Abstract
The phylogenetic relationships of Lasiosphaeriaceae are complicated in that the family is paraphyletic and includes Sordariaceae and Chaetomiaceae, as well as several polyphyletic genera. This study focuses on the phylogenetic relationships of the coprophilous genera, Anopodium, Apodospora, Arnium, Fimetariella and Zygospermella. They are traditionally circumscribed based on ascospore characters, which have proven homoplasious in other genera within the family. Our results based on LSU nrDNA and ß–tubulin sequences distinguish four lineages of Lasiosphaeriaceae taxa. Anopodium joins the clade of morphologically similar, yellow-pigmented species of Cercophora and Lasiosphaeria. Apodospora is monophyletic and joins a larger group of taxa with unclear affinities to each other, while Arnium is polyphyletic being scattered throughout three of the four major clades of Lasiosphaeriaceae. Fimitariella is represented by a single collection and joins the clade containing Cercophora scortea and Podospora appendiculata. Zygospermella shows affinities to the Lasiosphaeris clade. Based on a combination of morphological and molecular data, Echria stat. nov. is recognized at the genus level for the former Arnium section and two new combinations are proposed: E. gigantospora and E. macrotheca.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sordariales is a large group of microfungi that occur worldwide as degraders of dung, wood, plant material, and soil (Cannon and Kirk 2007). The order includes several well-known members including species of Chaetomium that are common indoor contaminants associated with high humidity, and the model organisms Neurospora crassa, Podospora anserina, Sordaria fimicola, and S. macrospora (Alexopoulos et al. 1996). Uniting morphological features of Sordariales species are the erumpent to superficial, perithecial ascomata with large-celled ascomal walls, as well as the predominantly one or two-celled ascospores formed in unitunicate asci which show a remarkable morphological diversity (Huhndorf et al. 2004). The ascospores have hyaline and/or dark brown cells that typically form an apical head and basal tail when two-celled, and commonly are surrounded by a gelatinous sheath or have appendages in various sizes and shapes (von Arx et al. 1986; Lundqvist 1972). Most species have smooth ascospores, but distinctive ornamentations occur in some taxa (Dettman et al. 2001; Garcia et al. 2004; Lundqvist 1967).
Huhndorf et al. (2004) reduced Sordariales to the present three families, Chaetomiaceae, Lasiosphaeriaceae and Sordariaceae, and the order forms a monophyletic clade in phylogenies based on molecular data (Cai et al. 2006; Miller and Huhndorf 2005; Zhang et al. 2006). However, Sordariales relationships at the family and generic level are complicated. Chaetomiaceae and Sordariaceae are nested within the largest family Lasiosphaeriaceae (Greif et al. 2009; Huhndorf et al. 2004; Réblová 2008), and several of the 31 genera that belong to Lasiosphaeriaceae are polyphyletic, i.e. Cercophora, Podospora, Triangularia, and Zopfiella (Cai et al. 2006; Chang et al. 2010; Miller and Huhndorf 2005).
The circumscriptions of genera have traditionally been based on the morphology of the ascospores, but molecular results have shown that ascospore characters clearly are unreliable for interpreting relationships at the family and generic level (Miller and Huhndorf 2005). Attempts have been made to group taxa based on other characters, such as those pertaining to the ascomal wall morphology, which show better agreement with molecular data for some clades (Miller and Huhndorf 2005). For example, members of Cercophora and Lasiosphaeria with an outer wall layer of hyphae forming a tomentum, group together with strong support. Unfortunately, a large number of the species have relatively simple ascomal walls without obvious characters with which to distinguish them. There are also several well-supported clades that include morphologically dissimilar species (Miller and Huhndorf 2005).
Delimiting genera within Lasiosphaeriaceae is obviously very difficult and developing a stable taxonomic outline of the group is a continuously evolving project. A number of genera currently placed in Lasiosphaeriaceae have not yet been studied with molecular methods and their addition to the Sordariales phylogeny would strengthen the understanding of intra- and interfamilial relationships. The primary purpose of this study is to further resolve the phylogenetic relationships of Lasiosphaeriaceae and its allied taxa, with a focus on five coprophilous genera: Anopodium, Apodospora, Arnium, Fimetariella, and Zygospermella.
