Abstract
Xerophilic Aspergillus molds isolated from halo-alkaliphilic and dry environments are attractive genetic resources for obtaining salt- and osmo-adaptive enzymes. A. sydowii MA0196 secreted the largest amount of γ-glutamyl transpeptidase (GGT) during solid-state fermentation at a low initial water activity (aw = 0.85). Gel filtration analysis revealed that the molecular mass of the purified native enzyme (MA0196 GGT) was 120 kDa. SDS-PAGE analysis showed that MA0196 GGT consists of two subunits with molecular masses of 56.4 and 33 kDa, indicating production from a proenzyme via autoproteolysis. Deglycosylation of the subunits by N-glycosidase F yielded 40.9 and 19.6 kDa species. MA0196 GGT retained transpeptidase and hydrolysis activities and their catalytic efficiency (kcat/Km) under high salt and low water activity. The enzyme displayed broad substrate specificity toward γ-glutamyl acceptors such as amino acids and the imidazole dipeptides, carnosine and anserine. Carnosine and L-glutamine were converted into γ-glutamyl-β-alanyl-L-histidine by MA0196 GGT with a 32.9% yield in the presence of 2% (v/v) dimethyl sulfoxide. Phylogenetic analysis indicated that MA0196 GGT forms a distinct lineage from A. oryzae and A. sojae GGTs. These excellent properties indicate that MA0196 GGT can be used in salted fermentation and for producing bioactive peptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
γ-Glutamyl transpeptidase (GGT, EC 2.3.2.2) is a ubiquitous enzyme that catalyzes a two-step reaction as follows: cleavage of the γ-glutamyl bond present in the donor substrate followed by transfer of the γ-glutamyl moiety to another amino acid (transpeptidation) and/or short peptide or to a water molecule (hydrolysis) (Castellano and Merlino 2012; Saini et al. 2021). The physiology, chemistry, and structural properties of bacterial GGTs have been studied extensively because of their various applications in different biotechnology sectors (Castellano and Merlino 2012; Saini et al. 2021). GGTs are encoded by a single gene and are translated as a unique polypeptide, which then undergoes auto-proteolytic cleavage into a large and a small subunit (Castellano and Merlino 2012). The phylogenetic tree of GGTs from human, animals, yeast, and bacteria are divided into two main branches, B1 and B2, based on the presence or absence of the lid-loop region (e.g., 438Pro to 449Gly in E. coli GGT) (Verma et al. 2015; Saini et al. 2021). Among GGTs, bacterial GGTs have also been studied in detail for applications in producing valuable γ-glutamyl compounds (Mu et al. 2015; Yang et al. 2019). In contrast, fungal GGTs are poorly characterized despite the abundance of genes annotated as GGTs in fungal genomes.
In traditional Japanese wheat- and soybean-fermentation under high salt conditions (~ 17‒18% NaCl), glutaminases, including GGTs secreted by koji molds A. oryzae and A. sojae, are vital enzymes for flavor enhancement because these enzymes convert l-glutamine to the umami amino acid l -glutamic acid (Ito and Matsuyama 2021). However, the GGT is unstable and cannot retain activity at high salt concentrations (Koibuchi et al. 2000). Although Bacillus GGTs have been categorized into a salt-tolerant group (Minami et al. 2003; Pica et al. 2013; Li et al. 2020; Wada et al. 2010), Bacillus GGT (e.g., NP_389723) shows low sequence identity (~ 30%) with A. oryzae extracellular GGTs (XP_023088840, XP_023090648, and BAE63091) annotated using whole genome information from the koji molds (Ito et al. 2013). Understanding the molecular mechanism of salt-adaptation for application and protein engineering of fungal GGTs based on the comparison of these sequences and structures is challenging. Therefore, solving this issue involves isolating and characterizing new GGTs from Aspergillus spp. adapted to halophilic and/or xerophilic environments.
Xerophilic molds, such as A. chevalieri, A. pseudoglaucus, and A. sydowii, and their extracellular hydrolytic enzymes involved in the fermentation and ripening of foods have been characterized (Gobbetti et al. 1997; Virgili et al. 2007; Lim et al. 2019; Takenaka et al. 2020, 2021). In addition, hydrolytic enzymes from A. glaucus and its related species that showed salt-adaptive properties are expected to be used in biomass degradation (Esawy et al. 2016; Takenaka et al. 2019; Jiménez-Gómez et al. 2020). Thus, xerophilic molds and their extracellular enzymes possess potential molecular adaptation mechanisms under high salt and/or low water active conditions and are therefore attractive genetic resources for isolation of salt- and/or osmo-adaptive enzymes (Williams and Hallsworth 2009; Musa et al. 2018; Qiu et al. 2020).
In this study, the biochemical and molecular characterization of a different type of salt-tolerant GGT from the xerophilic mold A. sydowii is reported. The GGT from A. sydowii (MA0196 GGT) showed high salt tolerance when compared with previously reported Aspergillus GGTs. Enzymatic conversion of a γ-glutamyl peptide from a major imidazole dipeptide in katsuobushi filets was observed. These unique properties of MA0196 GGT indicate that this enzyme is an ideal candidate for use in the salted and fermented food industry.
Materials and methods
Microorganisms and culture conditions
A. sydowii MA0196 (formerly A. glaucus MA0196) (Lim et al. 2019; Takenaka et al. 2019, 2021) was isolated from the surface of katsuobushi (karebushi) filets. The γ-glutamyl transpeptidase from strain MA0196 (MA0196 GGT) was purified, and its gene was cloned. Escherichia coli (E. coli) HST08 (Takara Bio Inc., Kusatsu, Japan) was used for constructing the plasmid and gene cloning of MA0196 GGT. E. coli HST08 was cultivated in Luria–Bertani (LB) medium [supplemented, if necessary, with ampicillin (50 µg/mL)] with shaking at 140 rpm and 37 °C.
Strain MA0196 and A. oryzae NBRC 100959 (other culture collection no. RIB 40) were cultivated on katsuobushi solid medium and wheat bran solid medium, respectively, according to previously described methods (Koibuchi et al. 2000; Takenaka et al. 2019). Briefly, 5 g katsuobushi powder and 5 g wheat bran powder were placed in separate 50 mL flasks, and 0.5 and 4.0 mL water were added, respectively. After autoclaving, mycelia and spore suspension (0.05 mL, 1.0–5.0 × 106 spores/mL) were added to the solid medium. The cultures were stirred with a sterilized bar every week. Extracellular enzymes, including GGT, were extracted from the solid cultures using 30 mL of 50 mM sodium acetate buffer (pH 5.0) and incubated for 3 h with shaking (Takenaka et al. 2021). The mixture was vacuum filtered through a Whatman No. 1 filter paper (Buckinghamshire, UK) on a Büchner funnel, and the filtered extracts were dialyzed against 20 mM Tris–HCl buffer (pH 7.5). The dialyzed solution (the crude enzyme preparation) was used for the enzyme assay and purification.
Effect of water activity on GGT production by A. sydowii MA0196 in solid-state fermentation and submerged fermentation
A. sydowii MA0196 was cultivated under solid and submerged culture conditions with various water activities to determine the production profile of the GGT. Culture conditions for solid-state fermentation using katsuobushi solid medium and for submerged fermentation using MY medium (0.3% malt extract, 0.3% yeast extract, 0.5% peptone, pH 5.5) are described in the footnote of Table S1. The aw reached 0.94 in the medium containing 40% (w/v) d-glucose (El Halouat and Debevere 1996).
Enzyme assays
Enzyme assays were carried out according to modifications of previously reported methods (Koibuchi et al. 2000). Transpeptidation activity was assayed as follows: the reaction mixture (1.1 mL) contained 100 mM Tris–HCl (pH 8.0), 0.5 mM l-γ-glutamyl-p-nitroanilide (γ-GpNA), 33 mM glycylglycine, and 100 µL enzyme solution. This reaction mixture was incubated for 20 min at 40 °C (within linear range). The absorbance of the mixture at 405 nm was measured. Hydrolysis activity was assayed as follows: the reaction mixture (1.1 mL) containing 100 mM Tris–HCl buffer (pH 8.0), 0.5 mM γ-GpNA, and 100 µL enzyme solution was incubated for 20 min at 40 °C. The absorbance of the mixture at 405 nm was measured. The presence of glycylglycine suppresses hydrolysis of γ-GpNA and the auto-transpeptidation reaction effectively (Meister et al. 1981). Enzyme activities were expressed in units of µmol min–1 and calculated by the following equation according to the previously reported calculation method (Meister et al. 1981). One unit of activity was defined as the amount of enzyme that catalyzed the release of 1 µmol p-nitroanilide per min under these conditions. Protein concentrations were measured using a Lowry protein assay kit (Nacalai Tesque, Inc., Kyoto, Japan).
