Abstract
Non-coding RNAs (ncRNAs) are important regulatory molecules involved in various physiological and pathological cellular processes. Small nucleolar RNAs (snoRNAs), subclass of small ncRNAs, have been considered important but unglamorous elements in the production of protein synthesis machinery of cells. However, recent evidence has indicated that these non-coding RNAs might have a crucial role also in controlling cell behavior, and snoRNAs dysfunction could significantly contribute to carcinogenesis. Here, we summarize the most important aspects of snoRNAs biology, their functioning in cancer cell, and potential usage in diagnosis or as a new class of therapeutic targets in cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The nucleolus, the most prominent organelle in the interphase nucleus, provides the cellular locale for synthesis and processing of cytoplasmic ribosomal RNAs (rRNAs) [1]. The biogenesis of cytoplasmic ribosome is a complex process. The 18S, 5.9S, and 25/28S rRNAs are generated from a large, common precursor RNA (pre-rRNA) containing long external and internal transcribed spacer sequences. To generate functional rRNA, spacer regions are removed from the primary transcript to produce stoichiometric amounts of mature small and large subunit rRNAs. Before its cleavage by endo- and exonucleases, the nascent pre-rRNA undergoes a complex pattern of nucleoside modification of its mature small subunits and large subunit sequences. There are two types of these modifications: 2-O-ribose methylation and pseudouridylation, each involving about 50–100 sites per eukaryotic ribosome [2–5] located within the conserved, functionally important domains of mature RNAs [6]. While these modifications are not essential, they are likely to fine-tune rRNA folding and interactions with ribosomal proteins, thereby modulating both biogenesis and activity of ribosomes [2].
Besides precursor rRNAs processing, the nucleolus also contains an enormous number of metabolically stable 60–300 nucleotide long RNAs, called small nucleolar RNAs (snoRNAs), which are associated with a set of proteins to form small nucleolar ribonucleoproteins (snoRNPs). During the last decade, snoRNAs studies have revealed many novel, unexpected cellular functions for non-coding RNAs and changed long-held beliefs about eukaryotic gene expression [1]. Several hundreds of snoRNAs have been reported in a variety of organisms, making snoRNAs the most abundant group of non-coding RNAs. The two types of eukaryotic rRNA modification are directed by two large families of snoRNAs which specify the sites to be modified, in both cases through the formation of a specific duplex at the rRNA modification site, while the catalytic function is provided by a common protein enzyme, methylase or pseudouridine synthase, associated with the snoRNA [2].
For many years, snoRNAs have been considered one of the best characterized classes of non-coding RNAs (ncRNAs) [5, 7–9]. Despite the common assumption that snoRNAs have only cellular housekeeping functions, recent independent reports have converged in implicating snoRNAs in control of cell fate and oncogenesis [10–16]. The snoRNA are well-conserved, abundant, short non-coding RNA molecules, 60–300 nucleotides in length, which localize to a specific compartment of the cell nucleus—the nucleolus. In vertebrates, the majority of snoRNAs are encoded in introns of protein-coding or non-coding genes and are transcribed simultaneously by RNA Pol II [17]. Most snoRNA host genes encode for proteins or transcripts essential for ribosome biogenesis or function and often belong to the 5′-terminal oligopyrimidine (TOP) family that undergoes growth-dependent translation regulation. The generation of most intronic snoRNAs involves splicing and debranching followed by subsequent exonucleolytic trimming by the 5′- to 3′-exoribonuclease XRN2 [18, 19]. Second, splicing-independent pathway that generates intronic snoRNAs requires endonucleolytic cleavage of the pre-mRNA host transcript by the RNase III-like enzyme Rnt1p in yeast [20]. In mammals and the nematode Caenorhabditis elegans, in contrast to flies and plants, snoRNAs are almost exclusively monocistronic [17]. Vertebrate snoRNAs are mostly of intronic origin with the exception of autonomously transcribed essential snoRNAs that direct pre-rRNA cleavage. Notably, many intronic snoRNAs are associated with genes coding for ribosomal and nucleolar proteins. Mature snoRNAs are usually trafficked to the nucleolus. This process requires the presence of conserved structural elements within the nucleotide sequence of snoRNA [21] and relies on a number of transport factors, such as the cap-binding complex (CBC), the phosphorylated export adapter (PHAX), and exportin CRM1 [22, 23].
Biogenesis of snoRNAs
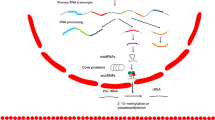
SnoRNAs associate with small nucleolar ribonucleoprotein particles (snoRNP) and serve as guide molecules of the snoRNPs in the post-transcriptional modification of rRNA and small nuclear RNA (snRNA). Based on the presence of conserved sequence motifs, snoRNAs are sub-categorized into H/ACA box (SNORA) that guides pseudouridylation, and C/D box (SNORD) that guides 2′O-ribose methylation of targeted RNA (Fig. 1a, b). The two types of snoRNAs associate with distinct sets of proteins, necessary for proper formation and enzymatic function of the snoRNP (Table 1). The snoRNA guide families have been identified in a wide spectrum of eukaryotes species (Table 2). In vertebrates, most snoRNAs are located within introns of protein-coding genes and are transcribed by RNA polymerase II (Fig. 2a). However, snoRNAs can also be processed from introns of long non-coding RNAs (lncRNAs) [24, 25]—for example, GAS5, a lncRNA, encodes 9 C/D box snoRNAs (snoRNDs74-81) [26]. The efficient processing of intronic snoRNAs requires active recruitment of snoRNPs to the nascent intronic snoRNAs during synthesis or before splicing of the host pre-mRNAs. The C/D box snoRNAs form functional snoRNPs by forming a complex with four evolutionary conserved and essential proteins (Fibrillarin (Nop1p), Nop56p, Nop58p, and 15.5-kDa/Snu13). On the other hand, the H/ACA box snoRNAs associate with dyskerin (Cbf5p, Gar1p, Nhp2p, and Nop10p) to form functional snoRNPs (Table 1). After liberation from introns, snoRNAs are processed to remove excess nucleotides from either end via exonuclease activity (Fig. 2a). Signal sequences within boxes C and D or H and ACA direct binding of protein interacting partners that represent the functional snoRNP complex. Within the H/ACA box and C/D box snoRNAs, there are two functionally distinct groups that are not presumably involved in pseudouridylation and methylation. The box H/ACA box snoRNA, snR30, and the C/D box snoRNAs, U3 and U14, are required for cleavage of the pre-ribosomal RNAs (rRNAs) at the early processing sites [27].
