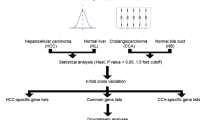
Abstract
Liver carcinomas have been classified into three types: hepatocellular carcinoma (HCC), cholangiocarcinoma (CC), and combined HCC-CC (CHC). We aim to find the common and different characteristic of these three types of liver cancer. The gene expression profiling of HCC, CC, and CHC were compared with each other, and enrichment pathways and processes in these three liver cancers were also identified. Using GSE15765 datasets downloaded from NCBI GEO database, the gene expression profiling of HCC, CC, and CHC were compared with each other (HCC compared with CC, HCC compared with CHC, and CC compared with HCC). Then, the differentially expressed genes (DEGs) were identified in these three groups respectively, and three PPI networks were constructed for DEGs in each group. Subsequently, the clusters in these networks were identified and further analyzed by ClusterONE and MCODE. Finally, gene set enrichment analysis enrichment analysis was performed to illustrate altered pathways and processes for each type of liver cancer. A total of 112, 530, and 64 DEGs were identified in three groups, respectively, and three PPI networks were constructed respectively for the corresponding group. Through the cluster analysis, we found some new differential marker genes for distinguishing the difference between these three types of liver cancer. We also indicated that we can distinguish HCC with CC through altered pathways and processes. Our findings develop new biomarkers for categorizing the primary liver cancer and may improve patient prognosis of these cancers. However, further validation is required since our results were based on microarray data derived from a small sample size.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver cancer is one of the most frequent cancer diseases, which accounts for the third most common cause of cancer-related deaths worldwide, especially in parts of Asia and Africa [1]. According to histology, the primary liver cancer usually can be categorized into: hepatocellular carcinoma (HCC), cholangiocarcinoma (CC), and combined HCC-CC (CHC) [2].
HCC is the most common type of liver cancer. Known risk factors for HCC include cirrhosis and infection caused by viral hepatitis account [3]. HCC often develops from the hepatocytes, which composed the main tissue of the liver and making up 70–80 % of the liver’s cytoplasmic mass. This type of liver cancer normally can be characterized by jaundice, bloating from ascites, and abdominal pain, especially in the upper right part [4].
CC is a type of cancer starts in the section of the bile ducts outside the liver. The bile ducts can drain bile from the liver, which helps digest fats in food [5]. In most of the case, CC is secondary to primary sclerosing cholangitis, infection with the parasites, and congenital liver malformations. CC may also be characterized by jaundice and abdominal pain. Sometimes, color of stool and urine may also change [6].
CHC is a rare form of liver cancer, which has the features of both HCC and CC. However, the tumors of CHC have its own characteristic. In most of patients, CHC shows an intermediate features between HCC and CC [7]. Moreover, the symptoms of CHC and CC are similar. Therefore, distinguishing the difference between HCC, CC, and CHC has become a difficult problem. To solve this problem, many biomarkers have been established by immunohistochemical and histological method. But research on this issue through bioinformatics way remains rare.
In this study, in order to find the common and different characteristic of these three types of liver cancer, gene expression profiling of HCC, CC, and CHC were compared with each other, and gene set enrichment analysis (GSEA) analysis was performed to identify altered pathways or processes for each type of liver cancer.
Materials and methods
Data source
Affymetrix microarray data
The transcription profile of GSE15765 [8] was obtained from NCBI GEO database (http://www.ncbi.nlm.nih.gov/geo/) which is based on Affymetrix Human Genome U133 Plus 2.0 Array. All biochips used for analysis were purchased from National Cancer Institute in Neurological Disorders and Stroke, Bethesda.
In this study, we collected gene expression profiling of HCC, CC, and CHC. A total of 90 biochips were analyzed, including 70 biochips for HCC, 13 biochips for CC, and 7 biochips for CHC.
Protein–protein interaction data
The Human Protein Reference Database (HPRD) [9] is a protein database accessible through the internet. The Biological General Repository for Interaction Datasets (BioGRID) [10] is a curated biological database of protein–protein and genetic interactions.
In this study, the protein–protein interaction (PPI) data were collected from the HPRD and BioGRID database. Total of 326,119 unique PPI pairs were collected, among which 39,240 pairs are from HPRD and 379,426 pairs are from BioGRID.
