned in T/A cloning vector (Fermentas, Lithuania). The vector was then digested with XhoI and NotI (Fermentas, Lithuania), and finally, the target sequences were cloned into luciferase psi-CHECK™-2 vector (Promega, Southampton, UK). All of the cloned fragments were verified by sequencing. The primer sequences used for plasmid construction have been reported in supplementary Table S1.
Cell culture
HEK 293T, MCF-7, and MDA-MB-231 cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10 % fetal bovine serum (FBS), 1 % penicillin/streptomycin, and 2 % glutamine at 37 °C in a humidified atmosphere and 5 % CO2.
MCF-10A (fastidious normal breast cell line), used as control, was cultured in a like manner as other cell lines, except that it was supplemented with 10 % horse serum (HS) instead of FBS.
mRNA and miRNA extraction, cDNA synthesis, and RT-qPCR
Total RNA was extracted using QIAzol (Qiagen, Germany) according to the manufacturer’s instructions and treated with DNaseI. The miRNA-specific stem-loop primer for miRNA reverse transcription was designed according to our previously described protocol [27]. Reverse transcription was performed using Expand™ Reverse Transcriptase (Sigma-Aldrich, USA). miRNA-specific forward primer, universal reverse primer, and TaqMan probe were employed for qPCR amplification. All qPCR reactions were performed in triplicate. GAPDH and SNORD 47(U47) genes were used as reference genes for mRNA and miRNA normalization, respectively. Fold change in gene expression was determined using the Relative Expression Software Tool (REST®) [28].
Transfection and Dual-Luciferase Reporter Assay
Cells were seeded in 96-well plates (2 × 104/well) and then transfected with pLJTG-miR340 or its control vector using Lipofectamine 2000. After puromycin treatment to select pLJTG-miR-340 -or pLJTG-control-transfected cells, either psi-CHECK™2-3′-UTRs or psi-CHECK™2-control vector was used to transfect the cells. After 48 h, cells were lysed and processed with Dual-Luciferase Reporter Assay system (Promega, Southampton, UK) according to the manufacturer’s instructions. The fold activation of Renilla luciferase was normalized against firefly luciferase. All the experiments were performed in triplicate.
Lentiviral-based miR-340-expressing vector transduction and MTT assay
MDA-MB-231 cells were seeded in six-well plates (1 × 106/ well), incubated overnight, and then separately transduced with pLJTG-miR340 or control vector-containing viral particles. The effect of miR-340 on the viability and proliferation of MDA-MB-231 cells was determined by 3-(4,5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide. Briefly, 5 × l04 stably transduced cells/well were plated in 96-well plates, and cell viability assay was performed after 24, 48, and 72 h according to the manufacturer’s protocol. All the experiments were performed in quadruplicate.
In vitro migration and invasion assays
Transwell insert with a pore size of 8 μm from SPL (Life Bioscience, Korea) was used to determine tumor cell migration and invasion capacity. MDA-MB-231 cells were stably transduced by viruses containing pLJTG-miR-340 or control vector. For cell migration assay, initial equilibrium of transwell was performed by adding 800 μL of DMEM containing 30 % FBS for positive control and 10 % FBS for test samples into the lower compartment of transwells as a chemo-attractant. In the upper chambers, 3 × 105 stably transduced cells were seeded. After 24 h, the media in the lower chambers were collected, and cells on the chambers were trypsinized and neutralized with 300 μL FBS. Lower chamber media were again collected, and finally, all of the transmitted cells were centrifuged and counted.
For cell invasion assay, the transwell inserts were coated with Extracellular Matrigel Matrix (ECM, Sigma-Aldrich, USA) and incubated at 37 °C to become gelatinous. After 4 h, MDA-MB-231 cells, transduced with miR-340 or control vector (1 × 105 cells/well), were trypsinized, resuspended in 0.1 mL of fresh medium containing 1 % FBS, placed into the transwell upper chamber on top of a Matrigel-coated filter, and cultured for 24 h at 37 °C. Subsequently, the Matrigel and non-invasive cells of the upper side of the filter were wiped off with cotton swabs. The invasive cells at the lower side of the filter were trypsinized and counted under a light microscope.
