Abstract
Early screening for bladder cancer (BC) holds the key to combat and control the increasing global burden of BC mortality. We presented a simple approach to characterize, analyze, and validate a panel of biomarkers in BC and their relationship to bilharziasis. We investigated voided urine and blood samples from patients with bladder cancer (n = 94), benign bladder lesions (n = 60), and age-matched normal controls (n = 56). This study was divided into the following phases. (1) We analyzed the expression of urinary Hyaluronoglucosaminidase 1 (HYAL1) protein in BC and control samples by zymography. (2) We performed bioinformatics analysis to retrieve a set of epigenetic regulators of HYAL1. (3) This set of three selected genes [long non-coding RNA-urothelial cancer associated 1(lncRNA-UCA1), microRNA-210, and microRNA-96] was then analyzed in the same urine samples used in phase I by quantitative real-time PCR. (4) A high reproducibility of gene selection results was also determined from statistical validation. The urinary expression of HYAL1 protein and its epigenetic regulators were higher in BC patients (P < .001). The receiver-operating characteristic curve analyses demonstrated that each one had good sensitivity and specificity for distinguishing BC patients from non-BC ones (HYAL1, 89.4 and 91.2 %; miR-210, 76.6 and 93 %; miR-96, 76.6 and 89.4 %; and lncRNA-UCA1, 91.5 and 96.5 %). There was a significant positive correlation between HYAL1 and the selected epigenetic biomarkers. The performance of this urine biomarker panel reached 100 % sensitivity and 89.5 % specificity for bladder cancer diagnosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although the current available screening modalities for bladder cancer (BC) like cystoscopy and urine cytology are inadequate because of their high cost and cumbersome preparatory procedures [1], it would be ideal, therefore, to develop an alternative modality based on urine biomarkers as the first line screening test. Diagnostic biomarkers have resulted in great advances, such as targeting specific molecules to inhibit tumor growth [2].
Identification of groups of markers and an understanding of their interactions allows for greater understanding of disease pathways and the biological functions of associated genes [3].
Hyaluronoglucosaminidase 1 (HYAL1) is an endoglycosidase enzyme that mainly degrades hyaluronic acid. Several studies point out that HYAL1 is involved in tumor growth and metastasis [4] especially in high-grade BC [5].
Tumor-specific molecular alterations, including both genetic and epigenetic aberrations, have been shown to play crucial roles in BC development. Impaired epigenetic regulation, including aberrant miRNA [6] and lncRNA [7], is frequently reported in BC. Such targets may represent attractive biomarkers for early detection [8].
In previous studies, associations between BC and several biomarkers such as lncRNA-UCA1 [9], miR-210 [10], miR-96 [11], miRNA-220, or Hyaluronoglucosaminidase 1 [12] were identified.
In the current research, we adopted a new approach to study a panel of genetic and epigenetic biomarkers in bladder cancer using urine samples. For polygenic diseases such as bladder cancer and a complex detection platform such as human urine, we recognize that a single protein biomarker approach will not suffice for the high performance requirement of bladder cancer diagnosis. Therefore, by enlisting multiple genes that are mechanistically linked to each other and to bladder cancer pathways or functional networks, we believe that the chance of success would be higher than the simpler conventional single-marker approach.
Patients and methods
Research ethics statement
This study was approved by the Research Ethics Committee of Ain Shams University, Egypt. An informed consent was obtained from all participants.
