Abstract
The three amino acid loop extension (TALE) class myeloid ecotropic viral integration site 1 (MEIS1) homeobox gene is known to play a crucial role in normal and tumor development. In contrast with its well-described cancer stemness properties in hematopoietic cancers, little is known about its role in solid tumors like esophageal squamous cell carcinoma (ESCC). Here, we analyzed MEIS1 expression and its clinical relevance in ESCC patients and also investigated its correlation with the SOX2 self-renewal master transcription factor in the ESCC samples and in the KYSE-30 ESCC cell line. MEIS1 mRNA and protein expression were significantly decreased in ESCC disease (P < 0.05). The inverse correlation between MEIS1 mRNA expression and tumor cell metastasis to the lymph nodes (P = 0.004) was significant. Also, MEIS1 protein levels inversely correlated to lymph node involvement (P = 0.048) and high tumor stage (stages III/IV, P = 0.030). The low levels of DNA methylation in the MEIS1 promoter showed that this suppression does not depend on methylation. We showed that downregulation of EZH2 restored MEIS1 expression significantly. Also, we investigated that MEIS1 downregulation is concomitant with increased SOX2 expression. To the best of our knowledge, this is the first report on the MEIS1 gene in ESCC. The inverse correlation of MEIS1 with metastasis, tumor staging, and the role of EZH2 in methylation, together with its correlation with stemness factor SOX2 expression, led us to predict cancer stemness properties for MEIS1 in ESCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Esophageal carcinoma (EC) is a considerable medical and public health challenge in different regions worldwide, especially in Asia. Globally, it is ranked as the sixth cause of cancer-related deaths [1]. The overall age-adjusted incidence rate (ASR) of EC for men and women in the highly developed areas of the world is 6.5 and 1.2 per 100,000 persons, respectively. In contrast, the related rates in the developing areas are 11.8 and 5.7 [2]. Based on histological features, EC has two major types: squamous cell carcinoma and adenocarcinoma. Esophageal squamous cell carcinoma (ESCC) is the most common type of EC in Asian countries, in a region defined as the “esophageal cancer belt” which extends from north Iran eastward to China [3]. Risk factors in ESCC etiology include lack of dietary fruits and vegetables, tobacco and opium consumption, and the drinking of hot beverages [4]. Diagnosis of ESCC in an early stage is strongly associated with improved outcome. However, most patients are diagnosed in advanced stages, and the 5-year survival rate after surgery is only about 35 % [5]. Different genetic as well as epigenetic processes contribute to the development and progression of tumors [6]. Increasing evidence suggests that tumors are maintained by cancer stem cells (CSCs). CSCs are found in ESCC, and several well-known CSC genes including CD133, NANOG, OCT4, SALL4, and SOX2 have been proposed as ESCC CSC and disease progression biomarkers (for a recent overview, see [7]). Therefore, exploring CSC marker expression in ESCC could pave the road for better therapy.
The myeloid ecotropic viral integration site 1 (MEIS1) transcription factor (TF) gene was originally identified as a common viral integration site involved in myeloid leukemia [8]. A member of the three amino acid loop extension (TALE) family of homeodomain proteins, it is an important developmental TF, both in its own right and as a protein cofactor to other (TALE or HOX) homeobox proteins. Also, another TALE family members, including the other MEIS genes (MEIS2 and 3) and the pre-B cell leukemia homeobox (PBX) genes (PBX1-4), are important in normal development [9]. Conversely, upon deregulated expression, these genes can cause severe developmental disorders and cancer. MEIS, PBX, and HOX genes regulate the expression of their complex target gene network as protein-DNA complexes, in which the MEIS, PBX, and HOX proteins display specific DNA binding properties [10]. MEIS1 has a distinct role in self-renewal and maintenance of stemness state of different stem cell types, including neural and hematopoietic [11–13]. In addition, it has been shown that high MEIS1 expression has a role in the self-renewing of neural stem cells in developing olfactory epithelium [14] and can regulate the transcription of the critical self-renewal gene, OCT4, in neural stem cells [15].
Since normal stem cells share different properties such as self-renewal with CSCs, significant roles can also be construed for developmental TFs in cancer progression and maintenance. Among these TFs, SOX2 is involved in normal development of different organs as well as maintenance of self-renewal capacity of embryonic stem cells (ESCs) [16, 17]. It has been shown that both SOX2 gene amplification and mRNA overexpression are correlated to poor prognosis in several malignancies. High SOX2 expression is associated with lymph node metastasis, depth of tumor invasion, and poor differentiation in ESCC and lung cancer [18–20], also with metastasis in brain, breast, colorectal, and prostate malignancies [21–23]. Furthermore, it is involved in tumor initiation and apoptosis resistance in ovarian cancer [24]. Although SOX2 expression and function have been shown in a variety of cancers, its upstream regulatory mechanisms are almost completely unknown. The clarification about all of these mechanisms will enrich our knowledge about CSC self-renewal.