Methods
Differentially expressed genes analysis
The limma package in R language [11] was applied to identify differentially expressed genes (DEGs). The gene expression profiling of HCC, CC, and CHC were compared with each other (HCC compared with CC, HCC compared with CHC, and CC compared with CHC). We defined p value < 0.05 and FDR >0.68 to be statistically significant to identify the differentially expressed genes.
Protein–protein interaction network construction
The PPI data were collected from the HPRD and BioGRID database. The Pearson correlation coefficient (PCC) [12] was performed to identify the significant interaction relationships. Finally, based on the significant relationships (PCC > 0.75 or PCC < −0.75), the PPI network for these DEGs was constructed by using Cytoscape [13]. The PPI network contained both the products of these DEGs and other proteins which interact with them.
Cluster identification
ClusterONE [14] strives to discover densely connected and possibly overlapping regions within the Cytoscape network you are working with. In protein–protein interaction networks, these dense regions usually correspond to protein complexes or fractions of them. ClusterONE works by “growing” dense regions out of small seeds (typically one or two vertices), driven by a quality function called cohesiveness. The parameters are grouped into basic and advanced ones. In most of the cases, the default values of the advanced parameters should be fine, but the basic parameters may need to be adjusted to your specific needs.
In this study, clusters were identified by using the ClusterONE in Cytoscape [15] (http://www.cs.rhul.ac.uk/home/tamas/assets/files/cl1/cl1-cytoscape-0.1.html).
Cytoscape MCODE analysis
MCODE [15] finds clusters (highly interconnected regions) in a network. Clusters mean different things in different types of networks. For instance, clusters in a protein–protein interaction network are often protein complexes and parts of pathways, while clusters in a protein similarity network represent protein families. The parameters are grouped into cluster and advanced ones. In most of the cases, the default values of the cluster and advanced parameters should be fine, but sometimes the cluster and advanced parameters may need to be adjusted to your specific needs.
GSEA enrichment analysis
GSEA [16, 17] is a statistical method (Kolmogorov–Smirnov statistic method) to determine if predefined sets of genes are differentially expressed in different classes. Predefined gene sets may be genes in a known metabolic pathway, located in the same cytogenetic band, sharing the same Gene Ontology category, or any user-defined set.
Here, we set the parameter as weighted enrichment statistic and meandiv normalization mode.
Result
Microarray data analysis
Publicly available microarray dataset GSE15765 were obtained from GEO which is based on Affymetrix Human Genome U133 Plus 2.0 Array. In our criterion (FDR >0.68 and p value <0.05), 112, 530, and 64 DEGs were identified in each groups (Table 1).
Co-expression PPI network
According to the HPRD and BioGRID database, three co-expression PPI networks with minimum size 4 were constructed for the products of DEGs in each group (Fig. 1). Shown in Fig. 1a–c are the PPI networks for HCC-CHC group, HCC-CC group, and CC-CHC group, respectively.
Co-expression PPI network. The nodes represent the products of genes exist in the network. The yellow nodes represent products of DEGs and the pink nodes represent products of genes which have same expression pattern. a The PPI co-expression network of (HCC) and (CHC).b The PPI co-expression network of (HCC) and (CC). c The PPI co-expression network of (CC) and (CHC)
Network clustering
To further analyze the structure of the PPI network, we used ClusterONE to identify the clusters in our PPI network. Our basic parameters were set as that minimum size is 6, minimum density is 0.3, and p value <0.01. Finally, one cluster was found in each group (Fig. 2). The clusters a, b, and c in Fig. 2 are corresponding to the PPI networks a, b, and c in Fig. 1, respectively. Four genes, including FGA, FGB, FGG, and SPERINA5 were most significantly overexpressed and predicted to be highly interconnected by MCODE. What is more, their products were common in clusters b and c (Fig. 3).
GSEA enrichment analysis
In order to compare with HCC, CC, and CHC in the point view of biology process and pathway, we performed GSEA to illuminate dysregulated pathways or processes for HCC, CC, and CHC, respectively. Tables 2 and 3 represent significant enrichment biology processes in CC and HCC (|NES| > 1.5, p value <0.01, FDR <0.25). No significant enrichment biology process was found in CHC.
We also analyzed significant enrichment biology pathways for each group. Table 4 represents significant enrichment biology pathways in HCC (|NES| > 1.5, p value <0.01, FDR <0.25), and no significant enrichment biology process was found in CC and CHC.