ɛ: molar extinction coefficient at 405 nm (8.3 × 103 M–1 cm–1) of p-nitroanilide (pH 8.0); 106: is used for converting moles into micromoles; 103: is used for converting U/mL into mU/mL.
Purification of γ-glutamyl transpeptidase from A. sydowii MA0196 (MA0196 GGT)
GGT production profiles in submerged culture and solid-state fermentation of strain MA0196 were examined (Table S1). The first purification step was challenging because the crude enzyme solution contained a large amount of d-glucose. Thus, strain MA0196 was cultivated by solid-state fermentation in a katsuobushi solid medium, and MA0196 GGT was purified from the crude enzyme preparation using previously modified procedures (Takenaka et al. 2020). The details about the procedures for purification are described in the Supplementary Information. Transpeptidation and hydrolysis activities were assayed, as described above. Purified MA0196 GGT was used for characterization. Unless otherwise stated, the protein amount corresponded to 65–115 mU/mL transpeptidase activity and 25–44 mU/mL hydrolysis activity.
SDS-PAGE and LC–MS-MS-Mascot analysis for protein identification
SDS-PAGE
Tris–glycine-SDS-PAGE was performed using a 12.5% polyacrylamide precast-gel (e-PAGEL precast gel E-T12.5L) with the electrophoresis buffer containing 25 mM Tris, 192 mM glycine, and 0.1% (w/v) SDS and an ATTO AE6530 gel electrophoresis system (ATTO Corp., Tokyo, Japan), according to the manufacturer’s instructions. The final purified preparation (0.5–3 µg protein) was denatured and loaded onto the precast gel. Protein molecular mass markers (broad range type: 9–200 kDa) were from Nacalai Tesque Inc.
Staining for protein visualization
The protein bands were detected by EzStain AQua AE-1340 and EzStain Silver AE-1360 (ATTO Corp.), according to the manufacturer’s instructions. MA0196 GGT was identified as a glycoprotein (see Fig. 1c). The purified enzyme was digested with glycopeptidase F (peptide: N-glycosidase F, Takara Bio Inc.), according to the manufacturer’s instructions, to compare the original protein with the deglycosylated protein by SDS-PAGE analysis.
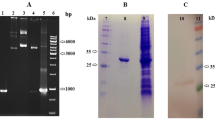
SDS-PAGE analysis of purified γ-glutamyl transpeptidase (MA0196 GGT) from A. sydowii MA0196. (a) Protein molecular mass markers (broad-range type: 9–200 kDa, Nacalai Tesque, Inc.). (b) Purified MA0196 GGT (approximately 3 µg of MA0196 GGT, Coomassie blue staining with EzStain Aqua (ATTO Co.). (c) Purified MA0196 GGT [approximately 3 µg of MA0196 GGT, Periodic acid Schiff staining (Zacharius et al. 1969)]. (d) MA0196 GGT deglycosylated using N-glycosidase F (approximately 0.5 µg of MA0196 GGT, silver staining). Bands representing the large and small subunits produced via autoprocessing of MA0196 GGT and N-glycosidase F are indicated by solid and open arrows, respectively
Staining by periodic acid Schiff (PAS) for glycoprotein visualization
PAS staining was carried out according to a reported method (Zacharius et al. 1969). Briefly, the gel was fixed with 12.5% (w/v) trichloroacetic acid and stored at room temperature for 30 min. The gel was washed with water and incubated in 1% (w/v) periodic acid—3% (v/v) acetic acid solution for 50 min. The gel was washed with water and placed in Schiff's reagent (Nacalai Tesque, Inc.) for 50 min in darkness and destained using 1 N HCl–10% (w/v) sodium sulfate solution to visualize a pink-colored band.
LC–MS-MS-Mascot analysis for protein identification
Main protein bands corresponding to the large- and small-subunits of MA0196 GGT (see Fig. 1b) were excised from the gel and digested with trypsin, as described previously (Abbas et al. 2005). The proteins were treated with in-gel digestion, and the molecular masses of the produced peptides were determined using LTQ Orbitrap Discovery (Thermo Fisher Scientific, Waltham, MA, USA), as described previously (Takahata et al. 2012). The whole-genome assembly sequence for A. sydowii CBS 593.65 (GCA_001890705.1), which is registered in the GenBank database, was used to identify GGTs from A sydowii by using the automated search function of Mascot v2.3 (Matrix Science Ltd. London, UK). The identified peptide sequences were identical to an uncharacterized protein (XP_040703591) in A. sydowii CBS 593.65. A set of specific primers for GGT gene cloning was designed using the nucleotide sequence of the unknown protein.
Characterization of MA0196 GGT
Determination of kinetic parameters
Kinetic parameters of purified MA0196 GGT were determined by performing steady-state kinetic studies with various concentrations of γ-GpNA with and without 33 mM glycylglycine, as described in the footnote of Table 1. The parameters were also determined in a buffer containing 20% (w/v) NaCl or 20% (v/v) glycerol.
Effect of NaCl and glycerol concentration on enzyme activity
Transpeptidation and hydrolysis assays were carried out using the purified MA0196 GGT in the reaction mixture containing NaCl (0–20% (w/v) as final concentrations), and these assays were also conducted in the reaction mixture containing glycerol (0–50% (v/v) as final concentrations) to reproduce low water activity conditions (Fig. 2).
Effect of NaCl and glycerol concentration on enzyme activity. The reaction mixture (1.1 mL) containing 50 mM Tris–HCl (pH 7.5), 0.5 mM L-γ-glutamyl-p- nitroanilide, and 100 µL of purified γ-glutamyl transpeptidase (MA0196 GGT) from A. sydowii MA0196 or crude enzyme preparation from A. oryzae NBRC 100959 in the presence (for transpeptidase assay, GGT_T) and absence (for hydrolysis assay, GGT_H) of 33 mM glycylglycine was incubated at 30 °C. (a) Effect of NaCl. The reaction mixture consisted of various concentrations of NaCl (0–20% (w/v) as a final concentration). (b) Effect of glycerol. The reaction mixture consisted of various glycerol concentrations (0%–50% (v/v) as a final concentration). The relative activities were calculated based on transpeptidase and hydrolysis activities in the absence of NaCl or glycerol. In MA0196 GGT, the specific activities of transpeptidation and hydrolysis without NaCl and glycerol were 5.0 and 0.93 U/mg, respectively. The relative activity was calculated based on these specific activities. The crude enzyme preparation from A. oryzae NBRC 100959 corresponded to 100 mU/mL transpeptidase activity and 50 mU/mL hydrolysis activity was used, and the relative activity was calculated
Effect of NaCl on stability
The purified MA0196 GGT was incubated in buffer A without and with 10% (w/v) NaCl and 20% (w/v) NaCl at 4 °C for 1 week, as described in the footnote of Table 2.
Effect of temperature on enzyme activity and stability
The effect of temperature on the transpeptidation and hydrolysis activities of the purified MA0196 GGT was determined over a temperature range of 20–70 °C. The purified MA0196 GGT was pre-incubated at various temperatures (10–70 °C) for 10 min, and residual activities were measured.
Effect of pH on enzyme activity and stability
The pH (3.5–11.0) of a 100 mM 3,3-dimethylglutaric acid (G), 100 mM Tris (T), and 100 mM 2-amino-2-methyl-1,3-propanediol (A) mixture (designated “GTA”) was adjusted by adding 1.0 N HCl and 1.0 N NaOH, according to the previously described procedure for preparing GTA buffer (Sekizaki et al. 2000). Enzyme activities were measured using the purified MA0196 GGT in 50 mM GTA buffer (pH 3.5‒11.0). The pH stability of MA0196 GGT was determined by mixing concentrated purified MA0196 GGT with 100 mM GTA buffer (pH 3.5–11.0) at 4 °C for 15 h. The enzyme solution was concentrated tenfold using an Amicon® Ultra-15 filter (Merck KGaA). The concentrated enzyme solution and each buffer solution (pH 3.5–11.0) were mixed at a ratio of 1:9 (v/v), and the residual activity was measured.