Methods
Taxon sampling
We assembled 62 newly generated sequences from 23 species and eight genera currently classified in the Lasiosphaeriaceae plus 126 GenBank (http://www.ncbi.nlm.nih.gov) sequences representing various members of Sordariomycetes. A representative of Xylariomycetidae (Eutypa sp.) was used as an outgroup based on results from previous phylogenetic analyses (Huhndorf and Miller 2011; Zhang et al. 2006). We compiled a combined data set with 117 LSU nrDNA sequences and 71 ß-tubulin sequences. Collection data and GenBank accession numbers for the sequenced specimens are listed in Table 1, Electronic Supplementary Material.
DNA extraction, PCR amplification and sequencing
DNA was extracted from ascomata or cultures using the EZNA Fungal DNA kit (Omega bio-tek, Doraville, Georgia, USA). The protocol was used according to the manufacturer’s instructions, except that the material was homogenized for 3 min using a Mini-Beadbeater (BioSpec Products, Bartlesville, Oklahoma, USA) with Buffer FG1 and 2.5 mm Zirconia/Silica beads (BioSpec Products, Bartlesville, Oklahoma, USA). Double-stranded copies of the LSU and ß-tubulin were obtained by polymerase chain reaction (PCR) amplifications using AccuPowerTM PCR premix (BioNeer, Alameda, California, USA) to which DNA extract, primers (10 μM) and water were added. The DNA was amplified with primers LR0R/LR7 (R. Vilgalys website), and BT1819R/BT2916 (Miller and Huhndorf 2005), while thermocycler conditions followed Miller and Huhndorf (2005). PCR-products were cleaned with MultiScreen HTS™ PCR plates (Millipore, Cork, Ireland). Macrogen Ltd (Seoul, Korea) performed sequencing reactions using the following primers: LR0R, LR5, LR3R, LRAM1, LR3, LR7; BT1819R, BT1283R, BTAM1R, BT2916 (Huhndorf et al. 2004; Miller & Huhndorf 2005; R. Vilgalys website). Sequence fragments were assembled and edited in Sequencher 4.8 (Gene Codes Corp., Ann Arbor, Michigan, USA).
Sequence alignment
Alignment of each set of sequences were done in Geneious Pro 5.6.5 (Biomatters Ltd, Auckland, New Zealand), using the MAFFT application (Katoh et al. 2002). We used algorithm E-INS-i, scoring matrix 100PAM/k = 2, gap open penalty 1.53 and offset value 0 for the ß-tubulin data set, and algorithm G-INS-i, scoring matrix 100PAM/k = 2, gap open penalty 1.53 and offset value 0.123 for the LSU data set. We excluded parts of the 5′ and 3′ ends, plus one intron from the analyses of the ß-tubulin data set (96 characters), and two ambiguous indel sections of the LSU data set (259 characters). The alignment is deposited in TreeBASE (accession number S14831).
Phylogenetic analyses
We used PartitionFinder 1.0.1 (Lanfear et al. 2012) to select the best partitioning scheme and best-fit models of nucleotide evolution. The partitioning scheme with the best likelihood, according to the Baysian information criterion, separated the data sets among genes and among nucleotide positions in the codons for the ß-tubulin gene. The best-fit evolutionary models were the SYM + I + Γ model for ß-tubulin codon position 1, the JC + I model for ß-tubulin codon position 2, and the GTR + I + Γ model for ß-tubulin codon position 3 and LSU. Bayesian Markov Chain Monte Carlo (MCMC) analyses were conducted in MrBayes 3.2 (Ronquist and Huelsenbeck 2003) and performed using resources at SNIC through Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX). MCMC sampling was performed with four parallel runs, six chains, sampling every 1000th tree, and the temperature set to 0.25. Sampling was halted when the critical value for the topological convergence diagnostic had reached 0.01. Log files from the Bayesian analyses were analyzed with Tracer v 1.5 (Rambaut and Drummond 2009) to determine the burn-in value, and a 50 % majority rule consensus tree was generated in mrBayes using the remaining trees. We evaluated the contribution of individual genes and the amount of incongruence among them by comparing significantly supported clades from separate Bayesian analyses with two parallel runs and eight chains (Mason-Gamer and Kellogg 1996; de Queiroz 1993). Significant support in this study refers to Bayesian posterior probability values (BPP) ≥95 % and bootstrap values (BS) ≥70 % (Alfaro et al. 2003; Hillis and Bull 1993). We also conducted a Maximum Likelihood search with rapid bootstrap analysis for the best tree with the GTR + Γ model for all partitions combined with 5,000 bootstrap replicates in RAxML 7.2.6 (Stamatakis 2006; Stamatakis et al. 2008).