snoRNAs. a Boxed sequences C and D are hallmarks of C/D box snoRNAs. b Boxed sequences H (hinge region) and ACA are hallmarks of the H/ACA box snoRNAs. Adapted from [59]
a snoRNAs predominantly located in introns. Following splicing, debranching, and trimming, mature snoRNAs are either exported, in which case they function in ribosomal RNA (rRNA) processing or remain in the nucleus, where they are involved in alternative splicing and additional yet unknown functions. Adapted from [100], b an example of a snoRNA gene independently annotated as the gene of miRNA. The gene of the human SNORA36B snoRNA is simultaneously annotated as the gene of miR-664 miRNA. The mature miRNA (miR-664-3p), its precursor (miR-664), and SNORA36B snoRNA are shown. Below small RNAs are shown detected during deep sequencing and corresponding to fragments of SNORA36B snoRNA. Adapted from [39], c the role of snoRNA in Prader-Willi syndrome (PWS). The loss of the snoRNA in PWS changes the alternative splicing of the serotonin receptor HTR2C precursor mRNA, resulting in a protein with reduced functions. Adapted from [100]
snoRNAs function
snoRNAs in processing of rRNA
The well-studied function of snoRNAs is their involvement in processing of rRNA. 18S, 5.8S, and 25/28S rRNAs are transcribed from the common precursor (pre-rRNA), which is cleaved into mature rRNA molecules. The snoRNAs have two basic mechanisms: 2′-O-methylation and pseudouridylation of rRNAs [28]. 2′-O-methylation of rRNAs is carried out by the C/D box snoRNA family, which is characterized by a kink-turn (stem-bulge-stem) structure containing two short sequence motifs. The box C sequence (RUGAUGA, where R stands for A or G) is typically located close to the 5′ terminus, whereas box D sequence (CUGA) is situated near the 3′ end (Fig. 1a). The two motifs are generally brought together in a typical 5′-3′ terminal stem-box structure involving 4–5 nucleotides at both termini, which is critical for snoRNA biogenesis and nuclear localization [2, 3, 27, 29–33]. Many C/D box snoRNAs contain a less conserved copy of the box C motif C′ in the central position [34] and an additional box D motif, termed box D′ in their 5′ half [35], where RNA-protein interactions occur to direct the proper assembly of the functional snoRNP complexes [36]. Upstream from box D and/or box D′, there are one or two of the so-called antisense elements, i.e., 10–21 nucleotide long sequences that are complementary to a site of rRNA allowing for proper alignment and methylation of appropriate nucleotides (Fig. 1a) [7, 37, 38]. The snoRNA component directs the snoRNP complex to the appropriate rRNA location, where the methyltransferase, fibrillarin, catalyzes the methylation reaction [2, 3, 19, 36]. Several C/D box snoRNAs (SNORD3, SONRD14, SNORD22, SNORD118, and probably SNORD13) are required for the pre-rRNA cleavage [37, 39–44]. Methylation guide function of box C/D antisense snoRNAs and the essential role of the CUGA box motif in determining the precise nucleotide to be methylated in the RNA duplex at the fifth position upstream from box D or box D′ have been experimentally demonstrated [7, 37]. Expression of an artificial C/D box snoRNA carrying an appropriate antisense element is sufficient to target a novel ribose methylation on the predicted pre-rRNA nucleotide and also, to a lesser extent, to RNA-polymerase II transcripts. This shows that the antisense element associated with box D (or D′) is the sole determinant of the site of methylation [2, 37].
The pseudouridylation of rRNAs is accomplished by the H/ACA box of snoRNAs family composed of two large hairpin domains linked by a hinge and followed by short tail [2, 36]. The conserved motif called box H (ANANNA, N stands for any nucleotide) and ACA (a trinucleotide always found three nucleotides away from the 3′ end) are located in the hinge and tail (Fig. 1b) [45, 46]. The H/ACA box of snoRNA contains an appropriate bipartite guide sequence in the internal loop of one (or both) of the two large hairpin domains [47]. The two stems forming the 9–13-bp bipartite guide duplex precisely flank the substrate uridine which remains accessible for isomerization. Guide sequences that direct the snoRNAs to the appropriate rRNA sequence are in one or both of the hairpin loop domains. After targeting the snoRNP complex to the appropriate site, the substrate rRNA uridine is positioned at the base of the upper stem. Reminiscent of the target/box-D spacing rule observed for methylation guides, the conserved distance (14–16 nucleotides) between the target uridine and the downstream H or ACA box of the snoRNA is a critical determinant of the pseudouridylation site (Fig. 1b) [47]. The core H/ACA RNP includes the pseudouridine synthase dyskerin (cbf5p in yeast) and proteins Nop10, Nhp2, and Gar1 [48, 49]. The H/ACA box snoRNA SNORA73, similarly to some C/D box snoRNAs, does not direct modification and is necessary for cleavage of pre-rRNA [39, 50]. The modifications are necessary for normal functioning of the ribosome and seem to be responsible for the correct packing of rRNA, stabilization of its structure, and for correct interaction of rRNA with other participants of translation [39, 51–54].
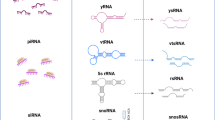
A combination type of snoRNAs that comprises features of both SNORAs and SNORDs localizes specifically to Cajal bodies, small sub-organelles in the nuclei of proliferative or metabolically active cells involved in the biogenesis of small nuclear RNPs. Cajal body-specific RNAs (SCARNAs) contain a specific element, called Cajal body box (CAB box) that is necessary for their retention within these bodies (Fig. 3) [55, 56]. SCARNAs can guide both methylation and pseudouridylation of the RNA Pol II transcribed spliceosomal RNAs [57]. The main function of these bodies is likely the assemblage of snRNP complexes, which are later transported into other compartments of the nucleus.
