Abstract
Background
Primordial dwarfism (PD) is a group of genetically heterogeneous disorders related to developmental disabilities occurring in the uterus and prolongs during all stages of life, resulting in short stature, facial deformities and abnormal brain.
Objective
To determine the exact cause of the disease in two Vietnamese patients priory diagnosed with PD by severe pre-and postnatal growth retardation with marked microcephaly and some bone abnormalities.
Methods
Whole-exome sequencing was performed for the two patients and mutations in genes related to PD were screened. Sanger sequencing was applied to examine the mutations in the patients of their families.
Results
Three novel mutations in the PCNT gene which have not been reported previously were identified in the two patients. Of which, two frameshift mutations (p.Thr479Profs*6 and p.Glu2742Alafs*8) were detected in patient I and one stop-gained mutation (p.Gln1907*) was detected in the patient II. These mutations may result in a truncated PCNT protein, leading to an inactivated PACT domain corresponding to residue His3138–Trp3216 of PCNT protein. Therefore, the three mutations may cause a deficiency of protein functional activity and result in the phenotypes of primordial dwarfism in the two patients.
Conclusions
Clinical presentations in combination with genetic analyses supported an accurate diagnosis of the two patients with microcephalic osteodysplastic primordial dwarfism type II (MOPD II). In addition, these results have important implications for prenatal genetic screening and genetic counseling for the families.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Primordial dwarfism (PD) is a group of disorders related to developmental disabilities occurring in the uterus and prolongs during all stages of life, resulting in short stature, facial deformities and abnormal brain (Vakili and Hashemian 2019). Most of PD individuals have head size-reduction features which are also known as a highly specified property of the PD group to distinguish with other dwarfism (Khetarpal et al. 2016). PD is extremely rare and presents in both sexes at the same rate (Vakili and Hashemian 2019). There are five main types of primordial dwarfism: Seckel syndrome, ear-patella-short stature syndrome (Meier–Gorlin); Russell–Silver syndrome; microcephalic osteodysplastic bird-head primordial dwarfism type I/III (MOPD I/ MOPD III); and microcephalic osteodysplastic primordial dwarfism type II.
MOPD II (OMIM # 210720) is typical with congenital disproportionate short stature, microcephaly, dysmorphic face and characteristics skeletal dysplasia (Brancati et al. 2005). Other signs and symptoms have been reported in MOPD II individuals, such as microdontia and cerebrovascular (Hall et al. 2004) and insulin resistance (Bober and Jackson 2017). The adult height of MOPD II individuals was less than 100 mm and the circumference of the head was around 40 cm or less (Pachajoa et al. 2014). Interestingly, compared to normal people, patients with MOPD II have a smaller brain size, but most of them have almost normal intelligence. Although the incidence of MOPD II has not been estimated, MOPD II is one of the most common types of the microcephalic primordial dwarfism group with more than 150 individuals have been reported (Bober and Jackson 2017). MOPD II is a genetically homogeneous condition caused by mutations in the PCNT gene (Chen et al. 1996; Bober and Jackson 2017). The PCNT gene spans 122 kb of genomic sequence on chromosome 21q22.3 with 47 exons and encodes a 3336-amino acid protein (Chen et al. 1996). It plays an important role in the process of mitosis phase of the cell cycle, therefore, mutations in this gene can cause disruption in cell division, followed by reduction of cell number as well as body size (Griffith et al. 2008; Willems et al. 2010; Kim and Rhee 2014). Hitherto, 53 distinct mutations have been identified in the PCNT gene in MOPD II patients, including 24 nonsense mutations, 20 frameshifts mutations, eight splice site mutations and one in-frame deletion mutation (Abdel-Salam et al. 2020; Waich et al. 2020).
In this study, we identify three novel mutations in the PCNT gene in two Vietnamese patients priory diagnosed with primordial dwarfism using whole-exome sequencing and Sanger sequencing.
