Abstract
Korean native cattle, known as Hanwoo, have been raised in the Korean Peninsula since 2000 B.C. for use as a draft animal. However, Hanwoo now have an important position in the Korean livestock industry as a meat source. Therefore, the breeding and selection of Hanwoo are crucial for the industry. Although many researchers have studied the genetic architecture of Hanwoo, no well-established Hanwoo-related databases exist. In order to better understand the genetic contents of Hanwoo, an integrated database is necessary. We constructed a genetic variants database including annotation information. HanwooGDB (http://hanwoogdb.snu.ac.kr) provides genetic variants (SNPs, INDELs, CNVs) in the Hanwoo genome produced by Next Generation Sequencing data collected from 23 cattle. The identified SNPs were integrated with SNP chip data and annotation information for checking the concordance of position of each SNP and inferring functional aspects. This database provides browsers to understand and visualize the comprehensive information of these variants and allows users to download data according to their preference from this database without limitation. This database will contribute to genetic research and development of Hanwoo breeding strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Korean native cattle, Hanwoo have been raised in the Korean Peninsula since 2000 B.C. During the ancient Agricultural Age, Hanwoo were primarily used as a draft animal or sometimes as an object of sacrificial rite, and rarely consumed as meat due to religious and political issues in Korea and the limited cattle population. The production of Hanwoo as meat cattle begun in the 1960s with the rapid growth of the Korean economy. Thus, eating Hanwoo beef as a main meat source has a short history (Jo et al. 2012). Despite their brief history, Hanwoo is now regarded as a premium beef and a national favorite, despite its relative high price, and the breed has been relatively well-maintained. Hanwoo, now, have substantial commercial value in the Korean livestock industry (Hwang et al. 2010).
In the livestock industry, genetic research has contributed to optimization of breeding, productivity, and product quality. Furthermore, the importance and utility of genetics have been increasing due to improved technologies and data analyses techniques. Since Hanwoo have maintained stable traits through pure breeding, the current blood lineage is valuable and spread out mainly in the Korean Peninsula (Kim and Lee 2000). Additionally, Hanwoo occupy a substantial proportion of the livestock industry in Korea making the genetic information important for the Korean livestock industry.
In the case of cattle (Bos Taurus), a whole genome database is already in existence; the Bovine Genome Database (BGD, http://bovinegenome.org) is a genome informatics resource for Bos Taurus which provides data through genome browsers, BLAST searching, and gene query interfaces. It also displays gene predictions, non-coding RNA, repeats, pseudogenes, SNPs, markers, QTL, and alignments of protein homologs and transcript sequences (Reese et al. 2010). Moreover, a SNP annotation tool for cattle also exists, and this tool provides annotation information for SNPs by integrating various sources of genomic information such as Entrez Gene and UniProt (Jiang et al. 2011).
Genetic specification of Hanwoo traits is needed to maintain the breed. Research on the genetic properties of Hanwoo has been conducted by various groups and fragments of information for genetic variants of Hanwoo have been produced. For example, a number of studies on expressed genes related to meat quality of Hanwoo have been conducted. Researchers have identified 14 candidate genes related to intramuscular fat development (Lee et al. 2007) and analyzed 11 candidate genes related to marbling in Hanwoo by RT-PCR (Lee et al. 2008a). Genetic variants (SNPs) of Hanwoo have been studied as well. A previous study attempted to identify SNP markers to estimate marbling score and meat quality grade for Hanwoo (Shin et al. 2012). This study revealed that fatty acid binding protein 4 (FABP4) gene known to be related to intramuscular fat has three SNPs and one of those was related to marbling score and meat quality for Hanwoo. Additionally, 44 SNPs related to carcass quality traits were identified using DNA chips in Hanwoo populations (Lee et al. 2010). In addition, to detect the origin of beef, a study was conducted to identify SNP markers that characterize the Hanwoo breed. In this study, researchers identified 90 SNP markers that characterize Hanwoo breed (Cheong et al. 2013). Although many studies on genetic information of Hanwoo have been conducted, a whole genome database of Hanwoo produced by Next Generation Sequencing (NGS) technology that offers easy access to genetic information for the researchers and breeders has yet to be developed.
