Abstract
While adriamycin (adr) offers improvement in survival for breast cancer (BCa) patients, unfortunately, drug resistance is almost inevitable. Mounting evidence suggests that exosomes act as a vehicle for genetic cargo and constantly shuttle biologically active molecules including microRNAs (miRNAs) between heterogeneous populations of tumor cells, engendering a resistance-promoting niche for cancer progression. Our recent study showed that exosomes from docetaxel-resistance BCa cells could modulate chemosensitivity by delivering miRNAs. Herein, we expand on our previous finding and explore the relevance of exosome-mediated miRNA delivery in resistance transmission of adr-resistant BCa sublines. We now demonstrated the selective packing of miRNAs within the exosomes (A/exo) derived from adr-resistant BCa cells. The highly expressed miRNAs in A/exo were significantly increased in recipient fluorescent sensitive cells (GFP-S) after A/exo incorporation. Gene ontology analysis of predicted targets showed that the top 30 most abundant miRNAs in A/exo were involved in crucial biological processes. Moreover, A/exo not only loaded miRNAs for its production and release but also carried miRNAs associated with Wnt signaling pathway. Furthermore, A/exo co-culture assays indicated that miRNA-containing A/exo was able to increase the overall resistance of GFP-S to adr exposure and regulate gene levels in GFP-S. Our results reinforce our earlier reports that adr-resistant BCa cells could manipulate a more deleterious microenvironment and transmit resistance capacity through altering gene expressions in sensitive cells by transferring specific miRNAs contained within exosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BCa) is the most common malignant tumor in women [1]. Chemotherapy comprises the major therapeutic strategy for clinical treatment; however, chemotherapy fails to eliminate all tumor cells because of drug resistance [2]. Exosomes have attracted much recent interest since the realization that these nanosized vesicles are not merely “cellular debris,” but rather miniature maps of their cells of origin with both physiological and pathological significance [3]. Accumulating evidence indicates that exosomes may play an important role in resistance regulation of BCa. By displaying human epidermal growth factor receptor-2 ligands on their surface, exosomes could competitively bind to monoclonal antibody (trastuzumab) and attenuate latter’s bioavailability [4]. Besides, increased proliferation, migration and invasion capacity, and enhanced angiogenesis, which all present with aggressive phenotype and therapy failure, were observed in recipient BCa cells treated with exosomes from triple-negative BCa cells [5].
One topic of considerable interest is that exosomes contain microRNAs (miRNAs) that mediate intercellular communication. miRNAs are single-stranded non-coding nucleic acids that serve as post-transcriptional regulators of gene expression and participate in almost every aspect of tumor biology including drug resistance [6]. We recently reviewed studies with respect to the role of exosome-mediated miRNA delivery in formatting drug resistance of BCa [7]. In particular, exosomes export selective tumor suppressor miRNAs and retain substantial oncomiRNAs to form a resistance-promoting intracellular circumstance; miRNA-rich exosomes from BCa cells regulate angiogenesis, immunosuppression, and invasion and metastasis after respectively taken by vascular endothelial cells, immunocytes, and macrophages; and stromal cells within the microenvironment communicate with each other to organize a large and ordered network for a tumor-protective niche. By using the ready-established cell lines, our group also discovered that docetaxel-resistant BCa cells may spread chemoresistance to sensitive cells by shedding abundant exosomes and that the effects could be partly attributed to the cell-to-cell transfer of specific miRNAs [8]. Since adriamycin (adr) represents the cornerstone of chemotherapy in BCa, it is a tendency to investigate the mechanism of adr resistance and reinforce the above discussed findings.
Materials and methods
Cell culture
The cell lines used in the present work were wild-type drug-sensitive human BCa MCF-7 (MCF-7/S), purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China), and a drug-resistant MCF-7/Adr variant, kindly provided by Wen-jing Li (Suzhou Municipal Hospital, Suzhou, China). MCF-7/Adr was established by stepwise selection after prolonged (over 14 months) treatment of MCF-7/S to increasing concentrations of adr and is known to express high levels of multidrug resistance associated protein 1 (MRP1) and breast cancer resistance protein (BCRP) with strong resistance to adr [9]. Parental MCF-7/S maintained synchronously in the absence of drug was used as a control. Both of these cell lines have been validated earlier by us as a good model for investigating mechanisms of chemotherapy failure in vitro [10, 11]. In selected experiments, MCF-7/S expressing green fluorescent protein (GFP-S) was generated and characterized as previously described [12]. Fluorescence expression in GFP-S and daughter GFP-S remained stable under prolonged culture.