Flow cytometric analysis of cell cycle
miR-340 stably transduced MDA-MB-231 cells (1 × 106) were harvested, rinsed with PBS, and fixed in 70 % ethanol. Then, the cells were stained with 1 mL of propidium iodide (PI) solution containing PI (50 mg/L), RNase A (1 g/L), and 0.1 % Triton X-100. The samples were examined by a fluorescence-activated cell sorting (FACS) flow cytometer (Partec, Germany). The results were analyzed using FlowJo software [29], and proliferation Index was determined.
Western blot analysis of protein expression
MDA-MB-231 cells (3.5 × 106) were seeded into 10-cm2 plates. Seventy-two hours after transduction with miR-340-containing or control vector, protein content was extracted using RIPA buffer containing protease inhibitor. The protein concentration was determined using a BCA Protein Assay kit (Pierce, Thermo Scientific, USA). Lysates from miR-340-over-expressing and control vector-containing cells (70 μg proteins) were resolved by SDS-PAGE and were transferred to nitrocellulose membranes. Following an overnight incubation with the primary antibodies against CTNNB1, c-MYC, and ROCK1 (Abcam, USA), specific reactive bands were detected using an anti-rabbit or anti-mouse IgG conjugated to horseradish peroxidase. The immune-reactive bands were visualized by the ECL western blot detection kit (Amersham Life Science, USA). Equal loading was verified using an anti-β-actin antibody.
Statistical analysis
The data were analyzed using one-way ANOVA test by SPSS 16.0 statistical software (Scarborough, Canada). All of the results were expressed as mean ± standard error. For all of the analyses, a two-sided p value of less than 0.05 was considered statistically significant. All RT-qPCR results were analyzed using REST® 2009 software.
Results
Expression of key Wnt pathway metastatic genes in MCF-7 and MDA-MB-231 cell lines
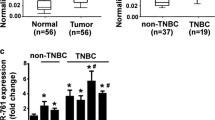
By bioinformatic analysis and sequence alignment, it was shown that miR-340 has at least one target site in the 3′-UTR of the target genes (Fig. 1a). RT-qPCR showed that over-expression of CTNNB1 was more significant in MDA-MB-231 metastatic cell line compared with non-metastatic and control cell lines. While the expression level of CTNNB1 was approximately three times higher in non-metastatic cancer cell line compared to its expression in the control, it was more than six times higher in metastatic cancer cell line than that of control (Fig. 1b). Interestingly, RT-qPCR assay showed that the expression level of miR-340 in the invasive breast cancer cell lines was originally lower than that of the control (Fig. 1c). In addition, compared to the control cell line, down-regulation of miR-340 was more significant in metastatic breast cancer cell line than that of non-metastatic one. This indicates a possible regulatory role of miR-340 in breast cancer metastasis. Remarkably, the expression of CTNNB1, ROCK1, and c-MYC was significantly suppressed in the metastatic cell line upon restoration of miR-340 expression (Fig. 2a).
Bioinformatic predictions, comparison of CTNNB1, ROCK1, and c-MYC expression in breast cancer cell lines and miR-340 expression in cell lines. a Schematic representation of the target 3′-UTRs with miR-340 binding sites and sequence alignment of predicted miR-340 binding sites on 3′-UTRs. b The expression of target genes was examined by RT-qPCR in MCF-7 and MDA-MB-231 cell lines compared to MCF-10A. The expression of each target gene was normalized to GAPDH. c The expression of miR-340 was investigated by RT-qPCR. Each bar represents the relative fold change compared to MCF-10A cell line. Each sample was analyzed in triplicate and was normalized to SNORD 47. In a–c, data are represented as mean ± SEM (n = 3). ***p value <0.0001; **p value <0. 001; *p value <0.05. Not significant, not shown
Expression of target genes after miR-340 induction and luciferase assay. a Gene expression results in MDA-MB-231 cell line after miR-340 transduction. The effect of miR-340 restoration in MDA-MB-231 on CTNNB1, ROCK1, and c-MYC was evaluated by RT-qPCR. b Constructs carrying 3′-UTRs, luciferase reporter, or scramble were transfected in HEK 293T cells expressing miR-340 or control vector. Cells were harvested, and luciferase activities were measured 48 h after second transfection. In a, b, data are represented as mean ± SEM (n = 3). ***p value <0.0001;**p value <0.001; *p value <0.05. Not significant, not shown
miR-340 directly targets 3′-UTR of Wnt pathway metastatic genes
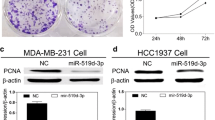
The results of luciferase assay indicated that miR-340 could directly target 3′-UTRs of CTNNB1, ROCK1, and c-MYC (Fig. 2b). Furthermore, upon ectopic miR-340 restoration, Western blot analysis on proteins extracted from invasive breast cancer cell line (MDA-MB-231) showed that miR-340 induction led to a slight reduction in protein levels of the desired target genes (Fig. 3a). To quantify the observed Western blot bands, we used TotalLab version 1.10 software. The quantification and normalization to β-actin showed remarkable decline in the level of target proteins upon miR-340 restoration (Fig. 3b).
