Abstract
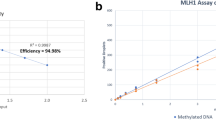
Detecting cfDNA in plasma or serum could serve as a ‘liquid biopsy’, for circulating tumor DNA with aberrant methylation patterns offer a possible method for early detection of several cancers which could avoid the need for tumor tissue biopsies. Bone Morphogenetic Protein 3 (BMP3) was identified as a candidate tumor suppressor gene putatively down-regulated in colorectal cancer (CRC). In this study, we aimed to assess the potential role of BMP3 promoter methylation changes in plasma DNA for detection of colorectal cancerous and precancerous lesions. Plasma DNA samples were extracted from 50 patients with histologically diagnosed polyps or tumor and 50 patients reported negative for polyps or tumors. The procedure consists of bisulfite conversion of the extracted DNA, purification of bis-DNA, and BMP3 methylation status analysis by using the bisulfite specific high resolution melting analysis. This study demonstrated that there was a significantly higher frequency of BMP3 methylated DNA in plasma in patients with polyps versus healthy controls with a sensitivity and specificity of 40 and 94%, respectively. In conclusion, our results demonstrated that BMP3 DNA methylation in plasma had not have sufficient sensitivity and it should be used in combination with other biomarkers for the detection of CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC), the major cause of morbidity and mortality, accounts for over 9% of all cancer incidences worldwide (Arnold et al. 2017; Haggar and Boushey 2009; Hessami Arani and Kerachian 2017; Lange and Laird 2013; Tanzer et al. 2010). Thus, early detection of CRC could have considerable clinical benefits including reducing mortality and morbidity (Lange et al. 2012). In this cancer, 5-year survival rates are 70 and 13% for regional and distant stages, respectively (Gonzalez-Pons and Cruz-Correa 2015). Due to the fact that CRC evolves primarily via the established adenoma-to-carcinoma pathway (Gonzalez-Pons and Cruz-Correa 2015), cancer screening could prevent cancerous formation and early treatment intervention for CRC patient (Levin et al. 2008). The American Cancer Society (ACS) has introduced CRC as a major priority since the application of current science and knowledge have such a great potential to prevent cancer, lower suffering and extend life expectancy (Levin et al. 2008). Although colonoscopy is the gold standard for CRC screening, this procedure is invasive, expensive, and patients suffer from inconvenience (Lange et al. 2012). Hence, there is a strong need for discovery of noninvasive detection method assays. In the recent years, several noninvasive tests have been developed for CRC screening (Ahlquist et al. 2012a). Noninvasive biomarkers are expected to be highly sensitive and specific to evaluate genetic, epigenetic or protein markers that can be detected in the stool or in plasma of patients (Kim et al. 2008; Lange et al. 2012).
One of the main processes causing the initiation and progression of CRC is the accumulation of a variety of genetic and epigenetic changes in colon epithelial cells (Okugawa et al. 2015). Aberrant DNA methylation in promoter region of predominantly tumor suppressor genes occurs in the early stages of tumor development in precancerous lesions. This is a well-characterized event in tumor biology and is relevant to CRC development and progression (Grützmann et al. 2008; Mansour 2014; Tanzer et al. 2010).
Changes in methylation could be feasibly detected in both stool and blood-based samples, making these biological markers ideal candidates for a noninvasive test for early detection of CRC (Gonzalez-Pons and Cruz-Correa 2015).
Epi proColon® 2.0 CE is based on methylated Septin9 (SEPT9) gene from the cfDNA in the plasma which is accessible in Europe and different nations such as china (Jin et al. 2015; Lamb and Dhillon 2017). Behrouz Sharif et al. represented that SEPT9 promoter hypermethylation may serve as a promising biomarker for the detection of CRC development (Behrouz Sharif et al. 2016).