All cell lines were grown in Dulbecco’s modified Eagle’s medium (DMEM) high glucose (HyClone, USA) supplemented with 10 % fetal bovine serum (FBS), 100 U/ml penicillin, and 100 μg/ml streptomycin at 37 °C in a 5 % CO2 atmosphere. In order to have exosome-free serum, FBS was ultracentrifuged at 100,000g overnight to spin down any preexisting vesicular content (Avanti J-30I, Beckman Coulter, USA). FBS depleted of exosomes was used for all studies.
Exosome isolation and characterization
Exosome isolation was conducted by using protocols that we recently described [8]. Exosomes from MCF-7/Adr and MCF-7/S were designated as A/exo and S/exo, respectively, for simplicity. Briefly, collected culture media were centrifuged at 300g for 10 min, 2000g for 15 min, and 12,000g for 30 min to remove floating cells and debris. Cell-free supernatants were then passed through a 0.22-μm filter. Filtrates were transferred to 8 ultracentrifuge tubes and spun down at 100,000g for 2 h at 4 °C. The supernatants were discharged, and the pellets were resuspended in PBS. All pellets were pooled in one single ultracentrifuge tube and submitted to a second ultracentrifugation in the same conditions. The final pellets containing purified exosomes were either lysed for protein/RNA extraction or diluted in PBS for incubation assays or labeled for cytometry analysis and confocal observation. Validation of the harvested exosomes was performed by transmission electron microscopy (JEM-1010, JEOL, Japan) as previously described [13].
RNA extraction and microarray
RNA was extracted from A/exo and MCF-7/Adr using the Total Exosome RNA and Protein Isolation Kit (Invitrogen, USA) and mirVana RNA Isolation Kit (Ambion, USA) in accordance with the manufacturer’s protocols. RNA yield was quantified spectrophotometrically (Nanodrop 2000, Thermo Scientific, USA) and the integrity was assessed by capillary electrophoresis (Agilent 2100 Bioanalyzer, Agilent Technologies, USA). miRNA profiles were then analyzed using the Affymetrix GeneChip miRNA Array 3.0 (CapitalBio Corporation, Beijing, China) according to the manufacturer’s instruction. Briefly, RNA samples were labeled using the Flash Tag RNA Labeling Kit (Genisphere, USA) and hybridized to miRNA microarray. Data files were processed with the Affymetrix miRNA QC Tool software and following the guided workflow as described in the user manual (http://www.affymetrix.com/support/technical/manuals.affx). The levels of miRNAs between A/exo and MCF-7/Adr were calculated as previously by Jaiswal [14].
Validation of miRNA microarray
Quantitative real-time polymerase chain reaction (qRT-PCR), applying SYBR green technique, was performed to validate the miRNA microarray results in this study [10]. cDNA for miRNA was synthesized using the BU-Script RT Kit (Biouniquer Technology, Nanjing, China) on an iCycler iQ system (Bio-Rad, USA). Specific stem-loop primers were designed for the detection of selected miRNAs using U6 as internal control (all primers were from Springen Biotechnology, Nanjing, China) (Table 1). SYBR green qRT-PCR amplifications (Light Cycler 480, Roche, Australia) were carried out as follows: 91 °C for 5 min followed by 45 cycles (91 °C for 15 s, 60 °C for 30 s), followed by melting curve detection. Relative miRNA expression was calculated by ΔΔCt method. PCR products were analyzed by agarose gel electrophoresis.
Visualization of exosome uptake
Exosomes were labeled with the PKH26 Red Fluorescent Cell Linker Kit (Sigma-Aldrich, USA) according to the manufacturer’s protocol with minor modifications. Exosomes resuspended in PBS were added to 1 ml Diluent C. In parallel, 4 μl PKH26 was added to 1 ml Diluent C and mixed with the exosome solution for 5 min. To bind excess dye, 5 ml 1 % bovine serum albumin was added. The stained exosomes were washed with PBS by ultracentrifugation at 100,000g for 2 h, diluted in complete culture medium, and used for subsequent experiments. For observation of interaction, GFP-S and dye-treated exosomes were co-cultured for 24 h. Fluorescence images were then acquired utilizing confocal laser scanning microscopy (LSM710, Carl Zeiss, Germany) based on the wavelength for GFP and PKH26 as we previously described [8].