In this study, we aimed to evaluate the regulatory role of MEIS1 expression in ESCC and elucidate a possible interaction between MEIS1 and SOX2, which may be involved in maintaining the stemness state and self-renewal of ESCC cells.
Materials and methods
Clinical samples
Primary tumor tissue samples, with adjoining nontumoral tissue margins, from 50 ESCC patients were freshly collected during therapeutic surgery at the Omid Oncology Hospital of the Mashhad University of Medical Sciences, Mashhad, Iran. The clinical characteristics of the patients are summarized in Table 1. All specimens were obtained before any chemotherapy and radiotherapy treatments to prevent influence of treatment on clinically relevant features of the tumor samples. All tumor and adjacent nontumoral tissues were histopathologically confirmed, and the clinical features of the tumor samples were defined based on the 7th edition of Union International Cancer TNM classification guidelines [25]. The size of tumor samples ranged from 5 to 12 cm (mean ± SD, 4.12 ± 1.96). The mean age ± SD of patients was 61.5 ± 11.7. The study was approved beforehand by the local ethics committee, and the declared consent of all patients for enrollment in this study is on record.
Cell lines and culture conditions
Human KYSE-30 ESCC cells [26] and human HEK293T embryonal kidney cells were cultured in RPMI-1640 medium (PAA, Pasching, Austria) and DMEM medium (Gibco, Grand Island, NY), respectively. Media were supplemented with 10 % heat-inactivated fetal bovine serum (FBS, Invitrogen, Carlsbad, CA), 10 mM glutamine, 100 U/mL, and 100 mg/mL penicillin-streptomycin (PAA) at 37 °C in a humidified atmosphere containing 5 % CO2. All cell lines were purchased from the Pasteur Institute Cell Bank of Iran (http://ncbi.pasteur.ac.ir/) and used at low passage numbers only. The KYSE-30 cell line was last successfully authenticated by short tandem repeat profiling at the Pasteur Institute Cell Bank of Iran on April 26, 2015, after completion of all experiments in this manuscript.
RNA extraction and qRT-PCR
RNA was extracted from ESCC cell lines and from tumor and adjacent nontumoral tissues of ESCC patients using TRIpure reagent (Roche, Nutley, NJ). Following cDNA synthesis, quantitative real-time PCR (qRT-PCR) to evaluate quantitative changes of MEIS1, EZH2, and SOX2 mRNA expression in ESCC samples using gene-specific primer sets (Table 2) with GAPDH as the reference mRNA was performed as described before [27]. Briefly, PCR was performed on 200 ng reverse-transcribed RNA in a total volume of 20 μL in 1× SYBR Green Real Time PCR Master Mix (Parstous, Mashhad, Iran) containing 0.5 μM of each primer. PCR consisted of an initial denaturation for 10 min at 94 °C, followed by 40 cycles of 15 s 94 °C, 30 s 60 °C, 30 s 72 °C, and was performed in an Mx-3000P real-time thermocycler (Stratagene, La Jolla, CA).
Immunohistochemistry
For MEIS1 protein tissue detection, we used the Novolink Polymer Detection kit (RE7200-CE, Leica Biosystems, Newcastle, UK) according to the manufacturer’s protocols. Briefly, immunohistochemistry was performed on formalin-fixed, paraffin-embedded esophageal tumor and adjacent nontumoral sections after deparaffinization and rehydration by xylene and ethanol dilutions, respectively. The sections were treated with antigen retrieval buffer for 30 min at 100 °C. Prediluted anti-MEIS1 antibody solution (ChIP Grade, ab19867, Abcam, Cambridge, UK) was applied to tissue sections for 30 min at room temperature. After washing with Tris-buffered saline, specific antigen-antibody binding was detected with Novolink polymer solution + DAB (Leica Biosystems, Buffalo Grove, IL). Tissue sections were immersed in hematoxylin-eosin for counterstaining and dehydrated using ethanol. Immunohistochemical staining was analyzed by light microscopy and scored according to Sincrope et al. based on either the percentage of cells with positive nuclear and cytoplasmic staining or on the overall cellular expression intensity, with scores <6 or ≥6 defined as low or high, respectively [28].
Public ESCC dataset analysis
For MEIS1 mRNA expression differences between ESCC and adjacent nontumoral tissue, all three ESCC mRNA profiling datasets available in the public domain were analyzed: Hu-34 (GSE20347) [29], Kimchi-24 (GSE1420) [30], and Su-106 (GSE23400) [31]. The transcript view genomic analysis and visualization tool (http://r2.amc.nl) were used to test whether the probe set selected uniquely mapped to an antisense position in an exon of the gene. The probe sets selected for MEIS1 in the Affymetrix Human Genome U133A arrays (204069_at) met all these criteria and showed the highest expression for MEIS1 in the datasets analyzed. The datasets were obtained from the Oncomine Web site (www.oncomine.org) and analyzed at standard settings. Other cohort details are available through GSE (www.ncbi.nlm.nih.gov/gds/) and PubMed (www.ncbi.nlm.nih.gov/pubmed/) links.
