Abstract
Osteosarcoma is the most common primary malignant bone tumor in children and adolescents. Unfortunately, treatment failures are common due to the metastasis and chemoresistance, but the underlying molecular mechanism remains unclear. Accumulating evidence indicated that the deregulation of DNA-binding protein high-mobility group box 1 (HMGB1) was associated with the development of cancer. This study aimed to explore the expression of HMGB1 in osteosarcoma tissues and its correlation to the clinical pathology of osteosarcoma and to discuss the role of HMGB1 in the development of osteosarcoma. The results from RT-PCR and Western blot showed that the expression rate of HMGB1 messenger RNA (mRNA) and the expression of HMGB1 in the osteosarcoma tissues were significantly higher than those in normal bone tissue (p < 0.05), the expression rate of HMGB1 mRNA and the expression of HMGB1 in the carcinoma tissues with positive lung metastasis were significantly higher than those without lung metastasis (p < 0.05), and with increasing Enneking stage, the expression rate of HMGB1 mRNA and the expression of HMGB1 also increased (p < 0.05). In order to explore the role of HMGB1 in osteosarcoma, the expression of HMGB1 in the human osteosarcoma MG-63 cell line was downregulated by the technique of RNA interference. Western blot results showed that the protein expression of HMGB1 was significantly decreased in the MG-63 cells from HMGB1-siRNA transfection group (p < 0.05), which suggested that HMGB1 was successfully downregulated in the MG-63 cells. Then the changes in proliferation, apoptosis, and invasion of MG-63 cells were examined by MTT test, PI staining, annexin V staining, and transwell chamber assay. Results showed that the abilities of proliferation and invasion were suppressed in HMGB1 knockdown MG-63 cells, and the abilities of apoptosis were enhanced in HMGB1 knockdown MG-63 cells. The expression of cyclin D1, MMP-9 was downregulated in HMGB1 knockdown MG-63 cells, and the expression of caspase-3 was upregulated in HMGB1 knockdown MG-63 cells. Taken together, the overexpression of HMGB1 in osteosarcoma might be related to the tumorigenesis, invasion, and metastasis of osteosarcoma, which might be a potential target for the treatment of osteosarcoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteosarcoma (OS), also known as osteogenic sarcoma, is the most bone-associated malignancy with high lethality [1]. The estimated incidence rate of OS worldwide is four million/year, and its incidence is bimodally distributed by age with peaks in adolescence [2–4]. OS can form both bone tissue and osteoid tissue during the development process, mostly in the long bones of the extremities. Due to the emergence of adjuvant and neoadjuvant chemotherapy, the survival rate has been greatly improved in individuals with localized OS, which has an average 5-year survival of about 80 %. However, the long-term survival rate for OS patients with metastatic or recurrent disease remains poor [5, 6]. OS is characterized by a high malignant and metastatic potential. The major cause of death in OS is the increase in metastatic potential [7, 8]. Approximately 90 % of patients with OS have varying degrees of metastasis on diagnosis [9]. Tumor metastasis is a complex multistep event influenced by multiple regulatory genes [10, 11]. Although many studies have focused on the metastasis of OS, the exact molecular mechanism remains poorly understood.
High-mobility group box 1 (HMGB1) is a widely existing DNA-binding nuclear protein participating in gene transcription, chromatin remodeling, DNA recombination and repair processes, and stabilizing nucleosome construction [12, 13]. Accumulating evidence indicates that its function now extends beyond the nucleus, notably its extracellular role in inflammation, immune response, autophagy, and cancer [14–16]. HMGB1, a highly conserved nuclear protein, can be actively secreted by nature immunocyte cells and passively released by injured cells or necrosis cells as a damage-associated molecular pattern (DAMP) [17], exerting pleiotropic biological effects by binding to receptor for advanced glycation and products (RAGE) and toll-like receptor (TLR), which lead to activation of mitogen-activated protein kinases (MAPKs) and NF-κB, thereby promoting angiogenesis, unlimited replicative potential, tissue invasion, and metastasis [18, 19]. HMGB1 is an evolutionarily ancient and critical regulator of cell death and survival, and released HMGB1 triggers extracellular damage [15]. Many studies suggest that HMGB1 plays a critical role in the development and progression of multiple malignancies [20].
