Abstract
Arsenic trioxide has shown remarkable biological activity against bladder cancer in some clinical studies. However, the mechanism of its action is unknown. Our aim was to find the relationship between miRNAs and arsenic trioxide treatment by using T24 human bladder carcinoma cells. By performing microRNA microarray and quantitative real-time PCR after ATO treatment, we found that expression levels of several miRNAs, in particular, miRNA-19a, were significantly decreased in T24 cell line. Furthermore, cell proliferation assay, flow cytometry analysis, prediction of miRNA targets, Western blot analysis, and luciferase reporter assay were performed to determine the role of mir-19a in affecting the biological behaviors of T24 cells. Several miRNAs were up-regulated or down-regulated in T24 cells treated with arsenic trioxide compared to their controls. If only changes above two folds were considered, two miRNAs were identified, miRNA-19a was down-regulated, while miRNA-222* was up-regulated. Among them, knockdown of miRNA-19a by anti-miRNA-19a transfection showed a positive therapeutic effect in bladder cancer cells by inhibiting cell growth and inducing cell apoptosis targeting PTEN through the PTEN/Akt pathway. Besides this, a synergy effect was detected between knockdown of miRNA-19a and arsenic trioxide. Arsenic trioxide altered miRNA expression profile in T24 cells. It seems miRNA-19a plays a critical role in the mechanism of arsenic trioxide treatment in bladder cancer. The synergy effect between miRNA-19a and arsenic trioxide that advocates targeting the mir-19a may represent a potential approach to enhance the efficacy and safety of ATO to treat bladder cancer by a decrease in dose.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arsenic trioxide (ATO) has been used primarily in the treatment as a therapeutic agent for acute promyelocytic leukemia (APL) [1]. ATO was approved to treat relapsed APL by the US Food and Drug Administration (FDA) in 2000 [2]. It has been extensively investigated in neoplastic hematologic disorder and solid tumor such as prostate, kidney, cervix, bladder, neuroblastoma, glioma, and gastric cancer [3–6], owing to its various cytological effects including inhibition of angiogenesis, promotion of apoptosis, and growth inhibition. Proposed mechanisms for the apoptotic effects include induction of apoptosis through caspases activation, down-regulation of BCL-2, reactive oxygen species (ROS) [7–9], and interactions with multiple signaling pathways, including MAPK, P38, and AKT [10–13]. Although apoptosis or growth inhibition induced by ATO was reported in several solid tumor cell lines, it was not accepted clinically because of the reported chronic toxicities, carcinogenicity, and other side effects associated with the unachievable dose. Therefore, the molecular mechanisms of apoptosis induced by ATO were important to be clarified in addition with the enhancement and replacement ATO.
MicroRNAs (miRNA) are small nucleotide noncoding RNA molecules that function to control gene expression by binding with imperfect complementarity to the 3′ untranslated region (3′UTR) of the target mRNA. In recent studies, more and more miRNAs have shown their potential of being oncogene or anti-oncogene, which emerge in cell death, proliferation, tumorigenesis, differentiation, etc. [14–17] Several studies suggest that miRNAs may influence the chemotherapy effect of some cancer cells [18, 19].
In this study, we have shown the alteration of miRNA expression profile in bladder cancer cells after ATO treatment using miRNA microarrays. It suggests that specific miRNAs expression altered in bladder cancer cells after ATO treatment. The miRNA that most dramatically down-regulated was miR-19a, which might play a decisive role. Furthermore, we have showed the target genes of miR-19a might be involved in the mechanisms of ATO therapy.
Materials and methods
Cell lines
The high-grade T24 human bladder carcinoma cells line was purchased from ATCC (Manassas, VA, USA) and was cultured following the manufacturer’s instructions. The cells were grown in McCoy’s 5A supplemented with 10% fetal bovine serum. McCoy’s 5A and FCS were purchased from Hyclone (Logan, UT, USA).