Bisulfite sequencing
Bisulfite conversion of genomic DNA for methylation detection was performed using the CpGenome DNA Modification kit (Chemicon International, Temecula, CA) according to the manufacturer’s protocol with the following exceptions: 2 μg DNA was resuspended in 0.3 M NaOH and heated at 50 °C for 10 min. The DNA was then incubated with reagent-I at 50 °C for 7 h with tube inverting every 30 min. Methylation-specific (MSP) PCR amplification of two CpG islands on the MEIS1 promoter region was performed using MSP primers (Table 2) on bisulfite-converted DNA from ESCC tissues showing low MEIS1 mRNA expression. Briefly, PCR was performed on 40 ng bisulfite-treated DNA in a total volume of 20 μL in 1× Hot Start Taq polymerase buffer (Finzymes, Espoo, Finland) containing 0.2 mM dNTP, 0.5 μM of each primer, and 0.5 U Taq Hot Start Polymerase (Takara, Shiga, Japan). PCR consisted of an initial denaturation for 12 min at 95 °C, followed by 35 cycles of 30 s 95 °C, 30 s 56 °C, 30 s 72 °C), and a final incubation for 15 min at 72 °C. The purified products were cloned into the pTZ57R/T vector using the T/A cloning kit (Fermentas, Vilnius, Lithuania) and sequenced using the M13 primer set (Macrogen, Seoul, Korea). The results were analyzed using the BiQAnalyser online tools (http://biq-analyzer.bioinf.mpi-inf.mpg.de/).
MEIS1 and EZH2 gene expression knockdown
The pLKO.1 lentiviral shRNA expression vector [32] encoding a validated shRNA sequence targeting human MEIS1 (TRCN0000015969) was obtained from Sigma-Aldrich (St. Louis, MO). The shc003 plasmid encoding GFP in the pLKO.1 backbone (Sigma) was used as a control. Lentiviral second-generation packaging plasmids psPAX2 and pMD2.G were purchased from Addgene (plasmids 12260 and 12259, respectively, Cambridge, MA). To produce lentiviral particles, the pLKO.1-MEIS1 plasmid was cotransfected into HEK293T cells along with the packaging plasmids according to the calcium phosphate-based Trono lab protocol [33]. Retrovirus vectors encoding EZH2-specific shRNA (RNAi-Ready pSIREN-RetroQ Vector, kindly provided by Yutaka Kondo, Nagoya, Japan) were used to target EZH2 expression. The shRNA vector was cotransfected with VSV-G and GP vectors into HEK293T cells as described above. After 48 h of transfection, the supernatant containing viral particles was harvested by ultracentrifugation (40-mL culture medium per 50-mL Beckman tube, ultracentrifugation for 120 min at 70,000×g, at 4 °C) and used to transduct KYSE-30 cells; the infected cells were selected by puromycin (Invitrogen Corporation, Carlsbad, CA) 48 h after infection. Quantification of MEIS1 and EZH2 mRNA knockdown was performed by qRT-PCR as described above.
Western blotting
Western blotting was performed as in reference [34] using BioRad equipment (Munich, Germany). The same quantity of protein from each sample was separated from 10 % SDS-PAGE gel, and then, protein was transferred to nitrocellulose membrane (N7892, Sigma-Aldrich). β-Actin was used as a loading control. The primary antibodies used were as follows: MEIS1 (ab19867, Abcam); SOX2 (NB110-37235, Novus Biologicals, Littleton, CO); β-actin (ab25894, Abcam) diluted at 1:1000, 1:400, and 1:2000, respectively. The secondary antibody used for MEIS1 and β-actin was anti-rabbit IgG Peroxidase (A0545, Sigma-Aldrich) diluted at 1:20,000; for SOX2, it was anti-mouse IgG1 (NBP1-51688, Novus Biologicals) diluted at 1:500. All antibodies were diluted in 2.5 % skim milk. Protein was incubated with specific primary antibodies at 4 °C overnight. After incubation with secondary antibody for 1 h at room temperature, the protein was detected by enhanced chemiluminescence (Clarity™ Western ECL Substrate kit #170-5060, BioRad).
Statistical analysis
Statistical analysis was performed using the SPSS 19.9 statistical package (SPSS, Chicago, IL). The correlations between gene expression and various histopathological features were assessed using both the χ2 and Fisher exact tests, and the correlation between MEIS1 and SOX2 expression was assessed using Pearson’s correlation (Table 1). To correlate gene expression levels (mRNA and protein), two-sided t tests were performed (Figs. 1, 4, and 5, and Supplemental Fig. 1). P < 0.05 was considered statistically significant.