Patient selection and clinical samples
We investigated sera and freshly voided urine samples obtained from 150 patients admitted to the Urology Department, Faculty of Medicine, Ain Shams University Hospitals, Cairo, Egypt, between May 2013and December 2014. Demographic data and medical history were obtained at the entry of each patient to the study. Patients who had undergone any previous chemotherapy, or with past history of bladder cancer, or any other malignancy within the past 5 years, were excluded from the study (Table 1). Study groups included 94 patients with current BC, as well as 56 cases who were age and sex-matched benign patients, and required cystoscopy for benign urinary conditions (e.g., bilharzial cystitis, benign prostatic hyperplasia). Tumor samples were staged according to the tumor-node-metastasis (TNM) classification by the American Joint Committee on Cancer-(AJCC) [13] and Union Internationale Contre le Cancer (UICC) [14]; in addition, the samples were histologically graded by a histopathologist [15]. A group of 60 healthy volunteers recruited from the hospital laboratory staff was also included in the study.
All subjects provided around 40–60 ml voided urine before instrumentation or bladder tumor removal. Each sample was centrifuged, and the supernatant was stored at −80 °C and the pellet was washed twice with phosphate-buffered saline. A portion of the pellet was used for cytological and microscopic examinations, while the other portion was treated with RNAlater (Qiagen, USA) and stored at −80 °C for further processing.
Schistosomal antibodies were detected in sera by Cellognost® Schistosomiasis H kit supplied by Dade Behring Marburg GmbH, Marburg, Germany [16].
Detection of urinary HYALA1 protein by semi-quantitative zymography
Urinary HYAL1 activity was examined using a HYAL1 zymography procedure, as described previously [17]. Briefly, the urine supernatants were electrophoresed under denaturing conditions on 8 % SDS-polyacrylamide gel containing 0.17 mg/mL HA. At the end of the electrophoresis, the gel was submerged in HAase assay buffer for enzymatic digestion at 37 °C for 16 h. Then, the gel was stained sequentially with Alcian blue to stain intact HA and with Coomassie blue to overstain the Alcian blue stained HA and the non-enzymatic protein bands. Then, the gel was destained with 10 % methanol/10 % acetic acid solution. The enzyme activity appeared as white bands in a dark blue background. Finally, semi-quantification of the enzymatic activity was achieved after scanning the gel by a digital scanner and analyzed by Gel-pro version 3.1 software (Media cybernetics, USA), thus determining the amount of HYL1 enzyme in different samples in relation to the standard.
Retrieval of candidate epigenetic regulators of HYAL1 in BC
We interrogated hyaluronidase gene as an input to produce a list of regulatory genes in a ranking order of best prediction using microRNA.Org Target and expression database (available at http://www.microrna.org/microrna/home.do), StarBase (http://starbase.sysu.edu.cn/), and lnCeDB Database (Available at http://gyanxet-beta.com/lncedb/). Since HYAL1 has many putative epigenetic regulator genes, the current bioinformatic analysis was focused on specific lncRNA and miRNAs which were published in BC and were targeting HYALA1. The three databases confirmed that the aberrant expression of the selected epigenetic biomarkers is consistent with their mechanism of action and correlates with HYAL1 protein target modulation in bladder cancer with higher ranking score (Supplementary Figs. 1s , 2s , 3s , and 4 s). Pathway enrichment analysis of miR210 and miR-96 was performed using the DIANA-mirPath software [18] and the KEGG pathway [18]. It revealed that the selected 2 miRNAs have got higher number of target genes related to carcinogenesis, e.g., focal adhesion, MAPK, Wnt signaling. Moreover, both miRs, lncRNA-UCA1 and HYAL1, are intersected in pathways related to glycerophospholipid degradation, TGFB pathway, and notch signaling pathway. In order to explore the evolutionary relationship between this, panel of genes and phylogenetic trees were generated by comparing the nucleotide sequences using AlignX (Vector NTI Advance v. 11.5.1, Life Technologies) which uses the ClustalW algorithm [19]. Results indicate close correlation between HYAL1 gene and the selected epigenetic regulators. The area of conserved regions with the multiple sequence alignment is shown in Supplementary Fig. 5s .