RNA isolation
Total RNAs were isolated using Trizol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions for miRNA microarray or real-time PCR analysis.
miRNA microarray
T24 cells were treated with 4 μM of ATO (Sigma, St. Louis, MO, USA) and incubated for 24 h, then, miRNA-enriched total RNA was extracted for miRNA microarray analysis. RNA of untreated T24 cells was extracted as control. miRNA microarray hybridization was performed by Beijing CapitalBio using fluorescein-labeled miRNAs, each miRNA microarray chip containing 1,320 probes, corresponding to 998 human (including 122 predicted miRNAs), 350 mouse, and 627 rat miRNAs found in the miRNA Registry (http://microrna.sanger.ac.uk/sequences/, miRBase12.0). SmartArray microarrayer (CapitalBio) was used for each probe, which was designed to be complementary to the full-length mature miRNA and printed in triplicate. Procedures were performed as described in detail (http://www.capitalbio.com); briefly, miRNAs were labeled using the T4 RNA ligase labeling method described by Thomson et al. before [20]. Arrays were scanned with a LuxScan 10 K-A double-channel laser scanner (CapitalBio), and images were analyzed using LuxScan 3.0 software (CapitalBio). Differentially expressed miRNAs were identified by Significance Analysis of Microarrays (SAM) software.
Quantitative real-time PCR for miRNA analysis
RT-PCR quantification of mature miRNAs was conducted using TaqMan miRNAssays and following the manufacturer’s supplied instructions (ABI, Foster City, CA, USA). Mean cycle threshold was determined in triplicate PCRs and U6 snRNA was used as the internal control.
miRNA transfection
Mature miRNA and the miRNA negative control were purchased from GenePharma (Shanghai, China). miRNA mimics or miRNA inhibitor was transfected using Lipofectamine 2000 (Invitrogen) following the manufacturer’s instructions. All transfections were carried out in triplicate. To transfect, we diluted miRNA mimics or miRNA inhibitor into 50 μl Opti-Mem into each well. Lipofectamine™ 2000 was mixed gently before use, then 1 μl was diluted in 50 μl Opti- MEM. This was mixed gently and incubated for 5 min at room temperature. After the 5-min incubation, the diluted oligomer was combined with the diluted Lipofectamine™ 2000. This was also mixed gently and incubated for 20 min at room temperature. The oligomer-Lipofectamine™ 2000 complexes were added to each well containing 10,000 cells and medium. After 6 h, the medium was changed and the samples were assayed after 48 h.
Cell viability assay (CCK-8 assay)
Cells (10,000 cells/well) were seeded in a 96-well plate, after a further 48 h transfection, 10 μM of CCK8 (Cell Counting Kit-8, Dojindo, Kumamoto, Japan) was added to each well and incubated for 2 h at 37°C. The OD450nm was measured by a microplate reader (BMG LABTECH). All the experiments were performed in triplicate independently.
Flow cytometry analysis
Detection of apoptotic cells after transfection was performed by using Annexin V/PI (Propidium Iodide) flow cytometric assay (BD Biosciences, San Jose, CA, USA) according to the manufacturer’s instructions. In brief, transfected T24 cells were washed twice after being harvested with cold PBS and resuspended in100 μL cold Annexin V binding buffer (1 × 105 cells). After addition of 5 μL of Annexin V and 5 μL of PI, cells were incubated in the dark for 15 to 30 min at room temperature. Four hundred microliters of binding buffer was added per sample. Samples were analyzed using a FACSort flow cytometer (BD Biosciences). Values are given as means of three independent experiments.
Prediction of miRNA targets
Potential miRNA targets were predicted and analyzed using miRecords [21] (http://mirecords.umn.edu/miRecords/), which contain 11 publicly available algorithms, including Pictar, miRanda, TargetScan, DIANA-microTest, Microinspector, miRDB, miTarget, NbmiRTar, PITA, RNA22, and RNAhybrid. To decrease the number of false-positive results, only putative target genes predicted by at least six programs were accepted.
Western blot analysis
Forty-eight hours after miRNA transfection, T24 cells were lysed and protein was extracted. Western blot analysis was performed as described before [22]. Primary antibody of PTEN, Phospho-Akt, Akt (1:2,000 dilution) and β-actin (1:1,000 dilution) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA).
Luciferase reporter assay
For luciferase assays, pGL3 Firefly Luciferase reporter vector (Promega, Madison, WI, USA) was transfected into T24 cells with the 3′UTR fragment of human PTEN cDNA containing the putative target site for miR-19a. Renilla luciferase construct was co-transfected considering normalization of transfection efficiency. Twenty-four hours post transfection, dual-luciferase assay (Promega) was carried out as described by the manufacturer. Relative firefly luciferase activity of the samples was normalized against the value of Renilla luciferase activity. Experiments were performed more than three times independently.