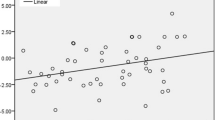
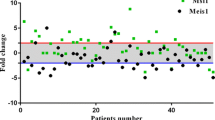
MEIS1 mRNA analysis in 50 ESCC and matched adjacent nontumoral tissue samples and mRNA profiling datasets in the public domain. a MEIS1 mRNA expression is significantly lower in tumor than in adjacent nontumoral tissue. MEIS1 mRNA expression was determined with qRT-PCR, with GAPDH as the reference gene. Shown are the 2log values, median centered. A t test was used to assess the statistical difference between ESCC and adjacent nontumoral tissue. b MEIS1 expression is significantly lower in ESCC than in adjacent nontumoral tissue samples in three different ESCC mRNA profiling datasets in the public domain. a Hu-34: 17 ESCC and matched adjacent nontumoral tissue samples. b Kimchi-24: 8 ESCC and 8 adjacent nontumoral tissue samples. c Su-106: 53 ESCC and matched adjacent nontumoral tissue samples. Shown are the 2log median centered mean values, the error bars represent the SD. A two-sided t test was performed that was used to assess the statistical difference in mRNA expression. See “Materials and methods” for further details on the datasets and analysis
Results
MEIS1 decreased expression at mRNA and protein level in ESCC tumor samples
To determine a possible role for MEIS1 expression in ESCC pathogenesis, we evaluated MEIS1 mRNA expression levels in 50 ESCC samples and compared these with the expression levels in the matched adjacent nontumoral margin tissues of esophageal epithelium, by qRT-PCR. As shown in Fig. 1a, MEIS1 mRNA expression in ESCC tissues was significantly lower than in the matched adjacent nontumoral tissue (P < 0.05). In 38 % of samples (19 of 50), ESCC MEIS1 mRNA expression was more than 2-fold lower than in the adjacent nontumoral esophageal tissue. To verify that our ESCC patient cohort is representative, and the results could be repeated in other ESCC series, we also analyzed ESCC mRNA expression profiling datasets in the public domain. Publicly available ESCC datasets are few in size and number. We found three datasets with MEIS1 expression data in ESCC and adjacent nontumoral samples. Two of these sets, Hu-34 and Kimchi-24, are quite small, and their analysis should be considered with some care. However, in all three sets, including the much larger Su-106 set, the MEIS1 expression was significantly lower in ESCC than in (matched) adjacent nontumoral esophageal tissue (Fig. 1b). We therefore tentatively concluded that our patient cohort was representative, and that the observation of lower MEIS1 ESCC expression is robust.
To extend this observation, we examined MEIS1 protein expression in 27 ESCC tissues and their adjacent nontumoral margin by immunohistochemical staining. Although both tissue types demonstrated immunoreactivity, in line with the mRNA expression pattern, the ESCC tissues showed significantly lower MEIS1 immunoreactivity than their adjacent nontumoral margin (P < 0.05). Only 1 out of 27 (3.7 %) tumor samples showed high MEIS1 protein expression, while 8 of 27 (29.6 %) adjacent nontumoral samples had similar high MEIS1 expression. Figure 2 shows representative images of tumor tissue (panel a) and matched adjacent nontumoral (panel b) with low and high MEIS1 immunostaining, respectively.
Downregulation of MEIS1 expression is correlated to metastasis, lymph node involvement, and tumor staging
To assess potential clinical consequences of MEIS1 downregulation in ESCC tumor samples, we examined the correlation between MEIS1 mRNA/protein expression and different clinically relevant tumor parameters in our ESCC cohort. Low MEIS1 mRNA expression was significantly correlated with different indices of poor prognosis: tumor metastasis (P = 0.027) and lymph node involvement (P = 0.004). Interestingly, 84.2 % (16 out of 19) of samples with low MEIS1 expression showed invasion of tumor cells to adventitia (stage T3). In patients with low stage (stages I/II) ESCC, low MEIS1 mRNA expression was significantly correlated with poor tumor differentiation (P = 0.002). At MEIS1 protein level, we observed a significant correlation between MEIS1 protein expression in tumor samples and the number of involved lymph node (P = 0.019). Furthermore, MEIS1 protein expression was significantly correlated to lymph node involvement (P = 0.048) and high tumor stage (stages III/IV, P = 0.030). There was no other significant association between the level of MEIS1 mRNA or protein expression and clinical data (Table 1).
MEIS1 underexpression is not due to promoter methylation
To assess the role of promoter hypermethylation in the decreased MEIS1 expression in ESCC tumor samples, we amplified and analyzed MEIS1 gene promoter sequences from ESCC samples with low MEIS1 expression (6 samples selected arbitrarily) for probable methylated CpG islands using the UCSC Genome Browser (http://genome.ucsc.edu). A total of 58 CpG sites exist within the 1320-bp region upstream of the MEIS1 transcription start site. Two distinct segments of the promoter containing 23 CpG sites were selected for methylation analysis. Genomic DNA was isolated, subjected to methylation-specific (MSP) PCR, and cloned (Fig. 3). For every tumor sample, 10 separate clones were selected and sequenced. HL-60 cell line DNA was used as a positive control for methylation of these CpG’s, as based on previous work [35]. The results indicated that the promoter was not significantly more often methylated in ESCC samples with low MEIS1 expression than in matched adjacent nontumoral tissue samples.