HanwooGDB provides SNP data for the whole genome of Hanwoo produced by NGS technology. Since this database also provides SNP chip data of reference sequences, it can be used to check the concordance of position of each SNP by NGS technology. It provides a number of statistics related to data production and validation, which can be accessed by all users, as well as includes annotation information of genes for each SNP and SNP data for each individual. Thus, it could provide inference to help understand the functional aspects of these variants and the specific characters of Hanwoo breed. Almost all of the information in this database can be downloaded in vcf format, which is easily applicable to other data for both researchers and breeders. Moreover, this database provides browsers to help understand the comprehensive information of these variants and visualize the database. This database will contribute to genetic researches and breeding strategies of Hanwoo.
Materials and Methods
Sample preparation
Blood samples were collected from 12 Hanwoo cattle from Kyungpook National University, Korea, and 11 Hanwoo cattle from the National Institute of Animal Science, Korea. Samples (10 ml) were drawn from the carotid artery and treated with heparin to prevent clotting. Genomic DNA samples were extracted from blood samples using a G-DEXTMIIb Genomic DNA Extraction Kit (iNtRoN Biotechnology, Seoul, Korea). Randomly sheared 3 μg of genomic DNA was used to generate DNA fragments about 300 bp by the Covaris System. The fragments of sheared DNA were passed through preparation processes for sequencing including end-repaired, A-tailed, adaptor ligated, and amplified using a TruSeq DNA Sample Prep. Kit (Illumina, San Diego, CA). For prepared samples, paired-end sequencing was conducted by National Instrumentation Center for Environmental Management (NICEM), Seoul, Korea, using the Illumina HiSeq 2000 platform with TruSeq SBS Kit v3-HS (Illumina). Here, we performed sequencing on one lane per one sample. As a result of the sequencing process, we acquired 42,534,069,646 paired-end reads in the average of 23 Hanwoo cattle and the mean coverage ratio was 15.93.
Sequence alignment
Obtained sequences were aligned to the Bos Taurus 6 (UMD 3.1) reference genome using Bowtie 2 (Langmead and Salzberg 2012). Next, four open-source software tools were used for further processing and variant calling for SNPs and INDELs: Picard tools (http://picard.sourceforge.net), SAMtools 0.1.18 (Li et al. 2009), VCFtools 4.0 (Danecek et al. 2011) and the Genome Analysis Toolkit 1.4 (McKenna et al. 2010). First, duplicate reads were removed using Picard tools MarkDuplicates. Then, to index the resulting bam format files and calculate the mapped read length with the flagstat option (Li et al. 2009) SAMtools was used. Realignments were performed using GATK (McKenna et al. 2010) Count-Covariates, RealignerTargetCreator, and IndelRealigner. VCFtools was used to treat the vcf file format (Danecek et al. 2011).
Genotype calling and phasing
Variant calling was conducted through GATK Select-Variant, VariantFiltration, and UnifiedGenotyper (McKenna et al. 2010). SNPs and INDELs with a Phred-scaled quality score less than 30 were removed and variants with MQ0 (median quality score zero) bigger than 4, MQ0/read depth bigger than 10 %, quality depth less than 5, and FS (Phred-scaled P value using Fisher’s exact test to detect strand bias) bigger than 200 were removed. After screening, the SNPs within 10 bp of INDELs were removed again (Fig. 1).
Procedure of producing HanwooGDB SNPs and INDELs. Re-sequencing and SNP calling procedures are shown. Sequences obtained from samples were aligned to the reference genome Bos taurus 6 (UMD 3.1) using Bowtie 2 (Langmead and Salzberg 2012). Removing duplicates and indexing were performed by Picard tools (http://picard.sourceforge.net) and SAMtools 0.1.18 (Li et al. 2009). Realignment and variant calling for SNPs and INDELs were performed by the Genome Analysis Toolkit 1.4 (McKenna et al. 2010). VCFtools 4.0 (Danecek et al. 2011) were used for handling vcf file format
We used BEAGLE (Browning and Browning 2007) to infer the haplotype phase and impute missing alleles for the entire set of cattle populations. CNVs in the population were discovered and genotyped by using Genome STRucture in Population (Genome STRiP). It utilizes an integrative approach detecting breakpoint-spanning reads, abnormal read-pair separation, and local variation in read depth as CNV signatures. It also combines these signatures with population-scale patterns that span many genomes (Handsaker et al. 2011). SNP chip data was gained from BovineHD Genotyping BeadChip data generated by Illumina (ftp.illumina.com/Whole%20Genome%20Genotyping%20Files/). This site is only available through password provided by Illumina customer service. Annotation information was obtained from Ensembl 67 database (UMD_3.1) (Flicek et al. 2012) with SnpEff version 3.0 (Cingolani et al. 2012). Statistics including Tajima D, Theta, Pi, and Het were obtained by Arlequin (Excoffier and Lischer 2010) and visualized through Gbrowser.