Circulating tumor DNA (ctDNA) in plasma or serum could serve as a ‘liquid biopsy’, for detecting abnormal methylation patterns. It offers a possible method for screening of several cancers and avoids the need for tumor tissue biopsies (Elshimali et al. 2013; Mansour 2014; Schwarzenbach et al. 2011; Zou et al. 2007). It has been shown cfDNA in cancer patients is a result of direct shedding from the primary tumor through apoptosis, necrosis or secretion, or potentially originates from free circulating tumor cells (Lange and Laird 2013; Schwarzenbach et al. 2011). DNA methylation patterns measured in peripheral blood not only have great potential to be informative biomarkers of cancer risk but also it is useful as a noninvasive test for CRC screening (Elshimali et al. 2013).
ColoGaurd™ kit is now a FDA approved DNA stool-based CRC screening test, combine aberrant BMP3 and NDRG4 promoter region methylation as well as Kras mutation and fecal immunochemical test (Ahlquist et al. 2012b).
Bone Morphogenetic Protein 3 belongs to the transforming growth factor-beta (TGFB) superfamily also known as osteogenin, which induces bone formation. It was identified as a candidate tumor suppressor gene putatively down-regulated in CRC (Loh et al. 2008). One of the first evidence for the importance of BMP3 inactivation through methylation process in early polyp formation and colorectal tumor development has been published by Loh et al. (2008). Their observation suggested that BMP3 is an attractive target for the future development of molecular blood and/or stool screening tests for the early detection of lesions with neoplastic potential.
The aim of this study was to investigate aberrant DNA hypermethylation of BMP3 gene in plasma of patients with precursor lesions of CRC.
Materials and methods
Study participants
This was a case-control study. Patients with sporadic CRC who participated in this study were recruited consecutively from September 2015 to March 2017. CRC tissues were collected during colonoscopy from 100 patients referred to Reza Radiotherapy and Oncology Center (RROC, Mashhad, Iran). In total, 50 polyp/tumor positive patients and 50 patients with normal colons diagnosed by colonoscopy were enrolled in this study. Histopathology reports were assessed to determine polyp/tumor characteristics. Patients with prior colorectal resection and history of any cancer or chemotherapy or radiation therapy were excluded from this study. In order to reduce bias, we designed this experiment as a blinded assay and samples were randomly coded before processing. All sample collection and preservation were taken care of by an individual who did not participate in the follow-up studies. All patients gave informed written consent to participate and to have their biologic specimens analyzed. The study was approved by the Ethical Committee of Tabriz University of Medical Sciences, Iran.
Collection of plasma
5 ml Peripheral blood was collected from patients and healthy individuals into EDTA tubes and kept at room temperature (18– 22 °C). Plasma was separated by double centrifugation (800 g; 10 min, separation, 1600×g; 10 min), no more than 2 h after blood draw. Plasma aliquots were immediately frozen at − 70 °C because of cfDNA instability.
Cell free DNA extraction
cfDNA purification was performed by the standard Triton/Heat/Phenol protocol (THP) method, which removes proteins from nucleic acids by mixture of phenol–chloroform–isoamyl alcohol. Briefly, in this method 500 μl of plasma was mixed with 5 μl Triton X-100 (Applichem, Germany) and heat denatured at 98 °C for 5 min. Samples were placed on ice for 5 min, then extracted with an equal volume of phenol–chloroform–isoamyl alcohol (25:24:1, v:v:v), saturated with 50 Mm Tris–Cl, pH 8.0 and centrifuged for 10 min at 14,000×g. The aqueous phase was precipitated for 2 h with X2.5 volume of 100% ethanol at − 70 °C. The DNA pellet was washed with 1 ml ethanol 70%, air-dried and re-suspended in 50 μl of AE buffer (10 mM Tris–Cl, 0.5 mM EDTA; pH 9.0) and incubated overnight at 37 °C.
Bisulfite treatment
20 μl extracted cfDNA undergone sodium bisulfite conversion and DNA recovery using the EpiTect Fast Bisulfite Conversion Kits (Qiagen, Germany) according to the manufacturer’s instructions.