Results
Sequence alignment and phylogenetic analyses
Our combined ß-tubulin and LSU alignment included 2,408 (984 + 1,424) characters. The number of parsimony informative sites compared to variable sites was 257/316 (81 %) for the ß-tubulin partition and 215/335 (64 %) for the LSU.
The MCMC analysis of the two combined genes proceeded for 17.45 × 106 generations, and resulted in 69 824 trees. The first 2 500 trees per run were discarded as burn-in, and the consensus tree was generated using the remaining 59 824 trees (Fig. 1). We found no significant incongruence in the tree topologies of the separate gene analyses. The MCMC analysis of LSU resulted in 34 051 trees per run and 48 102 were included in a consensus tree. The ß-tubulin analysis produced 104 401 trees per run and 158 802 were used to build the consensus tree. The significantly supported topology of the best ML tree coincides with the Bayesian tree (data not shown).
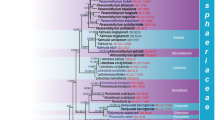
The Bayesian tree is a 50 % majority rule consensus of 59 824 combined trees of the ß-tubulin and LSU nrDNA for 119 taxa. Thickened branches have both significant BPP values (≥95 %) and BS values (≥70 %). Numbers above the branches indicate significant support values in the following order: BPP/BS, and support values below these thresholds are indicated by a dash. Outgroup is Eutypa sp. Sordariales include four clades of Lasiosphaeriaceae taxa (I-IV) a. Lasiosphaeriaceae clade I and II in focus. b. Lasiosphaeriaceae clade III and IV in focus
Phylogenetic relationships
Sordariales is monophyletic with high posterior probability support, while the largest family Lasiosphaeriaceae is paraphyletic and includes Sordariaceae and Chaetomiaceae (Fig. 1). We distinguish four core lineages of Lasiosphaeriaceae taxa (clades I-IV) with only clade IV having both significant BPP and BS support. Clades I, II and III are unsupported sister taxa. Clade I includes representatives of eleven genera of Lasiosphaeriaceae (Fig. 1a). In clade II, the monophyletic Sordariaceae and one member of Chaetomiaceae are nested among members of eight genera of Lasiosphaeriaceae. Clade III includes members of seven genera of Lasiosphaeriaceae with significant BPP support (Fig. 1b). Clade IV is sister group to the basal Chaetomiaceae and includes members of five Lasiosphaeriaceae genera with significant BS and BPP support.
The main focus of this study, the genera Anopodium, Apodospora, Arnium, Fimetariella and Zygospermella, belong to different clades in the tree. The 26 representatives of Arnium are scattered in clades I, II and IV. The three Apodospora species form a well-supported monophyletic group in clade II where the single representative of Fimetariella is also placed. Anopodium belongs to clade III and Zygospermella to clade I.
Taxonomy
Based on a combination of morphological and molecular data, Arnium section Echria is raised to genus level and two species are transferred as new combinations.
Echria (N. Lundq.) Kruys, Huhndorf and A.N. Mill., stat. nov. MycoBank #: MB808571.
≡ Arnium section Echria N. Lundq. Symb. Bot. Upsal. 20(1): 237. 1972 (basionym).
Echria gigantospora (Raja and Shearer) Kruys, Huhndorf and A.N. Mill., comb. nov. MycoBank #: MB808572.