Construction of database
HawooGDB was constructed not only to play a role as a repository of information but also to allow users to search and utilize the stored information. Hardware based on Intel Xeon CPU and openSUSE 12.2 O/S was used to construct this database. In order to manage a large amount of data, sqlite 3.7 and MySQL 5.5.28 were used. In addition to this, php5.3 and javascript were used to connect these with the web server (Apache2.2.22). Data was processed using python 2.7. We used Php-gd package and Gbrowser UCSC for visualization.
Results
Contents of the HanwooGDB
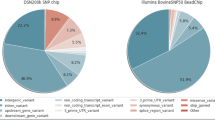
HanwooGDB is a web based database on genetic variants of Hanwoo and is available at http://hanwoogdb.snu.ac.kr. HanwooGDB provides four menus for genetic information of Hanwoo: ‘Data & Procedure’, ‘Statistics’, ‘Browser’ and ‘Download’. On the ‘Data & Procedure’ menu, users can see the summary of the database and an overview of the data production procedure with a figure of the work flow. It provides overall information about the database, samples, and schematic procedure. The ‘Statistics’ menu consists of ‘Variant’ and ‘Annotation’ pages. The ‘Variant’ page provides a table and graphs of statistical information of SNPs and INDELs: variant number of each chromosome, variant number, and density distribution according to chromosome length. The ‘Annotation’ page provides annotation information of variants, transition and transversion ratio of variant site, the number of known variant sites, and length distribution of small INDELs. The ‘Browser’ menu provides visualized and annotated information and statistics of genetic variant region produced by Arlequin with various filter options. Users can download annotation information of genetic variants in vcf format from the ‘Download’ menu (Fig. 2).
The home of HanwooGDB. HanwooGDB is available at http://hanwoogdb.snu.ac.kr. There are seven menus on the top of the webpage: Home, Data & Procedure, Statistics, Browser, Download, FAQ and Contact Us. Users can search data by typing the gene name, RSID or chromosome region. On the middle of the webpage, there is a brief summary of the database and other information including ‘Search example’, ‘What’s new’ and ‘Database information.’ On the bottom of the webpage, there is ‘Karyotype navigator’ by which users can directly select a chromosomal location they want to display in the browser page
Annotation information of SNPs on browser
The ‘Browser’ page under the ‘Browser’ menu provides visualized and annotation information of Hanwoo SNPs. This browser support visualizing only information of SNPs but not for INDELs and CNVs. Users can search a region, which contains SNPs by the gene name, SNP ID or chromosome region. Since it displays SNPs with a gene as a unit, one of genes which will be shown above the browser should be selected if the users search SNPs with a SNP ID and chromosome region. Furthermore, users can directly access Ensembl annotation database by clicking on the gene name. With the browser, users can see the location of the SNPs on a chromosome and a gene. By using the filter options, users can visualized different types of SNPs on the browser with different colors. In addition, SNP chip data can be also shown with or without SNP data by NGS technology. More information about a given region on Gbrowser can be accessed by clicking on the “View Detail Information by Gbrowse” button or the gene region in the figure. Detail information of SNPs, which are shown on browser are also provided in the form of tables: a position of SNPs, the difference between reference base and alternative base, Phred scaled probability (QUAL), allele count in the population (AC), effects predicted by snpEff, effects categorized by ‘impact’ (High, Moderate, Low, and Modifier), Transcript ID, and RSID. More information about each transcript and RSID with Ensembl and NCBI database can be accessed by clicking the IDs on the table. Users can directly access the browser page, which shows the chromosomal location they selected on the Karyotype navigator on ‘Home’ menu (Fig. 3a).