Statistical analysis
Data were expressed as means ± SD and compared via Student’s t test for paired samples. Statistical significance was accepted at P < 0.05.
Results
Altered expression of miRNAs in T24 cells with ATO treatment
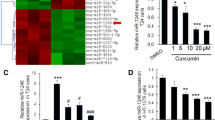
The relative Cy3 and Cy5 dye intensity levels are shown in Fig. 1. SAM software using miRNA microarray data revealed that 30 miRNAs were upregulated and 49 were down-regulated after ATO treatment compared with untreated controls. If only significant differential expressed miRNAs (greater than twofold difference) were considered, two miRNAs were identified; among them, miR-222* was up-regulated and miR-19a was down-regulated after ATO treatment (Table 1).
miRNA expression profiling on miRNA microarray. a RNAs of the group treated with ATO were labeled with cy3 (green), and RNAs of the control group were labeled with cy5 (red). b As a dye-swap, RNAs of the group treated with ATO were labeled with cy5 (red), and RNAs of the control group were labeled with cy3 (green). c The four significant differentially expressed miRNAs
The expression of mature miRNA-222* and miRNA-19a was confirmed by real-time PCR
To confirm the differential expression of miRNAs after ATO treatment (greater than twofold difference), the expressions of mature miRNA-222* and miRNA-19a were examined by qRT-PCR. Consistent with microarray results, the expression of miRNA-222* was up-regulated and miR-19a was down-regulated significantly with ATO treatment compared with untreated control samples (hsa-miR-222*: 2.32 ± 0.24; hsa-miR-19a: 0.42 ± 0.25) (Fig. 3b).
Anti-miRNA-19a inhibited T24 cells proliferation
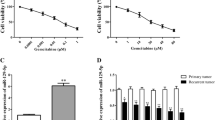
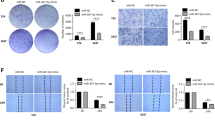
To investigate whether the two miRNAs contribute to cell viability, T24 cells were transfected with miRNA-222* mimic and anti-miRNA-19a, respectively, for 48 h. The cck8-based cell viability assay demonstrated anti-miRNA-19a obtained an inhibition rate above 20% (24.2%), while miRNA-222* mimic obtained 6.9%. Based on these results, we focus on miRNA-19a for our further studies. Interestingly, a synergy effect was detected when anti-miRNA-19a was transfected into T24 cells with ATO treatment, which may lead to equal efficacy and lower dose of ATO treatment (Fig. 2a). CDI is calculated according to the formula \( {\hbox{CDI}} = {\hbox{AB}}/\left( {A * B} \right) \), where AB is the ratio of cell viability index of the combination group of ATO treatment and anti-miR-19a inhibitor transfection to the control group and A or B is the ratio of viability index of the single agent groups to the control group. A value of CDI less than, equal to, or greater than 1 indicates that the drugs are synergistic, additive, or antagonistic, respectively.
a Comparison of ATO dose-dependent inhibition curves between cells transfected with anti-miR-19a inhibitor or control miRNA. Note that anti-miR-19a inhibitor transfection significantly enhanced the inhibition rate of ATO (p < 0.05) at a low dose. The inhibition rate of T24 cells transfected with anti-miR-19a inhibitor is 24.2%. A synergy effect was detected between anti-miRNA-19a transfection and ATO treatment in T24 cells. All the experiments were performed in triplicate independently. b Determination of apoptosis of T24 cells after different treatment by flow cytometry. a Control T24 cells. b Cells transfected with anti-miR-19a. c Cells treated with 2 μM ATO. d Cells treated with 2 μM ATO after anti-miR-19a inhibitor transfection. e Cells treated with 2 μM ATO after miR-19a mimic transfection. Significant difference was found when d or e compared with c (p < 0.05)
Apoptosis mediated by miR-19a
In our study, transfection of anti-miRNA-19a into T24 cells was found to induce cell apoptosis; furthermore, transfection with anti-miR-19a inhibitor enhanced the apoptosis effects induced by ATO, while the apoptosis effect of ATO attenuated by miR-19a mimic transfection on T24 cells (p < 0.05) (Fig. 2b).