miR-340 negatively regulates MDA-MB-231 proliferation and suppresses breast cancer cell motility and invasion
Cell proliferation significantly decreased in miR-340-transduced cells compared with control after 24 h (p value <0.01) (Fig. 4a). FlowJo analysis and proliferation index showed that miR-340 over-expression had no significant effect on cell proliferation index after 72 h, but surprisingly, the percentage of sub-G1 population was slightly higher in cells stably transduced with miR-340 (Fig. 4b).
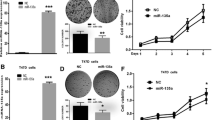
Proliferation assay by MTT and flow cytometry results after miR-340 induction in MDA-MB-231 cells. a MTT assay for analyzing the effects of miR-340 on viability of MDA-MB-231 cells. The results showed a significant difference between control vector and miR-340 after 24 h (**p value <0.001). b Representative of cell cycle distribution of cells transduced by miR-340-containing or control vectors. In comparison with MDA-MB-231 cells, the percentage of cells in sub-G1 step increased after miR-340 restoration (p value <0.05)
Cell migration showed almost 14 % decrease in the miR-340-transduced cells compared with control cells (Fig. 5a). According to our findings, the migratory behavior of the metastatic MDA-MB-231 cells was inversely correlated with the expression levels of miR-340.
The effect of miR-340 induction on cell migration and invasion. a Migration assay results for MDA-MB-231 cell line transfected with miR-340-containing or control vector. The results showed a significant difference in migration of the cells in PC vector and NC vector groups, PC vector and miR-340 groups, and also control vector and miR-340 groups. b Invasion assay results for MDA-MB-231 cell line transduced with lentiviruses containing miR-340 or control vector. The results showed a significant difference in invasion of MDA-MB-231 and miR-340-transduced cells and also control vector and miR-340-transduced groups (**p value = 0.001). PC positive control, NC negative control
As shown in Fig. 5b, a lower number of miR-340-transduced cells invaded the membrane pre-coated with Matrigel, compared with control cells, indicating that miR-340 was able to block the invasion of transduced cells. The induction of miR-340 prompted a significant decrease of 10.12 % in the penetration rate through the Matrigel-coated membrane in miR-340-transduced cells compared with the negative control (p value <0. 001).
Discussion
In the present study, bioinformatic approaches were applied to predict miRNAs that are capable of simultaneously targeting genes in both β-catenin-dependent and β-catenin-independent pathways with major focus on CTNNB1, c-MYC, and ROCK1 genes. Considering the innate multi-targeting potential of miRNAs, bioinformatics is a valuable tool to improve the accuracy of the final outcome with focus on Wnt pathway-blocking miRNAs. In our study, bioinformatic results were obtained based on seed region matches, preferably 8mer, 7mer-m8, and 7mer-1A ones.
Among the three target genes, c-MYC and CTNNB1 had a 7mer-m8 seed match. Furthermore, bioinformatic predictions showed that there are three binding sites for miR-340 in 3′-UTR of ROCK1 with one 8mer and two 7mer-1A seed matches. Although bioinformatic predictions detected three binding sites for miR-340 in ROCK1 3′-UTR, RT-qPCR and luciferase assays confirmed that miR-340 expression has a greater effect on CTNNB1 than ROCK1 (p value = 0.00).