≡ Arnium gigantosporum Raja and Shearer, Fungal Diversity 22: 220. 2006 (basionym).
Echria macrotheca (P. Crouan and H. Crouan) Kruys, Huhndorf and A.N. Mill., comb. nov.
MycoBank #: MB808573.
≡ Sphaeria macrotheca P. Crouan and H. Crouan, Florule Finistère (Paris): 24. 1,867 (basionym).
Note: Echria gigantospora and E. macrotheca are morphologically similar in that both have ascomata covered with long pointed tufts of agglutinated rigid hairs and striated gelatinous sheaths that swell in water and surround their ascospores. They differ in that E. gigantospora has larger (84–126 × 20–34 μm), fusiform spores with a roughened spore wall (Raja and Shearer 2006), while E. macrotheca (Fig. 3o-q) has smaller (43–54 × 20–29 μm), ellipsoidal spores (Lundqvist 1972). They also seem to prefer different substrates as E. gigantospora is described from submerged wood, while E. macrotheca is coprophilous.
Discussion
The main topology of our Sordariales tree is consistent with previous phylogenies in which Chaetomiaceae and Sordariaceae are nested within the paraphyletic Lasiosphaeriaceae (Miller and Huhndorf 2005; Réblová 2008). Although our study includes additional taxa, the clades that Miller and Huhndorf (2005) distinguished based on the morphology of the ascomal walls remain unchanged. We reveal several new significantly supported clades, but the major challenge is still to find morphological features to characterize them when monophyletic genera are mostly the exception rather than the rule.
Anopodium
Lundqvist (1964a) established the genus Anopodium for its unique uniseptate ascospores with one dark brown basal cell and a hyaline apSical pedicel (Fig. 2a-d). They resemble Podospora spores except the cells are in a reversed position (Lundqvist 1964a). Two collections of the generic type, Anopodium ampullaceum, were included in our molecular analyses and their closest relative is Cercophora sulphurella in clade III (Fig. 1b). Both species join the weakly supported clade containing C. sparsa, Corylomyces selenosporus, Bellojisia rhynchostoma and Podospora didyma. The sister taxa of this extended Anopodium clade are Lasiosphaeria species. The latter all have a similar three-layered ascomal wall in which the outer wall layer is composed of hyphae that form a tomentum (Miller and Huhndorf 2005). Cercophora sulphurella fits morphologically into this group with its typical yellowish-green tomentum, along with B. rhynchostoma (Réblová 2008) and C. selenosporus (Stchigel et al. 2006). Corylomyces selenosporus has olivaceous–yellow, tomentose perithecia and olivaceous-yellow mycelium (Stchigel et al. 2006), and B. rhynchostoma has perithecia with a yellowish inconspicuous tomentum and a subiculum of yellowish to pale olivaceous-brown hyphae around the base (Réblová 2008). Anopodium species on the other hand have glabrous or hairy perithecia, as do P. didyma and C. sparsa. A feature that is shared by both the extended Anopodium clade and the Lasiosphaeria clade is the presence of yellow pigment. It occurs in the membranaceous, transparent peridium of both A. ampullaceum (Lundqvist 1964b) and P. didyma (Mirza and Cain 1969). Cercophora sparsa and C. sulphurella both have yellow centrum pigments (Hilber and Hilber 1979).
Anopodium ampullaceum, a ascoma, b immature ascospores in the ascus, c ascospore, d ascospores position in the ascus. (Coll. Kruys 699, UPS). Apodospora simulans, e ascoma, f ascoma, g ascus, h ascospore with gelatinous sheath. (e Coll. TRTC156582, TRTC. f –h Coll. Kruys 701, UPS). Fimetariella rabenhorstii, i ascomata, j ascoma, k ascus with ascospores. (i–j Coll. F-644,827, UPS. k Coll. TRTC156796, TRTC). Zygospermella insignis. l ascoma, m ascospore with gelatinous appendages. (l Coll. Kruys 718, UPS. m Coll. TRTC156807, TRTC). Scale bars: a, e-f, i-j, l = 200 μm, b-d, g-h, k, m = 20 μm
Within this extended Anopodium clade are taxa exemplifying a range of ascospore types: Podospora didyma with the typical dark brown apical cell and a hyaline basal pedicel, Anopodium with inverted Podospora-like spores, Cercophora spores with a long basal pedicel, and the dark brown navicular, lunate to reniform ascospores of Corylomyces and Bellojisia.