Discussion
Our studies revealed that common and different characteristic of these three types of liver cancer. Besides, we also identify some important molecular biomarkers of these three types of liver cancer, including FGA, FGB, FGG, and SPERINA5. According to the HPRD and BioGRID database, three co-expression PPI networks with minimum size 4 were constructed for the products of DEGs in each group of liver cancer. To further analyze the structure of the PPI network, we used ClusterONE to identify the clusters in our PPI network.
We have identified three clusters in our network. Most of the proteins in these three clusters were involved in blood coagulation process, including coagulation factor family, serpins and fibrinogen. And some of the proteins have been proven to be related to HCC and CC.
For example, coagulation factor can be secreted by the liver, and it can form a complex with tissue factor to play an important role in the development of both HCC and CC [18, 19]. The coagulation factor can activate cancer cell migration and invasion [20].
Serpins are a group of protein inhibitors which can inhibit activation of proteases. Serpins are also involved in the process of blood coagulation. In addition, abnormal serpins in liver cells may lead to liver disease, ranging from cirrhosis to HCC and CC [21]. The expression of SERPINC1 is reported to be altered in liver cancer cell [22].
It indicates that both serpins and coagulation factor family were participated in the development of HCC and CC. But according to our finding, the expression level of SERPINC1, F9 (coagulation factor family IX), and F2 (coagulation factor family II) were significantly different between HC and CC. So, we can regard that these three genes as differential marker in diagnosis for HCH and CC.
Combined HCC and CC (CHC) forms a small but significant proportion of primary liver carcinomas. However, its diagnostic features are not well established. Here, we reported that FGA and FGG may be become a potential differential marker in diagnosis. According to our results, FGA and FGG were identified as DEGs in both group B (CHC compared with HCC) and group C (CHC compared with CC). Moreover FGA and FGG are proved to be related to the development of both CC and HCC by experimental verification [23–25]. Together with the previously studies, we are likely to indicate that FGA and FGG may be regarded as a new markers to distinguish CHC with CC and HCH.
Furthermore, we compared HCC, CC, and CHC in the point of view of biology process and pathway. The processes or pathways enriched in HCC were mainly involved in metabolism, including metabolism of fat and amino acid. It is expected that many literatures report that metabolism is mis-regulated in patients with HCC [26, 27]. Compared with HCC, positive regulation of binding and DNA binding were enriched in CC, and no significant enrichment pathways or processes were found in CHC. However, we were firstly reported that we can distinguish the difference between the CC and HCC through altered pathways or processes.
Conclusion
In this study, we compared the expression profiling of three types of liver cancer, and then identified the enrichment pathways and processes for them. Finally, we found that most blood coagulation process was related to HCC and CC. Moreover, SERPINC1, F9, and F2 can be regarded as differential markers in diagnosis to distinguish the difference between the HCH and CC, and FGA and FGG can be used as differential markers for comparing CHC with CC and HCC. What is more, we also indicated that we can distinguish HCC with CC through altered pathways and processes. Our findings develop new biomarkers for categorizing the primary liver cancer and may improve patient prognosis of these cancers. However, further validation is required since our results were based on microarray data derived from a small sample size.
Change history
26 June 2020
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s13277-015-3695-5
Reference
Yi X, Luk JM, Lee NP, Peng J, Leng X, Guan XY, et al. Association of mortalin (HSPA9) with liver cancer metastasis and prediction for early tumor recurrence. Mol Cell Proteomics. 2008;7(2):315–25.
Goodman ZD, Ishak KG, Langloss JM, Sesterhenn IA, Rabin L. Combined hepatocellular–cholangiocarcinoma. A histologic and immunohistochemical study. Cancer. 2006;55(1):124–35.
Kojiro M. Pathology of hepatocellular carcinoma. Mol Genet Liver Neoplasia. 2011;2011:37–48.
Ohishi W, Fujiwara S, Cologne JB, Suzuki G, Akahoshi M, Nishi N, et al. Impact of radiation and hepatitis virus infection on risk of hepatocellular carcinoma. Hepatology. 2011;53(4):1237–45.
Lau WY, Lai ECH, Lau SHY. Management of bile duct injury after laparoscopic cholecystectomy: a review. ANZ J Surg. 2010;80(1–2):75–81.
Lazaridis KN, Gores GJ. Cholangiocarcinoma. In: Hawley CJ, Bosch J, Richter JE, Garcia-Tsao G, Chan FKL, editors. Textbook of clinical gastroenterology and hepatology. 2nd ed. New York: Wiley; 2012. p. 580–5.