Materials and methods
Patients who participated in the study were all agreed on by their parents. Genomic DNA was extracted from the peripheral blood of the two patients and members of their families using the QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany), following the manufacturer’s instructions. Whole-exome sequencing was performed in the two patients. The DNA libraries were prepared according to the instructions of the Agilent SureSelect Target Enrichment Kit (Agilent Technologies, CA, USA) and sequenced on Illumina sequencer (Illumina, CA, USA). Burrows-Wheeler Aligner tool v0.7.12 (Li and Durbin 2009) was used to map the sequencing reads with the GRCh37/hg19 reference human genome. The duplicate reads were detected and deleted using Picard tool v1.130 (http://broadinstitute.github.io/picard/). The variants were called and annotated by using Genome Analysis Toolkit v3.4.0 (McKenna et al. 2010) and SnpEff v4.1 (http://snpeff.sourceforge.net/SnpEff.html). The population frequency of the variants was estimated by comparing with the Exome Sequencing Project (https://evs.gs.washington.edu/EVS/), 1000 Genome (http://browser.1000genomes.org) and Kinh Vietnamese (KHV) database (https://genomes.vn). Variants with a minor allele frequency > 0.01 were excluded. Variants were annotated by SnpEff v4.1. Then, candidate variants in the pathogenic genes related to primordial dwarfism (Supplementary Table S1) were screened. Selected pathogenic candidate variants were validated by Sanger sequencing and examined for whether the variants are de novo or inherited from the parents. Protein structural regions of PCNT protein were predicted by SMART (http://smart.embl-heidelberg.de/). The pathogenicity of mutations was evaluated according to the five-tier classification system of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (Richards et al. 2015).
Results
Clinical presentations
The patient I is the first child of non-consanguineous Vietnamese parents. He was born at 34 weeks of gestation by cesarean section with a birth weight of 600 g. He was transferred to the intensive care unit for of postpartum care for 3 months, in which, he received mechanical ventilation for one month. He could roll over at 7-month-old. At the age of 25 months, he suffered from a convulsion without fever and was referred to the Vietnam National Children’s Hospital. At the time of examination, he presented with severe growth retardation, with a height of 45 cm [13 standard deviations (SD) below the mean in normal subjects (− 13 SD)], a weight of 3.5 kg (< − 5.4 SD) and a head circumference of 33.8 cm (− 11.1 SD). He had dysmorphic features, including microcephaly, micrognathia, prominent nose, brachydactyly and hypotonia (Fig. 1a). He could not sit down and stand up. He has a high-pitched voice.
Clinical manifestations of two patients. Patient I presented with microcephaly, micrognathia, prominent nose and brachydactyly (a). Patient II showed dysmorphic features consisting of a prominent nose, micrognathia, low-set ears, brachydactyly and clinodactyly of the 5th finger, hypertrophy and metaphysis of the distal femurs and bilateral coxa vara (b)
The patient II is the third child of non-consanguineous Vietnamese parents. He was born preterm with a gestational age of 35 weeks with a birth weight of 1.2 kg. He was admitted to the hospital when he was 5-years-old. On examination at presentation, he had growth retardation with a height of 76 cm (– 7 SD), a weight of 10.2 kg (– 4.5 SD) and a head circumference of 40.5 cm (– 6 SD). His developmental quotient (DQ) score was 65%. His face was dysmorphic with a prominent nose, micrognathia and low-set ears (Fig. 1b). His primary teeth showed hypoplasia. He presented with brachydactyly, clinodactyly of the 5th finger and micropenis (Fig. 1b). His skeletal X-ray demonstrated hypertrophy and metaphysis of the distal femurs and bilateral coxa vara (Fig. 1b). No abnormalities were detected on examination of the heart and brain systems.