Liu S. et al. showed that ROCK1 suppression by ROCK inhibitors other than siRNA can inhibit anchorage-independent growth, migration, and invasion of MDA-MB-231 cells [22]. These data were in accordance with our finding. However, they did not consider CTNNB1 and c-MYC in their study [14]. We also showed that miR-340 can have a similar effect as a ROCK1 inhibitor on the ROCK1 gene. Fernandez S. et al. found that miR-340 inhibits tumor cell proliferation and induces apoptosis by targeting MET, ROCK1, RHOA, and CDH1 in non-small cell lung cancer [30]. Their result is in line with ours in breast cancer and can confirm our results. In another study, Cimino D. et al. found that miR-148b suppresses breast cancer metastasis by directly targeting players of integrin signaling such as ROCK1 [31]. By careful considerations [32], we hypothesized that the pioneer miRNA prediction algorithms, including TargetScan, have a relatively lower sensitivity than assumed and also predict fewer numbers of total miRNA: mRNA interactions in comparison with experimental studies.
Several cellular pathways, including Wnt signaling pathway, are related to cancer metastasis. Many reports have suggested that dysregulation of Wnt signaling can lead to cancer initiation and progression in a wide range of human tissues including breast [3, 8, 33–37]. There is a growing body of evidence indicating that downstream components of the Wnt signaling pathway are over-activated in many metastatic breast tumors. Despite our poor understanding of the role of Wnt signaling pathway in breast cancer metastasis, results obtained from some studies have showed an up-regulation of this pathway’s genes, including CTNNB1, c-MYC, and ROCK1, in breast cancer samples and cell lines [12, 22, 23, 38].
The role of miRNAs as regulators of tumor initiation, progression, and metastasis has come under intense scrutiny in recent years. Other studies have demonstrated that different tumor suppressor miRNAs such as miR-139-5p, 34a, and 145 [2, 5, 39] and oncogenic miRNAs like miR-374a [25] have the ability to regulate the downstream components of Wnt signaling pathway. In most of the cases, many tumors, including breast cancer, have shown an overall down-regulation in miRNA expression compared to normal tissues. Therapeutic strategies based on induction of miRNA expression are a new field that employs miRNAs for tumor suppression.
We observed higher expression levels of the target genes in the metastatic breast cancer cell line (MDA-MB-231) compared to non-metastatic one (MCF-7), and the expression of target genes had an inverse correlation with the expression of miR-340 in these cell lines. In accordance with our expectations, induction of miR-340 expression significantly decreased the expression of the target genes. Most miRNAs inhibit target genes by interacting with the 3′-UTRs of the mRNAs, and this was confirmed by evaluating the activity of separately cloned 3′-UTR of each individual gene downstream of luciferase gene in the vector. Accordingly, we demonstrated that miR-340 inhibits CTNNB1, c-MYC, and ROCK1 by direct interaction with their 3′-UTRs.
We observed no significant and continuous changes in the cell proliferation index after miR-340 induction, but an increase was seen in the percentage of apoptotic cells in the sub-G1 section.
Migration and invasion are two metastatic traits of cancer cells. Here, we confirmed that miR-340 over-expression regulates motility of cancer cells and decreases cell mobility and invasion in vitro. These results are consistent with down-regulation of cell mobility by miR-340 targeting c-Met, recently reported by Wu Z. et al. [40].
Of great importance, our results indicate that down-regulation of miR-340 is a key event in breast cancer metastasis and refers to an imminent invasive cancer. The restoration of miR-340 expression presents a novel therapeutic strategy for preventing breast cancer progression and metastasis. Nonetheless, there is a need for more comprehensive investigations and trials.
References
Rehmsmeier M, Steffen P Fau - Hochsmann M, Hochsmann M Fau - Giegerich R, Giegerich R. Fast and effective prediction of microRNA/target duplexes. (1355–8382 (Print).
Takahashi RU, Miyazaki H, Ochiya T The roles of microRNAs in breast cancer. (2072–6694 (Electronic)
Klarmann GJ, Decker A, Farrar WL. Epigenetic gene silencing in the Wnt pathway in breast cancer. Epigenetics. 2008;3(2):59–63.