Total RNA including miRNA extraction from urine samples
Total RNA, including small non-coding miRNA, was isolated from urine samples using miREasy RNA isolation kit (Qiagen, MD). Then, RNA quality was determined using Ultraspec 1000, UV/visible spectrophotometer, Amersham Pharmacia Biotech, Cambridge, England, and NanoDrop 2000 Thermoscientific. The ratio of absorbance at 260 and 280 nm was used to assess the purity of RNA. A ratio of ~2.0 is generally accepted as “pure” for RNA. Afterwards, the RNA was kept in −80 °C till its use in the reverse transcription polymerase chain reaction.
Quantitative real-time PCR analyses of urinary lncRNA-UCA1
cDNA was synthesized using a QuantiTect® Reverse Transcription kit (Qiagen, Germany) in Hybaid thermal cycler (Thermo Electron Waltham, MA) according to manufacturer instructions. The expression of lncRNAs-UCA1 was measured by TaqMan probe qPCR using the specific primer/probe combination provided with each QuantiFast Probe Assay (UCA1, Accession NR_015379.3 and GAPDH, AccessionNM_004360),(Qiagen, USA) and TaqMan Universal PCR Master Mix on StepOnePlus™ System (Applied Biosystems Inc., Foster, CA) according to the manufacturer’s instructions. Each sample was analyzed in triplicate (Supplementary Fig. 6s ).
qRT-PCR analyses of urinary miRNA-210 and miRNA-96
cDNA was synthesized using 1 μg miRNA with a miScript II RT Kit (Qiagen/SABiosciences Corporation, Frederick, MD). Quantitative RT-PCR was carried out using StepOnePlus™ System (Applied Biosystems Inc., Foster, CA) using RNU-6 as a control. The PCR primers for RNU-6, miR-210, and miR-96 were purchased from (Qiagen, MD). SYBR Green Master Mix (Qiagen/SABiosciences Corporation, Frederick, MD) was used in the real-time PCR reaction according to the manufacturer’s suggested protocol, along with the manufacturer-provided miScript Universal primer and miRNA-specific forward primer (details in supplementary material, Supplementary Figs. 6 s 7s ).
Data normalization and quantification of lncRNA-UCA1 and microRNAs in urine samples
Relative gene expression (fold change) was used to measure the relative changes in expression of the various RNAs by using the 2−△△Ct method [20].
Statistical analysis
Data analysis was performed using Statistical Package for the Social Sciences software (SPSS, Version 19, Chicago, IL, USA). Comparisons were performed using chi-square and t tests or ANOVA tests, as appropriate (details in supplementary material).
Results
Patient characteristics
The basic clinical and demographic data are summarized in Table 1.
Urinary hyaluronidase activity in bladder cancer
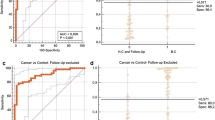
Urine samples were subjected to hyaluronan zymography. In the majority of samples, one lysis band was observed corresponding to 55 kDa. Semi-quantitative analysis of the zymograms revealed increased urinary Hyaluronoglucosaminidase 1 activity in BC samples compared to control samples (Tables 2 and 3; Fig. 1). The median levels for HYAL1 protein in benign and malignant groups were significantly increased by1.2- and 1.89-fold as compared to normal group, respectively (P < 0.0001) (Table 2). Using the ROC curve, the best cutoff point, for HYAL1 to discriminate between the malignant and the non-malignant groups, was 0.948 (Fig. 2a). Based on this cutoff value, 84 out of 94 (89.4 %) malignant patients, 6 out of 54 (10 %) benign patients, and 4 out of 60 (7.1 %) normal individuals were positive (P < 0.0001) (Table 3). No significant correlation was found between HYAL1 protein positivity rate and any of the studied clinicopathological factors (P > 0.05). The highest activity was observed at late stages and higher grades (Supplementary Table 1).