Target gene prediction of miR-29a
We used the united prediction algorithm miRecords to identify targets of miR-19a and found most promising 143 candidate genes (Table 2), which is predicted by six databases, except PTEN, which is predicted by seven databases. Then, PTEN was chosen for further studies because it is a well-known tumor suppressor gene associated with many kinds of cancers and apoptosis pathways; more importantly, it has been proven to be involved in ATO treatment.
PTEN is a direct target of miR-19a
The results showed a significant decrease in luciferase activity for the 3′UTR of PTEN transfected with miR-19a mimic compared with the control, whereas the 3′UTR luciferase showed much higher activity when transfected with miR-19a inhibitor in T24 cells (p < 0.05) (Fig. 3a). Therefore, miR-19a represses PTEN by binding to the 3′UTR of PTEN in a direct manner at the posttranscriptional level.
a Luciferase activity assay. T24 cells were co-transfected with the luciferase reporter vector, which contained PTEN 3′-UTR fragment, and miR-19a, anti-miR-19, or control sequence. Luciferase activity was measured to determine the effect of these miRNAs on luciferase translation. Firefly luciferase activity of cells was assayed 48 h after transfection and renilla luciferase was used as transfection efficiency correction. Negative control was set as 100%. PTEN 3′-UTR-luciferase activity was decreased significantly in T24 cells transfected with miR-19a mimic, while silencing miR-19a with anti-miR-19a inhibitor led to a higher luciferase activity in T24 cells (p < 0.05). b RT-PCR: the expression of miRNA-222* was up-regulated and miR-19a was down-regulated significantly with ATO treatment compared with untreated control samples (hsa-miR-222*: 2.32 ± 0.24; hsa-miR-19a: 0.42 ± 0.25)
Anti-miRNA-19a promotes protein PTEN
PTEN protein expression was significantly decreased in T24 cells transfected with miRNA-19a, while transfection of anti-miRNA-19a led to increased expression of PTEN protein in T24 cells. ATO treatment increased PTEN protein expression, while transfection with the miR-19a suppressed the function of ATO. These results indicate that miRNA-19a may play a crucial role in ATO treatment and PTEN regulation (Fig. 4).
Western blot analysis. a Transfection with the anti-miR-19a inhibitor increased the PTEN expression and induced a significant decrease in P-AKT (phosphorylation levels of Akt) protein expression in T24 cells. b Overexpression of miR-19a induced a significant decrease in PTEN protein expression and increased the P-AKT expression. c Arsenic increased the PTEN protein expression in T24 cells, while transfection with miR-29a counteracted ATO function
Discussion
miRNAs represent a class of small non-protein coding regulatory RNAs that have central roles in gene silencing and function as oncogenes or tumor suppressors. miRNAs were also found to be potentially useful for therapeutic purposes recently [23, 24]. ATO has been effectively used for many years to treat patients with cancers due to its remarkable biological effects on several cellular functions, including growth-inhibitory and proapoptotic effects. Previous data from our laboratory show that activation of several signaling pathways mediates the arsenic-induced apoptotic signal in T24 cells [22]. As miRNAs have been considered to be interacted with many well-recognized cell signal pathways, we hypothesized that ATO would alter the expression profile of miRNAs in human bladder cancer cells, which may reveal the role of miRNAs in ATO treatment.
The microarray analysis identified 67 miRNAs that were up- or down-regulated in T24 cells following ATO treatment. Among them, two miRNAs (miR-19a, miR-222*) were selected according to the twofold criterion. Furthermore, anti-miR-19a showed a strong inhibition of cell proliferation of T24 cells and an increased apoptotic rate compared with control after transfection. Unexpectedly, miR-222* transfection did not show clear positive results. These results indicate that miR-19a is the most potent growth suppressor or apoptosis inducer in the ATO treatment of T24 cells. Interestingly, we showed that ATO gets a satisfactory growth inhibition and apoptotic rate at a low dose of 1.0 μmol/L with transfection of anti-miR-19a. This result could effectively improve the clinical utility of ATO treatment for its lower dose, which, in turn, lowered toxic side effects.