HMGB1 overexpression has been reported in a variety of human cancers. Several clinical studies have shown that HMGB1 is a promising biomarker for a variety of cancer types [21]. Yildirim and his colleagues demonstrated that HMGB1 displayed an upregulation in colorectal cancers, which had a significant role in tumor progression and tumor ability to metastasize in colorectal cancers [14]. Xiao et al. suggested that HMGB1 displayed an overexpression in hepatocellular carcinoma (HCC), and that the HCC patients with HMGB1 overexpression had a significantly shorter overall survival time than those with a downregulated expression of HMGB1 [15]. HMGB1 knockdown could suppress the ability of proliferation, migration, and invasion of HCC cells. HMGB1 overexpression could strengthen the ability of proliferation, migration, and invasion of HCC cells [15, 22]. HMGB1 expression is significantly increased in laryngeal squamous cell carcinoma (LSCC) tissues, and HMGB1 overexpression is associated with a poorer prognosis [23]. Zhang et al. demonstrated that increased expression of HMGB1 was associated with tumor metastasis of gastric adenocarcinoma, and knockdown of HMGB1 suppressed growth and invasion of gastric adenocarcinoma cells through the NF-κB pathway [20]. Ko et al. showed that HMGB1 enhanced cell proliferation and suppresses apoptosis by enhancing Bcl-xL, Bcl-2, cyclin D1, and NF-κB expression, decreasing Bax and p53 expression [24]. HMGB1 may be involved in liver cancer development and progression through Ki-67 and MMP-2 [22]. HMGB1 was overexpressed in the highly invasive ovarian cancer. HMGB1 knockdown significantly inhibited ovarian cancer cell proliferation and migration accompanied by decreased cyclin D1, MMP-2, and MMP-9 [25, 26]. The expression of HMGB1 has been reported in many types of cancers, but no information is available to date regarding the function of HMGB1 in OS. In this study, we performed RT-PCR assay to determine HMGB1 expression in both OS tissue and normal bone tissue and to observe changes in the ability of proliferation, apoptosis, and invasion of MG-63 cells following the reduced expression of HMGB1 via the small interference RNA. The relationship between HMGB1 and the carcinogenesis, progression, invasion, and metastasis of OS was investigated in order to provide scientific information for prognosis prediction and a new therapeutic target for the treatment of OS in the future.
Materials and methods
Specimens
Fresh OS tissues were collected from 26 patients with OS performed resection operation in our hospital between May 2011 and April 2014. Without any preoperative treatment, all 26 cases were pathologically diagnosed with OS postoperatively. In addition, 20 normal bone tissues were collected as well. These tissue samples were frozen by immersion in liquid nitrogen immediately for subsequent analysis.
Reagents
RT-PCR kit was from Gibco-BRL (Gaithersburg, MD). MG-63 OS cell line was from Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China). Trizol kit, trypsin, and fetal bovine serum were from Invitrogen (Carlsbad, CA). RIPA [1 % NP-40, 1 % deoxycholate, 0.1 % SDS, 500 mM Tris, 150 mM NaCl, 1 mM PMSF, and 1× protease inhibitor cocktail] was from Roche (New Jersey, USA). Polyvinylidene difluoride (PVDF) membranes were from Amresco Company (Solon, OH). Bradford protein concentration assay kit was from Bio-Rad (Richmond, CA). Annexin V-FITC, propidium iodide (PI), methyl thiazolyl tetrazolium (MTT), and dimethyl sulfoxide (DMSO) were from Beyotime (Haimen, China). Transwell invasion chambers and cell culture plates were from Corning Corp. (Midland, MI). Matrigel was from BD Biosciences (San Jose, CA). HMGB1 siRNA oligonucleotides targeting human HMGB1, control oligonucleotides (HMGB1 siRNA negative control), and transfection reagent were from RiboBio (Guangzhou, China). HMGB1 (GI:118918424) forward primer: 5′-ATATGGCAAAAGCGGACAAG-3′, reverse primer: 5′-GCAACATCACCAATGGACAG-3′; GAPDH (GI: 182976) forward primer: 5′- TCAGTGGTGGACCTGACCT-3′, reverse primer: 5′-TGAGCTTGACAAAGTGGTCG-3′. All primers were synthesized by Shanghai Generay Biotech Co., Ltd (Shanghai, China). Rabbit anti-HMGB1 polyclonal antibody was from Abgent (San Diego, CA). Mouse anti-cyclin D1 polyclonal antibody was from Abcam (Cambridge, UK). Mouse anti-caspase-3 monoclonal antibody was from Life Technologies (Carlsbad, CA). Rabbit anti-human MMP-9 antibody was from Cell Signaling Technology (Beverly, MA). Rabbit anti-β-actin polyclonal antibody and horseradish peroxidase (HRP)-conjugated goat anti-rabbit/mouse IgG polyclonal antibodies were from Abbiotec Corp. (San Diego, CA). ECL luminescence solution was purchased from Pierce Corp. (Rockford, IL). PDQuest software was from Bio-Rad (Richmond, CA).