King TD, Suto Mj Fau - Li Y, Li Y The Wnt/beta-catenin signaling pathway: a potential therapeutic target in the treatment of triple negative breast cancer. J cell Biochem (1097–4644 (Electronic)
De Boer J, Wang Hj Fau - Van Blitterswijk C, Van Blitterswijk C Effects of Wnt signaling on proliferation and differentiation of human mesenchymal stem cells. (1076–3279 (Print))
Wang YZ, Han YS, Ma YS, Jiang JJ, Chen ZX, Wang YC, et al. (2012) Differential gene expression of Wnt signaling pathway in benign, premalignant, and malignant human breast epithelial cells. Tumour Biol.
Shtutman M, Zhurinsky J Fau - Simcha I, Simcha I Fau - Albanese C, Albanese C Fau - D'Amico M, D'Amico M Fau - Pestell R, Pestell R Fau - Ben-Ze'ev A, et al. The cyclin D1 gene is a target of the beta-catenin/LEF-1 pathway. (0027–8424 (Print)
Howe LR, Brown AM. Wnt signaling and breast cancer. Cancer Biol Ther. 2004;3(1):36–41.
Brabletz T, Jung A, Reu S, Porzner M, Hlubek F, Kunz-Schughart LA. Variable beta-catenin expression in colorectal cancers indicates tumor progression driven by the tumor environment. Proc Natl Acad Sci U S A. 2001;98(18):10356–61.
Takebe N, Warren RQ, Ivy SP. Breast cancer growth and metastasis: interplay between cancer stem cells, embryonic signaling pathways and epithelial-to-mesenchymal transition. Breast Cancer Res. 2011;13(3):211.
Ryo A, Nakamura M, Wulf G, Liou YC, Lu KP. Pin1 regulates turnover and subcellular localization of beta-catenin by inhibiting its interaction with APC. Nat Cell Biol. 2001;3(9):793–801.
Lin SY, Xia W, Wang JC, Kwong KY, Spohn B, Wen Y, et al. Beta-catenin, a novel prognostic marker for breast cancer: its roles in cyclin D1 expression and cancer progression. Proc Natl Acad Sci U S A. 2000;97(8):4262–6.
Jaffe AB, Hall A. Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol. 2005;21:247–69.
Liu S, Goldstein Rh Fau - Scepansky EM, Scepansky Em Fau - Rosenblatt M, Rosenblatt M Inhibition of rho-associated kinase signaling prevents breast cancer metastasis to human bone. (1538–7445 (Electronic)
Shenouda SK, Alahari SK. MicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastasis Rev. 2009;28(3–4):369–78.
Hurst DR, Edmonds MD, Welch DR. Metastamir: the field of metastasis-regulatory microRNA is spreading. Cancer Res. 2009;69(19):7495–8.
De Cecco L, Berardi M, Sommariva M, Cataldo A, Canevari S, Mezzanzanica D, et al. Increased sensitivity to chemotherapy induced by CpG-ODN treatment is mediated by microRNA modulation. PLoS One. 2013;8(3):e58849.
Presneau N, Eskandarpour M, Shemais T, Henderson S, Halai D, Tirabosco R, et al. MicroRNA profiling of peripheral nerve sheath tumours identifies miR-29c as a tumour suppressor gene involved in tumour progression. Br J Cancer. 2013;108(4):964–72.
Das S, Bryan K, Buckley PG, Piskareva O, Bray IM, Foley N, et al. Modulation of neuroblastoma disease pathogenesis by an extensive network of epigenetically regulated microRNAs. Oncogene. 2013;32(24):2927–36.
Wu ZS, Wu Q, Wang CQ, Wang XN, Huang J, Zhao JJ, et al. (2011) miR-340 inhibition of breast cancer cell migration and invasion through targeting of oncoprotein c-Met. Cancer.
Han J, Hendzel MJ, Allalunis-Turner J. Notch signaling as a therapeutic target for breast cancer treatment? Breast Cancer Res. 2011;13(3):210.
Liu S. The ROCK signaling and breast cancer metastasis. Mol Biol Rep. 2011;38(2):1363–6.
Yamazaki D, Kurisu S, Takenawa T. Involvement of Rac and Rho signaling in cancer cell motility in 3D substrates. Oncogene. 2009;28(13):1570–83.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120(1):15–20.
John B, Enright Aj Fau - Aravin A, Aravin A Fau - Tuschl T, Tuschl T Fau - Sander C, Sander C Fau - Marks DS, Marks DS Human microRNA targets. (1545–7885 (Electronic).