miR-19a belong to the miR-17–92 cluster, which is localized on chromosome 13q31.3 and is well known for its oncogenic role as reported [25–27]. In our study, we performed miR-19a overexpression and depletion in bladder cancer T24 cells and observed an obvious change in cell growth and apoptosis in this cell line. Indeed, we showed that transfection of T24 cells with anti-miR-19a resulted in decreased cell viability and induction of cell death. For further study, we focus on the targets of miR-19a because miRNAs exert their function through translational repression or degradation of mRNA targets. To predict the target genes of miR-19a, we used miRecords and finally chose PTEN for further studies as PTEN was a well-known anti-oncogene and has been confirmed to be involved in ATO treatment before [28]. Western blot results showed that over-expression of miR-19a decreased the expression of PTEN and depletion of miR-19a increased the expression of protein PTEN by analysis (Fig. 4). Furthermore, the phosphorylation levels of Akt, a major target of PTEN, increased by over-expression of miR-19a and decreased by depletion of miR-19a. Our luciferase reporter assays also confirmed that PTEN was a direct target for miR-19a in bladder cancer T24 cells as reported before in other cell lines [29]. ATO has been proved previously to upregulate the expression of PTEN [28], which coincided with our results that the expression of PTEN was increased in T24 cells treated with ATO. We reported that ATO treatment of T24 cells decreased the expression of miR-19a and also up-regulated the expression of its target gene PTEN with increased expression of protein PTEN and decreased expression of protein phospho-Akt, and the miR-19a mimics suppressed the effect of ATO on PTEN. The results indicate that miR-19a–PTEN–Akt was one of the important pathways of ATO on cancer cells.
Our study is the first to examine miRNAs expression alteration in human bladder cancer cells after ATO treatment. Whether the differential expressed miRNAs influence other biological activity against bladder cancer or in vivo remains to be investigated in future studies. However, our data provide evidence that miRNAs could serve as a potential approach to enhance the efficacy and safety of ATO treatment. We were overwhelmed to see the synergy effect between anti-miR-19a and ATO for its further clinical usefulness.
References
Shen ZX, Chen GQ, Ni JH, et al. Use of arsenic trioxide (As2O3) in the treatment of acute promyelocytic leukemia (APL): II. Clinical efficacy and pharmacokinetics in relapsed patients. Blood. 1997;89:3354–60.
Soignet SL, Frankel SR, Douer D, et al. United States multicenter study of arsenic trioxide in relapsed acute promyelocytic leukemia. J Clin Oncol. 2001;19:3852–60.
Chen X, Zhang M, Liu LX. The overexpression of multidrug resistance-associated proteins and gankyrin contribute to arsenic trioxide resistance in liver and gastric cancer cells. Oncol Rep. 2009;22:73–80.
Karlsson J, Pietras A, Beckman S, Pettersson HM, Larsson C, Pahlman S. Arsenic trioxide-induced neuroblastoma cell death is accompanied by proteolytic activation of nuclear Bax. Oncogene. 2007;26:6150–9.
Lin TH, Kuo HC, Chou FP, Lu FJ. Berberine enhances inhibition of glioma tumor cell migration and invasiveness mediated by arsenic trioxide. BMC Cancer. 2008;8:58.
Murgo AJ. Clinical trials of arsenic trioxide in hematologic and solid tumors: overview of the National Cancer Institute Cooperative Research and Development Studies. Oncologist. 2001;6 Suppl 2:22–8.
Bi X, Gu J, Guo Z, et al. Different pathways are involved in arsenic-trioxide-induced cell proliferation and growth inhibition in human keratinocytes. Skin Pharmacol Physiol. 2010;23:68–78.
Cai BZ, Meng FY, Zhu SL, et al. Arsenic trioxide induces the apoptosis in bone marrow mesenchymal stem cells by intracellular calcium signal and caspase-3 pathways. Toxicol Lett. 2010;193:173–8.
Han YH, Moon HJ, You BR, Kim SZ, Kim SH, Park WH. Effects of arsenic trioxide on cell death, reactive oxygen species and glutathione levels in different cell types. Int J Mol Med. 2010;25:121–8.
Han YH, Moon HJ, You BR, Kim SZ, Kim SH, Park WH. The effect of MAPK inhibitors on arsenic trioxide-treated Calu-6 lung cells in relation to cell death, ROS and GSH levels. Anticancer Res. 2009;29:3837–44.