Detection of HMGB1 mRNA by RT-PCR
Total RNA was extracted and quantified. The cDNA was synthesized with reverse transcription kit. The primers of HMGB1 and GAPDH were dissolved with ddH2O and stored at −20 °C in aliquots for later use. Then PCR reaction was performed in a 20-μl PCR reaction system under the following program: an initial denaturation step (3 min at 95 °C), 30 cycles of amplification (denaturing at 94 °C for 30 s, annealing at 56 °C for 30 s, and extension at 72 °C for 30 s), and the final extension step (10 min at 72 °C). The PCR products were detected by 1.5 % agarose gel electrophoresis.
MG-63 cells culture and HMGB1 siRNA transfection
MG-63 cells were cultured in minimum essential medium (MEM) containing 10 % fetal bovine serum (FBS), 2 mM L-glutamine, 0.1 mg/ml streptomycin, and 100 U/ml penicillin at 37 °C under a humidified atmosphere (95 % air, 5 % CO2). The recommended confluency for MG-63 cells at the day of transfection is 70–90 %, and then the complete medium was converted to serum-free MEM just before experiments. MG-63 cells were transfected with small interfering RNAs (siRNAs) targeting HMGB1 (50 nM) and negative control siRNAs (50 nM) according to the manufacturer’s instructions for Lipofectamine 2000. The protein expression levels of HMGB1 were determined to assess the effects of RNA interference.
Cell viability
MG-63 cell viability following HMGB1 knockdown was assessed by MTT test. MG-63 cells were cultured in 96-well plates. When the cell population reaches optimal densities, the medium was removed and was replaced with 100 μl of fresh culture medium. A 12 mM MTT stock solution was prepared by adding 1 ml of sterile PBS to one 5-mg vial of MTT. Ten microliter of the 12 mM MTT stock solution was added to each well. It was incubated for 4 h at 37 °C. The SDS-HCl solution was prepared by adding 10 ml of 0.01 M HCl to one tube containing 1 mg of SDS. Then 100 μl of the SDS-HCl solution was added to each well and was mixed thoroughly. The microplate was incubated at 37 °C for 4 h in a humidified chamber. Each sample was mixed again using a pipette and absorbance was read at 570 nm. With this procedure, only viable cells with functioning mitochondria can oxidize MTT to a violet-red reaction product.
Cell proliferation
MG-63 cell proliferation following HMGB1 knockdown was assessed by flow cytometry. MG-63 cells at 70–80 % confluency were cultured in serum-free medium for 24 h to synchronize and then cultured in complete medium for 24 h. Then MG-63 cells were trypsinized and harvested and were fixed into 0.5 ml 70 % EtOH precooled to −20 °C. The fixed MG-63 cells were stored on ice at least 1 h and for up to several days and were spin down at 4 °C for 2 min at 4,000 rpm. MG-63 cell pellet was resuspended in 0.5 ml of PBS buffer containing 0.25 % Triton X-100 and incubated on ice for 15 min. MG-63 cells were spin down for 2 min at 4,000 rpm. The supernatant was discarded, and the MG-63 cell pellet was resuspended in 0.5 ml of PBS buffer containing 10 μg/ml RNase A and 20 μg/ml propidium iodide. Then the MG-63 cells were transfered to fluorescence-activated cell sorting (FACS) tubes and incubated at room temperature for 30 min in the dark. For cell cycle analysis, data were expressed as fractions of cells in different cycle phases, and the experiment was repeated three times.
Detection of cell apoptosis by flow cytometry
MG-63 cell apoptosis following HMGB1 knockdown was assessed by flow cytometry. Apoptosis was assayed using the annexin V-fluorescein isothiocyanate (FITC) apoptosis detection kit (Haimen, China) according to the manufacturer’s instructions. Briefly, MG-63 cells were suspended and labeled with annexin V-FITC and propidium iodide for 15 min at 25 °C in the dark. Then MG-63 cells were observed on FACS and the data were analyzed by the CellQuest software. The cells that stained positive for annexin V were counted as apoptotic. Each experiment was repeated thrice.