MEIS1 promoter methylation status in ESCC. Bisulfite sequencing results of clonal MEIS1 promoter MSP-PCR products. Each horizontal line represents a different tumor sample. The CpG di-nucleotides are represented by circles, with open and closed circle for unmethylatedand methylated CpG, respectively. Shown are two representative tumor samples compared with their matched adjacent nontumoral tissue. HL60 cell line DNA was used as a positive control for methylation of these CpG’s as based on previous work [35]
Knocking down epigenetic factor EZH2 to assess its effect on MEIS1 expression
The absence of MEIS1 promoter CpG island methylation in cells with low MEIS1 expression inspired us to modulate EZH2 expression and assess its effect on MEIS1 regulation in KYSE 30 cells. A retroviral EZH2 shRNA construct efficiently silenced EZH2 expression as shown by qRT-PCR. Additional qRT-PCR analysis showed that MEIS1 expression increased after EZH2 silencing in KYSE 30, to more than 2-fold (Fig. 4).
EZH2 silencing mediated by EZH2-specific retrovirus shRNA in KYSE 30 cells. MEIS1 is overexpressed (b) as result of EZH2 underexpression (a) confirmed by qRT-PCR, with GAPDH as the reference gene. The experiment was performed in triplicate. Shown are the mean values, the error bars represent the SD. A two-sided t test was used to analyze the statistical difference in mRNA expression. The difference was significant, P = 0.04
Expression correlation of MEIS1 and SOX2 in ESCC
To analyze MEIS1 role as a stemness factor in ESCC, we performed SOX2 expression analysis in our ESCC cohort (Table 1), we found that SOX2 showed higher expression in ESCC than in matched adjacent nontumoral tissue (Supplemental Fig. 1). This led to a significant negative correlation between MEIS1 and SOX2 mRNA expression (P = 0.011, R = −0.790, Pearson test). To prove an actual, dynamic relationship between MEIS1 and SOX2 expression in ESCC cells, we performed lentiviral MEIS1 knockdown in KYSE-30 cells. The MEIS1 knockdown was confirmed on mRNA and protein levels by qRT-PCR and Western blot analysis (Fig. 5, panels a and b, respectively). Interestingly, we found that MEIS1 knockdown resulted in significant overexpression of SOX2 both at both mRNA (Fig. 5a, with over 3-fold SOX2 overexpression) and levels (Fig. 5b). Together, we take these results as a strong indication that MEIS1 is involved in ESCC cell differentiation, possibly in part through regulation of SOX2.
MEIS1 silencing mediated by MEIS1-specific lentivirus shRNA in KYSE-30 cells. MEIS1 knockdown was confirmed by a qRT-PCR and b Western blot. a Lower MEIS1 expression resulted in SOX2 mRNA overexpression, as established by qRT-PCR, with GAPDH as the reference gene. In a, the results of two separate qRT-PCR experiments are shown, with a, c showing MEIS1 and b, d SOX2 expression for the first and second experiment, respectively. The experiment was performed in triplicate, and repeated three times. Shown are the mean values, the error bars represent the SD. A two-sided t test was used to assess the statistical difference in mRNA expression. The difference was significant, P = 0.037. b MEIS1 knockdown and concomitant SOX2 overexpression was also confirmed on protein level using Western blot analysis. To confirm equal loading of the gel, β-actin was used as a reference
Discussion
In this study, we found that MEIS1 expression is inversely correlated to metastasis, lymph node involvement, and tumor staging in ESCC (Figs. 1 and 2). In addition, we provided evidence that MEIS1 downregulation during ESCC development is caused by EZH2, and not by methylation of CpG islands (Fig. 3). Last, we determined an inverse correlation between MEIS1 and SOX2 in ESCC tumor samples and showed that MEIS1 knockdown led to SOX2 overexpression in an ESCC cell line (Figs. 4 and 5).
MEIS1 is a developmentally conserved member of TALE family and HOX gene clusters. Although many studies have focused on the function of MEIS1 as cofactor of different transcription machineries, its exclusive role as an independent transcription factor remains to be determined. Recent evidence has demonstrated the role of MEIS1 in cancer stem cells (CSCs), self-renewal in myeloid/lymphoid or mixed lineage leukemia (MLL), and potential rate limiting determinant in leukemia stem cell (LSC) [12, 36]. K. Okumura et al. have shown two roles for MEIS1 in epidermis: regulator of stem cells in normal tissues and as proto-oncogenic in skin tumorigenesis [37]. The oncogenic role of stem cell factor MEIS1 has been extensively determined in hematopoietic disorders. In other malignancies, including neuroblastoma [38], ovarian cancer [39], and Wilms tumor [11], high expression of MEIS1 has potential oncogenic properties with direct or indirect effects on the tumor cell growth and resistance to chemotherapy. In spite of these published MEIS1 oncogenic roles, we found downregulation of MEIS1 in ESCC compared to adjacent nontumoral tissue both on mRNA and protein level. In line with our finding in ESCC, Crist et al. showed downregulation of a specific isoform of MEIS1 in colon cancer [40]. Furthermore, it has been shown that low level of MEIS1 expression is correlated to poor prognosis in prostate cancer [41].