Assessment of miRNA transfer
miRNA transfer experiments were conducted as previously reported [8]. In brief, GFP-S cells were seeded in 6-well plates, 24 h before the stimulation. A/exo was divided evenly between all groups and incubated with the pre-plated GFP-S for different times. Then, GFP-S cells were collected at time 0 and after 120 min, 12 and 24 h incubation with A/exo or with PBS (vehicle) alone as control. To eliminate any residual exosomes, GFP-S was washed with PBS twice, enzymatically dissociated, and washed two more times with PBS. After that, cellular RNA was extracted, and qRT-PCR was performed for a subset of miRNAs (Table. 1) as described above. As an indirect measure of miRNA transfer, difference in Ct values was evaluated between the negative control and each experimental time point, a positive value indicated transfer [15].
Analysis of transmitted chemoresistance
Effects of exosomes on cellular chemosensitivity were studied on GFP-S seeded in 6-well plates. When the cells reached ∼50 % confluence, the media were aspirated, and fresh media containing A/exo, S/exo, and vehicle were added for 48 h. Total GFP-S number was subsequently counted under a fluorescence microscope (ImagingA1, Carl Zeiss, Germany) in three non-consecutive microscopic fields (magnification ×200), after which the cells were treated with 250 nM adr. Twenty-four hours later, floating cells were discarded, and residual GFP-S cells were microscopically recorded in the same manner. As an indirect measure of cell viability, survival rate of GFP-S = residual GFP-S ÷ total GFP-S. For apoptosis assay, GFP-S cells pretreated for 48 h with A/exo, S/exo, and vehicle were also prepared. After 24 h exposure to 250 nM adr, all cells including both floating and attached cells were collected, washed in cold PBS, and stained according to the manufacturer’s instruction using the Annexin-V-FITC Apoptosis Detection Kit (BD Biosciences, USA) as we previously described [8]. The unstained cells, cells stained with propidium iodide (no Annexin-V-FITC), or labeled with Annexin-V-FITC (no propidium iodide) were used as controls. Apoptotic rate of GFP-S was analyzed by flow cytometry equipped with CellQuest software. While Annexin-V-FITC positive cells were marked as early apoptotic cells, cells positive for both Annexin-V-FITC and propidium iodide were treated as late apoptotic cells.
Evaluation of mRNA expression
Total RNA was extracted from GFP-S after a 48-h incubation with A/exo, S/exo, and vehicle in 6-well plates. cDNA synthesis and qRT-PCR were carried out using the BU-Script RT Kit with SYBR green. Expressions of Sprouty2, p27, and phosphatase and tensin homolog (PTEN) mRNA were evaluated. After an initial denaturation step at 94 °C for 2 min, 35 cycles were performed including a denaturation step at 94 °C for 30 s, annealing at 55 °C for 30 s, and extension at 72 °C for 30 s. The final extension step was continued for 5 min. β-actin amplification was used as a quantitative control. All reactions, along with the negative control (nuclease-free water as a template), were run in triplicate. Ct values of each sample were collected automatically, and data were expressed as previously described.
Target gene prediction
Three softwares PicTar (http://pictar.mdc-berlin.de/), TargetScan (http://www.targetscan.org/), and MicroCosm (http://www.ebi.ac.uk/enright-srv/microcosm/htdocs/targets/v5/) were employed to predict miRNA targets [16–18]. Only the genes predicted by three independent tools were taken into account. The Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were annotated using the online DAVID program (http://david.abcc.ncifcrf.gov/) [19–21].
Statistical methods
All experiments were done in triplicate, and the representative data were presented. Differences were determined by Student’s t test or by ANOVA with Newman-Keuls multicomparison test when appropriate. A value of p < 0.05 was considered significant.