Molecular analyses and pathogenic interpretation
The whole-exome sequencing resulted in a total of 111,665 variants in patient I and 109,176 variants in patient II (Supplementary Table S2). Among them, only two frameshift mutations (c.1435delA, p.Thr479Profs*6 and c.8223_8224delTG, p.Glu2742Alafs*8) in the patient I and a stop-gained mutation (c.5719 C > T, p.Gln1907*) in the patient II were determined as disease-causing mutations (Supplementary Table S3). All three of these mutations were predicted to cause disease by the Mutation Taster tool (http://www.mutationtaster.org/). Sanger sequencing revealed patient I carried a compound heterozygous mutation c.1435delA and c.8223_8224delTG (Fig. 2b). Maternal PCNT allele harbored a heterozygous mutation in exon 9 (Thr479Profs*6) and paternal PCNT one contained a point mutation located in exon 38 (Glu2742Alafs*8). The A nucleotide deletion at position cDNA 1435 switched threonine to proline at residue 479 and resulted in a frameshift, in which, PCNT protein may be truncated 6 amino acids downstream (Fig. 2a). The deletion of two GT nucleotides at position c.DNA 8223_8224 changed glutamic acid to alanine at residue 2742 in the PCNT protein. An early termination codon appeared at 8 amino acids downstream (Fig. 2a).
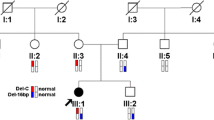
Mutations in the PNCT gene in the two families. Sanger sequencing revealed patient I carried a compound heterozygous mutation c.1435delA (p.Thr479Profs*6) in exon 9 and c.8223_8224delTG (p.Glu2742Alafs*8) in exon 38 of the PCNT gene. The maternal PCNT allele harbored the mutation c.1435delA in the heterozygous state and the paternal one contained the mutation c.8223_8224delTG in the heterozygous state (a). Molecular genetic analyses revealed patient II carried the mutation c.5719 C > T (p.Gln1907*) in exon 28 of the PCNT gene in the homozygous state with each mutant allele was inherited from his parents (b). Protein structural regions of PCNT protein were predicted by SMART, including the low complexity region (pink), coiled-coil regions (green) and PACT domain (yellow) (c)
Patient II carried the mutation c.5719 C > T in the homozygous state. He inherited one maternal mutant allele and one paternal mutant allele (Fig. 2b). The mutation c.5719 C > T resulted in the substitution of Glu1907 with a stop codon, leading to a truncated PCNT.
Three mutations, c.8223_8224delTG, c.1435_1435delA and c.5719 C > T were submitted to ClinVar database under accession numbers SCV001441536, SCV001441537 and SCV001441538, respectively. These three mutations are classified as pathogenic mutations with one very strong evidence of pathogenicity (PVS1), two moderate evidences of pathogenicity (PM2 and PM4) and one supporting evidence of pathogenicity (PP4) (Supplementary Table S4).
Discussion
Our patients were initially clinically diagnosed with a PD group along with other forms of microcephalic primordial dwarfism as severe prenatal and postnatal growth retardations with marked microcephaly. Due to the heterogeneity feature of the diseases in the PD group, it is difficult to diagnose patients based on the clinical features. Therefore, we conducted whole-exom sequencing in the probands and screened mutations in all gene-related to the PD group (Supplementary Table S1). Whole-exome sequencing identified three pathogenic mutations in the PCNT gene (p.Thr479Profs*6 and p.Glu2742Alafs*8 in patient I and p.Gln1907* in patient II). All these may inactivate the PACT domain corresponding to residues His3138 to Trp3216 of the PCNT (Fig. 2c). The PACT domain has a direct role in the reception of AKAP-450 and pericentrin to the centrosome (Gillingham and Munro 2000). This domain deputized as a coiled-coil region occupies close to the C terminus of centrosomal which were found in mammalian centrosomes, drosophila and fission yeast. PCNT serves as a central scaffold that anchors the complex of microtubule nuclei and other centrosomal proteins. The action of this protein ensures proper chromosome division during cell division. Truncating mutations were reported as a loss of functional PCNT, causing clinical conditions related to the disease (Rauch et al. 2008).