Detection of MG-63 cells invasion
MG-63 cell invasion following HMGB1 knockdown was assessed by transwell invasion chamber assay. The transwell invasion chamber was divided into upper and lower chambers by the polycarbonate membrane (8-μm pore size) covered by Matrigel. 1 × 105 cells cultured in 200 μl of serum-free MEM medium were seeded in the upper chamber of the transwell invasion system, while 600 μl MEM medium with 10 % FBS was added into the lower chamber. Then the transwell invasion system was placed into a cell culture incubator. After 24 h of incubation, the transwell invasion system was taken out, and the cells on the upper surface of the polycarbonate membrane were removed with a sterile cotton swab. Plugged cells in 8-μm pores or cells attached to the under-surface of the membrane were stained with 1 % crystal violet and counted. Eight fields were randomly selected for observation. Invasion capability was evaluated through the number of plugged cells in 8-μm pores and cells attached to the under-surface of the membrane. The results are presented as the mean ± SD, with three repeated experiments for each group.
Western blot analysis
Total protein was extracted from MG-63 cells with radioimmunoprecipitation assay (RIPA) lysis buffer, measured, separated by SDS-PAGE, and transferred onto a PVDF membrane. The membrane was blocked in TBS solution containing 5 % skim milk at room temperature for 1 h, followed by incubation with rabbit anti-HMGB1 antibody, mouse anti-cyclin D1 antibody, mouse anti-caspase-3 antibody, rabbit anti-MMP-9 antibody, and rabbit anti-β-actin antibody at 4 °C overnight. The membranes were washed with TBST, followed by incubation with HRP-conjugated goat anti-rabbit IgG polyclonal antibody or HRP-conjugated goat anti-mouse IgG polyclonal antibody at 37 °C for 1 h. After TBST washing, autoradiography was conducted with ECL chemiluminescence reagents. The results were analyzed with the QuantityOne software. The relative expression of the target protein content was valuated with the gray value ratio of target and β-actin.
Statistical analysis
The SPSS17.0 software was used to analyze the related data with a χ 2 test (Chi-square test) or one-way ANOVA. The results were considered statistically significant if p < 0.05.
Results
The positive expression rate of HMGB1 mRNA was higher in OS tissues
The results from RT-PCR indicated that the positive expression rate of HMGB1 messenger RNA (mRNA) in the OS tissues was 80.77 %, which was significantly higher than that in normal bone tissues (15.00 %) (p < 0.01). The positive expression rate of HMGB1 mRNA in the OS tissues from patients with positive lung metastasis (100.00 %) was significantly higher than that from patients with negative lung metastasis (68.75 %) (p < 0.05). With increasing Enneking stages, the HMGB1 mRNA expression rate in the OS tissues also increased (p < 0.05). However, the positive expression rate of HMGB1 mRNA in OS was independent of the patient’s gender, age, or tumor size (p > 0.05), as shown in Table 1.
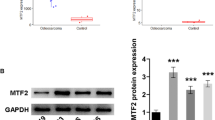
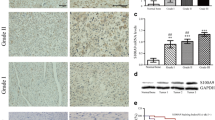
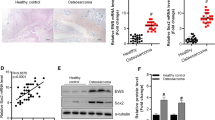
HMGB1 protein displayed upregulation in osteosarcoma tissues
The HMGB1 protein expression in osteosarcoma tissues was significantly higher than that in normal bone tissues (p < 0.05). The HMGB1 protein expression in the carcinoma tissues from patients with lung metastasis was significantly higher than that in the carcinoma tissues from patients with negative lung metastasis (p < 0.05). With increased Enneking staging, the HMGB1 protein expression in the osteosarcoma tissues was increased (p < 0.05) (Fig. 1).
I The HMGB1 protein expression in osteosarcoma tissues; II the relative expression of SASH1 protein in osteosarcoma tissues (A normal bone tissue, B osteosarcoma tissue, C Enneking stage I, D Enneking stage II, E Enneking stage III, F lung metastasis negative, G lung metastasis positive; asterisk indicates p < 0.05). These data were analyzed by t test or one-way ANOVA
HMGB1 improves MG-63 cell viability
To address the role of HMGB1 in osteosarcoma cells, MG-63 osteosarcoma cell line was chosen to study further. To suppress HMGB1 expression in MG-63 cells, MG-63 cells were transfected with HMGB1-siRNA, served as the knockdown group. MG-63 cells were transfected with control-siRNA or without any treatment, served as the control group or blank group, respectively. The results from qRT-PCR and western blot analysis demonstrated that HMGB1 displayed significant downregulation in knockdown group compared to control group or blank group (p < 0.05). The expression of HMGB1 did not display significant difference between control group and blank group. These data indicated that we effectively inhibit the expression of HMGB1 in MG-63 cells (Fig. 2).