Semi-quantitative analysis of hyaluronidase activity in bladder cancer using gel zymography that produced bands with 55 kDa. Lane 1 positive control, lane 2 protein standard b, lanes 3–4 normal urine samples, lanes 5–8 bladder cancer lesions, lane 9 benign bladder lesion, and lane 10 negative control
a ROC curve analysis for HYAL1 protein to calculate the best cutoff point that discriminated between malignant and non-malignant groups. Best cutoff point of HYAL1 protein was 107.5 [sensitivity = 89.4% and specificity = 91.2 %]. Area under the curve (AUC) [SE] = 0.948 [0.015], 95 % confidence limits range = 0.919–0.977, P < 0.001. b ROC curve analysis for miRNA-210 to calculate the best cutoff point that discriminated between malignant and non-malignant groups. Best cutoff point of miRNA-210 was 1.17 [sensitivity = 76.6 %, specificity = 93 %]. Area under the curve (AUC) [SE] = 0.874 [0.025], 95 % confidence limits range = 0.825–0.924, P < 0.001. c ROC curve analysis for miRNA-96 to calculate the best cutoff point that discriminated between malignant and non-malignant groups. Best cutoff point of miRNA-96 was 1.65 [sensitivity = 76.6 % and specificity = 89.4 %]. Area under the curve (AUC) [SE] = 0.864 [0.025], 95 % confidence limits range = 0.815–0.914, P < 0.001. d ROC curve analysis for lncRNA-UCA1 to calculate the best cutoff point that discriminated between malignant and non-malignant groups. Best cutoff point of lncRNA-UCA1 was 1.09 [sensitivity = 91.5% and specificity = 96.5 %]. Area under the curve (AUC) [SE] = 0.975[0.011], 95 % confidence limits range = 0.953–0.996, P < 0.001. e ROC curve analysis for the combined genes to calculate the best cutoff point that discriminated between malignant and non-malignant groups. Best cutoff point of the combined genes was 23.6 [sensitivity = 100% and specificity = 89.5 %]. Area under the curve (AUC) [SE] = 0.981 [0.009], 95 % confidence limits range = 0.963–0.998, P < 0.001
Urinary miRs and lncRNA-UCA1 levels in the investigated groups
Urinary miR levels, based on RQ values, are summarized in Table 2. Compared to control groups, the malignant group had higher expression levels of miR-210, miR-96, and lncRNA-UCA1 (P < .001). We further analyzed these epigenetic markers by the receiver operating characteristic curve. The areas under the curve of miR-210, miR-96, and lncRNA-UCA1 were 0.874, 0.864, and 0.975, respectively. At a cutoff level of 1.17, 1.65, and 1.09, respectively, urinary miR-210, miR-96, and lncRNA-UCA1 had a sensitivity of 76.6, 76.6, and 91.5 %, respectively, and specificity of 93, 89.4, and 96.5 %, respectively, for diagnosis of bladder cancer (Table 4, Fig. 2b–e). We found no significant difference between miRNAs and lncRNA expression and any of the clinicopathological factors (P > 0.05) in the malignant group except significant correlations between miR-210 with age, miR-96 with age, bilharziasis and stage, and lncRNA-UCA1 with bilharziasis and type (P < 0.05) (Supplementary Table 1). Authors have explored the concordance between miR-210, miR-96, and lncRNA-UCA1 expression patterns and urine cytology among all study population (n = 210); the concordance rate was 72.4, 70.5, and 75.2 %, respectively as shown in Supplementary Table 2s
Performance characteristics of investigated urinary biomarkers in bladder cancer diagnosis
The accuracy of investigated biomarkers (HYAL1 Protein, miR-210, miR-96, and lncRNA-UCA1) varied from 89.5, 93.3, 82.9, and 84.8 %, respectively. lncRNA-UCA1 had the best individual performance with specificity, positive and negative predictive values of 96.5, 89.4, and 91.2 %, respectively. lncRNA-UCA1 and HYAL1 protein showed the highest sensitivity and specificity even in early-stage bladder cancer. The performance of the whole biomarker panel reached 100 % sensitivity and 89.5 % specificity as shown in Table 4.