Apodospora
The genus comprises species that have one-celled, dark brown ascospores with an apical germ pore and a gelatinous sheath and asci with an apical ring (Fig. 2e-h) (Bell et al. 2008; Cain and Mirza 1970; Lundqvist 1972). The ascospore morphology is reminiscent of Sordaria but Apodospora differs in the apical location of the germ pore as well as an ascomatal peridium composed of smaller-sized cells (Lundqvist 1972). The species included in our study (A. gotlandica, A. peruviana and the generic type, A. simulans) have ascomata that are all more or less glabrous with a reddish-brown peridium. The genus forms a well-supported monophyletic group in clade II but without higher-level support (Fig. 1a). The unsupported clade II is an assemblage of taxa in a number of morphologically diverse genera that often have branch support for several species. Molecular data confirms the morphological viewpoint given by Lundqvist (1972) that Apodospora does not belong within the Sordariaceae. Additional taxa included in the genus based on morphology are A. viridis, a species with yellow-green peridium and papillae around the neck, as well as A. bulgarica, characterized by olive-brown peridium and rigid hairs around the neck. Their inclusion in the genus must be further investigated with molecular data.
Arnium
The genus Arnium is morphologically very diverse, and therefore not surprisingly, found in three of the four clades in the phylogenetic tree (Fig. 1, Fig. 3). The ascomatal peridium varies from membranaceous and light coloured to coriaceous or carbonaceous and opaque in different species. Various hairs or bristles may cover the ascomata, and short inconspicuous tubercles occur around the neck in certain species (Krug and Cain 1972; Lundqvist 1972). Some species have asci with an apical ring. The ascospores are dark brown, single-celled or with a transverse septum and often provided with one gelatinous appendage at each end or covered with a striated sheath (Lundqvist 1972).
Arnium olerum , a ascoma, b paraphyses, c asci, d immature hyaline ascospore with gelatinous appendages, e ascospores. (Coll. SMH3253 3, F). A. arizonense, f ascoma, g ascus with ascospores. (f Coll. F66864, S, g Coll. TRTC38138, TRTC). A .mendax , h ascoma, i immature ascospore with gelatinous appendages, j ascus, k ascospores, (h Coll. F66872, S, i-k Coll. TRTC156621, TRTC). A. leporinum , l ascoma, m asci and ascospores, n immature ascospore with gelatinous appendages. (l Coll. F-007,119, UPS, m-n Coll. TRTC156605, TRTC). Echria macrotheca , o ascoma, p asci, q ascospore with gelatinous sheath. (o Coll. F-644,828, UPS, p-q Coll. TRTC54023, TRTC). Scale bars: a, f, h, l, o = 200 μm, b-e, g,i-k, m-n, p-q = 20 μm
Included in the analyses are two Arnium species with perithecia forming a tomentum, the type species A. olerum (Fig. 3a-e) and A. tomentosum. Both species belong to clade IV (Fig. 1b) and are related to the tomentose species C. coprophila and C. grandiuscula. Arnium olerum is characterized by a greyish-whitish tomentum, which is also found in C. coprophila and C. grandiuscula, while A. tomentosum has an olivaceous-brown tomentum. We have included two representatives of A. olerum, one of which, (SMH3253.3) is a sister taxon to A. tomentosum. The molecular data suggest that either these two species may be one entity or one of these specimens is misidentified. Given the morphological and apparent molecular similarity of these two species, additional collections should be sought for sequencing, perhaps including the RPB2 gene in a potentially more-informative dataset. The other A. olerum (1), originating from CBS 120,012, groups with Cercophora grandiuscula (CBS 120,013).