MTT assay suggested that the viability of MG-63 cells in knockdown group was significantly lower compared to that in control group or blank group (p < 0.05), and that the viability of MG-63 cells did not display significant difference between control group and blank group (Fig. 3). These results indicated that the high level expression of HMGB1 might be related to the improvement of MG-63 cells viability, and that the knockdown of HMGB1 might be related to the suppression of MG-63 cells viability.
HMGB1 enhances MG-63 cell cycle progression
PI staining analysis by flow cytometry showed that the HMGB1-siRNA transfected knockdown group had significantly more MG-63 cells in G1 phase, significantly less MG-63 cells in S and G2 phase compared to the control group as well as the blank group (p < 0.05) (Fig. 4). These data suggested that the dysregulated HMGB1 affected the MG-63 cell cycle distribution and that HMGB1 knockdown induced G1 cell cycle arrest.
HMGB1 inhibits MG-63 cell apoptosis
The results from annexin V and PI staining demonstrated that HMGB1-siRNA transfection knockdown group had more apoptotic MG-63 cells compared to the control group as well as the blank group (p < 0.05); furthermore, the number of apoptotic cells did not differ significantly between the control group as well as the blank group (p > 0.05) (Fig. 5). These data indicated that HMGB1 knockdown could improve MG-63 cell apoptosis.
HMGB1 contributes to MG-63 cell invasion
The invasive ability of human OS cells MG-63 was evaluated through the transwell invasion chamber. The crystal violet staining results showed that the number of MG-63 cells invading the Matrigel membrane was significantly decreased in the HMGB1-siRNA transfected knockdown group compared to the control group as well as the blank group (p < 0.05). The number of MG-63 cells invading the Matrigel membrane displayed no significant difference between the control group and the blank group (p > 0.05) (Fig. 6). The results showed that the downregulation of HMGB1 protein was closely associated with the decreased invasive ability of MG-63 cells.
Effects of HMGB1 on cyclin D1, caspase-3, and MMP-9 expression
HMGB1 is an important regulatory factor for the expression of cyclin D1, caspase-3, and MMP-9 [20, 24, 27]. Cyclin D1, often amplified and overexpressed in malignancies, is a major regulator of the progression of cells into the proliferative stage of the cell cycle [28]. Caspase-3, the major effector caspase, is one of the key executioners of apoptosis [29]. MMP-9 plays important roles in tumor invasion and angiogenesis [30, 31]. So the protein expression of cyclin D1, caspase-3, and MMP-9 were detected in this work. The results showed that the expression of cyclin D1 and MMP-9 was significantly lower in MG-63 cells from HMGB1-siRNA transfected knockdown group than that from the control group and the blank group (p < 0.05). Caspase-3 displayed a significant upregulation in HMGB1-siRNA transfected knockdown group than that in the control group and the blank group (p < 0.05). The expression of cyclin D1, caspase-3, and MMP-9 did not show significant difference between the control group and the blank group (Fig. 7). These results indicated that HMGB1 knockdown could downregulate MMP-9 and cyclin D1 expression and upregulate caspase-3 expression in OS cells.
Discussion
Osteosarcoma (OS), a malignant tumor mainly occurring in children and adolescents, is characterized by its strong invasion and early metastasis, which are the major causes of both treatment failure and death [32]. Therefore, it is necessary to explore the molecular mechanisms of OS invasion and metastasis and the effective treatments for OS. HMGB1 is a widely existing DNA-binding nuclear protein, which can be actively secreted by nature immunocyte cells and passively released by injured cells or necrosis cells as a DAMP [17], exerting pleiotropic biological effects by binding to RAGE and TLR, thereby promoting angiogenesis, unlimited replicative potential, tissue invasion, and metastasis [18, 19]. Many studies suggest that HMGB1 overexpression plays a critical role in the development and progression of multiple malignancies [20], but no information is available to date regarding the function of HMGB1 in OS.
In this study, we found that the expression rate of HMGB1 mRNA in the OS tissues was significantly higher than that in normal bone tissue, the expression rate of HMGB1 mRNA in the carcinoma tissues with positive lung metastasis was significantly higher than that without lung metastasis, and with increasing Enneking stage, the expression rate of HMGB1 mRNA also increased. These data showed that HMGB1 overexpression might be correlated with the development and progression of OS. These observations also support previous conclusions that increased expression HMGB1 is associated with increased cell proliferation, angiogenesis, and metastasis during cancer progression [15, 33].