Results
Characterization of A/exo
To study the role of tumor-derived exosomes in resistance transmission, we first optimized conditions for their isolation. The protocol was based on differential sedimentation properties and used a series of centrifugation and ultracentrifugation steps. Exosome yield was confirmed by measuring protein content. A/exo yield (mean ± SD of 3 separate samples) was 2.49 ± 0.32 μg/106 cells, whereas S/exo yield was 2.38 ± 0.37 μg/106 cells (Fig. 1a). Electron microscopy showed that A/exo was homogeneous in appearance and ranged from 20 to 100 nm in size (Fig. 1b, c). S/exo displayed the similar morphology and size (not shown).
Isolation and characterization of A/exo. a Exosome protein analysis indicated similar quantities of exosomes to be expelled from MCF-7/Adr and MCF-7/S cells. b Transmission electron microscopy showed nanosized vesicles. c A/exo showed a size ranging from 30 to 100 nm. S/exo displayed the same morphology and size (not shown). For b and c, three A/exo preparations were analyzed with similar results
A/exo contains miRNAs
To determine whether the RNA constituents of A/exo were similar to those of MCF-7/Adr, we examined the exosomal and cellular RNA by capillary electrophoresis. Bioanalysis revealed that A/exo lacks the ribosomal RNA peaks characteristic of cellular RNA (Fig. 2a). To further characterize miRNA profile, microarray analysis was done. Among the 19,724 probe sets in Affymetrix Gene Chip miRNA 3.0 Array, 1733 probes were annotated as human mature miRNAs. The scatter plot of signal intensities of these 1733 miRNAs exhibited a connection between A/exo and MCF-7/Adr. Moreover, of the miRNAs examined, a number of miRNAs were detected exclusively in A/exo with respect to those in donor MCF-7/Adr cells (Fig. 2b). The relative expressions of several miRNAs in A/exo and MCF-7/Adr were validated by RT-PCR, supporting the data obtained from miRNA microarray (Fig. 2c).
Detection and validation of A/exo-contained miRNAs. a RNA from MCF-7/Adr and A/exo were analyzed using Agilent 2100 Bioanalyzer. The RIN value of the samples ranged between 6 and 9. Data is representative of a typical experiment. b The scatter plot of miRNA signal intensity showed a significant correlation between A/exo and MCF-7/Adr. c The validation of selected miRNAs indicated that the microarray was generally agreed well with RT-PCR results. Data are normalized to U6 and expressed as the mean ± SD, n = 3
A/exo delivers miRNAs
The presence of miRNAs in A/exo opens up the intriguing possibility that miRNAs may be delivered by exosomes. To examine the potential for uptake, A/exo was labeled by red dye and co-cultured with GFP-S for 24 h, after which localization of exosomes was imaged by confocal laser scanning microscopy. We observed a binding of PKH26-labeled A/exo on the cell membrane (arrows) and an internalization as endosome-like vesicles in the cytoplasm (arrowheads). Control GFP-S incubated with unstained A/exo did not show red fluorescence (Fig. 3a). Transfer of selected miRNAs was then determined by PCR, and the variation in Ct values in GFP-S stimulated with A/exo was evaluated with respect to GFP-S treated with vehicle. As shown in Fig. 3b, the abundance of several miRNAs (miR-20a, miR-23a, miR-24, miR-149, and miR-222) increased progressively in GFP-S along with the internalization of PKH26-labeled A/exo (Fig. 3c), suggesting transfer. Peak accumulation occurred at nearly the same time for all miRNAs tested in our study, between 2 and 12 h, but although at 24 h the level for miR-149 had not returned to the baseline, it displayed a downward trend. Moreover, the miRNAs that appeared to shuttle most efficiently (miR-20a and miR-23a) were those found rich in A/exo (Fig. 3b). Collectively, these indicated that the increase in specific miRNA content was due to their transport from A/exo to GFP-S.