Upstream regulators of MEIS1 and its ensuing promoter methylation state were first studied in leukemia. Xiang et al. revealed that ELF1 is an important positive regulator of MEIS1 expression in K562 erythroleukemia cells [35]; moreover, Lasa et al. showed that MEIS1 expression is downregulated through promoter hypermethylation in AML1-ETO acute myeloid leukemias [42]. Our data showed that MEIS1 gene silencing might be caused by DNA methylation-independent mechanism (Fig. 3). Kondo et al. have described that downregulation of genes involved in prostate cancer progression can occur through H3K27me3 by EZH2 [43]. MEIS1 was recently identified as a target of Polycomb genes in bladder cancer [44].
The crosstalk between MEIS1 and core pluripotency circuit genes, including OCT4, SOX2, NANOG, and KLF4/5, has been identified in developmental and neurogenesis studies. Yamada T et al. showed a reciprocal relationship between MEIS1 and OCT4 expression in neural differentiation and observed induction of SOX2 by ectopic expression of MEIS1 [15]. Tucker and others demonstrated that olfactory epithelium precursors have slowly dividing lateral precursors that are regulated by antagonistic expression of SOX2 and MEIS1 [14].
Regarding context-dependent mechanisms for regulation of gene expression (Fig. 6), an inverse correlation between MEIS1 and SOX2 both in vivo in ESCC tumor samples and in vitro in an ESCC cell line would propose a crosstalk between MEIS1 and SOX2, where MEIS1 may suppress SOX2 gene expression, leading to tumor cell differentiation in ESCC. Altogether, our finding, regarding this correlation between MEIS1 and SOX2, established a novel important point for designing a model network between these genes in different ESCC cell lines in the near future.
Schematic drawing for MEIS1 and SOX2 expression in different cancers, in relation to differentiation status. Poorly differentiated cells with low MEIS1 expression (e.g., ESCC MEIS1-knockdown cells or prostate cancer [41] along with SOX2 overexpression may help CSCs to maintain a self-renewal and stemness state). In contrast, in well-differentiated cancer cells such as hematopoietic malignancies [36], high MEIS1 and low SOX2 expression could cause CSC maintenance and cancer progression. Similarly in neuroblastoma, high MEIS1 expression causes cancer
In conclusion, the inverse correlation of MEIS1 with metastasis, tumor staging, and the role of EZH2 in methylation, together with its correlation with stemness factor SOX2 expression, led us to predict cancer stemness properties for MEIS1 in ESCC. These concepts will require deeper analysis.
Abbreviations
- EC:
-
Esophageal carcinoma
- ESCC:
-
Esophageal squamous cell carcinoma
- ESC:
-
Embryonic stem cells
- CSC:
-
Cancer stem cell
- MEIS1:
-
Myeloid ecotropic viral integration site 1
- MLL:
-
Myeloid/lymphoid or mixed lineage leukemia
- MSP-PCR:
-
Methylation-specific PCR
- LSC:
-
Leukemia stem cell
- PBX:
-
Pre-B cell leukemia homeobox
- TALE:
-
Three amino acid loop extension
References
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics. CA Cancer J Clin. 2005;55:74–108.
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
Kamangar F, Malekzadeh R, Dawsey SM, Saidi F. Esophageal cancer in Northeastern Iran: a review. Arch Iran Med. 2007;10:70–82.
Sadjadi A, Marjani H, Semnani S, Nasseri-Moghaddam S. Esophageal cancer in Iran: a review. Middle East J Cancer. 2010;1:11–20.
Chang AC, Ji H, Birkmeyer NJ, Orringer MB, Birkmeyer JD. Outcomes after transhiatal and transthoracic esophagectomy for cancer. Ann Thorac Surg. 2008;85:424–9.
Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. New Engl J Med. 2003;349:2042–54.
Islam F, Gopalan V, Wahab R, Smith RA, Lam AK. Cancer stem cells in oesophageal squamous cell carcinoma: identification, prognostic and treatment perspectives. Crit Rev Oncol Hematol. 2015. doi:10.1016/j.critrevonc.2015.04.007.
Moskow JJ, Bullrich F, Huebner K, Daar IO, Buchberg AM. Meis1, a PBX1-related homeobox gene involved in myeloid leukemia in BXH-2 mice. Mol Cell Biol. 1995;15:5434–43.