In order to explore the effect of HMGB1 on osteosarcoma cells, MG-63 osteosarcoma cell line was chosen to study further. MTT assay suggested that the viability of MG-63 cells in HMGB1 knockdown group was significantly lower compared to that in control group or blank group, which indicated that the knockdown of HMGB1 might be related to the suppression of MG-63 cells viability. Flow cytometry analysis suggested that HMGB1 knockdown induced G1 cell cycle arrest. These observations were consistent with other findings that downregulating HMGB1 could inhibit the proliferation of human lung cancer cell [34]. The ability of proliferation of HCC cells was strengthened when the expression endogenous HMGB1 was enhanced using HMGB1 DNA [15]. Knockdown of HMGB1 suppresses growth of GAC cells through the NF-κB pathway, suggesting that HMGB1 may serve as a potential therapeutic target for GAC. The ectopic expression of HMGB1 activates cell growth by decreasing Bax and p53 expression while enhancing Bcl-2, cyclin D1 [24]. In this study, cyclin D1, a critical cell cycle regulatory factor, was chosen to determine by Western blotting. The results showed that the expression of cyclin D1 was significantly lower in MG-63 cells from HMGB1 knockdown group. Cell cycle changes were accompanied by decreases in cyclin D1 expression. These might be responsible for the explanation that HMGB1 knockdown inhibited cell proliferation.
The apoptosis assay results from this study demonstrated that HMGB1 knockdown could improve MG-63 cell apoptosis. Suppression of HMGB1 expression, tumor cell apoptosis, and chemotherapeutic drug sensitivity were increased. Yin et al. found that HMGB1 promoted autophagy and inhibited anticancer drug-induced apoptosis [35]. HMGB1 knockdown sensitized cells to apoptosis that was mediated by the caspase-3 pathway [36]. Downregulation of HMGB1 expression resulted in the decreased cell number was due to transfected prostate cancer cells undergoing apoptosis via caspase-3-dependent pathways [37]. HMGB1 overexpression could inhibit ADM-induced apoptosis in leukemia K562 cells through regulating the protein level of Bcl-2 and the activities of caspase-3 [38]. Therefore, the expression of caspase-3 was determined, and the results demonstrated that caspase-3 displayed a significant upregulation in HMGB1 knockdown group than that in the control group and the blank group, which might be responsible for that HMGB1 knockdown group had more apoptotic MG-63 cells compared to the control group as well as the blank group.
The results of transwell invasive chamber experiment indicated that the invasion ability of MG-63 cells was significantly decreased in the HMGB1 knockdown group compared to the control group and the blank group. These findings suggested that HMGB1 might have stimulative effect on the migration of human OS cell MG-63, which was consistent with other findings. HMGB1 overexpression has a significant role in tumor progression, especially the migration of tumor cells [39]. HMGB1 activates RAGE signaling pathways and induces NF-кB activation to promote cellular invasion, and metastasis, in HCC cell lines [40]. Knockdown of HMGB1 suppresses invasion of GAC cells [20]. Silencing of HMGB1 significantly reduced cellular metastatic ability and MMP-9 expression in MGC-803 cells [36]. Silencing of HMGB1 can effectively inhibit the invasion and migration of gastric cancer cells, and this effect of HMGB1 may be partly due to its regulation of NF-κB and MMP-9 expressions [41]. Therefore, the expression of MMP-9 was determined, and the results demonstrated that MMP-9 displayed a significant downregulation in HMGB1 knockdown group, which might be responsible for that knockdown of HMGB1 resulted in the decreased invasive ability of MG-63 cells.
These results may suggest that the overexpression of HMGB1 may play an important role in the carcinogenesis, development, invasion, and metastasis of OS. HMGB1 knockdown suppresses proliferation and invasion and improve apoptosis of OS. From these studies, a deeper understanding of the incidence and progression of OS could be developed and might provide new strategies and targets for the treatment of OS.
References
Li R, Liu J, Wu H, Liu L, Wang L, Zhang S. TIKI2 suppresses growth of osteosarcoma by targeting Wnt/β-catenin pathway. Mol Cell Biochem. 2014;392:109–16.
PosthumaDeBoer J, Witlox MA, Kaspers GJ, van Royen BJ. Molecular alterations as target for therapy in metastatic osteosarcoma: a review of literature. Clin Exp Metastasis. 2011;28:493–503.
Bielack SS, Carrle D, Hardes J, Schuck A, Paulussen M. Bone tumors in adolescents and young adults. Curr Treat Options Oncol. 2008;9:67–80.