The webpages to provide Hanwoo genetic variants data a ‘Browser’ page. Users can see the SNP data they searched on this page. This page consists of three sections. On the filter option section, users can select types of SNPs and determine whether SNP chip data will be shown on the browser. On the browser panel, users can see visualized locations of SNPs. The SNP detail info part section provides tables that contain detail information of SNPs. b ‘Gbrowser’ page. This page consists of five tabs: Browser, Select tracks, Snapshots, Custom tracks, Preferences. On the ‘Browser’ tab, users can see graphs of statistics related to genomic feature of a certain region users have searched. The ‘Select tracks’ tab and ‘Preferences’ tab allow users to customize Gbrowser. On the ‘Snapshots’ tab, users can take and load a snapshot of graphs resulted from a search. ‘Custom tracks’ tab allows users to upload custom track data in a variety of quantitative and non-quantitative formats. c ‘Download’ menu. On this menu, users can export data on genetic variants in vcf file format. Users can assign the location and the type of genetic variant. Moreover, four filter options provide criteria of filtering: General option, Quality option, Sample option and Output option
Genomic features on Gbrowser
The ‘Gbrowser’ page within the ‘Browser’ menu provides visualized data and interactive interface for statistics related to a certain region users have searched. If users type a certain chromosome range and click the ‘search’ button, general genomic features including CDS and Transcript region and statistics produced by Arlequin (Tajima D, Theta, Pi and Het values) are exhibited on each track of Gbrowser with distinctive colors. On each track, users can check the details of each statistic by clicking the graph and move to another region of interests using the scroll interface or drag. In addition, it is possible to control the scale of the region user are interested in by the zoom interface and change the direction of the region by clicking the ‘Flip’ button. The sequence files can be downloaded with a few options: decorated FASTA file and raw sequence file. Presenting annotating restriction sites based on selected restriction enzyme and downloading track data are also possible.
There are a few tabs including options that can be used for customizing obtained results. On the ‘Select tracks’ and ‘Preferences’ tabs, users can select tracks which will be shown on the browser and control visualization options such as grid option and image width. On the ‘Snapshot’ tab, a snapshot of the browser, which contains tracks about statistics can be taken and downloaded in image files. Moreover, users can load snapshots they have taken before. The ‘Custom tracks’ tab allows users to upload custom track data in a variety of quantitative and non-quantitative formats. Users can also import data sets that reside on network-connected servers by referring to their URLs (Fig. 3b).
Download data files with various filters
HanwooGDB provides data files of genetic variants in.vcf format. When users download data files, they can select a location and type of genetic variants (SNP, INDEL and CNV). In filter option section, there are four steps that users can determine. Step 1 provides options for the kind and effect of SNP. In Step 2, users can filter out genetic variants that didn’t satisfy the criteria they assigned. In Step 3, users can select individual samples that would be included in data files. Finally, Step 4 determines the separator by which the data set will be separated. It is possible to see the data files on the web and download the data files simultaneously (Fig. 3c).
Discussions
Values of HanwooGDB
HanwooGDB is a web-based database for genetic variants information of Korean native cattle, Hanwoo. This is the first NGS technology database produced for the Hanwoo breed. Therefore, this database itself is of immeasurable value in Korea. Furthermore, SNP data can be used as a detector of economic traits and breed. Studies on these which has been conducted (Edea et al. 2012; Lee et al. 2008a, b) will progress further in future. Therefore, to provide SNP data files as well as the raw sequence files can be a considerable help to researchers who are studying Hanwoo. Researchers will be able to obtain these data files through HanwooGDB easily.
SNP chip data and annotation information, also provided by HanwooGDB, help to check the concordance of position of each SNP and infer functional aspects. Statistics related to these data can also help to acquire genomic feature information. Moreover, interactive and visualized interface will be able to help researchers utilize this database conveniently. Finally, unlike other databases, this database provides data sets of individual samples. This feature can guide researchers who need specific data.
Future work
Since this database is limited to Hanwoo, it is necessary to have more information to characterize Hanwoo breed. Although, in the ‘Browser’ tab, result tables give reference bases and allele count score; however, these are insufficient. Statistics or visualized charts for entire samples for Hanwoo breed are in need. In addition, updating data of new Hanwoo samples will enhance the confidence of this database.