A recent study (Waich et al. 2020) also revealed that compound heterozygosity for two novel truncating PCNT mutations, c.1345_1864del (p.Leu449Thrfs*5) and c.3239_3240insCTGG (p.Gln1081Trpfs*114), reduced 30% PCNT mRNA expression in patient’s fibroblast as compared to healthy control. However, 20% of patient’s fibroblasts showed normal microtubule and spindle organization in all mitotic phases, suggesting some residual function of PCNT protein which might be associated with attenuated growth restriction in MOPD II in the patients (Waich et al. 2020). In another recent study, a pathogenic frame-shift variant in PCNT gene (c.7511delA, p.Lys2504Serfs*27) was discovered in an Iranian female patient with MOPD II (Dehghan Tezerjani et al. 2020). The mutation c.7511delA, p.Lys2504Serfs*27 lost the PACT domain and a region binding to Nek2A, leading to the premature centrosome splitting in interphase (Dehghan Tezerjani et al. 2020). Therefore, the three truncating mutations found in our patients may result a loss of PCNT protein, leading to their phenotype of MOPD II. As a result, the two patients were accurately diagnosed with MOPD II. The two patients in this study also shared the common characteristics of MOPD II such as growth retardation, dysmorphic features and skeletal abnormalities. Our patients showed a severe postnatal growth failure which is consistent with the findings of the previous studies (Bober et al. 2012; Willems et al. 2010). Dysmorphic features such as micrognathia, prominent nose and brachydactyly which were reported in the previous studies (Willems et al. 2010; Bober and Jackson 2017; Vakili and Hashemian 2019) also observed in our patients. Interestingly, patient II presented the rare skeletal abnormalities of MOPD II such as bilateral coxa vara and clinodactyly of the 5th finger. Coxa vara occurred in eight MOPD II patients with PCNT mutations (Fukuzawa et al. 2002; Karatas et al. 2014; Bober and Jackson 2017; Abdel-Salam et al. 2020), however, only one patient showed bilateral coxa vara (Bober and Jackson 2017). Clinodactyly has been reported in six patients with MOPD II (Kozlowski et al. 1993; Hall et al. 2004; Willems et al. 2010; Unal et al. 2014; Bober and Jackson 2017). Of which, clinodactyly of the 5th finger was reported in the two individuals (Kozlowski et al. 1993; Bober and Jackson 2017).
Motor delay may be seen early in MOPD II patients (Hall et al. 2004). An Iranian female patient with MOPD II could not walk until the age of 2.5 years (Vakili and Hashemian 2019). An Israeli male patient could not sit or stand on his own at the age of 14 months, however, at the age of 40 months, he could walk independently (Weiss et al. 2020). In our study, patient I also could not sit and stand up at the age of 25 months. However, his conditions may improve with age.
Seven Asian patients with MOPD II carrying loss-of-function mutations have been reported to date (Rauch et al. 2008; Kantaputra et al. 2011; Li et al. 2015). In comparison with these seven patients, our patients shared similar symptoms such as microcephaly, micrognathia, prominent nose and brachydactyly (Table 1). However, our patients showed incapacity to sit down and stand up, bilateral coxa vara, dysplasia of the neck and head of femurs, hypertrophy of the distal femurs, humerus, forearms and clinodactyly of the 5th finger (Table 1).
Conclusions
In this study, the detection of mutations in the PCNT gene in two patients provided convincing evidence for a precise diagnosis for both patients with MOPD II. These results may contribute significantly to prenatal screening tests to help the doctors to genetic counsel for the patient’s family.