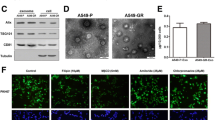
A/exo uptake by GFP-S and miRNA transfer. a Representative confocal microscopic images of GFP-S exposed to PKH26-labeled A/exo for 24 h. (a) PKH26 signal from stained A/exo. (b) GFP signal from recipient cells. (c) Merging of a and b. (d) No PKH26 signal was detected in unlabeled A/exo. (e) GFP signal from control cells with unlabeled A/exo. (f) Overlay of d and e. b GFP-S were incubated with A/exo for 2, 12, and 24 h, and transfer of selected miRNAs was determined by qRT-PCR. Time point 0 represents GFP-S without A/exo. The difference in Ct values between A/exo-treated GFP-S and control cells added with vehicle was shown for each miRNA. Data are expressed as the mean ± SD, n = 3. c Micrograph representative of the absorption of PKH26-labeled A/exo in GFP-S observed by confocal microscopy after incubation for 2 and 12 h
Target prediction of A/exo-loaded miRNAs
To characterize the biological processes modulated by the top 30 highly expressed miRNAs present in A/exo, we analyzed their targets by PicTar, TargetScan, and MicroCosm algorithms. Only the genes listed by three independent tools were taken into account in this paper. We detected 97 genes associated with these miRNAs. Then, we generated a network derived from the union of predicted genes, and we performed a GO enrichment analysis using DAVID, a popular online programme. As shown in the figure, we found a strong overrepresentation of terms belonging to crucial biological processes such as signaling pathway, biosynthesis regulation, differentiation and development, kinase activity, and metabolic process. Moreover, several functions related to exosome biogenesis were detected, in particular membrane organization, endocytosis, membrane invagination, and endosome organization were present (Fig. 4 and Table 2). These reveal that A/exo also carried miRNAs which may be potentially involved in its production and release. Next, we used DAVID to explore the predominant pathways. Based on KEGG analysis, the predicted genes were suggested to participate in Wnt signaling pathway. In BCa, this pathway is upregulated and generally considered to promote drug resistance [22, 23].
A/exo transmits chemoresistance
To answer the question whether A/exo is responsible for spreading chemoresistance, we next compared its effects with S/exo. Confocal microscopic images of the localization of stained exosomes in recipient cells after incubation for 24 h at 37 °C showed that A/exo and S/exo could be absorbed by GFP-S in equal manner (not shown). Transmitted chemoresistance was then assessed in exosome-treated GFP-S in the presence of 250 nM adr. A/exo, but not S/exo, significantly increased survival rate and decreased apoptotic rate of co-cultured GFP-S, with respect to S/exo as well as to vehicle (Fig. 5a, b; *p < 0.05). Besides, incubation of GFP-S with A/exo resulted in significantly more surviving cells after drug exposure, as compared to those incubated with S/exo or vehicle (Fig. 5c, d; *p < 0.05).
Resistance transmission by A/exo. Evaluation of GFP-S incubated with vehicle, S/exo, and A/exo for 48 h. a Survival rate of GFP-S was then determined after 24 h exposure to 250 nM adr. b Apoptotic rate of GFP-S was then evaluated after 24 h exposure to 250 nM adr. c Quantitative evaluation of residual GFP-S cells treated with or without 250 nM adr. d Representative micrographs showing the residual GFP-S cells treated with or without 250 nM adr. For a, b, and c, data are expressed as the mean, n = 3, *p < 0.05, + A/exo vs. + S/exo
To test possible alterations of gene expression in GFP-S as a result of exosome treatment, the following mRNAs were evaluated: Sprouty2 which was reported to be targeted by miR-23a, p27 which was directly regulated by miR-24, and PTEN which was negatively related to miR-222 [10, 24, 25]. They all showed a trend for higher expression in GFP-S receiving S/exo, while the levels were significantly decreased when GFP-S cells were co-cultured with A/exo (Fig. 6; *p < 0.05).
Discussion
Drug resistance accounts for most of the innate insensitivity or disease recurrence following an initial “positive” response in adjuvant chemotherapy of BCa. Although several mechanisms have been proved to play a vital role in such phenotype, recent studies of exosomes and the shuttled miRNAs provided us a new point of view to better understand treatment failure [7, 26]. Our previous work showed that docetaxel-resistant BCa cells were able to spread resistance capacity to sensitive cells via exosomes and that the modulatory effects were partly attributed to the intercellular shuttle of specific miRNAs [8]. Since adr represents the cornerstone of chemotherapy in BCa, it is a tendency to investigate the mechanism of adr resistance and to improve the clinical efficiency of adr-based regimens.