Effect of inhibitors
Purified MA0196 GGT was pre-incubated at 4 °C for 30 min in 20 mM Tris–HCl buffer (pH 7.5) containing inhibitors at concentrations of 0.5 or 5 mM (see Table 2), and the residual activity was measured.
Effect of organic solvents
Enzyme activities were measured using the purified MA0196 GGT in 100 mM Tris–HCl buffer (pH 8.0) in the presence of various organic solvents at final concentrations of 10% (v/v) (see Table 2).
Substrate specificity toward γ-glutamyl acceptors
Substrate specificity toward γ-glutamyl acceptors (see Table 3) was measured by the transpeptidation activity assay using the acceptors instead of glycylglycine.
γ-Glutamylation of carnosine by MA0196 GGT
Synthetic conditions
MA0196 GGT showed the highest transpeptidation activity toward carnosine (see Table 4). In addition, the activity was promoted in the presence of a small amount of dimethyl sulfoxide (DMSO) (see Table S4). Enzymatic synthesis of γ-glutamyl carnosine using MA0196 GGT as a biocatalyst was examined.
The reaction for γ-glutamylation was performed with reference to reported conditions (Saini et al. 2021). Thus, the reaction mixture (1.0 mL) containing 100 mM GTA buffer (pH 8.5–10), 50 mM l-glutamine, 50 mM carnosine (β-alanyl-l-histidine), and 0.1 mL purified MA0196 GGT (0.03 U/mL) was incubated at 30 °C for 12 h. In addition, the enzymatic γ-glutamylation was carried out in 2% (v/v) dimethyl sulfoxide-50 mM GTA buffer (pH 8.5–10).
Product analysis by HPLC
A Hitachi L-7100 HPLC system equipped with a 5C18-PAQ column (4.6 mm I.D. × 150 mm, Nacalai Tesque, Inc.) was used for measuring substrates, L-glutamine and carnosine, and produced γ-glutamyl derivatives. The flow rate at room temperature was 1.0 min/mL. Samples were eluted with 5 mM sodium pentanesulfonate–20 mM phosphoric acid with monitoring at 210 nm. The retention times of l-glutamine, l-glutamate, γ-glutamyl-glutamine (Toronto Research Chemicals Inc., Toronto, ON), carnosine, and γ-glutamyl-carnosine (γ-glutamyl-β-alanyl-l-histidine, Cosmo Bio Co., Ltd., Tokyo, Japan) were 2.7, 3.2, 3.8, 8.6, and 10.4 min, respectively.
Gene cloning and sequence analysis of γ-glutamyl transpeptidase from A. sydowii MA0196
Gene cloning of MA0196 GGT from genomic DNA and cDNA
Preparation of genomic DNA and total RNA from strain MA0196 and gene cloning were carried out as described in the Supplementary Information.
Sequence analysis of MA0196 GGT
The obtained sequence data were compared with appropriate reference data using the ClustalW algorithm and GENETYX-win software (Version 14.0; GENETYX, Tokyo, Japan). Signal peptide searches and predictions were performed using the Signal P-6.0 server. N-glycosylation sites were predicted using the online tool of NetNGly 1.0. Structural modeling of MA0196 GGT was performed using the SWISS-MODEL web server. The modeling template research indicated that the global model quality estimate (GMQE) value of GGT from Bacillus licheniformis ATCC 14580 (PDB ID, 4Y23; GMQE, 0.67) for MA0196 GGT was higher than that from E. coli K-12 (2E0W; 0.64). However, the GGT from B. licheniformis ATCC 14580, like GGT from B. subtilis 168 (Wada et al. 2010), lacks the lid loop in the amino acid sequence (see Fig. 3). Thus, the crystallographic structure of GGT from E. coli K-12 was used as the template for homology modeling and comparison with human GGT (see Fig. S6). Structures were visualized using PyMOL version 2.5 and for comparison with human GGT [PDB ID: 4GG2; identity with MA0196 GGT, 37% (198/533 amino acids); GMQE value with MA0196 GGT, 0.43] (see Fig. S6).
Multiple sequence alignment of γ-glutamyl transpeptidases (GGTs) from several representative organisms. Identical residues are indicated by an asterisk, whereas similar residues are indicated by a period or colon. Residues of the catalytic nucleophile, stabilizing the nucleophile, substrate binding, and comprising the oxyanion hole are indicated with the letters N, S, B, and O, respectively. Cleavage sites for auto-processing are shown in yellow. The extra sequence (391Gln to 402Gln) in B. subtilis GGT and the lid-loop region in E. coli GGT (339Val to 450Asp) and human GGT (429Ile to 439Pro) are shown in yellow (Saini et al. 2021; Minami et al. 2003; Wada et al. 2010). N-glycosylation sites in human GGT and conserved sites for N-glycosylation in A. sydowii MA0196 GGT are shown in green and gray, respectively. The secondary structure α-helices (orange bar) and β-strands (blue arrow) are shown above the alignment and based on the human GGT structure (West et al. 2011, 2013). Abbreviations: Bs_GGT, GGT from B. subtilis 168 (NP_389723); Ec_GGT, GGT from E. coli (WP_160532025); Hs_GGT, GGT from human (NP_001275762); As_GGT, GGT from A. sydowii MA0196 (this study)
Statistical analysis
All experiments were performed in duplicate and replicated at least three times. All data were calculated as the mean ± standard deviation. Analysis of variance (ANOVA) was selected to test value differences (p < 0.05). For comparison of different groups, the data were analyzed using one-way ANOVA Tukey’s multiple comparison. All calculations were conducted using GraphPad Prism software (ver. 6.02 for Windows, La Jolla, CA, USA).
Results and discussion
Production profile of γ-glutamyl transpeptidase in A. sydowii MA0196
The crude enzyme preparation from strain MA0196 showed the highest transpeptidation and hydrolysis activities (21.7 ± 0.63 and 11.8 ± 1.0 mU/mL) among laboratory stock molds. After fractionating a crude enzyme preparation from A. oryzae by DEAE-Toyopearl column chromatography, the transpeptidation activities were 6.5–20 mU/mL (Koibuchi et al. 2000). The amount of GGT production by A. sydowii in katsuobushi powder solid medium and by A. oryzae in wheat bran solid medium was comparable. In contrast, GGT production by Bacillus spp. is quite high with transpeptidation activity of culture supernatants from B. licheniformis ER15 (Bindal et al. 2016), B. subtilis MH2308 (Minami et al. 2003), and B. subtilis SK11.004 (Shuai et al. 2011) yielding 5.2, 0.19, and 22.4 U/mL, respectively.
GGT production profiles of strain MA0196 were investigated under solid-state fermentation and submerged fermentation conditions (Table S1). Production of GGT was promoted with decreasing aw in solid-state fermentation. In solid-state fermentation, the optimal aw value for GGT production by strain MA0196 was 0.85 (Table S1). The crude enzyme preparation from strain MA0196 showed transpeptidation and hydrolysis activities (61.7 ± 2.2 and 16.4 ± 0.55 mU/mL) (Table S1). The crude enzyme solution prepared from a submerged culture in MY liquid medium containing 25% (w/v) D-glucose showed the highest activity (36.4 ± 1.01 and 6.64 ± 0.67 mU/mL, Table S1). This observation indicated that strain MA0196 does not grow well in MY medium containing more than 30% (w/v) d-glucose, as reported previously (Takenaka et al. 2019). The aw in solid-state fermentation influences the growth of (Astoreca et al. 2007) and enzyme production (Sapna 2014; Demir and Tari 2016; Santos et al. 2018) by Aspergillus spp. Among the tested conditions, the crude enzyme preparation from solid-state fermentation of strain MA0196 at aw 0.85 showed the highest activity (Table S1). Other fungi prefer higher aw conditions when compared with that of xerophilic A. sydowii for extracellular enzyme production. For example, the optimal aw values for production of xylanase and β-glucosidase by A. niger, polygalacturonase by A. sojae, and phytase by A. oryzae were reported to be 0.86–0.90 (Santos et al. 2018), 0.99 (Demir and Tari 2016), and 0.95 (Sapna 2014), respectively.