Arnium arizonense also belongs to clade IV (Fig. 1b, 3f-g) and four representatives of the species form a well-supported clade. It clusters within an internally unresolved but significantly supported clade of Apiosordaria backusii, A. verruculosa, Cercophora samala, C. striata, Podospora austroamericana, P. comata, and P. pauciseta. This mixture of taxa contains a variety of morphological characteristics with no discernable pattern of similarities. The well-supported sister group to this overall clade is a clade comprising two Cercophora species and the type species of Podospora, P. fimiseda. Miller and Huhndorf (2005) distinguished this group of taxa with ascomata possessing pseudo-bombardioid walls. Arnium ontariense, a taxon sharing a similar wall structure, most likely belongs in this clade, however, no sequences and no recent collections of this species were available for inclusion in this study.
Several Arnium species have ascomata that are either nearly glabrous or have flexuous hairs, and the ones included in our study (A. caballinum, A. inaequilaterale, A. japonense, A. mendax, A. sudermanniae) all occur in an unsupported group in clade II (Fig. 1a). Within this clade, Arnium mendax (Fig. 3h-k) and A. inaequilaterale form a moderately-supported sister clade to two taxa distinguished by their areolate peridia, Cercophora ambigua and C. areolata, although this distinctive wall structure is not found in these two Arnium species. Arnium caballinum and A. sudermanniae occur in a well-supported clade with three collections of Arnium leporinum and a collection named as A. japonense. Arnium leporinum differs morphologically from the former two Arnium species by ascomata that are covered with rigid hairs around the neck and multi-spored asci with a range of 64–128 ascospores (Fig. 3l-n). The original description gives A. japonense as having ascomata without neck hairs, 256 spores, and occurring on hare dung (Furuya and Udagawa 1976). From the molecular data it appears that the A. japonense specimen from mouse dung is actually A. leporinum.
Four collections of Arnium hirtum are found on two separate branches within clade I (Fig. 1a). The two representatives from Scotland are strongly supported in a clade with Immersiella caudata and I. immersa. They fit Lundqvist’s (1972) description of A. hirtum well by having ascospores with the same large variation in size, rigid ascomatal hairs and a few swollen cells in the neck region, unlike the palisade of slightly swollen cells found in the neck region of I. caudata and I. immersa, which give these two species a sulcate appearance (Miller and Huhndorf 2004). The two North American specimens of A. hirtum are sister taxa to Podospora intestinacea and differ from the reported description of A. hirtum by having ascomata with longer necks that lack hairs, but are similar in having ascospores with inflated or distally swelling appendages. In both of these characteristics, the North American specimens of A. hirtum are similar to P. intestinacea. To clarify which of these two sets of A. hirtum specimens represent the taxon and which characteristics indicate relatedness, additional collections from the type locality should be sequenced. Arnium cirriferum also belongs to clade I, but due to low support values its closest relatives are uncertain (Fig. 1a).
The majority of species in Arnium have spores with two gelatinous appendages, one at each end of the spore. Lundqvist (1972) established the name Echria for a section of Arnium species that instead have spores covered with a fringed sheath or numerous filaments. We include two representatives of this group, which resolve in clade I (Fig. 1a). Since these two species are separate from the generic type species, Arnium olerum (clade IV), we recognize a separate genus utilizing the existing section name. Therefore we raise Section Echria to genus level and transfer the two species as new combinations. Comparing the two species, Echria gigantospora has larger, fusiform spores, with a roughened spore wall, and is described from submerged wood, while E. macrotheca (Fig. 3o-q) has smaller, ellipsoidal spores, and is coprophilous (Raja and Shearer 2006). The two species form a strongly supported monophyletic clade, with E. gigantospora nested within E. macrotheca, suggesting they may constitute a single species. They are morphologically similar in that both have ascomata covered with long pointed tufts of agglutinated rigid hairs. Additional species like A. ditremum may belong to Echria based on morphological similarities, but these taxonomic changes await the future addition of molecular data.