For exosome detection and characterization, a combination of methods is often recommended, as there are as yet no known specific markers according to the current version of the popular exosome database, Exocarta (http://www.exocarta.org) [27]. As reported, transmission electron microscopy and western blotting confirmed successful isolation of vesicles of the expected size, shape, and markers indicative of exosomes. We also demonstrated the presence of various RNAs within A/exo and in particular enrichment for miRNAs. Evidence for circulating miRNAs is increasing, and recent studies have detected miRNAs in patients’ different body fluids in a highly stable and cell-free form [28]. As a matter of fact, exosomes defend miRNAs from hostile ribonuclease-rich environment and serve as vehicles for these extracellular miRNAs.
In the present work, a thorough miRNA profile of A/exo and MCF-7/Adr was analyzed by microarray and further validated by RT-PCR. Heat map demonstrating the expression profile for MCF-7/Adr and the corresponding exosomes is unavailable here (data will be released in our next paper), but the scatterplot of miRNA signal intensity of A/exo and MCF-7/Adr displayed similar pattern with that of D/exo and MCF-7/Doc as we previously described [8]. Specifically, A/exo contained a series of miRNAs shared with their cells of origin; however, selected miRNAs were detectable only in the isolated A/exo but absent in MCF-7/Adr cells from which they derived. Therefore, the preliminary screening of miRNA content of A/exo and MCF-7/Adr reinforced the theory of a non-random and organized package of miRNAs in exosomes before their secretion.
Generally, toxic insults kill only a few malignant cells, leaving insensitive subpopulations alive, which would gradually spread resistance traits to residual cancer cells. Within BCa microenvironment, it is possible for adr-resistant cells to communicate with sensitive cells by transferring miRNAs via exosomes. For this study, we utilized a co-culture system to mimic the circumstance. Confocal microscopy analysis of the labeled A/exo indicated their successful interaction and internalization by recipient cells. While we cannot explain precisely what may be contributing to exosome absorption, it is noteworthy that the ready-made molecules on A/exo surfaces, specific structures on cellular membranes, or the involvement of endocytosis and phagocytosis would be responsible for uptake process. In the present work, we also confirmed that miRNAs highly expressed within A/exo were progressively accumulated in GFP-S concomitantly with the uptake of A/exo. This finding, along with our previous observation, is consistent with that of Yang et al. who demonstrated that invasion-potentiating miRNAs delivered by macrophages could be transferred to BCa cells [29]. Our results also suggest that only certain miRNAs are efficiently shuttled to target cells and that the efficiency of miRNA delivery may be different. Interestingly, recent study showed that specific miRNAs were sorted in exclusive exosomes. While miR-451 was loaded in exosomes with the normal shape and size, some other miRNAs, like miR-16 and miR-21, were released in CD44-positive, large exosome-like vesicles referred to as L-exosomes [30]. We have not tested and compared such exosomes at this time; the next fundamental step will therefore be the full characterization of A/exo, both for the ability and procedure of miRNA transfer and for the selective package of miRNA as discussed above.
The examination of predictive targets regulated by the top 30 highly expressed miRNAs in A/exo demonstrated that many of these miRNAs were involved in crucial biological processes such as signaling pathway, biosynthesis regulation, kinase activity, and metabolic process. Besides, of the interesting functions identified, four of them namely membrane organization, endocytosis, membrane invagination, and endosome organization were all related to exosome biogenesis. Moreover, the pathway analysis of the predicted targets of the top 30 most abundant miRNAs in our study showed the maximum percentage of target genes to be significantly associated with Wnt signaling pathway. The Wnt signaling pathway is an important cascade for cell migration, invasion, and survival that is frequently upregulated in BCa and generally considered to induce drug resistance [22, 23]. These results collectively suggest that A/exo could not only load miRNAs for its production and release but also carry miRNAs implicating traits to therapy resistance. We further confirmed that A/exo, but not S/exo, was able to significantly reduce drug sensitivity of GFP-S to adr treatment. In a separated experiment, A/exo conferred less changes of apoptotic and survival rate of GFP-S than D/exo (unpublished data). We speculate that A/exo might be lower resistance-promoting with respect to D/exo because the MCF-7 resistant sublines selected at 500 nM adr was 61-fold resistant to adr and 3-fold to docetaxel whereas the MCF-7 resistant sublines selected at 200 nM docetaxel was 37-fold resistant to docetaxel and 32-fold to adr as we previously reported [9].