Purification of γ-glutamyl transpeptidase from A. sydowii MA0196
γ-Glutamyl transpeptidase from strain MA0196 (MA0196 GGT) was purified by sequential column chromatography procedures, and transpeptidation and hydrolysis activities of the collected fractions were measured (Table S2). Specific activities for transpeptidation and hydrolysis were 2400 and 500 mU/mg, respectively (Table S2). The gel-phase distribution coefficient of MA0196 GGT on a Cosmosil Diol-300-II column gave a molecular mass of 120 kDa for native GGT (Fig. S1). GGTs are generally synthesized as a single polypeptide (propeptide) that undergoes autocatalytic cleavage, which leads to the formation of the large and small subunits that compose the mature enzyme (Saini et al. 2021). Purified GGT from B. subtilis shows a single band on a native-PAGE gel by active staining using γ-l-glutamyl-α-naphthylamide (Shuai et al. 2011) and γ-GpNA with glycylglycine (Saini et al. 2017). In this study, active staining of native PAGE gels was attempted; however, the active band derived from the purified and concentrated MA0196 GGT sample was not detected on the gel. The specific activities of the purified Bacillus GGT are quite high [683 U/mg (Shuai et al. 2011) and 90 U/mg (Saini et al. 2017)] when compared with that of MA0196 GGT. In addition, the protein band of MA0196 GGT was broad because of a high level of glycosylation. Thus, detection of a distinct color band on the gel is unlikely. SDS-PAGE analysis with staining dye and PAS reagent indicated that MA0196 GGT is composed of a large subunit of 56.4 kDa and a small subunit of 33 kDa, and both polypeptides are glycosylated (Fig. 1b, c). Treatment of the purified enzyme with N-glycosidase F yielded 40.9 kDa- and 19.6 kDa-protein bands in the SDS-PAGE gel (Fig. 1d). Thus, strain MA0196 secreted the GGT as a highly glycosylated protein. This high glycosylation of MA0196 GGT is because of the numerous potential N-glycosylation sites in the sequences of both subunits. Based on the LC–MS–MS and Mascot analysis, the small and large subunits (Fig. 1b) showed identity with an uncharacterized protein from A. sydowii CBS 593.65 (coverage: 18.4%; accession no. XP_040703591.1), and gene cloning of MA0196 GGT was carried out with reference to sequence data from strain CBS 593.65 described above.
Characterization of the γ-glutamyl transpeptidase from A. sydowii MA0196
Effect of NaCl and glycerol concentrations on transpeptidase and hydrolysis activities
Glutaminases, including GGT from soy sauce Aspergillus koji molds, are non-salt-tolerant; see Introduction. Activity is inhibited by ~ 80% in the presence of 18% (w/v) NaCl when compared with activity in the absence of salt (Koibuchi et al. 2000). This salt concentration is equivalent to that found in the soy sauce fermentation process (Koibuchi et al. 2000; Ito and Matsuyama 2021). MA0196 GGT retained transpeptidase and hydrolysis activities under high salt (Fig. 2a) and low water activity (Fig. 2b) when compared with the enzyme preparation from A. oryzae. The relative activity of the preparation from A. oryzae decreased dramatically to less than 18% under high-salt and low-water activity. MA0196 GGT was slightly less salt-tolerant than GGTs from several Bacillus spp. (Minami et al. 2003; Bindal and Gupta 2016; Li et al. 2020; Cho et al. 2021) in the presence of 18% (w/v) NaCl, with transpeptidase and hydrolysis activities retained at approximately 43% and 55%, respectively, relative to the activity in the absence of NaCl (Fig. 2a). In addition, purified MA0196 GGT was stable in buffer containing 10% and 20% (w/v) NaCl after incubating for a week (Table 2). GGT from B. licheniformis shows more than 80% residual activity even after one month-incubation in optimal buffer (Lin et al. 2018). Long storage period and stability profiles will be necessary to determine the suitability of MA0196 GGT in salting-out fermentation applications.
Glycerol not only increases osmosis in solutions but also reduces aw, like NaCl (Gekas et al. 1998; Nakagawa and Oyama 2019). The effect of the glycerol concentration on GGT activity was examined to determine the activity of MA0196 GGT under low aw. In the presence of 50% (v/v) glycerol, transpeptidation and hydrolysis activities of MA0196 GGT were retained at approximately 20% and 40%, respectively, relative to the activity in the absence of glycerol (Fig. 2b). The relationship between salt and glycerol concentrations in aqueous solution and aw have been clarified experimentally (Gekas et al. 1998; Nakagawa and Oyama 2019). A 20% (w/v) NaCl aqueous solution and a 40% (v/v) glycerol solution correspond to an aw of 0.8. The relative activities with 20% (w/v) NaCl (Fig. 2a) and 40% (v/v) glycerol (Fig. 2b) were similar. However, kinetic parameters of transpeptidation and hydrolysis in the absence and presence of NaCl and glycerol differed (Table 1).
Although GGTs from Bacillus spp. have been categorized as salt-tolerant enzymes (Minami et al. 2003; Bindal and Gupta 2016; Li et al. 2020; Cho et al. 2021), kinetic parameters in the absence and presence of NaCl and glycerol have not been compared, except for conformational stability studies (Lo et al. 2018; Cho et al. 2021). GGT catalyzes a two-step reaction: cleavage of the γ-glutamyl bond present in the donor substrate followed by transfer of the γ-glutamyl moiety to another amino acid (transpeptidation) and/or short peptide or to a water molecule (hydrolysis) (Saini et al. 2021). In the presence of glycylglycine (transpeptidation in Table 1), the Km values for γ-GpNA by MA0196 GGT were similar to those of GGTs from Bacillus spp. (Bindal and Gupta 2016; Saini et al. 2017). The Km value for γ-GpNA was only slightly affected by the presence of NaCl and glycerol; however, the Vmax and kcat decreased noticeably for MA0196 GGT. The steps after forming the enzyme–substrate complex, such as cleavage of the γ-glutamyl bond and transfer of the γ-glutamyl moiety to glycylglycine, may be affected by NaCl and glycerol. In the absence of glycylglycine (hydrolysis in Table 1), the kinetic parameters, especially the Km for γ-GpNA, were much more affected by glycerol than when NaCl was present. After forming the enzyme–substrate complex, the cleavage of the γ-glutamyl bond of γ-GpNA was probably not affected by the presence of NaCl and glycerol. Here, kinetic parameters were determined under presumably acceptor substrate saturation (33 mM glycylglycine), according to conditions reported previously (Shuai et al. 2011; Saini et al. 2017). Further investigation of the kinetic parameter profiles with various concentrations of glycylglycine in the absence and presence of NaCl and glycerol is required.
Effects of pH and temperature on transpeptidase and hydrolysis activities and stability
The properties of fungal GGTs from A. nidulans and A. sojae were partially studied (Table S3). Although the subunit structure and N-glycosylation of these Aspergillus GGTs have not been resolved, their optimal temperature, pH, and stabilities are similar to those of MA0196 GGT (Table S3). The transpeptidase and hydrolysis activities of MA0196 GGT were highest at 45 °C (Table 2; Fig. S2a). The enzyme was stable up to ~ 45 °C (> 80%), but activity was essentially lost at 55 °C (Fig. S2b). The optimal temperature and thermostability of MA0196 GGT were comparable to those of reported GGTs from A. oryzae (Koibuchi et al. 2000) and Aspergillus spp. (Table S3). The enzyme was stable over a wide pH range (5.0–9.5) with maximum activity at pH 8.5 (Table 2; Fig. S3). Bacterial GGTs function optimally over pH values ranging from 8.0 to 11.0 with different pH optima for transpeptidation and hydrolysis reactions (Saini et al. 2021). For a GGT from B. subtilis, transpeptidation and hydrolysis activities reached a maximum at pH 10 and 9, respectively (Morelli et al. 2014). The ratios of the transpeptidase and hydrolysis activities at pH 9.5 and 10 for MA0196 GGT were 3.2 and 4.9, respectively. The ratio at pH 9.5 was negligible. The pH profiles for hydrolysis and the transpeptidation reaction of MA0196 GGT were more pronounced in the γ-glutamylation reaction of carnosine with L-glutamate, as described below.