Mandegary A, Hosseini R, Ghaffari SH, et al. The expression of p38, ERK1 and Bax proteins has increased during the treatment of newly diagnosed acute promyelocytic leukemia with arsenic trioxide. Ann Oncol. 2010;21:1884–90.
Mann KK, Colombo M, Miller Jr WH. Arsenic trioxide decreases AKT protein in a caspase-dependent manner. Mol Cancer Ther. 2008;7:1680–7.
Wen J, Feng Y, Huang W, et al. Enhanced antimyeloma cytotoxicity by the combination of arsenic trioxide and bortezomib is further potentiated by p38 MAPK inhibition. Leuk Res. 2010;34:85–92.
Friedland DR, Eernisse R, Erbe C, Gupta N, Cioffi JA. Cholesteatoma growth and proliferation: posttranscriptional regulation by microRNA-21. Otol Neurotol. 2009;30:998–1005.
Guo L, Ding ZH. Advances in the studies of miRNAs and cell apoptosis. Sheng Li Ke Xue Jin Zhan. 2007;38:331–5.
Li Y, Guessous F, Zhang Y, et al. MicroRNA-34a inhibits glioblastoma growth by targeting multiple oncogenes. Cancer Res. 2009;69:7569–76.
Vecchione A, Croce CM. Apoptomirs: small molecules have gained the license to kill. Endocr Relat Cancer. 2010;17:F37–50.
Ren Y, Zhou X, Mei M, et al. MicroRNA-21 inhibitor sensitizes human glioblastoma cells U251 (PTEN-mutant) and LN229 (PTEN-wild type) to taxol. BMC Cancer. 2010;10:27.
Zhong M, Ma X, Sun C, Chen L. MicroRNAs reduce tumor growth and contribute to enhance cytotoxicity induced by gefitinib in non-small cell lung cancer. Chem Biol Interact. 2010;184:431–8.
Thomson JM, Parker J, Perou CM, Hammond SM. A custom microarray platform for analysis of microRNA gene expression. Nat Methods. 2004;1:47–53.
Xiao F, Zuo Z, Cai G, Kang S, Gao X, Li T. miRecords: an integrated resource for microRNA-target interactions. Nucleic Acids Res. 2009;37:D105–110.
Wang Y, An R, Dong X, Pan S, Duan G, Sun X. Protein kinase C is involved in arsenic trioxide-induced apoptosis and inhibition of proliferation in human bladder cancer cells. Urol Int. 2009;82:214–21.
Akao Y, Nakagawa Y, Hirata I, et al. Role of anti-oncomirs miR-143 and -145 in human colorectal tumors. Cancer Gene Ther. 2010;17:398–408.
Li Y, Kong D, Wang Z, Sarkar FH. Regulation of microRNAs by Natural Agents: an emerging field in chemoprevention and chemotherapy research. Pharm Res. 2010;27:1027–41.
Doebele C, Bonauer A, Fischer A, et al. Members of the microRNA-17-92 cluster exhibit a cell intrinsic anti-angiogenic function in endothelial cells. Blood. 2010;115:4631–3.
Loven J, Zinin N, Wahlstrom T, et al. MYCN-regulated microRNAs repress estrogen receptor-alpha (ESR1) expression and neuronal differentiation in human neuroblastoma. Proc Natl Acad Sci U S A. 2010;107:1553–8.
Pezzolesi MG, Platzer P, Waite KA, Eng C. Differential expression of PTEN-targeting microRNAs miR-19a and miR-21 in Cowden syndrome. Am J Hum Genet. 2008;82:1141–9.
Wang R, Li H, Guo G, et al. Augmentation by carnosic acid of apoptosis in human leukaemia cells induced by arsenic trioxide via upregulation of the tumour suppressor PTEN. J Int Med Res. 2008;36:682–90.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. Prediction of mammalian microRNA targets. Cell. 2003;115:787–98.
Conflict of interest statement
None declared
Author information
Authors and Affiliations
Corresponding author
Additional information
Yan Cao and Shi-Liang Yu are contributed equally to this work.
Rights and permissions
About this article
Cite this article
Cao, Y., Yu, SL., Wang, Y. et al. MicroRNA-dependent regulation of PTEN after arsenic trioxide treatment in bladder cancer cell line T24. Tumor Biol. 32, 179–188 (2011). https://doi.org/10.1007/s13277-010-0111-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-010-0111-z