Involvement of snoRNAs in various cellular processes. Adapted from [39]
snoRNAs as precursor of miRNAs
It has been believed that the central function of snoRNAs is to modify, mature, and stabilize rRNAs. These post-translation modifications are important for the production of efficient and accurate ribosomes [58]. In addition, snoRNAs cellular functions continue to expand outside of the nucleolus. Processed RNAs could hold a key to some of the newly found effects of snoRNAs [59]. Processing of snoRNAs into smaller fragments has been reported with convincing evidence suggesting that some snoRNAs act as precursors for functional miRNA [60–63]. The first report that identified fragments of the H/ACA box snoRNA ACA45 in association with Ago proteins from the miRNA RISC complex showed that processing of ACA45 is dependent on Dicer. Furthermore, resulting 20–22 nucleotide fragments were validated to function as a miRNA that represses expression of an identified target (CDC2L6) [60]. While only a small fraction of the total cellular ACA45 is processed by Dicer to act as miRNA, this clearly represents a role for snoRNA derivatives in the cytoplasm. Moreover, some miRNAs found in independent studies correspond to fragments of snoRNAs (Table 3). Genome browsers clearly demonstrate these findings (Fig. 2b). Recently, 11 C/D box snoRNAs that are processed to miRNA-sized species were identified and verified to indicate gene silencing activity [61]. A subset of miRNAs shared functional H/ACA box snoRNA characteristics suggesting that these miRNAs might have evolved from snoRNAs [64]. Moreover, some snoRNAs are processed to produce small RNAs. The small RNAs processed from the snoRNAs could function as miRNAs. Such processing could be of crucial importance, since miRNAs have essential roles in a spectrum of regulatory processes, such as the control of cell survival and proliferation. The small RNAs produced from snoRNAs—sno-miRNAs—could have a dual function; the same transcript could function both as snoRNA and as a precursor for microRNAs [61, 65]. The vertebrate telomerase RNAs contain a typical H/ACA domain that contributes to the synthesis of telomere DNA. Since the box H/ACA motif of the human telomerase reverse transcriptase (hTER) is required for its association with four proteins (dyskerin, NHP2, NOP10, and GAR1) that are common in snoRNAs, these molecules might be involved in telomere functioning [66–68].
snoRNAs for alternate splicing
The snoRNA transcripts serve as precursors of miRNA-like small RNAs and as regulators of alternative splicing [60, 69]. The 18-nucleotide-conserved target recognition element of a HBII-52 C/D box snoRNA is complementary to the serotonin receptor 5-HT (2C) mRNA [70]. The snoRNA HBII-52 can control processing of mRNA expression of the serotonin receptor 2C by regulating its alternative splicing and hence contributes to the Prader-Willi syndrome (Fig. 2c) [70, 71]. The mechanism appears to involve further processing of the snoRNA to remove 5′ and 3′ inverted repeats that usually stabilize the C/D box snoRNA stem. MBII-52 snoRNA that has been processed in this way loses its ability to associate with C/D box proteins (e.g., fibrillarin) and instead associates with hnRNPs that are involved with splice-site selection [69].
Since none of brain-specific snoRNAs show sequence complementarity to rRNAs, snRNAs, or other non-coding RNAs, only one brain-specific snoRNA was found to have a potential target mRNA. HBII-52 exhibits an 18-nucleotide complementarity to the mRNA for the human 5-HT2C receptor subunit which also holds true for the mouse homologs (MBII-52) of these two genes [71]. This region on the 5-HT2C receptor subunit message is subject to alternative splicing [72]. The MBII-52 snoRNA could direct 2′-O-methylation of an adenosine residue that is known to be partially deaminated to inosine in the serotonin receptor mRNA [73]. Since 2′-O-methylation inhibits adenosine deamination, the MBII-52 snoRNA might have an important regulatory function. Interestingly, the second intron of the human serotonin receptor 5-HT2C gene also encodes a brain-specific box H/ACA box snoRNA of unknown function.
snoRNAs in stress response
The nucleolus is one of the key participants of the cell response to stress. Various stress conditions can cause changes in the nucleolus and even its destruction. Under stress conditions, some ribosomal proteins of mammalian cells are moved into the cytoplasm and interact there with ubiquitin ligase MDM2. Under normal conditions, MDM2 ubiquitinates the transcriptional factor p53 that leads to its degradation. Interaction with ribosomal proteins inhibits MDM2 that stabilizes p53. As a result, the cell stops dividing or enters apoptosis (Fig. 5) [59, 61, 62, 64, 74, 75].
It was observed that snoRNA contribute to stress response. The expression of SNORD14A and SNORD83B was significantly increased under hypoxial conditions [76]. SNORD14A is required for normal translation, as it is involved in pre-rRNA cleavage, but the function of SNORD83B is unknown. The expression of SNORDs 32A, 33, and 35A was shown to significantly increase under oxidative stress upon treatment of cells with palmitate [77]. These SNORDs are encoded within introns of the gene for ribosomal proteins RPL13A and rRNA nucleotides serve as their targets, but involvement of these SNORDs in methylation has not been described [38]. Resistance against palmitate is due to the suppression of SNORD32A, SNORD33, and SNORD35A, which mediate cell death induction in response to an inducer [77]. Under stress conditions, full-length forms of these snoRNAs are accumulated not in the nucleus but in the cytoplasm, which is not accompanied by an increase in the extent of methylation of rRNA; therefore, their effect seems not to be associated with the modification of rRNA [77]. The snoRNAs in the cytoplasm are observed only during the early processing stages. For instance, independently transcribed snoRNAs (SNORD3, SNORD13, and SNORD118) [19] additionally target mRNAs; under metabolic stress, they complementarily interact and regulate their translation in the cytoplasm. The pathways of snoRNA involvement in the regulation of cellular processes can be even more diverse. Thus, there are some indications that the H/ACA box snoRNA ACA11 inhibits the cell response under conditions of oxidative stress [39, 78].
snoRNAs in cancer
snoRNAs involvement in molecular pathology of cancer
snoRNAs are the class of non-coding RNAs which have been well-characterized [5, 7–9]. I assumed that they only have cellular housekeeping functions, but recent reports have indicated the important role of these snoRNA in controlling cell fate and carcinogenesis [10–16]. One of the first implications of snoRNAs in carcinogenesis is from the study of B cell lymphoma which identified C/D box snoRNAU50 and its host gene U50HG at the breakpoint of chromosomal translocation t (3;6)(q21;q15) [79]. To identify prostate-associated genes in chromosome 6q14-q22, which is often deleted in human cancers, Dong et al. narrowed the common region of the deletion to a 2.5-Mb location at 6q14-15. Of the 11 genes located in this minimally deleted region, only snoRNAU50 was discovered to be mutated in prostate cancer cells [11]. A homozygous 2-bp deletion in the snoRNAU50 was found in prostate cancer cell lines/xenografts and in 9 out of 89 localized prostate cancer cases, but not in the 104 normal controls. Ectopic expression of snoRNAU50 significantly reduced colony formation in prostate cancer cells [11]. In addition, heterozygous genotype of the same deletion was observed in 8 of 32 breast cancer cell lines [10].