Effects of various compounds on transpeptidase and hydrolysis activities
The addition of 6-diazo-5-oxo-l-norleucine (DON, 0.5 and 5.0 mM), a GGT specific substrate analog, caused the complete inactivation of MA0196 GGT (Table 2). MA0196 GGT activity decreased when l-Ser (5 mM) and borate (5 mM) were present. Among the tested organic solvents, only n-butanol inhibited the transpeptidation and hydrolysis activities of MA0196 GGT strongly, even under 2% (v/v) (Table S4). GGT from B. amyloliquefaciens also showed weak tolerance toward organic solvents (Li et al. 2020). The hydrolysis activity of MA0196 GGT increased significantly in the presence of polar aprotic solvents (DMSO, DMF, and acetone) and polar protic solvents (methanol and ethanol), and the hydrolysis activity was not inhibited by adding 10% (v/v) of these solvents when compared with the transpeptidase activity (Tables 2 and S4). Only the addition of 2% (v/v) DMSO and N,N-dimethylformamide increased the transpeptidase activity (Table S4). These results indicated that the transpeptidase activity was promoted in the presence of a small amount of organic solvent, which may facilitate peptide synthesis applications, as described below.
Substrate specificity for γ-glutamyl acceptors
The transfer of γ-glutamyl from γ-GpNA to various amino acids, amines, and peptides by MA0196 GGT was studied (Table 3). The transfer activity with glycylglycine as the substrate was taken as 100%. The specificity for γ-glutamyl acceptors is shown in Table 3. MA0196 GGT showed broad substrate specificity. Katsuobushi is rich in specific peptides, amines, and L-histidine (1,992 mg/100 g of katsuobushi (Fuke and Konosu 1991)). For example, imidazole dipeptides, l-carnosine (β-alanyl-l-histidine, 107 mg/100 g) and anserine (β-alanyl-3-methyl-l-histidine, 1250 mg/100 g) and amines, creatine (2-(carbamimidoyl(methyl)amino]acetic acid, 540 mg/100 g) and creatinine (2-amino-1-methyl-5H-imidazol-4-one, 1,150 mg/100 g), are present in katsuobushi (Fuke and Konosu 1991). Interestingly, MA0196 GGT showed high activity toward carnosine (Table 3). MA0196 GGT preferred anserine (relative activity compared with glycylglycine was 47.0%) and carnosine (64.8%) over β-alanine (27.1%) (Table 3). Carnosine and its methylated form, anserine, exist as zwitterions and carry a net positive charge (Wu 2020). The pKa (9.66) of the amino group of the β-alanyl residue of carnosine is similar to the same moiety on anserine (9.49) (Wu 2020). In bacterial GGTs, the putative acceptor binding site is proximal to the donor binding site, located in the deep groove, and is highly variable and flexible when compared with the highly conserved donor binding site (Castellano and Merlino 2012; Saini et al. 2021). The methyl group located on the imidazole ring of anserine may affect the affinity of MA0196 GGT toward this compound, although the kinetic parameters of MA0196 GGT for carnosine and anserine should be compared.
Creatine and anserine improve human nutrition and health, including metabolic, retinal, immunological, muscular, cartilage, neurological, and cardiovascular health (Wu 2020). In addition, the tripeptide, γ-glutamyl-β-alanyl-histidine, which is most likely produced from carnosine via the γ-glutamyl transpeptidation step, was isolated in a beef soup stock solution, and the tripeptide, γ-glutamyl-carnosine is a meaty flavor enhancer (Kuroda et al. 2000). This tripeptide is a promising compound for controlling and preventing Influenza A (H1N1) virus infections (Babizhayev and Deyev 2012). The bioconversion of carnosine to this tripeptide by MA0196 GGT was investigated.
γ-Glutamyl translation of carnosine by γ-glutamyl transpeptidase from A. sydowii
The enzymatic reaction products in the reaction mixture containing 50 mM carnosine and 50 mM l-glutamine as substrates were determined by HPLC to be l-glutamic acid (via hydrolysis), γ-glutamyl-glutamine (via auto-transpeptidation), and γ-glutamyl-carnosine (via transpeptidation), and conversion ratios were calculated based on the γ-glutamyl-carnosine produced (Table 4). γ-Glutamylation of l-glutamine and production of l-glutamic acid were detected at pH 8.5 and 9.0. The optimal pH for γ-glutamyl translation of carnosine was 9.5, with a 29.6% molar yield. Conversion of γ-glutamyl was carried out in the presence of DMSO with reference to the results described above (Table S4). Clearly, the addition of 2% (v/v) DMSO to the reaction mixture (pH 9.5) promoted a 1.1-fold increase in γ-glutamylation of carnosine (Table 4). However, l-glutamine was used for γ-glutamyl translation to carnosine and l-glutamine in the reaction mixture containing DMSO (Table 4).
Among skipjack tuna meats, red meat contains as much carnosine and anserine as dorsal and abdominal parts (Hosokawa et al. 1990). A similar waste is generated in the manufacturing process of katsuobushi (Takenaka et al. 2020). Carnosine and anserine have diverse biological properties, as described above. Extracting valuable bioactive peptides from the waste parts of skipjack tuna or enzymatically converting them to more valuable γ-glutamyl compounds using GGTs are important potential applications. Enzymatic synthesis of valuable γ-glutamyl peptides has been optimized by varying the amount of l-glutamine, substrate, and GGT, as well as the reaction conditions (Saini et al. 2021). Future efforts will focus on factors that improve the yield of γ-glutamyl-carnosine produced by MA0196 GGT.
Cloning and characterization of the γ-glutamyl transpeptidase gene from A. sydowii MA0196
Gene cloning and sequence analysis of γ-glutamyl transpeptidase from A. sydowii
2.1 kb- and 1.8 kb-fragments were amplified with primers GGT_sydowii_F and GGT_sydowii_R, using genomic DNA and a cDNA library, respectively (Fig. S4). The genomic DNA and cDNA of the MA0196 GGT gene were composed of 2136 bp and 1782 bp, respectively. Six introns (55, 63, 61, 64, 55, and 56 bp) interrupt the γ-glutamyl transpeptidase coding sequence. SignalP analysis indicated the presence of an N-terminal signal peptide composed of amino acid residues 1‒27. The mature protein of γ-glutamyl transpeptidase contains 566 residues with a calculated molecular mass of 60.9 kDa. The signal peptide cleavage site between 27Phe and 28Ser and the new N-terminal sequence derived from the small subunit were confirmed by Mascot analysis based on the deduced amino acid sequences from 28Ser to 402Gly and from 403Thr to 593Val. The predicted molecular masses of the large and small subunits are 40.6 kDa and 20.3 kDa, which are consistent with the values of the deglycosylated polypeptides determined by SDS-PAGE (Fig. 1d).
In the A. oryzae RIB40 genome, four γ-glutamyl transpeptidase genes (GGTA (locus tag no., AO090005000169; accession no., XP_023088840); GGTB (AO090023000537; XP_023090648); GGTC (AO090113000029; BAE63091), and GGTD (AO090009000211; BAE54702)) were found (Ito et al. 2013). The deduced amino acid sequence of MA0196 GGT has the highest identity (339/442 a.a. (76%)) with GGTA among the four GGTs from A. oryzae (Fig. 4; Table S5). The identity among the small subunit sequences was higher than that observed among the large subunit sequences in GGTs from A. oryzae and A. sydowii (Table S5). Primary structure analysis of various microbial GGTs has shown that the intact polypeptide is generally composed of 500 ± 100 amino acid residues, and the large subunit is less conserved and consists of 300–405 residues, whereas the small subunit is 170–195 amino acids in length and is relatively conserved. The small subunit contains amino acid residues involved in donor substrate binding and the formation of the γ-glutamyl enzyme intermediate, as described below (Saini et al. 2021).
