Abstract
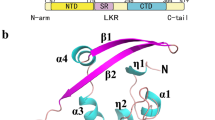
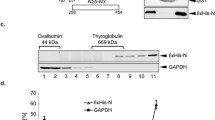
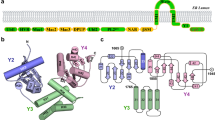
Coronaviruses are the causative agent of respiratory and enteric diseases in animals and humans. One example is SARS, which caused a worldwide health threat in 2003. In coronaviruses, the structural protein N (nucleocapsid protein) associates with the viral RNA to form the filamentous nucleocapsid and plays a crucial role in genome replication and transcription. The structure of Nterminal domain of MHV N protein also implicated its specific affinity with transcriptional regulatory sequence (TRS) RNA. Here we report the crystal structures of the two proteolytically resistant N- (NTD) and C-terminal (CTD) domains of the N protein from murine hepatitis virus (MHV). The structure of NTD in two different crystal forms was solved to 1.5 Å. The higher resolution provides more detailed structural information than previous reports, showing that the NTD structure from MHV shares a similar overall and topology structure with that of SARS-CoV and IBV, but varies in its potential surface, which indicates a possible difference in RNA-binding module. The structure of CTD was solved to 2.0-Å resolution and revealed a tightly intertwined dimer. This is consistent with analytical ultracentrifugation experiments, suggesting a dimeric assembly of the N protein. The similarity between the structures of these two domains from SARS-CoV, IBV and MHV corroborates a conserved mechanism of nucleocapsid formation for coronaviruses.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Almazán, F., Galán, C., and Enjuanes, L. (2004). The nucleoprotein is required for efficient coronavirus genome replication. J Virol 78, 12683–12688.
Almazán, F., González, J.M., Pénzes, Z., Izeta, A., Calvo, E., Plana-Durán, J., and Enjuanes, L. (2000). Engineering the largest RNA virus genome as an infectious bacterial artificial chromosome. Proc Natl Acad Sci U S A 97, 5516–5521.
Bailey, S., and the Collaborative Computational Project, Number 4. (1994). The CCP4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr 50, 760–763.
Baric, R.S., Nelson, G.W., Fleming, J.O., Deans, R.J., Keck, J.G., Casteel, N., and Stohlman, S.A. (1988). Interactions between coronavirus nucleocapsid protein and viral RNAs: implications for viral transcription. J Virol 62, 4280–4287.
Bednar, V., Verma, S., Blount, A., and Hogue, B.G. (2006). Importance of MHV-CoVA59 nucleocapsid protein COOH-terminal negative charges. Adv Exp Med Biol 581, 127–132.
Bost, A.G., Carnahan, R.H., Lu, X.T., and Denison, M.R. (2000). Four proteins processed from the replicase gene polyprotein of mouse hepatitis virus colocalize in the cell periphery and adjacent to sites of virion assembly. J Virol 74, 3379–3387.
Casais, R., Thiel, V., Siddell, S.G., Cavanagh, D., and Britton, P. (2001). Reverse genetics system for the avian coronavirus infectious bronchitis virus. J Virol 75, 12359–12369.
Chang, C.K., Hsu, Y.L., Chang, Y.H., Chao, F.A., Wu, M.C., Huang, Y. S., Hu, C.K., and Huang, T.H. (2009). Multiple nucleic acid binding sites and intrinsic disorder of severe acute respiratory syndrome coronavirus nucleocapsid protein: implications for ribonucleocapsid protein packaging. J Virol 83, 2255–2264.
Chen, C.Y., Chang, C.K., Chang, Y.W., Sue, S.C., Bai, H.I., Riang, L., Hsiao, C.D., and Huang, T.H. (2007). Structure of the SARS coronavirus nucleocapsid protein RNA-binding dimerization domain suggests a mechanism for helical packaging of viral RNA. J Mol Biol 368, 1075–1086.
Coley, S.E., Lavi, E., Sawicki, S.G., Fu, L., Schelle, B., Karl, N., Siddell, S.G., and Thiel, V. (2005). Recombinant mouse hepatitis virus strain A59 from cloned, full-length cDNA replicates to high titers in vitro and is fully pathogenic in vivo. J Virol 79, 3097–3106.
Cologna, R., Spagnolo, J.F., and Hogue, B.G. (2000). Identification of nucleocapsid binding sites within coronavirus-defective genomes. Virology 277, 235–249.
DeLano, W. (2002). The PyMOL Molecular Graphics System. DeLano Scientific, Palo Alto, CA, USA.