Methylation analysis
Methylation analysis was performed by bisulfite specific high resolution melting analysis (BS-HRM). The BS-HRM protocol consists of PCR amplification of bisulfite-modified DNA (Wojdacz and Dobrovic 2007). The primers used to amplify bisulfite-treated DNA were BMP3-F, 5′-GGGTTAGYGTAGTAAGTGGGGTTGG-3′ and BMP3-R, 5′- AACCTACTCRCCCCAACCATAACTAAATACCC-3′, designed to amplify both methylated and unmethylated bisulfite-treated DNA that did not amplify unmodified genomic DNA. These primers located in CpG island region and consisted of 24 CpG sites. Genomic amplicon region of BMP3 is shown in Fig. 1.
PCR amplification and HRM analysis were carried out sequentially on a light Cycler® 96 System (Roche, Germany). PCR was carried out in a 10 µl total volume using HiFiSYBR Green Master Mix (Farabin, Tehran), consisting of 300 nM of each primer, 0.2 µg/µl BSA and 2.5 µl of bisulfite modified template. The amplification run was 15 min at 95 °C, followed by 45 cycles of 20 s 95 °C, 15 s at the primer annealing temperature (60 °C) and 15 s at 72 °C.
HRM analyses were performed at the temperature ramping from 65 to 97 °C. Florescence acquisition setting was carried out at temperature recommended by the manufacturer. The melting curves were normalized by calculation of the ‘line of best fit’ in between two normalization regions before and after the major fluorescence decrease representing the melting of the PCR product using the software version 1.1 provided with the LightCycler® 96 System.
Statistical analysis
The sensitivity and specificity [with 95% confidence interval (CI)] of the BMP3 hyper methylation of cfDNA plasma were calculated. To compare characteristics of the different groups of patients and samples, t test for continue variables, Chi square test and Fisher exact test were used for categorical variables. Statistical analyses were performed using SPSS version 13.0. All values were two-sided and P value < 0.05 was considered to indicate a statistically significant difference.
Results
Patient and lesion characteristics
The clinical characteristics of the 100 patients included in this study was shown in Table 1. There was no significant difference with respect to gender and bone mass index (BMI) between cases and controls (gender: P = 0.54; BMI: P = 0.80). Among the 50 polyps: 26% were located at proximal (ascending colon, hepatic flexure and transverse colon) and 74% were located at distal colon (descending colon, sigmoid, rectum, anal).
BMP3 methylation status
Amongst the 100 cfDNA only five samples were excluded in this study since they were not amplified properly by real time PCR.
Figure 2 illustrates the comparison of the melting profiles of PCR products from samples with profiles specific for PCR products derived from methylated and unmethylated control DNAs.
Our results showed methylated BMP3 test identified18 out of 45 patient plasma samples with a sensitivity of 40% and overall specificity of 94%. Statistical test analysis revealed that BMP3 methylation in plasma was significantly different in patients with control groups (P < 0.05) as shown in Table 2.
Discussion
In this study, we aimed to assess the potential role of aberrant BMP3 promoter methylation changes in cfDNA released by tumor cells in different forms and at different levels in the blood circulation of CRC patients.
We demonstrated that there was significantly a higher frequency (P value < 0.05) of BMP3 methylated DNA in plasma of patients with polyps/ tumor versus healthy individuals with a sensitivity and specificity of 40 and 94%, respectively.
Sensitivity is the main characteristic for screening tests because the major role of a screening test is to rule out diseases such as cancer or precancerous lesions. Although high sensitivity is the most important characteristic of a cancer-screening test, specificity is also important, since it affects the number of individuals who have positive test results (Berger et al. 2016).