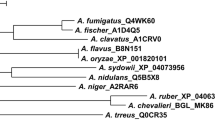
Phylogenetic analysis of γ-glutamyl transpeptidase from A. sydowii MA0196 and γ-glutamyl transpeptidases from Aspergillus spp. classified in clade γ-glutamyl transpeptidase 1. Multiple alignment of amino acid sequences of Aspergillus spp. was analyzed using GENETYX ver. 14.0 (Genetyx Corp.) and the unrooted phylogenetic tree was constructed using the UPGMA algorithm. In this study, GGT from strain MA0196 was analyzed with GGT sequences from Aspergillus spp. classified in clade GGT1 (a Pezizomycotina-only sub-clade within the Pezizomycotina clade (Bello and Epstein 2013). In addition, MA0196 GGT was also analyzed with GGT sequences from Aspergillus spp. classified in clades GGT1 and 3 (Bello and Epstein 2013) (see Fig. S4). Abbreviations: A_sydowii_MA0196_GGT, GGT from A. sydowii MA0196 (this study); A_cavatus ACLA 006,420, GGT from A. clavatus NRRL 1 [accession no. EAW11885; amino acids (a.a.), 575 a.a.; identity (%) with GGT_A_sydowii, 414/540 (76%)]; A_fischeri NFIA 025,700, GGT from A. fischeri NRRL 181 [XP_001261394; 552 a.a.; 375/495 (75%)]; A_flavus_AFL2G 00188, GGT from A. flavus NRRL 21882 [RAQ55454; 584 a.a.; 435/593 (73%)]; A_flavus_AFL2G 04378, GGT from A. flavus NRRL 3357 [XP_041144336; 629 a.a.; 370/594 (62%)]; A_flavus_AFL2G 08520, GGT from A. flavus NRRL 3357 [XP_041148340; 591 a.a.; 318/537 (59%)]; A_fumigatus_Afu7g04760, GGT from A. fumigatus Af293 [EAL86996; 492 a.a.; 365/503 (72%)]; A_fumigatus_AFUB 090300, GGT from A. fumigatus Af293 [EAL86996; 492 a.a.; 365/503 (72%)]; A_nidulans_ANIA 10444, GGT from A. nidulans FGSC A4 [CBF75434; 605 a.a.; 464/605 (76%)]; A_niger_An13g01300, GGT from A. niger CBS 513.88 [CAK41530; 580 a.a.; 435/593 (73%)]; A_oryzae_GGTA, GGTA from A. oryzae RIB40 [XP_023088840 (locus tag no., AO090005000169); 531 a.a.; 339/442 (76%)]; A_oryzae_GGTB, GGTB from A. oryzae RIB40 [XP_023090648 (AO090023000537); 626 a.a.; 365/594 (61%)]; A_oryzae_GGTC, GGTC from A. oryzae RIB40 [BAE63091 (AO090113000029); 591 a.a.; 317/532 (59%)]; A_sojae_GGTA, GGTA from A. sojae BA-104 [JP4651203B2; 585 a.a.; 429/593 a.a (72%)]; A_terreus_ATEG 04362, GGT from A. terreus NIH2624 [EAU34809; 585 a.a.; 420/595 (70%)]
Comparison of amino acid sequences between γ-glutamyl transpeptidases from A. sydowii and Aspergillus spp
Phylogenetic analysis, according to a reported phylogenetic analysis of γ-glutamyl transpeptidase clades, was performed (Bello and Epstein 2013) (Figs. 4 and S5). High bootstrap values separated the Aspergillus GGTs into four distinct clusters with γ-glutamyl transpeptidase from A. sydowii closely related to A. nidulans ANIA 10444 (identity with MA0196 GGT, 464/605 a.a. (76%)). MA0196 GGT forms a distinct lineage from three GGTs of A. oryzae (Figs. 4 and S5). In addition, the small subunit of MA0196 GGT showed a much higher identity than that of ANIA 10444 GGT (164/191 a.a., 85%) (Table S5). This higher identity was also seen with the small subunit of MA0196 GGT and those of other Aspergillus GGTs, with a few exceptions (Table S5).
The donor binding site that interacts with the γ-glutamyl moiety in the donor substrate is well known and conserved among known GGTs, whereas residues constituting the acceptor binding site have not been identified with certainty (Saini et al. 2021). Multiple sequence alignment of MA0196 GGT with bacterial and human GGTs revealed key amino acid residues involved in catalysis, such as the catalytic nucleophile (403Thr in MA0196 GGT), stabilizing nucleophile (421Thr), donor substrate recognition, and binding residues (125Arg, 423Asn, 442Glu, 445Asp, 473Ser, and 474Ser), and residues that form and stabilize the oxyanion hole (495Gly and 496Gly) (Fig. 3). There are two disulfide bonds (50Cys and 74Cys, 192Cys and 196Cys) in the large subunit of human GGT (West et al. 2011, 2013). The disulfide bond (50Cys and 74Cys) and N-glycosylation of 95Asn are essential for the autocatalytic cleavage of human GGT (West et al. 2011, 2013). 50Cys and 74Cys are located in the 1st and 2nd α-helices of human GGT, respectively (Fig. 3). These cysteines are conserved in MA0196 GGT (69Cys and 93Cys) and GGT from A. sojae (62Cys and 86Cys), whereas cysteines (26Cys and 56Cys) in GGTA from A. oryzae are clearly in different positions. Although these cysteine residues must form an intramolecular disulfide bond in MA0196 GGT for autocatalytic cleavage and enzyme activity, it is unclear whether this disulfide bond contributes to structural stability and salt-tolerance of the large subunit. In future research efforts, sequence comparison of the large subunits among Aspergillus GGTs, including MA0196 GGT, and protein engineering (e.g., site-direct mutagenesis) will be used to enhance stability and halo-tolerance of GGTs potentially.
Some members of the GGT protein family possess an additional flexible loop covering the substrate-binding cleft, known as the lid-loop region in the small subunit (Verma et al. 2015). Phylogenetic analysis of GGTs indicates that GGTs can be classified as Gram-negative bacteria- and eukaryote-GGTs with the lid-loop region and Bacillus- and extremophile bacteria-GGTs without the lid-loop region (Verma et al. 2015). This lid-loop consists of a small stretch of ~ 12 amino acid residues (e.g., 438Pro to 449Gly in E. coli GGT, 428Ser to 438Ser in human GGT, Fig. 3) and may affect the transpeptidation reaction in GGTs (Okada et al. 2006; Calvio et al. 2018). MA0196 GGT also possesses this additional flexible loop (450Asn to 460Ser). Bacillus GGTs possess an extra sequence composed of 14 consecutive residues (e.g., 389Tyr to 402Gln) (Saini et al. 2021), whereas MA0196 GGT and GGTs from E. coli and human lack this sequence.
Prediction of the glycosylation sites in γ-glutamyl transpeptidase from A. sydowii by homology modeling
N-glycosylation contributes to protein stability, activity, and protease resistance (Cavalcanti et al. 2018; Liu et al. 2018; Hu et al. 2019; Lim et al. 2019). N-glycosylation of human GGT is not a requisite feature for enzyme kinetics of the mature enzyme (West et al. 2011), but N-glycosylation of 95Asn facilitates proper folding of the human GGT propeptide into a conformation that induces autocatalytic cleavage into the mature heterodimeric enzyme (West et al. 2013). Six N-glycosylation sites on the surface of the large subunit of human GGT have been identified (95Asn, 120Asn, 230Asn, 266Asn, 297Asn, and 344Asn), whereas only one site (511Asn) has been identified on the small subunit (Fig. 3) (West et al. 2013). The heterologous expression of human GGT by a yeast recombinant has been optimized for protein structure and glycan analyses (West et al. 2013). However, purification of native MA0196 GGT in sufficient quantities was not possible for such structural analyses, and a heterologous overexpression system has not been established. MA0196 GGT has twelve potential N-glycosylation sites based on the Asn-X-Thr/Ser consensus sequence (Fig. 3). Thus, sequence and protein structure homology analyses were performed to predict N-glycosylation sites in MA0196 GGT (Figs. 3 and S6). LC–MS–MS Mascot analysis of glycosylated and deglycosylated MA0196 GGTs suggested that there were ten predicted N-glycosylated sites, except at 450Asn and 484Asn. A comparison of the primary structures of MA0196 and human GGTs yielded a sequence identity of 37% (198/533 a.a.). Structural comparison of MA0196 GGT with human GGT revealed that 19 α-helices and 20 β-strands are conserved (Figs. 3 and S6). In addition, 95Asn, 120Asn, and 266Asn in human GGT and 116Asn, 139Asn, and 287Asn in MA0196 GGT are located at similar positions in each structure (Figs 3 and S6). 450Asn and 484Asn of MA0196 GGT are located in loop regions (Fig. S6).