Drosten, C., Günther, S., Preiser, W., van der Werf, S., Brodt, H.R., Becker, S., Rabenau, H., Panning, M., Kolesnikova, L., Fouchier, R.A., et al. (2003). Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med 348, 1967–1976.
Emsley, P., and Cowtan, K. (2004). Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, 2126–2132.
Fan, H., Ooi, A., Tan, Y.W., Wang, S., Fang, S., Liu, D.X., and Lescar, J. (2005). The nucleocapsid protein of coronavirus infectious bronchitis virus: crystal structure of its N-terminal domain and multimerization properties. Structure 13, 1859–1868.
Fleischauer, A.T., and the Centers for Disease Control and Prevention (CDC). (2003). Outbreak of severe acute respiratory syndrome—worldwide, 2003. MMWR Morb Mortal Wkly Rep 52, 226–228.
Grossoehme, N.E., Li, L., Keane, S.C., Liu, P., Dann, C.E. 3rd, Leibowitz, J.L., and Giedroc, D.P. (2009). Coronavirus N protein Nterminal domain (NTD) specifically binds the transcriptional regulatory sequence (TRS) and melts TRS-cTRS RNA duplexes. J Mol Biol 394, 544–557.
Huang, Q., Yu, L., Petros, A., Gunasekera, A., Liu, Z., Xu, N., Hajduk, P., Mack, J., Fesik, S., and Olejniczak, E. (2004). Structure of the N-Terminal RNA-Binding domain of the SARS CoV nucleocapsid protein. Biochemisty 43, 6059–6063.
Hurst, K.R., Kuo, L., Koetzner, C.A., Ye, R., Hsue, B., and Masters, P. S. (2005). A major determinant for membrane protein interaction localizes to the carboxy-terminal domain of the mouse coronavirus nucleocapsid protein. J Virol 79, 13285–13297.
Janin, J., and Chothia, C. (1990). Minireview:The Structure of Protein-Protein Recognition Sites. J Biol Chem 265, 16221–16224.
Jayaram, H., Fan, H., Bowman, B.R., Ooi, A., Jayaram, J., Collisson, E.W., Lescar, J., and Prasad, B.V.V. (2006). X-ray structures of the N- and C-terminal domains of a coronavirus nucleocapsid protein: implications for nucleocapsid formation. J Virol 80, 6612–6620.
Ksiazek, T.G., Erdman, D., Goldsmith, C.S., Zaki, S.R., Peret, T., Emery, S., Tong, S., Urbani, C., Comer, J.A., Lim, W., et al, and the SARS Working Group. (2003). A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med 348, 1953–1966.
Kuo, L., and Masters, P.S. (2002). Genetic evidence for a structural interaction between the carboxy termini of the membrane and nucleocapsid proteins of mouse hepatitis virus. J Virol 76, 4987–4999.
Lai, M.M., and Stohlman, S.A. (1978). RNA of mouse hepatitis virus. J Virol 26, 236–242.
Laskowski, R.A., Moss, D.S., and Thornton, J.M. (1993). Main-chain bond lengths and bond angles in protein structures. J Mol Biol 231, 1049–1067.
Luo, H., Chen, J., Chen, K., Shen, X., and Jiang, H. (2006). Carboxyl terminus of severe acute respiratory syndrome coronavirus nucleocapsid protein: self-association analysis and nucleic acid binding characterization. Biochemisty 45, 11827–11835.
Macneughton, M.R., and Davies, H.A. (1978). Ribonucleoprotein-like structures from coronavirus particles. J Gen Virol 39, 545–549.
Masters, P.S. (1992). Localization of an RNA-binding domain in the nucleocapsid protein of the coronavirus mouse hepatitis virus. Arch Virol 125, 141–160.
McCoy, A.J., Grosse-Kunstleve, R.W., Adams, P.D., Winn, M.D., Storoni, L.C., and Read, R.J. (2007). Phaser crystallographic software. J Appl Cryst 40, 658–674.
Mir, M.A., and Panganiban, A.T. (2006). Characterization of the RNA chaperone activity of hantavirus nucleocapsid protein. J Virol 80, 6276–6285.
Molenkamp, R., and Spaan, W.J.M. (1997). Identification of a specific interaction between the coronavirus mouse hepatitis virus A59 nucleocapsid protein and packaging signal. Virology 239, 78–86.