Zou et al. (2007) evaluated BMP3 gene methylated on 74 colorectal cancers, 62 adenomas, and 70 normal epithelia tissues. Methylation status was analyzed quantitatively and qualitatively and confirmed by bisulfite genomic sequencing. Methylation of BMP3 was detected in 66 of cancers; 74% of adenomas; and 7% of normal epithelia (P < 0.01, cancer or adenoma versus normal). Loh et al. (2008) observed BMP3 methylation in colorectal polyps and cancers, but not in normal mucosa samples, suggests that this may be an attractive target for the future development of molecular blood and/or stool screening tests for the early detection of lesions with neoplastic potential. Zou et al. (2012) used the QuARTS technology to quantify methylated BMP3 gene on 91 DNA samples extracted from colorectal tissues, including 37 cancers, 25 adenomas, and 29 healthy epithelia. Comparing cancer/adenoma to healthy epithelia, AUC values were 0.89 for BMP3 DNA methylation. Ashktorab et al. (2014) analyzed more than 21,500 CpG Islands methylation status using reduced representation bisulfite sequencing (RRBS) technique in colorectal cancer and adenoma tissues that were compared with DNA methylome from a healthy subject’s colon tissue and peripheral blood DNA. They represented novel DNA methylation in six genes including BMP3 that could be involved in CRC progression as it was significantly hypermethylated in tumor versus normal tissues. Houshmand et al. (2017) studied the methylation status of bone morphogenetic protein 3 (BMP3) gene, in tissue samples from patients with CRC obtained from colorectal surgery. They revealed that BMP3 methylation was detected with a sensitivity of 56.66% and a specificity of 93.3%, indicates its potentiality for early detection of CRC.
These tissue-based studies suggest that BMP3 DNA methylation could be a potential biomarker for early detection of CRCs. Since released cfDNA in blood and DNA extracted from exfoliated gastrointestinal epithelial cells in stool reflects genomic alterations, using blood and stool samples could be beneficial sources to detect cancer (Galanopoulos et al. 2017; Park et al. 2017).
In Park et al. (2017) study, bisulfate-modified stool DNA obtained from 36 patients with advanced adenoma; 35 patients with CRC; and 40 endoscopically diagnosed healthy controls using CRC screening colonoscopy. Methylated BMP3 were detected in 40.0% of CRC samples and in 33.3% of advanced adenoma samples with the specificity of 85%.
Ahlquist et al. reported sensitivity at 87% and specificity at 93% for methylated BMP3/NDRG4/VIM/TFPI2 in stool DNA samples. In other study, Ahlquist et al. showed the sensitivity 85% and specificity 89% for VIM/NDRG4/BMP3/TFPI2 genes for stool specimens Based on these studies and further investigations ColoGaurd™ kit was designed (Zhai et al. 2016).
The specificity degree of BMP3 methylation (of colorectal cancerous cells) in stool or plasma is influenced by the BMP3 methylation status of background normal DNA. The specificity and sensitivity of BMP3 methylation (of cancerous cells) has been only studied in stool and tissue and there is no evidence for plasma. To address this issue, the authors studied the plasma BMP3 DNA methylation.
The relatively low sensitivity in this study could be due to different reasons. First, the very low concentration and fragmented cfDNA in plasma. Second, cfDNA with genetic and epigenetic alterations can be mixed by normal free DNA, released in the bloodstream (1.0% of total cfDNA) (Danese et al. 2015; Diaz and Bardelli 2014). Third, tumors, themselves are a mixture of different cancer cell clones (inter-tumoral heterogeneity) which could lead to more complexity (Kamat et al. 2006). Fourth, having a large number of potential heteroduplexes generated by heterogeneous methylated CpG-rich amplicons is a challenge in BSP-HRM. It is difficult to compare the multifaceted melting HRM profile of heterogeneous methylated DNA samples with homogenous methylated and unmethylated controls.
In conclusion, our results demonstrated that BMP3 DNA methylation in plasma had not have sufficient sensitivity and it should be used in combination with other biomarkers for the detection of CRC.