The putative oncogene snoRNA42 (SNORA42) is associated with carcinogenesis. snoRNA42 is located on chromosome 1q22, a genomic region frequently subjected to amplification in lung carcinomas [80]; over-expression of snoRNA42 is frequently observed in non-small-cell-lung carcinoma (NSCLC) [81, 82]. SNORA42 exhibited a close correlation between increase of copy number and expression level, suggesting that SNORA42 over-expression could be activated through its amplification. Repression of SNORA42 in NSCLC cells caused a marked decrease in lung cancer growth in vitro and in vivo; enforced SNORA42 expression in bronchial epithelium increases cell growth and colony formation. It was also demonstrated that SNORA42 expression in lung tumor tissue samples inversely correlates with survival of NSCLC patients [81]. These independent studies on snoRNAU50 and SNORA42 provide evidence for the functional importance of snoRNAs in cancer. They show that snoRNAs can promote, as well as suppress, tumor development. Subsequently, Chang et al. proved that h5sn2, a box H/ACA box snoRNA, was significantly down-regulated in human meningiomas compared to normal brain tissue [83]. Genome-wide approaches identified six snoRNAs (SNORD33, SNORD66, SNORD73B, SNORD76, SNORD78, and SNORA42), which are differentially expressed in NSCLC compared to noncancerous samples [13]. Notably, all identified snoRNAs displayed up-regulation in lung tumor samples. Majority of these snoRNAs are located in frequently amplified genomic regions in lung cancer [84, 85]. SNORD33 is located on chromosome 19q13.3 that contains potential oncogenes implicated in lung cancer [86, 87], while SNORD66 and SNORD76 are situated in chromosomal regions 3q27.1 and 1q25.1, respectively. 3q27.1 and 1q25.1 are two of the most frequently amplified chromosomal segments in solid tumors, particularly NSCLC [84–88]. Four snoRNAs (RNU44, RNU48, RNU43, and RNU6B), often used as internal control genes to analyze cancer-related miRNAs, are associated with poor prognosis of cancer patients [12]. Recently, Xu et al. [89] showed that SNORD113-1 gene, which is located at chromosome region 14q32 within the intron of the small nucleolar RNA host gene 23 (SNHG23), functions as a tumor suppressor in hepatocellular carcinoma (HCC). SNORD113-1 exhibited a strong ability to inhibit cell growth and proliferation in human HCC cells. In addition, the results from a xenograft animal model provided direct evidence that SNORD113-1 can regulate HCC tumor cell growth and that the loss of SNORD113-1 gene function may be directly associated with tumor development. The activity of SNORD113-1 caused growth suppression [89].
The growth arrest-specific transcript 5 (GAS5) gene is not detectable in actively growing cells but is highly expressed in cells under serum starvation or growth arrest [84, 90]. GAS5 can control mammalian apoptosis and cell population growth. Interestingly, GAS5 does not have a protein-coding potential but encodes nine C/D box snoRNAs in its introns and exhibits some characteristics which belong to the class of genes encoding 5′ terminal oligopyrimidine (5′TOP) mRNA [91]. 5′TOP mRNAs have been largely restricted to proteins involved in translation, such as ribosomal proteins and translation factors. The translation of 5′TOP mRNAs is modulated to allow maximal translation when increased production of the protein synthesis machinery is required—in actively growing cells (Fig. 4a). The mTOR pathway controls the 5′TOP mRNA translation. For GAS5, and potentially for other snoRNA host gene transcripts, this arrangement has additional consequences [59]. The first evidence that GAS5 is related to cancer comes from a study on breast cancer samples where the expression the GAS5 transcript level is significantly reduced when compared to normal breast epithelial samples [16]. Other studies also showed that GAS5 reduced expression is associated with poor prognosis in both breast cancer and head and neck squamous cell carcinoma [12]. GAS5 has been also identified as a novel partner of BCL6 in a patient with diffuse large B cell lymphoma, harboring translocation t (1;3) (q25;q27) [92]. GAS5 mRNA contains a small and poorly conserved open reading frame. The termination codon for open reading frame is found in an early exon, when the GAS5 mRNA is translated in growing cells. When the mTOR pathway is active, GAS5 mRNA is degraded, but when the mTOR pathway is not active, the GAS5 mRNA is no longer actively translated and no longer degraded [26, 93]. Therefore, the GAS5 mRNA accumulates in cells that are undergoing growth arrest, and this mechanism might account for its initial isolation as transcript that is associated with growth arrest (Fig. 4b) [26, 59, 90].
Potential functions of the GAS5 host gene. a The GAS5 mRNA contains a small open reading frame (ORF) followed by multiple stop codons. GAS5 mRNA contains binding sites for glucocorticoid receptors, and other nuclear hormone receptors, indicating that it may function as a decoy glucocorticoid response element (GRE). The translation of GAS5 mRNA is stimulated in actively growing cells, and is mediated, in part, by the protein kinase mTOR. In growing cells, mTOR-stimulated translation of GAS5 mRNA promotes its degradation by the nonsense-mediated RNA decay pathway, levels of GAS5 mRNA in actively growing cells are very low, and the glucocorticoid hormone pathway can operate freely. b In growth-arrested cells, the reduced translation of 5′TOP mRNAs facilitates the accumulation of GAS5 mRNA and consequently produces higher levels of the GRE decoy. The GRE decoy binds to and sequesters the glucocorticoid receptor, preventing its interaction with genomic glucocorticoid response elements. In cells that are dependent on the glucocorticoid hormone pathway for growth, cell cycle arrest occurs. Adapted from [59]
Another spliced transcript which functions independently of snoRNAs is Zfas1 [94]. Zfas1 hosts three C/D box snoRNAs (SNORD12, SNORD12b, and SNORD12c) and has been identified as one of the most differentially expressed gene during mouse mammary development. Down-regulation of Zfas1 increased cell proliferation and differentiation without affecting the levels of the snoRNA [94]; these findings indicate that snoRNA host genes might have important functions in regulating cellular homeostasis and cancer biology. There are several reports showing that H/ACA box and C/D box snoRNAs are nucleolytically processed to produce smaller products. Ender et al. [60] found that the processing of H/ACA box snoRNA ACA45 produced 20–25 nucleotides-long RNAs that just like miRNAs associate with Ago (Argonaute) proteins, and target specific mRNA, including CDK11 within the cell. Some of the processed snoRNAs can function as miRNAs [62, 64]. Such processing may be of crucial importance, as miRNAs are known to have important roles in many aspects of cell behavior, including the control of cell survival and proliferation, and they have been implicated in the development of many cancers. Recently, Xiao et al. [95] have reported that an H/ACA box snoRNA-derived miRNA, miR-605, has a key role in stress-induced stabilization of the p53 tumor suppressor protein [95]. p53 transcriptionally activates its negative regulator, MDM2, in addition to miR-605, miR-605 counteracts MDM2 through post-transcriptional repression; under conditions of stress, this snoRNA-derived miRNA offsets the MDM2 negative-feedback loop, generating a positive-feedback loop to enable rapid accumulation of p53 (Fig. 5). However, whether this regulation of p53 by miR-605 is relevant to cancer biology has not yet been addressed [59, 74, 95].