Concordance and correlation between the four urinary biomarkers
The overall concordance rate between the HYAL1 and (miR-210, miR-96, incRNA-UCA1) was 87.6, 93.3, and 88.57 %, respectively (Supplementary Table 3s ). In addition, the overall concordance rate between different parameters is shown in Supplementary Tables 4s and 5s . Pearson’s correlation analysis was performed to determine whether there were correlations between the urinary levels of biomarkers among the groups of the study. There was a significant positive correlation between HYAL1, lncRNA-UCA1, and miRs. There was significant correlation between miRNA-210 and lncRNA-UCA1 (Table 5).
Relation of urinary biomarkers expression to bilharziasis
This study investigated urinary (HYAL1 protein, miR-210, miR-96, and lncRNA-UCA1) expression level relative to bilharziasis, and there is no significant difference between bilharzial and non-bilharzial cancer cases except in lncRNA-UCA1, yet a significant difference between bilharzial and non-bilharzial benign was observed in miR-210 and miR-96. Interestingly, all bilharzial positive cases (n = 38) were positive lncRNA-UCA1; 30 bilharzial malignant cases showed positive miR-210, 26showed positive miR-96, and 32 showed positive HYAL1 protein, as shown in (Supplementary Table 6s ).
Discussion
Cytoscopy and biopsies are the gold standard approaches for detecting BC. However, cystoscopy requires a degree of invasiveness, resulting in iatrogenic injury to the urethra and bladder, and there is a risk of infection too [1, 2]. The FDA-approved biomarkers for detecting BC rely on a single quantitative parameter, which may result in many false-negative or false-positive results. Hence, it does not provide satisfactory information for clinicians [3].
Urinary HA and HYA1 levels are diagnostic markers for BC, and they promote growth, invasion, and angiogenesis in BC [4, 5]. Recent work carried out by our research team showed that direct detection of hyaluronidase in urine using cationic gold nanoparticles is a promising diagnostic test for BC [21].
Understanding the interactome of BC will enable the development of novel approaches to tackle its occurrence, progression, and metastasis [3]. We presented a conceptually simple approach that enables reliable integration of differential HYAL1 protein expression and its epigenetic regulators. This approach generates a novel biomarkers panel (HYAL1 protein, miR-210, miR-96, and lncRNA-UCA1) for BC studies. In our study, we used the Hyaluronoglucosaminidase 1 gene as a starting point to generate a list of regulatory genes, in a ranking order of highest prediction, using relevant data bases. In addition, we selected lncRNA and miRNAs which were previously published in BC [9–11] and were reported to target HYAL1. Then, we performed pathway enrichment analysis and multiple sequence alignment to explore the intersected signaling pathways and their evolutionary relationship. Finally, we validated these in silico results and investigated their significance in BC, by measuring this biomarkers panel in the same urine samples obtained from a cohort of bladder cancer patients and controls.
We found a significant positive correlation between HYAL1 protein and the selected epigenetic regulators (P < 0.001), indicating their interactome in BC pathogenesis. This concurs with the multiple sequence alignment results and the close correlation in the phylogenic tree (Supplementary Fig. 5s).
Functional analysis of ln-coding RNA and miRNAs indicated that they activate the transcription of HYAL1 gene, by forming double-stranded RNA that match the promoter of HYAL1 gene, thus removing any transcription inhibitory factors [22].
Lower sensitivity of urinary cytology (only 51.1 %) was shown in the study, whereas HYAL1, miR-210, miR-96, and lncRNA-UCA1 sensitivity (71.6, 76.6, 91.5, and 89.5 %, respectively) was superior to urine cytology. The selected panel reduces urine cytology false negative results from 48.9 to 0 %. We strongly believe that the diagnostic accuracy for BC would be improved by a concurrent measurement of urinary miR-210, miR-96, and lncRNA-UCA1 as well as HYAL1 protein (approximately 93.3 % accuracy in the present study), especially in early-stage and low-grade bladder cancer.