Moens CB, Selleri L. Hox cofactors in vertebrate development. Dev Biol. 2006;291:193–206.
Penkov D, San Martín DM, Fernandez-Díaz LC, Rosselló CA, Torroja C, Sánchez-Cabo F, et al. Analysis of the DNA-binding profile and function of TALE homeoproteins reveals their specialization and specific interactions with Hox genes/proteins. Cell Rep. 2013;3:1321–33.
Dekel B, Metsuyanim S, Schmidt-Ott KM, Fridman E, Jacob-Hirsch J, Simon A, et al. Multiple imprinted and stemness genes provide a link between normal and tumor progenitor cells of the developing human kidney. Cancer Res. 2006;66:6040–9.
Cai M, Langer EM, Gill JG, Satpathy AT, Albring J, Wumesh KC, et al. Dual actions of Meis1 inhibit erythroid progenitor development and sustain general hematopoietic cell proliferation. Blood. 2012;120:335–46.
Hisa T, Spence SE, Rachel RA, Fujita M, Nakamura T, Ward JM, et al. Hematopoietic, angiogenic and eye defects in Meis1 mutant animals. EMBO J. 2004;23:450–9.
Tucker ES, Lehtinen MK, Maynard T, Zirlinger M, Dulac C, Rawson N, et al. Proliferative and transcriptional identity of distinct classes of neural precursors in the mammalian olfactory epithelium. Development. 2010;137:2471–81.
Yamada T, Urano-Tashiro Y, Tanaka S, Akiyama H, Tashiro F. Involvement of crosstalk between Oct4 and Meis1a in neural cell fate decision. PLoS One. 2013;8, e56997.
Graham V, Khudyakov J, Ellis P, Pevny L. SOX2 functions to maintain neural progenitor identity. Neuron. 2003;39:749–65.
Adachi K, Suemori H. Yasuda Sy, Nakatsuji N, Kawase E. Role of SOX2 in maintaining pluripotency of human embryonic stem cells. Genes Cells. 2010;15:455–70.
Bass AJ, Watanabe H, Mermel CH, Yu S, Perner S, Verhaak RG, et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet. 2009;41:1238–42.
Gen Y, Yasui K, Zen Y, Zen K, Dohi O, Endo M, et al. SOX2 identified as a target gene for the amplification at 3q26 that is frequently detected in esophageal squamous cell carcinoma. Cancer Genet Cytogenet. 2010;202:82–93.
Forghanifard MM, Khales SA, Javdani-Mallak A, Rad A, Farshchian M, Abbaszadegan MR. Stemness state regulators SALL4 and SOX2 are involved in progression and invasiveness of esophageal squamous cell carcinoma. Med Oncol. 2014;31:1–8.
Alonso MM, Diez-Valle R, Manterola L, Rubio A, Liu D, Cortes-Santiago N, et al. Genetic and epigenetic modifications of Sox2 contribute to the invasive phenotype of malignant gliomas. PLoS One. 2011;6, e26740.
Han X, Fang X, Lou X, Hua D, Ding W, Foltz G, et al. Silencing SOX2 induced mesenchymal-epithelial transition and its expression predicts liver and lymph node metastasis of CRC patients. PLoS One. 2012;7, e41335.
Li X, Xu Y, Chen Y, Chen S, Jia X, Sun T, et al. SOX2 promotes tumor metastasis by stimulating epithelial-to-mesenchymal transition via regulation of WNT/β-catenin signal network. Cancer Lett. 2013;336:379–89.
Bareiss PM, Paczulla A, Wang H, Schairer R, Wiehr S, Kohlhofer U, et al. SOX2 expression associates with stem cell state in human ovarian carcinoma. Cancer Res. 2013;73:5544–55.
Sobin LH, Gospodarowicz MK, Wittekind C. TNM classification of malignant tumours. 7th ed. Oxford: Wiley-Blackwell; 2009.
Shimada Y, Imamura M. Prognostic significance of cell culture in carcinoma of the oesophagus. Br J Surg. 1993;80:605–7.
Forghanifard MM, Gholamin M, Farshchian M, Moaven O, Memar B, Forghani MN, et al. Cancer-testis gene expression profiling in esophageal squamous cell carcinoma: Identification of specific tumor marker and potential targets for immunotherapy. Cancer Biol Ther. 2011;12:191–7.
Sinicrope FA, Ruan S, Cleary KR, Stephens LC, Lee JJ, Levin B. bcl-2 and p53 oncoprotein expression during colorectal tumorigenesis. Cancer Res. 1995;55:237–41.
Hu N, Clifford RJ, Yang HH, Wang C, Goldstein AM, Ding T, et al. Genome wide analysis of DNA copy number neutral loss of heterozygosity (CNNLOH) and its relation to gene expression in esophageal squamous cell carcinoma. BMC Genomics. 2010;11:576.