Comparison of salt-tolerant MA0196 GGT with non-salt-tolerant GGT from A. oryzae
The effect of NaCl on the activity indicated that GGTs from B. subtilis and E. coli are categorized as salt-tolerant and non-salt-tolerant types, respectively (Yang et al. 2011; Wada et al. 2010). There is a notable difference in the electrostatic surface potentials between these GGTs in a high-salt solution (Wada et al. 2010). The Bacillus GGT and Escherichia GGT have a negatively charged surface, consistent with their theoretical isoelectric point (pI) values (4.95 and 5.38, respectively) calculated from the amino acid sequences (Yang et al. 2011). The negative charge was eliminated from the surface of E. coli GGT in the presence of 3 M NaCl, whereas Bacillus GGT maintains its water-binding capacity and solubility, which results in its salt tolerance (Yang et al. 2011; Wada et al. 2010). A comparative sequence and structural analysis between salt-tolerant and non-salt-tolerant GGTs indicated that solvent-exposed negatively charged residues should play a major role in halotolerance (Pica et al. 2013). Here, MA0196 GGT from A. sydowii and its homologous GGTA from A. oryzae are probably salt-tolerant and non-salt-tolerant types, respectively (Fig. 2a). The physicochemical parameters of these two GGTs were compared (Table S6). The negatively charged protein surface of GGTA from A. oryzae is higher than that of MA0196 GGT, and the ratio of acidic amino acids (Glu and Asp) to basic amino acids (Lys, His, and Arg), which is termed the AB ratio, for GGTA from A. oryzae (1.23) is higher than that of MA0196 GGT (1.03) (Table S6). Consequently, the pI value of GGTA from A. oryzae (pI = 4.72) is lower than that of MA0196 GGT (pI = 5.26). The difference in the electrostatic surface potentials of these enzymes may explain the salt adaptation mechanism. Only the number of N-glycosyl consensus sequences (Asn-Xaa-Thr/Ser) in MA0196 GGT is higher than that in GGTA from A. oryzae. In an N-glycosylated aspartic protease, N-glycosylation probably contributes to thermostability and water-activity tolerance (Lim et al. 2019). Disulfide bond(s) and/or high N-glycosylation levels in MA0196 GGT may contribute to the stability of the protein structure under high salt conditions rather than the negatively charged protein surface. Characterization of MA0196 GGT deglycosylated by N-glycosidase and deglycosylated mutants will reveal the functional role of N-glycosylation in MA0196 GGT.
Conclusions
γ-Glutamyl transpeptidase and its gene from A. sydowii MA0196 (MA0196 GGT) were characterized. MA0196 GGT that retained transpeptidation and hydrolysis activities and was stable in 20% (w/v) NaCl can be categorized as a salt-tolerant enzyme when compared with non-salt-tolerant GGTA from A. oryzae. MA0196 GGT showed high γ-glutamyl translation activity toward β-alanyl-L-histidine dipeptides, carnosine, and anserine. The enzymatic conversion of the γ-glutamyl tripeptide from carnosine by MA0196 GGT was enhanced at pH 9.5 in the presence of a small amount of DMSO. These enzymatic characteristics make MA0196 GGT potentially suitable for applications in salting-out fermentation and food processing and the production of the γ-glutamyl tripeptide from carnosine in fish waste materials. For this purpose, it is important to establish heterologous overexpression of MA0196 GGT. The phylogenetic relationships of MA0196 GGT with GGTs from Aspergillus spp. including koji molds indicated that MA0196 GGT is a different type of GGT than GGTs from A. oryzae. However, the small subunit of MA0196 GGT showed ~ 80% identity with those of several Aspergillus GGTs. Future sequence and structural analyses between salt-tolerant and non-salt-tolerant fungal GGTs should facilitate our understanding of the salt adaptation mechanism of MA0196 GGT.
References
Abbas A, Koc H, Liu F, Tien M (2005) Fungal degradation of wood: initial proteomic analysis of extracellular proteins of Phanerochaete chrysosporium grown on oak substrate. Curr Genet 47:49–56. https://doi.org/10.1007/s00294-004-0550-4
Astoreca A, Magnoli C, Ramirez ML, Combina M, Dalcero A (2007) Water activity and temperature effects on growth of Aspergillus niger, A. awamori and A. carbonarius isolated from different substrates in Argentina. Int J Food Microbiol 119(3):314–318. https://doi.org/10.1016/j.ijfoodmicro.2007.08.027
Babizhayev MA, Deyev AI (2012) Management of the virulent influenza virus infection by oral formulation of nonhydrolized carnosine and isopeptide of carnosine attenuating proinflammatory cytokine-induced nitric oxide production. Am J Therap 19(1):e25-47. https://doi.org/10.1097/MJT.0b013e3181dcf589
Bello MH, Epstein L (2013) Clades of γ-glutamyltransferases (GGTs) in the ascomycota and heterologous expression of Colletotrichum graminicola CgGGT1, a member of the pezizomycotina-only GGT clade. J Microbiol 51:88–99. https://doi.org/10.1007/s12275-013-2434-0
Bindal S, Gupta R (2016) Thermo- and salt-tolerant chitosan cross-linked γ-glutamyl transpeptidase from Bacillus licheniformis ER15. Int J Biol Macromol 91:544–553. https://doi.org/10.1016/j.ijbiomac.2016.05.106
Calvio C, Romagnuolo F, Vulcano F, Speranza G, Morelli CF (2018) Evidences on the role of the lid loop of γ-glutamyltransferases (GGT) in substrate selection. Enzyme Microbial Technol 114:55–62. https://doi.org/10.1016/j.enzmictec.2018.04.001
Castellano I, Merlino A (2012) γ-Glutamyltranspeptidases: sequence, structure, biochemical properties, and biotechnological applications. Cell Mol Life Sci 69:3381–3394. https://doi.org/10.1007/s00018-012-0988-3
Cavalcanti RMF, Jorge JA, Guimarães LHS (2018) Characterization of Aspergillus fumigatus CAS-21 tannase with potential for propyl gallate synthesis and treatment of tannery effluent from leather industry. 3 Biotech 8:270. https://doi.org/10.1007/s13205-018-1294-z
Cho HB, Ahn JH, Yang HG, Lee J, Park WJ, Kim YW (2021) Effects of pH and NaCl on hydrolysis and transpeptidation activities of a salt-tolerant γ-glutamyltranspeptidase from Bacillus amyloliquefaciens S0904. Food Sci Biotechnol 30:853–860. https://doi.org/10.1007/s10068-021-00928-6
Demir H, Tari C (2016) Bioconversion of wheat bran for polygalacturonase production by Aspergillus sojae in tray type solid-state fermentation. Int Biodeterior Biodegrad 106:60–66. https://doi.org/10.1016/j.ibiod.2015.10.011
El Halouat A, Debevere JM (1996) Influence of modified atmosphere and preservatives on the growth of Zygosaccharomyces rouxii isolated from dried fruits. Int J Food Microbiol 33(2–3):219–229. https://doi.org/10.1016/0168-1605(96)01158-0
Esawy MA, Awad GEA, Wahab WAA, Elnashar MMM, El-Diwany A, Easa SMH, El-beih FM (2016) Immobilization of halophilic Aspergillus awamori EM66 exochitinase on grafted k-carrageenan-alginate beads. 3 Biotech 6:29. https://doi.org/10.1007/s13205-015-0333-2
Fuke S, Konosu S (1991) Taste-active components in some foods: a review of Japanese research. Physiol Behavior 49(5):863–868. https://doi.org/10.1016/0031-9384(91)90195-T
Gekas V, Gonzalez C, Sereno A, Chiralt A, Fito P (1998) Mass transfer properties of osmotic solutions. I. Water activity and osmotic pressure. Int J Food Properties 1(2):95–112. https://doi.org/10.1080/10942919809524570
Gobbetti M, Burzigotti R, Smacchi E, Corsetti A, De Angelis M (1997) Microbiology and biochemistry of gorgonzola cheese during ripening. Int Dairy J 7(8–9):519–529. https://doi.org/10.1016/S0958-6946(97)00047-2