Lo RCL, Ng IOL. Hepatocellular tumors: immunohistochemical analyses for classification and prognostication. Chin J Cancer Res. 2011;23(4):245–53.
Woo HG, Park ES, Cheon JH, Kim JH, Lee JS, Park BJ, et al. Gene expression-based recurrence prediction of hepatitis B virus-related human hepatocellular carcinoma. Clin Cancer Res. 2008;14(7):2056–64.
Keshava Prasad TS, Goel R, Kandasamy K, Keerthikumar S, Kumar S, Mathivanan S, et al. Human Protein Reference Database-2009 update. Nucleic Acids Res. 2009;37:D767–72. doi:10.1093/nar/gkn892 (Database issue).
Stark C, Breitkreutz BJ, Chatr-Aryamontri A, Boucher L, Oughtred R, Livstone MS, et al. The BioGRID Interaction Database. Nucleic Acids Res. 2011;39:D698–704. doi:10.1093/nar/gkq1116 (Database issue).
Smyth GK. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3. doi:10.2202/1544-6115.1027. Article3.
Derrick TR, Bates BT, Dufek JS. Evaluation of time-series data sets using the Pearson product–moment correlation coefficient. Med Sci Sports Exerc. 1994;26(7):919–28.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504.
Bader GD, Hogue CWV. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics. 2003;4:2.
Nepusz T, Yu H, Paccanaro A. Detecting overlapping protein complexes in protein–protein interaction networks. Nat Methods. 2012;9(5):471–2.
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–73.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545.
Bieker R, Kessler T, Schwöppe C, Padró T, Persigehl T, Bremer C, et al. Infarction of tumor vessels by NGR-peptide-directed targeting of tissue factor: experimental results and first-in-man experience. Blood. 2009;113(20):5019–27.
Papatheodoridis GV, Chung S, Keshav S, Pasi J, Burroughs AK. Correction of both prothrombin time and primary haemostasis by recombinant factor VII during therapeutic alcohol injection of hepatocellular cancer in liver cirrhosis. J Hepatol. 1999;31(4):747–50.
Koizume S, Jin MS, Miyagi E, Hirahara F, Nakamura Y, Piao JH, et al. Activation of cancer cell migration and invasion by ectopic synthesis of coagulation factor VII. Cancer Res. 2006;66(19):9453–60.
Van Steenbergen W. Alpha 1-antitrypsin deficiency: an overview. Acta Clin Belg. 1993;48(3):171.
Cheng P, Cheng Y, Su MX, Li D, et al. Bicluster and pathway enrichment analysis of HCV-induced cirrhosis and hepatocellular carcinoma. Asi P J Cancer Prevent. 2012;13:3741–5.
Seimiya M, Tomonaga T, Matsushita K, Sunaga M, Oh-ishi M, Kodera Y, et al. Identification of novel immunohistochemical tumor markers for primary hepatocellular carcinoma; clathrin heavy chain and formiminotransferase cyclodeaminase. Hepatology. 2008;48(2):519–30.
Chaerkady R, Harsha H, Nalli A, Gucek M, Vivekanandan P, Akhtar J, et al. A quantitative proteomic approach for identification of potential biomarkers in hepatocellular carcinoma. J Proteome Res. 2008;7(10):4289–98.
Comunale MA, Lowman M, Ronald E, Krakover J, Philip R, Seeholzer S, et al. Proteomic analysis of serum associated fucosylated glycoproteins in the development of primary hepatocellular carcinoma. J Proteome Res. 2006;5(2):308–15.
Ockner RK, Kaikaus RM, Bass NM. Fatty-acid metabolism and the pathogenesis of hepatocellular carcinoma: review and hypothesis. Hepatology. 2005;18(3):669–76.
Fan ST, Lo CM, Lai E, Chu KM, Liu CL, Wong J. Perioperative nutritional support in patients undergoing hepatectomy for hepatocellular carcinoma. New Engl J Med. 1994;331(23):1547–52.
Additional information
The Publisher and Editor retract this article in accordance with the recommendations of the Committee on Publication Ethics (COPE). After a thorough investigation we have strong reason to believe that the peer review process was compromised.
About this article
Cite this article
Wang, L. RETRACTED ARTICLE: Comparison of hepatocellular carcinoma (HCC), cholangiocarcinoma (CC), and combined HCC-CC (CHC) with each other based on microarray dataset. Tumor Biol. 34, 1679–1684 (2013). https://doi.org/10.1007/s13277-013-0702-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-013-0702-6