References
Ahlquist DA, Taylor WR, Mahoney DW, Zou H, Domanico M, Thibodeau SN, Boardman LA, Berger BM, Lidgard GP (2012a) The stool DNA test is more accurate than the plasma septin 9 test in detecting colorectal neoplasia. Clin Gastroenterol Hepatol 10:272–277.e271
Ahlquist DA, Zou H, Domanico M, Mahoney DW, Yab TC, Taylor WR, Butz ML, Thibodeau SN, Rabeneck L, Paszat LF et al (2012b) Next-generation stool DNA test accurately detects colorectal cancer and large adenomas. Gastroenterology 142:248–226
Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F (2017) Global patterns and trends in colorectal cancer incidence and mortality. Gut 66:683–691
Ashktorab H, Daremipouran M, Goel A, Varma S, Leavitt R, Sun X, Brim H (2014) DNA methylome profiling identifies novel methylated genes in African American patients with colorectal neoplasia. Epigenetics 9:503–512
Behrouz Sharif S, Hashemzadeh S, Mousavi Ardehaie R, Eftekharsadat A, Ghojazadeh M, Mehrtash AH, Estiar MA, Teimoori-Toolabi L, Sakhinia E (2016) Detection of aberrant methylated SEPT9 and NTRK3 genes in sporadic colorectal cancer patients as a potential diagnostic biomarker. Oncol Lett 12:5335–5343
Berger BM, Levin B, Hilsden RJ (2016) Multitarget stool DNA for colorectal cancer screening: a review and commentary on the United States preventive services draft guidelines. World J Gastrointest Oncol 8:450–458
Danese E, Minicozzi AM, Benati M, Montagnana M, Paviati E, Salvagno GL, Lima-Oliveira G, Gusella M, Pasini F, Lippi G et al (2015) Comparison of genetic and epigenetic alterations of primary tumors and matched plasma samples in patients with colorectal cancer. PLoS ONE 10:e0126417
Diaz LA Jr, Bardelli A (2014) Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol 32:579–586
Elshimali YI, Khaddour H, Sarkissyan M, Wu Y, Vadgama JV (2013) The clinical utilization of circulating cell free DNA (CCFDNA) in blood of cancer patients. Int J Mol Sci 14:18925–18958
Galanopoulos M, Tsoukalas N, Papanikolaou IS, Tolia M, Gazouli M, Mantzaris GJ (2017) Abnormal DNA methylation as a cell-free circulating DNA biomarker for colorectal cancer detection: a review of literature. World J Gastrointest Oncol 9:142–152
Gonzalez-Pons M, Cruz-Correa M (2015) Colorectal cancer biomarkers: where are we now?. Biomed Res Int 2015:149014
Grützmann R, Molnar B, Pilarsky C, Habermann JK, Schlag PM, Saeger HD, Miehlke S, Stolz T, Model F, Roblick UJ et al (2008) Sensitive detection of colorectal cancer in peripheral blood by septin 9 DNA methylation assay. PLoS ONE 3:e3759
Haggar FA, Boushey RP (2009) Colorectal cancer epidemiology: incidence, mortality, survival, and risk factors. Clin Colon Rectal Surg 22:191–197
Hessami Arani S, Kerachian MA (2017) Rising rates of colorectal cancer among younger Iranians: is diet to blame? Curr Oncol 24:e131–e137
Houshmand M, Abbaszadegan MR, Kerachian MA (2017) Assessment of bone morphogenetic protein 3 methylation in Iranian patients with colorectal cancer. Middle East J Dig Dis 9:158–163
Jin P, Kang Q, Wang X, Yang L, Yu Y, Li N, He Y, Han X, Hang J, Zhang J (2015) Performance of a second-generation methylated SEPT9 test in detecting colorectal neoplasm. J Gastroenterol Hepatol 30:830–833
Kamat AA, Bischoff FZ, Dang D, Baldwin MF, Han LY, Lin YG, Merritt WM, Landen CN Jr, Lu C, Gershenson DM et al (2006) Circulating cell-free DNA: a novel biomarker for response to therapy in ovarian carcinoma. Cancer Biol Ther 5:1369–1374
Kim HJ, Yu MH, Kim H, Byun J, Lee C (2008) Noninvasive molecular biomarkers for the detection of colorectal cancer. BMB Rep 41:685–692
Lamb YN, Dhillon S (2017) Epi proColon® 2.0 CE: a blood-based screening test for colorectal cancer. Mol Diagn Ther 21:225–232