microRNA and snoRNA-mediated p53 positive-feedback loop. p53-mediated activation of MDM2 transcription in dividing cells is normally limited by a negative-feedback loop: transcriptional activation of the MDM2 gene produces increased levels of MDM2 protein, resulting in MDM2-mediated ubiquitylation (Ub) and proteasomal degradation of p53. In cells undergoing stress, this negative-feedback loop is broken: increased levels of p53 produce p53-mediated transcriptional activation of mir-605. The mir-605 and snoRNA genes produce both an H/ACA box small nucleolar RNA (snoRNA) and microRNA, as do several other snoRNA genes. miR-605 inhibits MDM2 translation, and this increases p53 stability, resulting in a progressive increase in p53 protein levels, inhibition of cell cycle progression, and induction of apoptosis. Adapted from [59]
An unbiased functional screen for genes that respond to metabolic stress identified three C/D box snoRNA: SNORD32A, SNORD33, and SNORD35A [77], which are encoded within the introns of the gene for ribosomal proteins RPL13A. In this study, mutagenesis was induced by retroviral promoter trap and followed by palmitate selection. The resistance against palmitate is due to the suppression of SNORD32A, SNORD33, and SNORD35A, which mediate cell death induction in response to inducer. Further studies tried to understand the molecular mechanisms involved in cell death in diabetes and other metabolic syndromes. It was shown that snoRNAs are involved in the response to general oxidative stress during cancerogenesis [77].
The association between snoRNAs and tumorigenesis is represented by the involvement of their associated proteins in cancer. Dyskerin, an enzyme that associates with H/ACA box snoRNAs and catalyses pseudouridylation, is also an essential component of mammalian telomerase [96]. A point mutation in the dyskerin gene (DKC1) causes a rare X-linked recessive disease, the dyskeratosis congenital (DC) [97, 98]. Individuals with DC display features of premature aging, as well as nail dystrophy, mucosal leukoplakia, interstitial fibrosis of the lung, and increased susceptibility to cancer. DKC1 codes for dyskerin, a putative pseudouridine synthase, which carries out two separate functions, both fundamental for proliferating cells. One function is the pseudouridylation of ribosomal RNA (rRNA) molecules as a part of the H/ACA ribonucleoprotein complex, and the other is the stabilization of the telomerase RNA component necessary for telomerase activity. Like dyskerin, NHP2 and NOP10 proteins, both components of the H/ACA snoRNPs are also significantly up-regulated in sporadic cancers and their high levels may be associated with poor clinical prognosis. Moreover, germline NHP2 and NOP10 mutations give rise to autosomal recessive forms of dyskeratosis congenital, and cancer susceptibility is also a feature of these genetic forms of the disease.
snoRNAs as potential diagnostic and prognostic biomarkers in cancer
The role of snoRNAs in cells is diverse, and expression of cancer-specific snoRNAs should be exploited for the development of cancer biomarkers. It has been shown that snoRNAs are stable and measurable in peripheral blood plasma and serum samples [12, 13, 27, 99]. snoRNAs have the potential to become circulating biomarkers of cancer (Table 4). Increased levels of C/D box snoRNAs, SNORD33, SNORD66, and SNORD76, are detected not only in tumors but also in plasma with observed sensitivity and specificity in distinguishing NSCLC patients. Measuring circulating plasma snoRNAs early will serve as potential non-invasive approach for the diagnosis of NSCLC [13]. Increased expression of H/ACA snoRNA SNORA42 in tissues correlated the unfavorable outcome of the disease and can be used for predicting the disease course [81]. Recent observation further supported the development of snoRNAs as a potential biomarker [12]. The three C/D box snoRNAs, SNORD43, SNORD44, and SNORD48, seem to be putative suppressors of breast cancer and head and neck squamous cell carcinomas [12]. snoRNAU50 encoded by introns of the non-coding host gene (U50HG) seems to be suppressor of breast and prostate cancer (Table 4) [10, 11]. SNORD113-1 shown to be down-regulated in hepatocellular carcinoma (HCC) and functions as a tumor suppressor in HCC [89]. Taken together, snoRNAs may serve as potential biomarkers for both diagnosis and prognosis of malignancies.
snoRNAs as potential therapeutic targets in cancer
The molecular mechanism of snoRNA functioning in cancer is still not fully understood. However, some snoRNAs are ideal candidates for therapeutic intervention. Silencing of transcriptional gene pathway mediated by snoRNA may be of therapeutic benefit; also, the use of RNAi-mediated gene silencing could be used for selective silencing of oncogenic snoRNAs. For example, snoRA42 is expressed highly in lung tumor tissues; silencing the expression of snoRA42 was tested in a lung cancer cell line [81]. Knockdown of snoRA42 significantly decreased proliferation and viability in NSCLC cells. These findings provide an evidence to indicate the potential in developing snoRNA-mediated therapies. However, many challenges need to be overcome for using siRNA-mediated knockdown of snoRNA before snoRNAs could become potential targets for therapy.
Conclusion and future directions
In conclusion, in the last years, numerous and unexpected results were obtained on the role of snoRNAs in the cell biology. On one hand, new molecular mechanisms of the involvement of snoRNAs in cellular processes were described and, on the other hand, changes in the expression level of snoRNAs were shown to be significant for potential clinical application. However, comprehensive genomic approaches and large cohorts of patients are required to further evaluate potential usage of snoRNAs in cancer diagnostics indicated in pilot studies. Dysfunction of several snoRNAs was associated with the development of cancer and a few of concrete molecular mechanisms were described, but much more is waiting for elucidation. These snoRNAs after in vivo verification present new class of therapeutic targets in oncology.
References
Kiss T. Small nucleolar RNAs: an abundant group of noncoding RNAs with diverse cellular functions. Cell. 2002;109:145–8.
Bachellerie JP, Cavaille J, Huttenhofer A. The expanding snoRNA world. Biochimie. 2002;84:775–90.
Cavaille J, Bachellerie JP. SnoRNA-guided ribose methylation of rRNA: structural features of the guide RNA duplex influencing the extent of the reaction. Nucleic Acids Res. 1998;26:1576–87.
Maden BE. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog Nucleic Acid Res Mol Biol. 1990;39:241–303.
Tollervey D, Kiss T. Function and synthesis of small nucleolar RNAs. Curr Opin Cell Biol. 1997;9:337–42.
Brimacombe R, Mitchell P, Osswald M, Stade K, Bochkariov D. Clustering of modified nucleotides at the functional center of bacterial ribosomal RNA. FASEB J. 1993;7:161–7.