In the present study, we found that miRNAs delivered by A/exo were functional because able to downregulate mRNAs such as Sprouty2, p27, and PTEN, known to be respectively targeted by miR-23a, miR-24, and miR-222 [10, 24, 25]. Sprouty2 was recognized to be deregulated in BCa and served as an important regulator of pathways for cancer invasion and metastasis [24]. The cyclin dependent kinase inhibitor p27 plays a key role in cell cycle arrest and participate in several malignant progression of BCa including autophagy and angiogenesis. PTEN, as a tumor suppressor gene, was reported to regulate multidrug resistance of BCa [10]. We did not show that endogenous mRNAs or proteins are truly affected by these shuttled miRNAs and did not check the downstream signaling pathway; however, as proposed by many authors including us, exosome-contained miRNAs from immune cells, cancer cells, and stem cells could be delivered into recipient cells and serve as physiologically functional molecules to exert gene silencing through the same mechanism as endogenous miRNAs [31–33]. Given the emerging nature of this field, we are currently trying to find the association between miR-222, PTEN, and Wnt signaling pathway and verify their effects using gain- and loss-of-function assays [34].
In summary, our study expands on previous findings and provides the first evidence suggesting that, at least for our established cell lines, adr-resistant BCa cells could manipulate a more deleterious microenvironment and transmit resistance capacity through altering gene expressions in sensitive cells by transferring specific miRNAs contained in exosomes.
References
DeSantis C, Ma J, Bryan L, Jemal A. Breast cancer statistics, 2013. CA Cancer J Clin. 2014;64:52–62.
Gottesman MM. Mechanisms of cancer drug resistance. Annu Rev Med. 2002;53:615–27.
Azmi AS, Bao B, Sarkar FH. Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer Metastasis Rev. 2013;32:623–42.
Ciravolo V, Huber V, Ghedini GC, Venturelli E, Bianchi F, Campiglio M, et al. Potential role of her2-overexpressing exosomes in countering trastuzumab-based therapy. J Cell Physiol. 2012;227:658–67.
O’Brien K, Rani S, Corcoran C, Wallace R, Hughes L, Friel AM, et al. Exosomes from triple-negative breast cancer cells can transfer phenotypic traits representing their cells of origin to secondary cells. Eur J Cancer (Oxford, England 1990). 2013;49:1845–59.
Bartel DP. Micrornas: target recognition and regulatory functions. Cell. 2009;136:215–33.
Chen WX, Zhong SL, Ji MH, Pan M, Hu Q, Lv MM, et al. Micrornas delivered by extracellular vesicles: an emerging resistance mechanism for breast cancer. Tumour Biol J Int Soc Oncod Biol Med. 2014;35:2883–92.
Chen WX, Cai YQ, Lv MM, Chen L, Zhong SL, Ma TF, et al. Exosomes from docetaxel-resistant breast cancer cells alter chemosensitivity by delivering micrornas. Tumour Biol J Int Soc Oncod Biol Med. 2014;35:9649–59.
Li WJ, Zhong SL, Wu YJ, Xu WD, Xu JJ, Tang JH, et al. Systematic expression analysis of genes related to multidrug-resistance in isogenic docetaxel- and adriamycin-resistant breast cancer cell lines. Mol Biol Rep. 2013;40:6143–50.
Zhong S, Li W, Chen Z, Xu J, Zhao J. Mir-222 and mir-29a contribute to the drug-resistance of breast cancer cells. Gene. 2013;531:8–14.
Hu Q, Chen WX, Zhong SL, Zhang JY, Ma TF, Ji H, et al. Microrna-452 contributes to the docetaxel resistance of breast cancer cells. Tumour Biol J Int Soc Oncod Biol Med. 2014;35:6327–34.
Miot S, Gianni-Barrera R, Pelttari K, Acharya C, Mainil-Varlet P, Juelke H, et al. In vitro and in vivo validation of human and goat chondrocyte labeling by green fluorescent protein lentivirus transduction. Tissue Eng Part C Methods. 2010;16:11–21.
Corcoran C, Rani S, O’Brien K, O’Neill A, Prencipe M, Sheikh R, et al. Docetaxel-resistance in prostate cancer: evaluating associated phenotypic changes and potential for resistance transfer via exosomes. PLoS One. 2012;7, e50999.