References
Abdel-Salam GMH, Sayed ISM, Afifi HH, Abdel-Ghafar SF, Abouzaid MR, Ismail SI, Aglan MS, Issa MY, El-Bassyouni HT et al (2020) Microcephalic osteodysplastic primordial dwarfism type II: additional nine patients with implications on phenotype and genotype correlation. Am J Med Genet A 182(6):1407–1420. https://doi.org/10.1002/ajmg.a.61585
Bober MB, Jackson AP (2017) Microcephalic osteodysplastic Primordial dwarfism, type II: a clinical review. Curr Osteoporos Rep 15(2):61–69. https://doi.org/10.1007/s11914-017-0348-1
Bober MB, Niiler T, Duker AL, Murray JE, Ketterer T, Harley ME, Alvi S, Flora C, Rustad C, Bongers EMHF et al (2012) Growth in individuals with Majewski osteodysplastic primordial dwarfism type II caused by pericentrin mutations. Am J Med Genet 158A(11):2719–2725. https://doi.org/10.1002/ajmg.a.35447
Brancati F, Castori M, Mingarelli R, Dallapiccola B (2005) Majewski osteodysplastic primordial dwarfism type II (MOPD II) complicated by stroke: clinical report and review of cerebral vascular anomalies. Am J Med Genet A 139(3):212–215. https://doi.org/10.1002/ajmg.a.31009
Chen H, Gos A, Morris MA, Antonarakis SE (1996) Localization of a human homolog of the mouse pericentrin gene (PCNT) to chromosome 21qter. Genomics 35(3):620–624. https://doi.org/10.1006/geno.1996.0411
Dehghan Tezerjani M, Vahidi Mehrjardi MY, Hozhabri H, Rahmanian M (2020) A novel PCNT frame shift variant (c.7511delA) causing osteodysplastic Primordial dwarfism of majewski Type 2 (MOPD II). Front Pediatr. https://doi.org/10.3389/fped.2020.00340
Fukuzawa R, Sato S, Sullivan MJ, Nishimura G, Hasegawa T, Matsuo N (2002) Autopsy case of microcephalic osteodysplastic primordial “dwarfism” type II. Am J Med Genet 113(1):93–96. https://doi.org/10.1002/ajmg.10716
Gillingham AK, Munro S (2000) The PACT domain, a conserved centrosomal targeting motif in the coiled-coil proteins AKAP450 and pericentrin. EMBO Rep 1(6):524–529. https://doi.org/10.1093/embo-reports/kvd105
Griffith E, Walker S, Martin C-A, Vagnarelli P, Stiff T, Vernay B, Al Sanna N, Saggar A, Hamel B, Earnshaw WC et al (2008) Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling. Nat Genet 40(2):232–236. https://doi.org/10.1038/ng.2007.80
Hall JG, Flora C, Scott CI, Pauli RM, Tanaka KI (2004) Majewski osteodysplastic primordial dwarfism type II (MOPD II): natural history and clinical findings. Am J Med Genet 130A(1):55–72. https://doi.org/10.1002/ajmg.a.30203
Kantaputra P, Tanpaiboon P, Porntaveetus T, Ohazama A, Sharpe P, Rauch A, Hussadaloy A, Thiel CT (2011) The smallest teeth in the world are caused by mutations in the PCNT gene. Am J Med Genet A 155A(6):1398–1403. https://doi.org/10.1002/ajmg.a.33984
Karatas AF, Bober MB, Rogers K, Duker AL, Ditro CP, Mackenzie WG (2014) Hip pathology in Majewski osteodysplastic primordial dwarfism type II. J Pediatr Orthop 34(6):585–590. https://doi.org/10.1097/BPO.0000000000000183
Khetarpal P, Das S, Panigrahi I, Munshi A (2016) Primordial dwarfism: overview of clinical and genetic aspects. Mol Genet Genom 291(1):1–15. https://doi.org/10.1007/s00438-015-1110-y
Kim S, Rhee K (2014) Importance of the CEP215-pericentrin interaction for centrosome maturation during mitosis. PLoS One 9(1):e87016. https://doi.org/10.1371/journal.pone.0087016
Kozlowski K, Donovan T, Masel J, Wright RG (1993) Microcephalic, osteodysplastic, primordial dwarfism. Australas Radiol 37(1):111–114. https://doi.org/10.1111/j.1440-1673.1993.tb00029.x
Li FF, Wang XD, Zhu MW, Lou ZH, Zhang Q, Zhu CY, Feng HL, Lin ZG, Liu SL (2015) Identification of two novel critical mutations in PCNT gene resulting in microcephalic osteodysplastic primordial dwarfism type II associated with multiple intracranial aneurysms. Metab Brain Dis 30(6):1387–1394. https://doi.org/10.1007/s11011-015-9712-y