Lange CP, Laird PW (2013) Clinical applications of DNA methylation biomarkers in colorectal cancer. Epigenomics 5:105–108
Lange CP, Campan M, Hinoue T, Schmitz RF, van der Meulen-de Jong AE, Slingerland H, Kok PJ, van Dijk CM, Weisenberger DJ, Shen H et al (2012) Genome-scale discovery of DNA-methylation biomarkers for blood-based detection of colorectal cancer. PLoS ONE 7:e50266
Levin B, Lieberman DA, McFarland B, Smith RA, Brooks D, Andrews KS, Dash C, Giardiello FM, Glick S, Levin TR et al. (2008) Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: a joint guideline from the american cancer society, the US multi-society task force on colorectal cancer, and the american college of radiology. CA Cancer J Clin 58:130–160
Loh K, Chia JA, Greco S, Cozzi SJ, Buttenshaw RL, Bond CE, Simms LA, Pike T, Young JP, Jass JR et al (2008) Bone morphogenic protein 3 inactivation is an early and frequent event in colorectal cancer development. Genes Chromosomes Cancer 47:449–460
Mansour H (2014) Cell-free nucleic acids as noninvasive biomarkers for colorectal cancer detection. Front Genet 5:182
Okugawa Y, Grady WM, Goel A (2015) Epigenetic alterations in colorectal cancer: emerging biomarkers. Gastroenterology 149:1204–1225.e1212
Park SK, Baek HL, Yu J, Kim JY, Yang HJ, Jung YS, Choi KY, Kim H, Kim HO, Jeong KU et al (2017) Is methylation analysis of SFRP2, TFPI2, NDRG4, and BMP3 promoters suitable for colorectal cancer screening in the Korean population? Intest Res 15:495–501
Schwarzenbach H, Hoon DS, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 11:426–437
Tanzer M, Balluff B, Distler J, Hale K, Leodolter A, Rocken C, Molnar B, Schmid R, Lofton-Day C, Schuster T et al (2010) Performance of epigenetic markers SEPT9 and ALX4 in plasma for detection of colorectal precancerous lesions. PLoS ONE 5:e9061
Wojdacz TK, Dobrovic A (2007) Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res 35:e41
Zhai RL, Xu F, Zhang P, Zhang WL, Wang H, Wang JL, Cai KL, Long YP, Lu XM, Tao KX et al (2016) The diagnostic performance of stool DNA testing for colorectal cancer: a systematic review and meta-analysis. Medicine (Baltimore) 95:e2129
Zou H, Harrington JJ, Shire AM, Rego RL, Wang L, Campbell ME, Oberg AL, Ahlquist DA (2007) Highly methylated genes in colorectal neoplasia: implications for screening. Cancer Epidemiol Biomark Prev 16:2686–2696
Zou H, Allawi H, Cao X, Domanico M, Harrington J, Taylor WR, Yab T, Ahlquist DA, Lidgard G (2012) Quantification of methylated markers with a multiplex methylation-specific technology. Clin Chem 58:375–383
Acknowledgements
This study was supported financially by Tabriz University of Medical Sciences, and Reza Radiotherapy and Oncology Center, Mashhad, Iran. Our sincere thanks also go to Mr. Ebrahim Pouladin and Mrs. Nafiseh Shalchi for their close support in CRC research programs. We would also give special thanks to Dr. Abdorasoul Hayatbakhsh, Miss. Maryam Yassi and Adeleh Rezaie, Mrs. Neda Ziafati, and Zohreh Alizadeh for their cooperation in this study.
Funding
This study was supported financially by Tabriz University of Medical Sciences (Grant # 54125388), and Reza Radiotherapy and Oncology Center, Mashhad, Iran.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consents were obtained from all individual participants included in this study.
Rights and permissions
About this article
Cite this article
Rokni, P., Shariatpanahi, A.M., Sakhinia, E. et al. BMP3 promoter hypermethylation in plasma-derived cell-free DNA in colorectal cancer patients. Genes Genom 40, 423–428 (2018). https://doi.org/10.1007/s13258-017-0644-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-017-0644-2