Fimetariella
Lundqvist (1964b) and Krug (1995) circumscribed the genus mainly based on the characteristic one-celled, dark brown ascospores with a gelatinous sheath, two terminal germ pores, and several scattered smaller pores (Fig. 2i-k). The ascospores are four or eight in the ascus and frequently develop with half the number of spores reversed in position with respect to the germ pore. Our phylogeny places Fimetariella rabenhorstii on a long branch in clade II with significant support as a sister taxon to Podospora appendiculata and Cercophora scortea (Fig. 1a). Both P. appendiculata and C. scortea have pseudo-bombardioid walls with short, brown, hyaline-tipped setae (Miller and Huhndorf 2005), and they show no obvious similarity to the sometimes white-powdered, ± glabrous, membranaceous peridium of F. rabenhorstii. Also found in clade II is Bombardioidea, which occurs on an unsupported sister clade to F. rabenhorstii. Bombardioidea also possesses small, minor ascospore pores similar to those of F. rabenhorstii (Lundqvist 1972) making their putative, relative positions in the phylogeny intriguing. Lundqvist (1972) speculated of a potential relationship between the two taxa based on similarities in long-stipitate asci, ascospore pores and filiform paraphyses and the data suggests this possibility.
Zygospermella
The characteristics of Zygospermella are perithecia with pointed, verrucose setae around the neck, a pseudoparenchymatous peridium with scattered reddish crust in the outer layer, and dark brown, two-celled ascospores with gelatinous caudae (Fig. 2l-m) (Lundqvist 1969). Similar neck setae are found in a few other taxa including Zygospermella striata and Cercophora muskokensis and on the surface of the ascomata in Arnium imitans, none of which were available for sequencing. Three collections of the type species, Z. insignis were included in our analyses and they find placement in clade I grouping with moderate support with Cercophora sordarioides (Fig. 1a). This clade is a moderately supported sister group to the Lasiosphaeris clade. While ascomata with pointed setae are found in Lasiosphaeris hispida, they differ from the setae in Zygospermella by being smooth and mostly covering the entire ascomata. The sister taxon, C. sordarioides, lacks setae and shows no obvious similarities.
References
Alexopoulos CJ, Mims CW, Blackwell M (1996) Introductory mycology. Wiley, New York
Alfaro ME, Zoller S, Lutzoni F (2003) Bayes or bootstrap? A simulation study comparing the performance of Bayesian Markov Chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Mol Biol Evol 20:255–266
von Arx JA, Guarro J, Figueras MJ (1986) The ascomycete genus Chaetomium. Beih Nova Hedwig 84:1–162
Bell A, Mahoney D, Debuchy R (2008) A new record of Apodospora from Australia, a rarely collected coprophilous ascomycete. Aust Mycol 27:136–140
Cai L, Jeewon R, Hyde KD (2006) Molecular systematics of Zopfiella and allied genera: evidence from multi-gene sequence analysis. Mycol Res 110:359–368
Cain RF, Mirza JH (1970) Apodospora, a new genus of the Sordariaceae. Can J Bot 48:891–896
Cannon PF, Kirk PM (2007) Fungal families of the world. CABI, Wallingford
Chang J-H, Kao H-W, Wang Y-Z (2010) Molecular phylogeny of Cercophora, Podospora, Schizothecium (Lasiosphaeriaceae, Pyrenomycetes). Taiwania 55:110–116
Dettman JR, Harbinski FM, Taylor JW (2001) Ascospore morphology is a poor predictor of the phylogenetic relationships of Neurospora and Gelasinospora. Fungal Genet Biol 34:49–61
Furuya K, Udagawa S (1976) Coprophilous pyrenomycetes from Japan IV. T Mycol Soc Jpn 17:248–261
Garcia D, Stchigel AM, Cano J, Guarro J, Hawksworth DL (2004) A synopsis and re-circumscription of Neurospora (syn. Gelasinospora) based on ultrastructural and 28S rDNA sequence data. Mycol Res 108:1119–1142