Kimchi ET, Posner MC, Park JO, Darga TE, Kocherginsky M, Karrison T, et al. Progression of Barrett's metaplasia to adenocarcinoma is associated with the suppression of the transcriptional programs of epidermal differentiation. Cancer Res. 2005;65:3146–54.
Su H, Hu N, Yang HH, Wang C, Takikita M, Wang Q-H, et al. Global gene expression profiling and validation in esophageal squamous cell carcinoma and its association with clinical phenotypes. Clin Cancer Res. 2011;17:2955–66.
Root DE, Hacohen N, Hahn WC, Lander ES, Sabatini DM. Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat Methods. 2006;3:715–9.
Barde I, Salmon P, Trono D. Production and titration of lentiviral vectors. Curr Protoc Neurosci. 2010;4:4.21.
Mahmood T, Yang P-C. Western blot: Technique, theory, and trouble shooting. N Am J Med Sci. 2012;4:429–34.
Xiang P, Lo C, Argiropoulos B, Lai CB, Rouhi A, Imren S, et al. Identification of E74-like factor 1 (ELF1) as a transcriptional regulator of the Hox cofactor MEIS1. Exp Hematol. 2010;38:798–808.
Wong P, Iwasaki M, Somervaille TC, So CW, Cleary ML. Meis1 is an essential and rate-limiting regulator of MLL leukemia stem cell potential. Genes Dev. 2007;21:2762–74.
Okumura K, Saito M, Isogai E, Aoto Y, Hachiya T, Sakakibara Y, et al. Meis1 Regulates Epidermal Stem Cells and Is Required for Skin Tumorigenesis. PLoS One. 2014;9, e102111.
Geerts D, Revet I, Jorritsma G, Schilderink N, Versteeg R. MEIS homeobox genes in neuroblastoma. Cancer Lett. 2005;228:43–50.
Crijns APG, de Graeff P, Geerts D, Ten Hoor KA, Hollema H, Van Der Sluis T, et al. MEIS and PBX homeobox proteins in ovarian cancer. Eur J Cancer. 2007;43:2495–505.
Crist RC, Roth JJ, Waldman SA, Buchberg AM. A conserved tissue-specific homeodomain-less isoform of MEIS1 is downregulated in colorectal cancer. PLoS One. 2011;6, e23665.
Chen JL, Li J, Kiriluk KJ, Rosen AM, Paner GP, Antic T, et al. Deregulation of a hox protein regulatory network spanning prostate cancer initiation and progression. Clin Cancer Res. 2012;18:4291–302.
Lasa A, Carnicer M, Aventin A, Estivill C, Brunet S, Sierra J, et al. MEIS 1 expression is downregulated through promoter hypermethylation in AML1-ETO acute myeloid leukemias. Leukemia. 2004;18:1231–7.
Kondo Y, Shen L, Cheng AS, Ahmed S, Boumber Y, Charo C, et al. Gene silencing in cancer by histone H3 lysine 27 trimethylation independent of promoter DNA methylation. Nat Genet. 2008;40:741–50.
Beukers W, Hercegovac A, Vermeij M, Kandimalla R, Blok AC, van der Aa MM, et al. Hypermethylation of the polycomb group target gene PCDH7 in bladder tumors from patients of all ages. J Urol. 2013;190:311–6.
Acknowledgments
The authors gratefully acknowledge the scientific and technical support of colleagues at the Division of Human Genetic, Avicenna Research Institute (Mashhad University) and the Department of Biology, Ferdowsi University of Mashhad. In addition, we thank Dr. Heydar Parsaee (Department of Pharmacology, Mashhad University) and Dr. Thomas Mikeska (Department of Pathology, Peter MacCallum Cancer Centre, East Melbourne, Australia) for their technical assistances. This study was a MSc thesis and supported by a grant from Mashhad University of Medical Sciences (# 89751).
Conflict of interest
None.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The study was approved beforehand by the local Ethics Committee. Informed, declared consent was obtained from all individual participants included in the study, and is on record. This article does not contain any studies with animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Abolfazl Rad and Moein Farshchian contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Figure 1
SOX2 mRNA detection in 50 ESCC and matched adjacent non-tumoral tissue samples. SOX2 mRNA expression is higher in tumor than in adjacent non-tumoral tissue, but this difference does not reach significance. SOX2 mRNA expression was determined with qRT-PCR, with GAPDH as the reference gene. Shown are the 2log median centered mean values, the error bars represent the SD. A two-sided t-test was used to assess the statistical difference between ESCC and adjacent non-tumoral tissue. (TIFF 329 kb)
Rights and permissions
About this article
Cite this article
Rad, A., Farshchian, M., Forghanifard, M.M. et al. Predicting the molecular role of MEIS1 in esophageal squamous cell carcinoma. Tumor Biol. 37, 1715–1725 (2016). https://doi.org/10.1007/s13277-015-3780-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3780-9