Given the considerable economic value of Hanwoo as livestock in Korea, annotation information related to economic traits such as body weights and fat development is extremely valuable. An integrated database between information related to economic traits based on up-to-date studies and genomic database (gene, SNPs, INDEL etc.) will support future research. Consequently, this database will have further value if it can provide integrated information regarding economic traits of Hanwoo with the genomic information. For example, when users search a certain gene containing SNPs, it will provide information of economic traits related to that gene by listing related articles.
References
Browning SR, Browning BL (2007) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81:1084–1097
Cheong HS, Kim LH, Namgoong S, Shin HD (2013) Development of discrimination SNP markers for Hanwoo (Korean native cattle). Meat Sci 94:355–359
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6:80–92
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Edea Z, Dadi H, Kim SW, Dessie T, Kim KS (2012) Comparison of SNP variation and distribution in indigenous Ethiopian and Korean Cattle (Hanwoo) populations. Genomics Inform 10:200–205
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Flicek P, Amode MR, Barrell D, Beal K, Brent S, Carvalho-Silva D, Clapham P, Coates G, Fairley S, Fitzgerald S (2012) Ensembl 2012. Nucleic Acids Res 40:D84–D90
Handsaker RE, Korn JM, Nemesh J, McCarroll SA (2011) Discovery and genotyping of genome structural polymorphism by sequencing on a population scale. Nat Genet 43:269–276
Hwang YH, Kim GD, Jeong JY, Hur SJ, Joo ST (2010) The relationship between muscle fiber characteristics and meat quality traits of highly marbled Hanwoo (Korean native cattle) steers. Meat Sci 86:456–461
Jiang J, Jiang L, Zhou B, Fu W, Liu JF, Zhang Q (2011) Snat: a SNP annotation tool for bovine by integrating various sources of genomic information. BMC Genet 12:85
Jo C, Cho SH, Chang J, Nam KC (2012) Keys to production and processing of Hanwoo beef: a perspective of tradition and science. Anim Front 2:32–38
Kim JB, Lee C (2000) Historical look at the genetic improvement in Korean cattle. Rev Asian Australas J Anim Sci 13:1467–1481
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Lee SH, Park EW, Cho YM, Kim SK, Lee JH, Jeon JT, Lee CS, Im SK, Oh SJ, Thompson JM (2007) Identification of differentially expressed genes related to intramuscular fat development in the early and late fattening stages of hanwoo steers. J Biochem Mol Biol 40:757–764
Lee SH, Cho YM, Lee SH, Kim BS, Kim NK, Choy YH, Kim KH, Yoon DH, Im SK, Oh SJ et al (2008a) Identification of marbling-related candidate genes in M. longissimus dorsi of high- and low marbled Hanwoo (Korean Native Cattle) steers. BMB Rep 41:846–851
Lee YS, Lee JH, Lee JY, Kim JJ, Park HS, Yeo JS (2008b) Identification of candidate SNP (single nucleotide polymorphism) for growth and carcass traits related to QTL on chromosome 6 in Hanwoo (Korean cattle). Asian Aust J Anim Sci 21:1703–1709
Lee YM, Han CM, Li Yi, Lee JJ, Kim LH, Kim JH, Kim DI, Lee SS, Park BL, Shin HD (2010) A whole genome association study to detect single nucleotide polymorphism for carcass traits in Hanwoo populations. Asian Australas J Anim Sci 23:417–424
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Reese JT, Childers CP, Sundaram JP, Dickens CM, Childs KL, Vile DC, Elsik CG (2010) Bovine Genome Database: supporting community annotation and analysis of the Bos taurus genome. BMC Genomics 11:645
Shin SC, Heo JP, Chung ER (2012) Genetic variants of the FABP4 gene are associated with marbling scores and meat quality grades in Hanwoo (Korean cattle). Mol Biol Rep 39:5323–5330
Acknowledgments
This study was supported by grants from Research Program (Project No: PJ008487) of the National Livestock Research Institute and Next Generation BioGreen 21 Program (Project No: PJ008191 and PJ00995401) of Rural Development Administration, Republic of Korea.
Conflict of Interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Kwondo Kim and Woori Kwak have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Kim, K., Kwak, W., Sung, SS. et al. A novel genetic variant database for Korean native cattle (Hanwoo): HanwooGDB. Genes Genom 37, 15–22 (2015). https://doi.org/10.1007/s13258-014-0224-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-014-0224-7