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinf Oxf Engl 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M et al (2010) The genome analysis toolkit: a map reduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
Pachajoa H, Ruiz-Botero F, Isaza C (2014) A new mutation of the PCNT gene in a Colombian patient with microcephalic osteodysplastic primordial dwarfism type II: a case report. J Med Case Rep 8:191. https://doi.org/10.1186/1752-1947-8-191
Rauch A, Thiel CT, Schindler D, Wick U, Crow YJ, Ekici AB, van Essen AJ, Goecke TO, Al-Gazali L, Chrzanowska KH et al (2008) Mutations in the pericentrin (PCNT) gene cause primordial dwarfism. Science 319(5864):816–819. https://doi.org/10.1126/science.1151174
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med Off J Am Coll Med Genet 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Shaheen R, Maddirevula S, Ewida N, Alsahli S, Abdel-Salam GMH, Zaki MS, Tala SA, Alhashem A, Softah A, Al-Owain M et al (2019) Genomic and phenotypic delineation of congenital microcephaly. Genet Med Off J Am Coll Med Genet 21(3):545–552. https://doi.org/10.1038/s41436-018-0140-3
Unal S, Alanay Y, Cetin M, Boduroğlu K, Utine E, Cormier-Daire V, Huber C, Ozsurekci Y, Kılıç E, Simsek-Kiper P et al (2014) Striking hematological abnormalities in patients with Microcephalic osteodysplastic Primordial dwarfism type II (MOPD II): a potential role of pericentrin in hematopoiesis. Pediatr Blood Cancer. https://doi.org/10.1002/pbc.24783
Vakili R, Hashemian S (2019) Primordial dwarfism: a case series from North East of Iran and literature review. J Pediatr Rev 7(2):113–120. https://doi.org/10.32598/jpr.7.2.113
Waich S, Janecke AR, Parson W, Greber-Platzer S, Müller T, Huber LA, Valovka T, Vodopiutz J (2020) Novel PCNT variants in MOPD II with attenuated growth restriction and pachygyria. Clin Genet. https://doi.org/10.1111/cge.13797
Weiss K, Ekhilevitch N, Cohen L, Bratman-Morag S, Bello R, Martinez AF, Hadid Y, Shlush LI, Kurolap A, Paperna T et al (2020) Identification of a novel PCNT founder pathogenic variant in the Israeli Druze population. Eur J Med Genet 63(2):103643. https://doi.org/10.1016/j.ejmg.2019.03.007
Willems M, Geneviève D, Borck G, Baumann C, Baujat G, Bieth E, Edery P, Farra C, Gerard M, Héron D et al (2010) Molecular analysis of pericentrin gene (PCNT) in a series of 24 Seckel/microcephalic osteodysplastic primordial dwarfism type II (MOPD II) families. J Med Genet 47(12):797–802. https://doi.org/10.1136/jmg.2009.067298
Acknowledgements
We thank all the patients and their families for their time and support.
Funding
This work was supported by a grant from the Vietnam Academy of Science and Technology (KHCBSS.02/18-20).
Author information
Authors and Affiliations
Contributions
THN performed experiments, analyzed data and wrote the manuscript; NLN analyzed data; CDV, CTBN and NKN diagnosed and interpreted the patients’ data; NHN designed the study, reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Thu Hien Nguyen, Ngoc-Lan Nguyen, Chi Dung Vu, Can Thi Bich Ngoc, Ngoc Khanh Nguyen, Huy Hoang Nguyen declare that they have no conflict of interest.
Ethics approval
This study was conducted in accordance with the Declaration of Helsinki and the protocol was approved by the Ethics Committee of the Institute of Genome Research (Institute of Genome Research Institutional Review Board, Hanoi, Vietnam, Reference no: 18/QD-NCHG on 22 March 2018).
Consent to participate
Written informed consent were obtained from the parents.
Consent for publication
The parents provided informed consent for publication of the images in Figs. 1 and 2a and b.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nguyen, T.H., Nguyen, NL., Vu, C.D. et al. Identification of three novel mutations in PCNT in vietnamese patients with microcephalic osteodysplastic primordial dwarfism type II. Genes Genom 43, 115–121 (2021). https://doi.org/10.1007/s13258-020-01032-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-01032-5