Jaiswal R, Luk F, Gong J, Mathys JM, Grau GE, Bebawy M. Microparticle conferred microrna profiles—implications in the transfer and dominance of cancer traits. Mol Cancer. 2012;11:37.
Yuan A, Farber EL, Rapoport AL, Tejada D, Deniskin R, Akhmedov NB, et al. Transfer of micrornas by embryonic stem cell microvesicles. PLoS One. 2009;4, e4722.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, et al. Combinatorial microrna target predictions. Nat Genet. 2005;37:495–500.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microrna targets. Cell. 2005;120:15–20.
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ. Mirbase: tools for microrna genomics. Nucleic Acids Res. 2008;36:D154–8.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat Genet. 2000;25:25–9.
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, et al. Kegg for linking genomes to life and the environment. Nucleic Acids Res. 2008;36:D480–4.
Huang DW, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, et al. David bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007;35:W169–75.
Blagodatski A, Poteryaev D, Katanaev VL. Targeting the wnt pathways for therapies. Mol Cell Ther. 2014;2:28.
Polakis P. Wnt signaling in cancer. Cold Spring Harbor perspectives in biology. 2012, 4
Li X, Liu X, Xu W, Zhou P, Gao P, Jiang S, et al. C-myc-regulated mir-23a/24-2/27a cluster promotes mammary carcinoma cell invasion and hepatic metastasis by targeting sprouty2. Hematol J Biol Chem. 2013;288:18121–33.
Giglio S, Cirombella R, Amodeo R, Portaro L, Lavra L, Vecchione A. Microrna mir-24 promotes cell proliferation by targeting the cdks inhibitors p27kip1 and p16ink4a. J Cell Physiol. 2013;228:2015–23.
Xiao X, Yu S, Li S, Wu J, Ma R, Cao H, et al. Exosomes: decreased sensitivity of lung cancer a549 cells to cisplatin. PLoS One. 2014;9, e89534.
Mathivanan S, Fahner CJ, Reid GE, Simpson RJ. Exocarta 2012: database of exosomal proteins, rna and lipids. Nucleic Acids Res. 2012;40:D1241–4.
Kosaka N, Iguchi H, Ochiya T. Circulating microrna in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010;101:2087–92.
Yang M, Chen J, Su F, Yu B, Su F, Lin L, et al. Microvesicles secreted by macrophages shuttle invasion-potentiating micrornas into breast cancer cells. Mol Cancer. 2011;10:117.
Palma J, Yaddanapudi SC, Pigati L, Havens MA, Jeong S, Weiner GA, et al. Micrornas are exported from malignant cells in customized particles. Nucleic Acids Res. 2012;40:9125–38.
Kosaka N, Iguchi H, Hagiwara K, Yoshioka Y, Takeshita F, Ochiya T. Neutral sphingomyelinase 2 (nsmase2)-dependent exosomal transfer of angiogenic micrornas regulate cancer cell metastasis. J Biol Chem. 2013;288:10849–59.
Morel L, Regan M, Higashimori H, Ng SK, Esau C, Vidensky S, et al. Neuronal exosomal mirna-dependent translational regulation of astroglial glutamate transporter glt1. J Biol Chem. 2013;288:7105–16.
Mittelbrunn M, Gutierrez-Vazquez C, Villarroya-Beltri C, Gonzalez S, Sanchez-Cabo F, Gonzalez MA, et al. Unidirectional transfer of microrna-loaded exosomes from t cells to antigen-presenting cells. Nat Commun. 2011;2:282.
Ma F, Zhang J, Zhong L, Wang L, Liu Y, Wang Y, et al. Upregulated microrna-301a in breast cancer promotes tumor metastasis by targeting pten and activating wnt/beta-catenin signaling. Gene. 2014;535:191–7.
Acknowledgments
We would like to acknowledge the funding body for supporting this work: the National Natural Science Foundation of China provided to Jin-hai Tang (81272470).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
None
Additional information
Ling Mao, Jian Li and Wei-xian Chen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Mao, L., Li, J., Chen, Wx. et al. Exosomes decrease sensitivity of breast cancer cells to adriamycin by delivering microRNAs. Tumor Biol. 37, 5247–5256 (2016). https://doi.org/10.1007/s13277-015-4402-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4402-2