Aberrant miRNAs are produced by the host in response to schistosomiasis infection; thereby, they mediate the pathogenesis of schistosomiasis and its carcinogenesis and can be used as potential biomarkers [23]. This was shown in the current study where we had significant difference between bilharzial malignant and benign cases regarding urinary miR-96 and lncRNA-UCA1.
Conclusion
We used a genetic-epigenetic approach to identify a panel of lncRNA- and miRNA-based biomarkers targeting HYAL1 that may improve the diagnosis of BC. This can overcome poor reliability issues, from single-gene biomarker experiment, while maintaining high accuracies by combining true signals from multiple genetic levels. Moreover, they can detect both bilharzial and non-bilharzial bladder cancer at high accuracy.
Abbreviations
- BC:
-
Bladder cancer
- HYAL1:
-
Hyaluronoglucosaminidase 1
- kDa:
-
Kilodalton
- LncRNA:
-
Long non-coding RNA
- MiR:
-
Micro-RNA
- qPCR:
-
Quantitative polymerase chain reaction
- RT-PCR:
-
Reverse transcription polymerase chain reaction
- SCC:
-
Squamous cell carcinoma
- TCC:
-
Transitional cell carcinoma
- UCA1:
-
Urothelial cancer associated 1
References
Siegel R, Jiemin M, Zhaohui Z, Jemal A. Cancer statistics 2014. A Cancer Journal for Clinicians. 2014;64(1):15422–4863.
Sanchez C, Lachaize C, Janody F, Bellon B, Roder L, et al. Grasping at molecular interactions and genetic networks in Drosophila melanogaster using FlyNets, an Internet database. Nucleic Acids Res. 1999;27:89–94.
Lancashire LJ, Lemetre C, Ball GR. An introduction to artificial neural networks in bioinformatics—application to complex microarray and mass spectrometry datasets in cancer studies. Brief Bioinform. 2009;10:315–29.
Fraser JR, Laurent TC, Laurent UB. Hyaluronan: its nature, distribution, functions and turnover. J Intern Med. 1997;242:27–33.
Eissa S, Shehata H, Mansour A, Esmat M, El-Ahmady O. Detection of hyaluronidase RNA and activity in urine of schistosomal and non-schistosomal bladder cancer. Med Oncol. 2012;29:3345–51.
Miah S, Dudziec E, Drayton RM, et al. An evaluation of urinary microRNA reveals a high sensitivity for bladder cancer. Br J Cancer. 2012;107(1):123–8. doi:10.1038/bjc.2012.221.
Han Y, Liu Y, Nie L, et al. Inducing cell proliferation inhibition, apoptosis, and motility reduction by silencing long noncoding ribonucleic acid metastasis-associated lung adenocarcinoma transcript 1 in urothelial carcinoma of the bladder. Urology. 2013;81(209):e1–7.
Noon AP, Catto JW. Noncoding RNA in bladder cancer: a specific focus upon high-risk nonmuscle invasive disease. Curr Opin Urol. 2014;24(5):506–11.
Srivastava AK, Singh PK, Rath SK, Dalela D, Goel MM, Bhatt ML. Appraisal of diagnostic ability of UCA1 as a biomarker of carcinoma of the urinary bladder. Tumour Biol. 2014;15.
Qin Q, Furong W, Baosheng L. Multiple functions of hypoxia-regulated miR-210 in cancer. Cancer Res J Exp Clin. 2014;33:50. doi:10.1186/1756-9966-33-50.
Wang Y, Luo H, Li Y, Chen T, Wu S, Yang L. hsa-miR-96 up-regulates (MAP4K1 and IRS1 and may function as a promising diagnostic marker in human bladder urothelial carcinomas. Mol Med. 2012;5(1):260–5.