Kiss-Laszlo Z, Henry Y, Bachellerie JP, Caizergues-Ferrer M, Kiss T. Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell. 1996;85:1077–88.
Weinstein LB, Steitz JA. Guided tours: from precursor snoRNA to functional snoRNP. Curr Opin Cell Biol. 1999;11:378–84.
Williams GT, Hughes JP, Stoneman V, Anderson CL, McCarthy NJ, Mourtada-Maarabouni M, et al. Isolation of genes controlling apoptosis through their effects on cell survival. Gene Ther Mol Biol. 2006;10:255–62.
Dong XY, Guo P, Boyd J, Sun X, Li Q, Zhou W, et al. Implication of snoRNA U50 in human breast cancer. J Genet Genomics. 2009;36:447–54.
Dong XY, Rodriguez C, Guo P, Sun X, Talbot JT, Zhou W, et al. SnoRNA U50 is a candidate tumor-suppressor gene at 6q14.3 with a mutation associated with clinically significant prostate cancer. Hum Mol Genet. 2008;17:1031–42.
Gee HE, Buffa FM, Camps C, Ramachandran A, Leek R, Taylor M, et al. The small-nucleolar RNAs commonly used for microRNA normalisation correlate with tumour pathology and prognosis. Br J Cancer. 2011;104:1168–77.
Liao J, Yu L, Mei Y, Guarnera M, Shen J, Li R, et al. Small nucleolar RNA signatures as biomarkers for non-small-cell lung cancer. Mol Cancer. 2010;9:198.
Martens-Uzunova ES, Jalava SE, Dits NF, van Leenders GJ, Moller S, Trapman J, et al. Diagnostic and prognostic signatures from the small non-coding RNA transcriptome in prostate cancer. Oncogene. 2012;31:978–91.
Mourtada-Maarabouni M, Hedge VL, Kirkham L, Farzaneh F, Williams GT. Growth arrest in human T-cells is controlled by the non-coding RNA growth-arrest-specific transcript 5 (GAS5). J Cell Sci. 2008;121:939–46.
Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F, Williams GT. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene. 2009;28:195–208.
Dieci G, Preti M, Montanini B. Eukaryotic snoRNAs: a paradigm for gene expression flexibility. Genomics. 2009;94:83–8.
Filipowicz W, Pogacic V. Biogenesis of small nucleolar ribonucleoproteins. Curr Opin Cell Biol. 2002;14:319–27.
Watkins NJ, Bohnsack MT. The box C/D and H/ACA snoRNPs: key players in the modification, processing and the dynamic folding of ribosomal RNA. Wiley Interdiscip Rev RNA. 2012;3:397–414.
Giorgi C, Fatica A, Nagel R, Bozzoni I. Release of U18 snoRNA from its host intron requires interaction of Nop1p with the Rnt1p endonuclease. EMBO J. 2001;20:6856–65.
Reichow SL, Hamma T, Ferre-D’Amare AR, Varani G. The structure and function of small nucleolar ribonucleoproteins. Nucleic Acids Res. 2007;35:1452–64.
Boulon S, Verheggen C, Jady BE, Girard C, Pescia C, Paul C, et al. PHAX and CRM1 are required sequentially to transport U3 snoRNA to nucleoli. Mol Cell. 2004;16:777–87.
Pradet-Balade B, Girard C, Boulon S, Paul C, Azzag K, Bordonne R, et al. CRM1 controls the composition of nucleoplasmic pre-snoRNA complexes to licence them for nucleolar transport. EMBO J. 2011;30:2205–18.
Bortolin ML, Kiss T. Human U19 intron-encoded snoRNA is processed from a long primary transcript that possesses little potential for protein coding. RNA. 1998;4:445–54.
Smith CM, Steitz JA. Sno storm in the nucleolus: new roles for myriad small RNPs. Cell. 1997;89:669–72.
Smith CM, Steitz JA. Classification of GAS5 as a multi-small-nucleolar-RNA (snoRNA) host gene and a member of the 5′-terminal oligopyrimidine gene family reveals common features of snoRNA host genes. Mol Cell Biol. 1998;18:6897–909.
Mannoor K, Liao J, Jiang F. Small nucleolar RNAs in cancer. Biochim Biophys Acta. 2012;1826:121–8.
Panse VG, Johnson AW. Maturation of eukaryotic ribosomes: acquisition of functionality. Trends Biochem Sci. 2010;35:260–6.
Cavaille J, Bachellerie JP. Processing of fibrillarin-associated snoRNAs from pre-mRNA introns: an exonucleolytic process exclusively directed by the common stem-box terminal structure. Biochimie. 1996;78:443–56.
Caffarelli E, Fatica A, Prislei S, De Gregorio E, Fragapane P, Bozzoni I. Processing of the intron-encoded U16 and U18 snoRNAs: the conserved C and D boxes control both the processing reaction and the stability of the mature snoRNA. EMBO J. 1996;15:1121–31.
Lange TS, Borovjagin A, Maxwell ES, Gerbi SA. Conserved boxes C and D are essential nucleolar localization elements of U14 and U8 snoRNAs. EMBO J. 1998;17:3176–87.
Samarsky DA, Fournier MJ, Singer RH, Bertrand E. The snoRNA box C/D motif directs nucleolar targeting and also couples snoRNA synthesis and localization. EMBO J. 1998;17:3747–57.
Villa T, Ceradini F, Bozzoni I. Identification of a novel element required for processing of intron-encoded box C/D small nucleolar RNAs in Saccharomyces cerevisiae. Mol Cell Biol. 2000;20:1311–20.
Kiss-Laszlo Z, Henry Y, Kiss T. Sequence and structural elements of methylation guide snoRNAs essential for site-specific ribose methylation of pre-rRNA. EMBO J. 1998;17:797–807.
Tycowski KT, Shu MD, Steitz JA. A mammalian gene with introns instead of exons generating stable RNA products. Nature. 1996;379:464–6.
Kiss T. Small nucleolar RNA-guided post-transcriptional modification of cellular RNAs. EMBO J. 2001;20:3617–22.
Cavaille J, Nicoloso M, Bachellerie JP. Targeted ribose methylation of RNA in vivo directed by tailored antisense RNA guides. Nature. 1996;383:732–5.
Nicoloso M, Qu LH, Michot B, Bachellerie JP. Intron-encoded, antisense small nucleolar RNAs: the characterization of nine novel species points to their direct role as guides for the 2′-O-ribose methylation of rRNAs. J Mol Biol. 1996;260:178–95.
Makarova JA, Ivanova SM, Tonevitsky AG, Grigoriev AI. New functions of small nucleolar RNAs. Biochemistry (Mosc). 2013;78:638–50.