Greif MD, Stchigel AM, Miller AN, Huhndorf SM (2009) A re-evaluation of genus Chaetomidium based on molecular and morphological characters. Mycologia 101:554–564
Hilber R, Hilber O (1979) Einige Anmerkungen zu der Gattung Cercophora Fuckel (Lasiosphaeriaceae). Z Mykol 45:209–233
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192
Huhndorf SM, Miller AM, Fernández FA (2004) Molecular systematics of the Sordariales: the order and the family Lasiosphaeriaceae redefined. Mycologia 96:368–387
Huhndorf SM, Miller AM (2011) A molecular re-appraisal of taxa in the Sordariomycetidae and a new species of Rimaconus from New Zealand. Stud Mycol 68:203–210
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Krug JC (1995) The genus Fimetariella. Can J Bot 73:1905–1916
Krug JC, Cain RF (1972) Additions to the genus Arnium. Can J Bot 50:367–373
Lanfear R, Calcott B, Ho S, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701
Lundqvist N (1964a) Anopodium, a new genus of coprophilous Pyrenomycetes with apically pedicellate spores. Bot Notiser 117:355–365
Lundqvist N (1964b) Fimetariella, a new genus of coprophilous Pyrenomycetes. Bot Notiser 117:238–248
Lundqvist N (1967) On spore ornamentation in the Sordariaceae, exemplified by the new cleistocarpous genus Copromyces. Ark Bot, Ser 2, 6:327–337
Lundqvist N (1969) Zygopleurage and Zygospermella (Sordariaceae s. lat., Pyrenomycetes). Bot Notiser 122:353–374
Lundqvist N (1972) Nordic Sordariaceae s. lat. Symb Bot Ups 20:1–374
Mason-Gamer RJ, Kellogg EA (1996) Testing for phylogenetic conflict among molecular data sets in the tribe Triticeae (Gramineae). Syst Biol 45:524–545
Miller AN, Huhndorf SM (2004) A natural classification of Lasiosphaeria based on nuclear LSU rDNA sequences. Mycol Res 108:26–34
Miller AN, Huhndorf SM (2005) Multi-gene phylogenies indicate ascomal wall morphology is a better predictor of phylogenetic relationships than ascospore morphology in the Sordariales (Ascomycota, Fungi). Mol Phylogenet Evol 35:60–75
Mirza JH, Cain RF (1969) Revision of the genus Podospora. Can J Bot 47:1999–2048
de Queiroz A (1993) For consensus (sometimes). Syst Biol 42:368–372
Raja HA, Shearer CA (2006) Arnium gigantosporum, a new ascomycete species from fresh water in Florida. Fungal Divers 22:219–225
Rambaut A, Drummond AJ (2009) Tracer v1.5.0. Available from http://beast.bio.ed.ac.uk
Réblová M (2008) Bellojisia, a new sordariaceous genus for Jobellisia rhynchostoma and a description of Jobellisiaceae fam. nov. Mycologia 100:893–901
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Stamatakis A, Hoover P, Rougemont J (2008) A Rapid Bootstrap Algorithm for the RAxML Web Servers. Syst Biol 57:758–771
Stchigel AM, Cano J, Miller AN, Calduch M, Guarro J (2006) Corylomyces: a new genus of Sordariales from plant debris in France. Mycol Res 110:1361–1368
Vilgalys R. Conserved primer sequences for PCR amplification and sequencing from nuclear ribosomal RNA. Duke University, USA. Internet address: http://www.biology.duke.edu/fungi/mycolab/primers.htm
Zhang N, Castlebury LA, Miller AN, Huhndorf SM, Schoch CL, Seifert KA, Rossman AY, Rogers JD, Kohlmeyer J, Volkmann-Kohlmeyer B, Sung GH (2006) An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny. Mycologia 98:1076–1087
Acknowledgment
This study is financed by the Swedish Taxonomy Initiative (dha 34/07 1.4) and the Helge Ax:son Johnson foundation. This work was supported in part by a NSF PEET Grant (DEB–9,521,926) to SMH and some sequences were generated in the Pritzker Laboratory for Molecular Systematics and Evolution at The Field Museum of Natural History.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kruys, Å., Huhndorf, S.M. & Miller, A.N. Coprophilous contributions to the phylogeny of Lasiosphaeriaceae and allied taxa within Sordariales (Ascomycota, Fungi). Fungal Diversity 70, 101–113 (2015). https://doi.org/10.1007/s13225-014-0296-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-014-0296-3