Mirabello L, Troisi RJ, Savage SA. International osteosarcoma incidence patterns in children and adolescents, middle ages and elderly persons. Int J Cancer. 2009;125:229–34.
Mirabello L, Troisi RJ, Savage SA. Osteosarcoma incidence and survival rates from 1973 to 2004: data from the surveillance, epidemiology, and end results program. Cancer. 2009;115:1531–43.
He H, Ni J, Huang J. Molecular mechanisms of chemoresistance in osteosarcoma (review). Oncol Lett. 2014;7:1352–62.
Chiappetta C, Leopizzi M, Censi F, Puggioni C, Petrozza V, Rocca CD, et al. Correlation of the Rac1/Rhoa pathway with ezrin expression in osteosarcoma. Appl Immunohistochem Mol Morphol. 2014;22:162–70.
Hou CH, Lin FL, Tong KB, Hou SM, Liu JF. Transforming growth factor alpha promotes osteosarcoma metastasis by ICAM-1 and PI3K/Akt signaling pathway. Biochem Pharmacol. 2014;89:453–63.
Shen A, Zhang Y, Yang H, Xu R, Huang G. Overexpression of ZEB1 relates to metastasis and invasion in osteosarcoma. J Surg Oncol. 2012;105:830–4.
Tumbar T, Guasch G, Greco V, Blanpain C, Lowry WE, Rendl M, et al. Defining the epithelial stem cell niche in skin. Science. 2004;303:359–63.
Spradling A, Drummond-Barbosa D, Kai T. Stem cells find their niche. Nature. 2001;414:98–104.
Hu YH, Yang L, Zhang CG. HMGB1-a as potential target for therapy of hematological malignancies. Zhongguo shi yan xue ye xue za zhi. 2014;22:560–4.
Srinivasan M, Banerjee S, Palmer A, Zheng G, Chen A, Bosland MC, et al. HMGB1 in hormone-related cancer: a potential therapeutic target. Horm Cancer. 2014;5:127–39.
Yildirim M, Suren D, Demirpence O, Kaya V, Alikanoglu AS, Bulbuller N, et al. The role of high mobility group box 1 (HMGB1) in colorectal cancer. Med Sci Monit. 2014;20:530–7.
Xiao J, Ding Y, Huang J, Li Q, Liu Y, Ni W, et al. The association of HMGB1 gene with the prognosis of HCC. PLoS ONE. 2014;9:e89097.
Liu K, Huang J, Xie M, Yu Y, Zhu S, Kang R, et al. MIR34a regulates autophagy and apoptosis by targeting HMGB1 in the retinoblastoma cell. Autophagy. 2014;10:442–52.
Jia L, Clear A, Liu FT, Matthews J, Uddin N, McCarthy A, et al. Extracellular HMGB1 promotes differentiation of nurse-like cells in chronic lymphocytic leukemia. Blood. 2014;123:1709–19.
Weng H, Deng Y, Xie Y, Liu H, Gong F. Expression and significance of HMGB1, TLR4 and NF-κB p65 in human epidermal tumors. BMC Cancer. 2013;13:311.
Moser B, Janik S, Schiefer AI, Mullauer L, Bekos C, Scharrer A, et al. Expression of rage and HMGB1 in thymic epithelial tumors, thymic hyperplasia and regular thymic morphology. PLoS ONE. 2014;9:e94118.
Zhang J, Kou YB, Zhu JS, Chen WX, Li S. Knockdown of HMGB1 inhibits growth and invasion of gastric cancer cells through the NF-κB pathway in vitro and in vivo. Int J Oncol. 2014;44:1268–76.
Srinivasan M, Banerjee S, Palmer A, Zheng G, Chen A, Bosland MC, et al. HMGB1 in hormone-related cancer: a potential therapeutic target. Horm Cancer. 2014;5(3):127–39.
Dong YD, Cui L, Peng CH, Cheng DF, Han BS, Huang F. Expression and clinical significance of HMGB1 in human liver cancer: knockdown inhibits tumor growth and metastasis in vitro and in vivo. Oncol Rep. 2013;29:87–94.
Liu Y, Xie CL, Qiu YZ, Tian YQ, Zhang X, Huang DH, et al. Expression of HMGB1 protein in laryngeal squamous cell carcinoma and its clinical significance. Zhonghua zhong liu za zhi. 2012;34:132–6.
Ko YB, Kim BR, Nam SL, Yang JB, Park SY, Rho SB. High-mobility group box 1 (HMGB1) protein regulates tumor-associated cell migration through the interaction with BTB domain. Cell Signal. 2014;26:777–83.