Borovjagin AV, Gerbi SA. U3 small nucleolar RNA is essential for cleavage at sites 1, 2 and 3 in pre-rRNA and determines which rRNA processing pathway is taken in Xenopus oocytes. J Mol Biol. 1999;286:1347–63.
Enright CA, Maxwell ES, Eliceiri GL, Sollner-Webb B. 5′ETS rRNA processing facilitated by four small RNAs: U14, E3, U17, and U3. RNA. 1996;2:1094–9.
Tycowski KT, Shu MD, Steitz JA. Requirement for intron-encoded U22 small nucleolar RNA in 18 s ribosomal RNA maturation. Science. 1994;266:1558–61.
Peculis BA, Steitz JA. Disruption of U8 nucleolar snRNA inhibits 5.8s and 28s rRNA processing in the Xenopus oocyte. Cell. 1993;73:1233–45.
Cavaille J, Hadjiolov AA, Bachellerie JP. Processing of mammalian rRNA precursors at the 3′ end of 18s rRNA. Identification of cis-acting signals suggests the involvement of U13 small nucleolar RNA. Eur J Biochem. 1996;242:206–13.
Balakin AG, Smith L, Fournier MJ. The RNA world of the nucleolus: two major families of small RNAs defined by different box elements with related functions. Cell. 1996;86:823–34.
Ganot P, Caizergues-Ferrer M, Kiss T. The family of box ACA small nucleolar RNAs is defined by an evolutionarily conserved secondary structure and ubiquitous sequence elements essential for RNA accumulation. Genes Dev. 1997;11:941–56.
Ganot P, Bortolin ML, Kiss T. Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell. 1997;89:799–809.
Lafontaine DL, Bousquet-Antonelli C, Henry Y, Caizergues-Ferrer M, Tollervey D. The box H + ACA snoRNAs carry Cbf5p, the putative rRNA pseudouridine synthase. Genes Dev. 1998;12:527–37.
Rashid R, Liang B, Baker DL, Youssef OA, He Y, Phipps K, et al. Crystal structure of a Cbf5-Nop10-Gar1 complex and implications in RNA-guided pseudouridylation and dyskeratosis congenita. Mol Cell. 2006;21:249–60.
Morrissey JP, Tollervey D. Yeast Snr30 is a small nucleolar RNA required for 18s rRNA synthesis. Mol Cell Biol. 1993;13:2469–77.
Baxter-Roshek JL, Petrov AN, Dinman JD. Optimization of ribosome structure and function by rRNA base modification. PLoS ONE. 2007;2:e174.
Blanchard SC, Puglisi JD. Solution structure of the a loop of 23s ribosomal RNA. Proc Natl Acad Sci U S A. 2001;98:3720–5.
Liu B, Liang XH, Piekna-Przybylska D, Liu Q, Fournier MJ. Mis-targeted methylation in rRNA can severely impair ribosome synthesis and activity. RNA Biol. 2008;5:249–54.
Basu A, Das P, Chaudhuri S, Bevilacqua E, Andrews J, Barik S, et al. Requirement of rRNA methylation for 80s ribosome assembly on a cohort of cellular internal ribosome entry sites. Mol Cell Biol. 2011;31:4482–99.
Henras AK, Dez C, Henry Y. RNA structure and function in C/D and H/ACA s(no) RNPs. Curr Opin Struct Biol. 2004;14:335–43.
Darzacq X, Jady BE, Verheggen C, Kiss AM, Bertrand E, Kiss T. Cajal body-specific small nuclear RNAs: a novel class of 2′-o-methylation and pseudouridylation guide RNAs. EMBO J. 2002;21:2746–56.
Tycowski KT, You ZH, Graham PJ, Steitz JA. Modification of U6 spliceosomal RNA is guided by other small RNAs. Mol Cell. 1998;2:629–38.
Decatur WA, Fournier MJ. rRNA modifications and ribosome function. Trends Biochem Sci. 2002;27:344–51.
Williams GT, Farzaneh F. Are snoRNAs and snoRNA host genes new players in cancer? Nat Rev Cancer. 2012;12:84–8.
Ender C, Krek A, Friedlander MR, Beitzinger M, Weinmann L, Chen W, et al. A human snoRNA with microRNA-like functions. Mol Cell. 2008;32:519–28.
Brameier M, Herwig A, Reinhardt R, Walter L, Gruber J. Human box C/D snoRNAs with miRNA like functions: expanding the range of regulatory RNAs. Nucleic Acids Res. 2011;39:675–86.
Scott MS, Avolio F, Ono M, Lamond AI, Barton GJ. Human miRNA precursors with box H/ACA snoRNA features. PLoS Comput Biol. 2009;5:e1000507.
Taft RJ, Glazov EA, Lassmann T, Hayashizaki Y, Carninci P, Mattick JS. Small RNAs derived from snoRNAs. RNA. 2009;15:1233–40.
Ono M, Scott MS, Yamada K, Avolio F, Barton GJ, Lamond AI. Identification of human miRNA precursors that resemble box C/D snoRNAs. Nucleic Acids Res. 2011;39:3879–91.
Scott MS, Ono M. From snoRNA to miRNA: dual function regulatory non-coding RNAs. Biochimie. 2011;93:1987–92.
Dez C, Henras A, Faucon B, Lafontaine D, Caizergues-Ferrer M, Henry Y. Stable expression in yeast of the mature form of human telomerase RNA depends on its association with the box H/ACA small nucleolar RNP proteins Cbf5p, Nhp2p and Nop10p. Nucleic Acids Res. 2001;29:598–603.
Mitchell JR, Cheng J, Collins K. A box H/ACA small nucleolar RNA-like domain at the human telomerase RNA 3′ end. Mol Cell Biol. 1999;19:567–76.
Pogacic V, Dragon F, Filipowicz W. Human H/ACA small nucleolar RNPs and telomerase share evolutionarily conserved proteins Nhp2 and Nop10. Mol Cell Biol. 2000;20:9028–40.
Kishore S, Khanna A, Zhang Z, Hui J, Balwierz PJ, Stefan M, et al. The snoRNA MBII-52 (Snord 115) is processed into smaller RNAs and regulates alternative splicing. Hum Mol Genet. 2010;19:1153–64.
Kishore S, Stamm S. The snoRNA HBII-52 regulates alternative splicing of the serotonin receptor 2c. Science. 2006;311:230–2.
Cavaille J, Buiting K, Kiefmann M, Lalande M, Brannan CI, Horsthemke B, et al. Identification of brain-specific and imprinted small nucleolar RNA genes exhibiting an unusual genomic organization. Proc Natl Acad Sci U S A. 2000;97:14311–6.