Dweep H, Sticht C, Pandey P, Gretz N. miRWalk—database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform. 2011;44(5):839–47.
Mohammadi-Yeganeh S, Paryan M, Mirab Samiee S, Soleimani M, Arefian E, Azadmanesh K, et al. (2013) Development of a robust, low cost stem-loop real-time quantification PCR technique for miRNA expression analysis. Mol Biol Rep.
Pfaffl MW, Horgan GW, Dempfle L (2002) Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30 (9).
Chen Y, Zou H, Yang LY, Li Y, Wang L, Hao Y, et al. ER81-shRNA inhibits growth of triple-negative human breast cancer cell line MDA-MB-231 in vivo and in vitro. Asian Pac J Cancer Prev. 2012;13(5):2385–92.
Fernandez S, Risolino M, Mandia N, Talotta F, Soini Y, Incoronato M, et al. Verde P miR-340 inhibits tumor cell proliferation and induces apoptosis by targeting multiple negative regulators of p27 in non-small cell lung cancer.
Cimino D, De Pitta C, Orso F, Zampini M, Casara S, Penna E, et al. miR148b is a major coordinator of breast cancer progression in a relapse-associated microRNA signature by targeting ITGA5, ROCK1, PIK3CA, NRAS, and CSF1. FASEB J. 2013;27(3):1223–35.
Kang L, Mao J Fau - Tao Y, Tao Y Fau - Song B, Song B Fau - Ma W, Ma W Fau - Lu Y, Lu Y Fau - Zhao L, et al. MiR-34a suppresses the breast cancer stem cell-like characteristics by downregulating Notch1 pathway.
Wang Y. Wnt/Planar cell polarity signaling: a new paradigm for cancer therapy. Mol Cancer Ther. 2009;8(8):2103–9.
Reya T, Clevers H. Wnt signalling in stem cells and cancer. Nature. 2005;434(7035):843–50.
Smalley MJ, Dale TC. Wnt signalling in mammalian development and cancer. Cancer Metastasis Rev. 1999;18(2):215–30.
Boras-Granic K, Wysolmerski JJ. Wnt signaling in breast organogenesis. Organogenesis. 2008;4(2):116–22.
Matsuda Y, Schlange T, Oakeley EJ, Boulay A, Hynes NE. WNT signaling enhances breast cancer cell motility and blockade of the WNT pathway by sFRP1 suppresses MDA-MB-231 xenograft growth. Breast Cancer Res. 2009;11(3):R32.
Ridley AJ. Rho GTPases and cell migration. J Cell Sci. 2001;114(Pt 15):2713–22.
Lee DY, Jeyapalan Z Fau - Fang L, Fang L Fau - Yang J, Yang J Fau - Zhang Y, Zhang Y Fau - Yee AY, Yee Ay Fau - Li M, et al. Expression of versican 3'-untranslated region modulates endogenous microRNA functions. (1932–6203 (Electronic)).
Wu ZS, Wu Q, Wang CQ, Wang XN, Huang J, Zhao JJ, et al. miR-340 inhibition of breast cancer cell migration and invasion through targeting of oncoprotein c-Met. Cancer. 2011;117(13):2842–52. doi:10.1002/cncr.25860.
Acknowledgments
The authors should thank colleagues in Cellular and Molecular Biology Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran, Orleigh Bogle, and Vahid Kia for kindly revising of the manuscript and also Stem Cell Technology Research Center, Tehran, Iran, for technical help. This work was supported by Pasteur Institute of Iran, Tehran, Iran [grant number BP-8698], and Stem Cell Technology Research Center, Tehran, Iran [grant number 302].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
None
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
The sequence of primers for 3’-UTRs cloning in psi-CHECK2® vector. Overhangs of MluI and NotI enzymes are underlined. (DOC 28 kb)
Rights and permissions
About this article
Cite this article
Mohammadi-Yeganeh, S., Paryan, M., Arefian, E. et al. MicroRNA-340 inhibits the migration, invasion, and metastasis of breast cancer cells by targeting Wnt pathway. Tumor Biol. 37, 8993–9000 (2016). https://doi.org/10.1007/s13277-015-4513-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4513-9