Nelson, G.W., Stohlman, S.A., and Tahara, S.M. (2000). High affinity interaction between nucleocapsid protein and leader/intergenic sequence of mouse hepatitis virus RNA. J Gen Virol 81, 181–188.
Otwinowski, Z., and Minor, W. (1997). Processing of X-ray diffraction data collected in the oscillation mode. Methods Enzymol 276, 307–326.
Perrakis, A., Morris, R.M., and Lamzin, V.S. (1999). Automated protein model building combined with iterative structure refinement. Nat Struct Biol 6, 458–463.
Robbins, S.G., Frana, M.F., McGowan, J.J., Boyle, J.F., and Holmes, K.V. (1986). RNA-binding proteins of coronavirus MHV: detection of monomeric and multimeric N protein with an RNA overlayprotein blot assay. Virology 150, 402–410.
Rota, P., Oberste, M., Monroe, S., Nix, W., Campagnoli, R., Icenogle, J., Peñaranda, S., Bankamp, B., Maher, K., Chen, M., et al. (2003). Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 300, 1394–1399.
Saikatendu, K.S., Joseph, J.S., Subramanian, V., Neuman, B.W., Buchmeier, M.J., Stevens, R.C., and Kuhn, P. (2007). Ribonucleocapsid formation of severe acute respiratory syndrome coronavirus through molecular action of the N-terminal domain of N protein. J Virol 81, 3913–3921.
Sawicki, S.G., Sawicki, D.L., Younker, D., Meyer, Y., Thiel, V., Stokes, H., and Siddell, S.G. (2005). Functional and genetic analysis of coronavirus replicase-transcriptase proteins. PLoS Pathog 1, e39.
Schuck, P. (2000). Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys J 78, 1606–1619.
Stohlman, S.A., Baric, R.S., Nelson, G.N., Soe, L.H., Welter, L.M., and Deans, R.J. (1988). Specific interaction between coronavirus leader RNA and nucleocapsid protein. J Virol 62, 4288–4295.
Sturman, L.S., and Holmes, K.V. (1983). The molecular biology of coronaviruses. Adv Virus Res 28, 35–112.
Takeda, M., Chang, C.K., Ikeya, T., Güntert, P., Chang, Y.H., Hsu, Y. L., Huang, T.H., and Kainosho, M. (2008). Solution structure of the c-terminal dimerization domain of SARS coronavirus nucleocapsid protein solved by the SAIL-NMR method. J Mol Biol 380, 608–622.
Terwilliger, T.C. (2000). Maximum-likelihood density modification. Acta Crystallogr D Biol Crystallogr 56, 965–972.
Terwilliger, T.C., and Berendzen, J. (1999). Automated MAD and MIR structure solution. Acta Crystallogr D Biol Crystallogr 55, 849–861.
van der Meer, Y., Snijder, E.J., Dobbe, J.C., Schleich, S., Denison, M. R., Spaan, W.J., and Locker, J.K. (1999). Localization of mouse hepatitis virus nonstructural proteins and RNA synthesis indicates a role for late endosomes in viral replication. J Virol 73, 7641–7657.
Verma, S., Bednar, V., Blount, A., and Hogue, B.G. (2006). Identification of functionally important negatively charged residues in the carboxy end of mouse hepatitis coronavirus A59 nucleocapsid protein. J Virol 80, 4344–4355.
You, J.-H., Reed, M.L., and Hiscox, J.A. (2007). Trafficking motifs in the SARS-coronavirus nucleocapsid protein. Biochem Biophys Res Commun 358, 1015–1020.
Yu, I.M., Oldham, M.L., Zhang, J., and Chen, J. (2006). Crystal structure of the severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein dimerization domain reveals evolutionary linkage between corona- and arteriviridae. J Biol Chem 281, 17134–17139.
Zhang, X., Liao, C.-L., and Lai, M.M. (1994). Coronavirus leader RNA regulates and initiates subgenomic mRNA transcription both in trans and in cis. J Virol 68, 4738–4746.
Zúñiga, S., Sola, I., Moreno, J.L., Sabella, P., Plana-Durán, J., and Enjuanes, L. (2007). Coronavirus nucleocapsid protein is an RNA chaperone. Virology 357, 215–227.
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to the work.
Rights and permissions
About this article
Cite this article
Ma, Y., Tong, X., Xu, X. et al. Structures of the N- and C-terminal domains of MHV-A59 nucleocapsid protein corroborate a conserved RNA-protein binding mechanism in coronavirus. Protein Cell 1, 688–697 (2010). https://doi.org/10.1007/s13238-010-0079-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-010-0079-x