Jamshidian H, Hashemi M, Nowroozi MR, Ayati M, Bonyadi M, NajjaranTousi V. Sensitivity and specificity of urinary hyaluronic acid and hyaluronidase in detection of bladder transitional cell carcinoma. Urol J. 2014;11(1):1232–7.
Edge SB, Byrd DR, Compton CC, et al. AJCC cancer staging manual. 7th ed. New York: Springer; 2010. p. 497–505.
NCCN Clinical Practice Guidelines in Oncology: Bladder Cancer V. 2.2013. Available at http://bit.ly/jXgCvZ.Accessed September 5, 2013.
Eble JN, Sauter G, Epstein JI, Sesterhenn I, editors. World Health Organization classification of tumors pathology and genetics: tumors of the urinary system and male genital organs. Lyon: IARC Press; 2004.
Gui M, Idris MA, Shi YE, Muhling A, Ruppel A. Reactivity of Schistosoma japonicum and S. mansoni antigen preparations in indirect haemagglutination (IHA) with sera of patients with homologous and heterogonous schistosomiasis. Ann Trop Med Parasitol. 1991;85:599–604.
Papageorgakopoulou N, Vynios DH, Karayanni K, Maras A, Papapetropoulou M. Electrophoretic analysis of hydrolytic enzymes of Escherichia coli cells starved in seawater and drinking water: comparison of gelatinolytic, caseinolytic, phosphohydrolytic and hyaluronolytic activities. Microbiol Res. 1997;152:299–305.
Paraskevopoulou MD, Georgakilas G, Kostoulas Net al. DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res. 2013 Jul;41(Web Server issue):W169-73. Available at http://diana.cslab.ece.ntua.gr/pathways/
Thompson JD, Higgins DG, Gibson TJ. Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–80. CLUSTAL W.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−delta delta C (T)) method. Methods. 2001;25:402–8.
Nossier AI, Eissa S, Ismail MF, Hamdy MA, Azzazy HM. Direct detection of hyaluronidase in urine using cationic gold nanoparticles: a potential diagnostic test for bladder cancer. Biosens Bioelectron. 2014;54:7–14. doi:10.1016/j.bios.2013.10.024.
Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A. 2008;105(5):1608–13. doi:10.1073/pnas.0707594105.
Zhu L, Liu J, Cheng G. Role of microRNAs in schistosomes and schistosomiasis. Front Cell Infect Microbiol. 2014;4:165. doi:10.3389/fcimb.2014.00165. eCollection 2014.
Acknowledgments
This work was supported by Ain Shams University Research Projects 2013–14. The authors are grateful to Dr. Nahla M. Awad, Ass. Prof. of Pathology at Early Cancer Detection Unit, Faculty of Medicine, Ain Shams University, for her help in the cytological examinations of all investigated urine samples. All authors have read the journal’s policy on disclosure of potential conflicts of interest. The authors have no conflict of interest. All authors have read the journal’s authorship agreement and that the manuscript has been reviewed by and approved by all named authors.
Conflicts of interest
None.
Authors’ contributions
Eissa S has participated in the design of the study, carried out data analysis, involved in drafting the manuscript or revising and has given final approval of the version to be published. Matboli M has performed bioinformatic analysis, practical work, participated in the design of the study, and performed the statistical analysis. Essawy N participated in the study design and involved in drafting the manuscript or revising. Youssef M. Kotb has provided us with urine, blood samples, and patient data and has given final approval of the version to be published. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM 1
(DOC 1082 kb).
Rights and permissions
About this article
Cite this article
Eissa, S., Matboli, M., Essawy, N.O.E. et al. Integrative functional genetic-epigenetic approach for selecting genes as urine biomarkers for bladder cancer diagnosis. Tumor Biol. 36, 9545–9552 (2015). https://doi.org/10.1007/s13277-015-3722-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3722-6