Canton H, Emeson RB, Barker EL, Backstrom JR, Lu JT, Chang MS, et al. Identification, molecular cloning, and distribution of a short variant of the 5-hydroxytryptamine2C receptor produced by alternative splicing. Mol Pharmacol. 1996;50:799–807.
Burns CM, Chu H, Rueter SM, Hutchinson LK, Canton H, Sanders-Bush E, et al. Regulation of serotonin-2C receptor G-protein coupling by RNA editing. Nature. 1997;387:303–8.
Suzuki A, Kogo R, Kawahara K, Sasaki M, Nishio M, Maehama T, et al. A new picture of nucleolar stress. Cancer Sci. 2012;103:632–7.
Scott MS, Ono M. From snoRNA to miRNA: dual function regulatory non-coding RNAs. Biochimie. 1987;93:1987–92.
Liu ZH, Yang G, Zhao T, Cao GJ, Xiong L, Xia W, et al. Small ncRNA expression and regulation under hypoxia in neural progenitor cells. Cell Mol Neurobiol. 2011;31:1–5.
Michel CI, Holley CL, Scruggs BS, Sidhu R, Brookheart RT, Listenberger LL, et al. Small nucleolar RNAs U32a, U33, and U35a are critical mediators of metabolic stress. Cell Metab. 2011;14:33–44.
Chu L, Su MY, Maggi Jr LB, Lu L, Mullins C, Crosby S, et al. Multiple myeloma-associated chromosomal translocation activates orphan snoRNA ACA11 to suppress oxidative stress. J Clin Invest. 2012;122:2793–806.
Tanaka R, Satoh H, Moriyama M, Satoh K, Morishita Y, Yoshida S, et al. Intronic U50 small-nucleolar-RNA (snoRNA) host gene of no protein-coding potential is mapped at the chromosome breakpoint t (3;6) (q27;q15) of human B-cell lymphoma. Genes to cells : devoted to mol & cell mech. 2000;5:277–87.
Goeze A, Schluns K, Wolf G, Thasler Z, Petersen S, Petersen I. Chromosomal imbalances of primary and metastatic lung adenocarcinomas. J Pathol. 2002;196:8–16.
Mei YP, Liao JP, Shen J, Yu L, Liu BL, Liu L, et al. Small nucleolar RNA 42 acts as an oncogene in lung tumorigenesis. Oncogene. 2012;31:2794–804.
Thompson DM, Lu C, Green PJ, Parker R. tRNA cleavage is a conserved response to oxidative stress in eukaryotes. RNA (New York, NY). 2008;14:2095–103.
Chang LS, Lin SY, Lieu AS, Wu TL. Differential expression of human 5s snoRNA genes. Biochem Biophys Res Commun. 2002;299:196–200.
Li R, Wang H, Bekele BN, Yin Z, Caraway NP, Katz RL, et al. Identification of putative oncogenes in lung adenocarcinoma by a comprehensive functional genomic approach. Oncogene. 2006;25:2628–35.
Jiang F, Yin Z, Caraway NP, Li R, Katz RL. Genomic profiles in stage I primary non small cell lung cancer using comparative genomic hybridization analysis of cDNA microarrays. Neoplasia (New York, NY). 2004;6:623–35.
Gebhart E. Double minutes, cytogenetic equivalents of gene amplification, in human neoplasia—a review. Clin Transl Oncol. 2005;7:477–85.
Schwab M. Oncogene amplification in solid tumors. Semin Cancer Biol. 1999;9:319–25.
Bell DW. Our changing view of the genomic landscape of cancer. J Pathol. 2010;220:231–43.
Xu G, Yang F, Ding CL, Zhao LJ, Ren H, Zhao P, et al. Small nucleolar RNA 113–1 suppresses tumorigenesis in hepatocellular carcinoma. Mol Cancer. 2014;13:216.
Schneider C, King RM, Philipson L. Genes specifically expressed at growth arrest of mammalian cells. Cell. 1988;54:787–93.
Amaldi F, Pierandrei-Amaldi P. Top genes: a translationally controlled class of genes including those coding for ribosomal proteins. Prog Mol Subcell Biol. 1997;18:1–17.
Nakamura Y, Takahashi N, Kakegawa E, Yoshida K, Ito Y, Kayano H, et al. The GAS5 (growth arrest-specific transcript 5) gene fuses to BCL6 as a result of t (1;3) (q25;q27) in a patient with B-cell lymphoma. Cancer Genet Cytogenet. 2008;182:144–9.
Yamashita A, Izumi N, Kashima I, Ohnishi T, Saari B, Katsuhata Y, et al. SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. Genes Dev. 2009;23:1091–105.
Askarian-Amiri ME, Crawford J, French JD, Smart CE, Smith MA, Clark MB, et al. SNORD-host RNA Zfas1 is a regulator of mammary development and a potential marker for breast cancer. RNA (New York, NY). 2011;17:878–91.
Xiao J, Lin H, Luo X, Wang Z. Mir-605 joins p53 network to form a p53:Mir-605:Mdm2 positive feedback loop in response to stress. EMBO J. 2010;30:524–32.
Montanaro L. Dyskerin and cancer: more than telomerase. The defect in mRNA translation helps in explaining how a proliferative defect leads to cancer. J Pathol. 2010;222:345–9.
Gupta V, Kumar A. Dyskeratosis congenita. Adv Exp Med Biol. 2010;685:215–9.
Ruggero D, Grisendi S, Piazza F, Rego E, Mari F, Rao PH, et al. Dyskeratosis congenita and cancer in mice deficient in ribosomal RNA modification. Sci (New York, NY). 2003;299:259–62.
Appaiah HN, Goswami CP, Mina LA, Badve S, Sledge Jr GW, Liu Y, et al. Persistent upregulation of U6:Snord44 small RNA ratio in the serum of breast cancer patients. Breast Cancer Res. 2011;13:R86.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12:861–74.
Di Leva G, Garofalo M (2013) Non-coding RNAs and cancer. In: Oncogene and cancer—from bench to clinic. License InTech.
Acknowledgments
This work was supported by the project “Employment of Best Young Scientists for International Cooperation Empowerment” (CZ.1.07/2.3.00/30.0037), co-financed from the European Social Fund and the state budget of the Czech Republic, by the project “CEITEC—Central European Institute of Technology” (CZ.1.05/1.1.00/02.0068) and by the project MZ CR–RVO (MOU, 00209805). The authors would like to thank Andrej Besse for preparation of the figures and John B. Smith for proofreading the manuscript.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Thorenoor, N., Slaby, O. Small nucleolar RNAs functioning and potential roles in cancer. Tumor Biol. 36, 41–53 (2015). https://doi.org/10.1007/s13277-014-2818-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2818-8