Chen J, Xi B, Zhao Y, Yu Y, Zhang J, Wang C. High-mobility group protein B1 (HMGB1) is a novel biomarker for human ovarian cancer. Gynecol Oncol. 2012;126:109–17.
Chen J, Liu X, Zhang J, Zhao Y. Targeting HMGB1 inhibits ovarian cancer growth and metastasis by lentivirus-mediated RNA interference. J Cell Physiol. 2012;227:3629–38.
Yu Y, Xie M, Kang R, Livesey KM, Cao L, Tang D. HMGB1 is a therapeutic target for leukemia. Am J Blood Res. 2012;2:36–43.
Monga SP. Role and regulation of β-catenin signaling during physiological liver growth. Gene Expr. 2014;16:51–62.
Mazumder S, Plesca D, Almasan A. Caspase-3 activation is a critical determinant of genotoxic stress-induced apoptosis. Methods Mol Biol. 2008;414:13–21.
Xu YB, Du QH, Zhang MY, Yun P, He CY. Propofol suppresses proliferation, invasion and angiogenesis by down-regulating ERK-VEGF/MMP-9 signaling in Eca-109 esophageal squamous cell carcinoma cells. Eur Rev Med Pharmacol Sci. 2013;17:2486–94.
Cock-Rada AM, Medjkane S, Janski N, Yousfi N, Perichon M, Chaussepied M, et al. SMYD3 promotes cancer invasion by epigenetic upregulation of the metalloproteinase MMP-9. Cancer Res. 2012;72:810–20.
Zwaga T, Bovee JV, Kroon HM. Osteosarcoma of the femur with skip, lymph node, and lung metastases. Radiographics. 2008;28:277–83.
Gatla HR, Singha B, Persaud V, Vancurova I. Evaluating cytoplasmic and nuclear levels of inflammatory cytokines in cancer cells by Western blotting. Methods Mol Biol. 2014;1172:271–83.
Zhang P, Liu Y, Wu ZH, Chen J, Chen G, Zhou QH. Effects of high mobility group box-1 silencing upon the invasion and proliferation in human lung cancer cell L9981 by RNA inhibition. Zhonghua yi xue za zhi. 2009;89:3156–9.
Yin Y, Li W, Deng M, Zhang P, Shen Q, Wang G, et al. Extracellular high mobility group box chromosomal protein 1 promotes drug resistance by increasing the expression of P-glycoprotein expression in gastric adenocarcinoma cells. Mol Med Rep. 2014;9:1439–43.
Song B, Song WG, Li ZJ, Xu ZF, Wang XW, Wang CX, et al. Effect of HMGB1 silencing on cell proliferation, invasion and apoptosis of MGC-803 gastric cancer cells. Cell Biochem Funct. 2011. doi: 10.1002/cbf.1811.
Gnanasekar M, Thirugnanam S, Ramaswamy K. Short hairpin RNA (shRNA) constructs targeting high mobility group box-1 (HMBG1) expression leads to inhibition of prostate cancer cell survival and apoptosis. Int J Oncol. 2009;34:425–31.
Yu Y, Xie M, He YL, Xu WQ, Zhu S, Cao LZ. Role of high mobility group box 1 in adriamycin-induced apoptosis in leukemia K562 cells. Ai zheng. 2008;27:929–33.
Suren D, Yildirim M, Demirpence O, Kaya V, Alikanoglu AS, Bulbuller N, et al. The role of high mobility group box 1 (HMGB1) in colorectal cancer. Med Sci Monit. 2014;20:530–7.
Chen RC, Yi PP, Zhou RR, Xiao MF, Huang ZB, Tang DL, et al. The role of HMGB1-rage axis in migration and invasion of hepatocellular carcinoma cell lines. Mol Cell Biochem. 2014;390:271–80.
Li ZJ, Song B, Liu J, Han JJ, Wang CX, Zhu YX, et al. Inhibitory effect of silencing of HMGB1 gene expression on the invasive and metastatic abilities of MGC-803 gastric cancer cells. Zhonghua zhong liu za zhi. 2013;35:244–8.
Acknowledgments
This study was supported by National Natural Science Foundation of China (No. 81402226).
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Qingbing Meng and Jie Zhao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Meng, Q., Zhao, J., Liu, H. et al. HMGB1 promotes cellular proliferation and invasion, suppresses cellular apoptosis in osteosarcoma. Tumor Biol. 35, 12265–12274 (2014). https://doi.org/10.1007/s13277-014-2535-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2535-3