Hosokawa M, Sakakibara H, Yajima I, Hayashi K (1990) Non-volatile flavor components in dorsal, abdominal and red meat parts of dried skipjack (katsuobushi). Nippon Shokuhin Kogyo Gakkaishi 37(11):856–861. https://doi.org/10.3136/nskkk1962.37.11_856
Hu X, Yuan X, He N, Zhuang TZ, Wu P, Zhang G (2019) Expression of Bacillus licheniformis α-amylase in Pichia pastoris without antibiotics-resistant gene and effects of glycosylation on the enzymic thermostability. 3 Biotech 9:427. https://doi.org/10.1007/s13205-019-1943-x
Ito K, Matsuyama A (2021) Koji molds for Japanese soy sauce brewing: characteristics and key enzymes. J Fungi 7(8):658. https://doi.org/10.3390/jof7080658
Ito K, Hanya Y, Koyama Y (2013) Purification and characterization of a glutaminase enzyme accounting for the majority of glutaminase activity in Aspergillus sojae under solid-state culture. Appl Microbiol Biotechnol 97:8581–8590. https://doi.org/10.1007/s00253-013-4693-4
Jiménez-Gómez I, Valdés-Muñoz G, Moreno-Perlin T, Mouriño-Pérez RR, Sánchez-Carbente MR, Folch-Mallol JL, Pérez-Llano Y, Gunde-Cimerman N, Sánchez NC, Batista-García RA (2020) Haloadaptative responses of Aspergillus sydowii to extreme water deprivation: Morphology, compatible solutes, and oxidative stress at NaCl saturation. J Fungi 6(4):316. https://doi.org/10.3390/jof6040316
Koibuchi K, Nagasaki H, Yuasa A, Kataoka J, Kitamoto K (2000) Molecular cloning and characterization of a gene encoding glutaminase from Aspergillus oryzae. Appl Microbiol Biotechnol 54:59–68. https://doi.org/10.1007/s002530000329
Kuroda M, Ohotake R, Suzuki E, Harada T (2000) Investigation on the formation and the determination of γ-glutamyl-β-alanyl histidine and related isopeptide in the macromolecular fraction of beef soup stock. J Agric Food Chem 48(12):6317–6324. https://doi.org/10.1021/jf001095r
Li Z, Zhu R, Liu Y, Li J, Gao H, Hu N (2020) γ-Glutamyltranspeptidase from Bacillus amyloliquefaciens: transpeptidation activity enhancement and l-theanine production. Enzyme Microb Technol 140:109644. https://doi.org/10.1016/j.enzmictec.2020.109644
Lim L, Senba H, Kimura Y, Yokota S, Doi M, Yoshida K, Takenaka S (2019) Influences of N-linked glycosylation on the biochemical properties of aspartic protease from Aspergillus glaucus MA0196. Proc Biochem 79:74–80. https://doi.org/10.1016/j.procbio.2018.12.017
Lin LL, Chi MC, Lan YJ, Lin MG, Juang TY, Wang TF (2018) Facile immobilization of Bacillus licheniformis γ-glutamyltranspeptidase onto graphene oxide nanosheets and its application to the biocatalytic synthesis of γ-l-glutamyl peptides. Int J Biol Macromol 117:1326–1333. https://doi.org/10.1016/j.ijbiomac.2017.11.153
Liu Y, Yang S, Yan Q, Liu J, Jiang Z (2018) High-level expression of a novel protease-resistant α-galactosidase from Thielavia terrestris. Proc Biochem 71:82–91. https://doi.org/10.1016/j.procbio.2018.05.025
Lo HF, Chi MC, Lin MG, Lan YG, Wang TF, Lin LL (2018) Protective effect of biological osmolytes against heat- and chaotropic agent-induced denaturation of Bacillus licheniformis γ-glutamyl transpeptidase. J Microbiol Biotechnol 28(9):1457–1466. https://doi.org/10.4014/jmb.1805.05005
Meister A, Tate SS, Griffith OW (1981) γ-Glutamyl transpeptidase. Method Enzymol 77:237–253. https://doi.org/10.1016/S0076-6879(81)77032-0
Minami H, Suzuki H, Kumagai H (2003) Salt-tolerant γ-glutamyltranspeptidase from Bacillus subtilis 168 with glutaminase activity. Enzyme Microb Technol 32(3–4):431–438. https://doi.org/10.1016/S0141-0229(02)00314-9
Morelli CF, Calvio C, Biagiotti M, Speranza G (2014) pH-Dependent hydrolase, glutaminase, transpeptidase and autotranspeptidase activities of Bacillus subtilis γ-glutamyltransferase. FEBS J 281(1):232–245. https://doi.org/10.1111/febs.12591
Mu W, Zhang T, Jiang B (2015) An overview of biological production of l-theanine. Biotechnol Adv 33(3–4):335–342. https://doi.org/10.1016/j.biotechadv.2015.04.004
Musa H, Kasim FH, Gunny AAN, Gopinath SCB (2018) Salt-adapted moulds and yeasts: potentials in industrial and environmental biotechnology. Proc Biochem 69:33–44. https://doi.org/10.1016/j.procbio.2018.03.026
Nakagawa H, Oyama T (2019) Molecular basis of water activity in glycerol–water mixtures. Front Chem 7:731. https://doi.org/10.3389/fchem.2019.00731
Okada T, Suzuki H, Wada K, Kumagai H, Fukuyama K (2006) Crystal structures of γ-glutamyltranspeptidase from Escherichia coli, a key enzyme in glutathione metabolism, and its reaction intermediate. PNAS 103(17):6471–6476. https://doi.org/10.1073/pnas.0511020103
Pica A, Krauss IR, Castellano I, La Cara F, Graziano G, Sica F (2013) Effect of NaCl on the conformational stability of the thermophilic γ-glutamyltranspeptidase from Geobacillus thermodenitrificans: Implication for globular protein halotolerance. Biochim Biophys Acta 1834:149–157. https://doi.org/10.1016/j.bbapap.2012.09.014
Qiu W, Li J, Wei Y, Fan F, Jiang J, Liu M, Han X, Tian C, Zhang S, Zhuo R (2020) Genome sequencing of Aspergillus glaucus ‘CCHA’ provides insights into salt-stress adaptation. Peer J 8:e8609. https://doi.org/10.7717/peerj.8609
Saini M, Bindal S, Gupta R (2017) Heterologous expression of γ-glutamyl transpeptidase from Bacillus atrophaeus GS-16 and its application in the synthesis of γ-d-glutamyl-l-tryptophan, a known immunomodulatory peptide. Enzyme Microbial Technol 99:67–76. https://doi.org/10.1016/j.enzmictec.2017.01.003
Saini M, Kashyap A, Bindal S, Saini K, Gupta R (2021) Bacterial gamma-glutamyl transpeptidase, an emerging biocatalyst: insights into structure–function relationship and its biotechnological applications. Front Microbiol 12:641251. https://doi.org/10.3389/fmicb.2021.641251
Santos TC, Reis NS, Silva TP, Bonomo RCF, Aguiar-Oliveira E, de Oliveira JR, Franco M (2018) Production, optimisation and partial characterisation of enzymes from filamentous fungi using dried forage cactus pear as substrate. Waste Biomass Valoriz 9:571–579. https://doi.org/10.1007/s12649-016-9810-z
Sapna SB (2014) Phytase production by Aspergillus oryzae in solid-state fermentation and its applicability in dephytinization of wheat ban. Appl Bioechm Biotechnol 173:1885–1895. https://doi.org/10.1007/s12010-014-0974-3
Sekizaki H, Murakami M, Itoh K, Toyota E, Tanizawa K (2000) Chum salmon trypsin-catalyzed peptide synthesis with inverse substrates as acyl donor components at low temperature. J Mol Cat B: Enzymatic 11(1):23–28. https://doi.org/10.1016/S1381-1177(00)00192-2
Shuai Y, Zhang T, Mu W, Jiang B (2011) Purification and characterization of γ-glutamyltranspeptidase from Bacillus subtilis SK11.004. J Agric Food Chem 59(11):6233–6238. https://doi.org/10.1021/jf2003249
Takahata Y, Inoue M, Kim K, Iio Y, Miyamaoto M, Masui R, Ishihama Y, Kuramitsu S (2012) Close proximity of phosphorylation sites to ligand in the phosphoproteome of the extreme thermophile Thermus thermophilus HB8. Proteomics 12(9):1414–1430. https://doi.org/10.1002/pmic.201100573
Takenaka S, Lim L, Fukami T, Yokota S, Doi M (2019) Isolation and characterization of an aspartic protease able to hydrolyze and decolorize heme proteins from Aspergillus glaucus. J Sci Food Agric 99(4):2042–2047. https://doi.org/10.1002/jsfa.9339
Takenaka S, Nakabayashi R, Ogawa C, Kimura Y, Yokota S, Doi M (2020) Characterization of surface Aspergillus community involved in traditional fermentation and ripening of katsuobushi. Int J Food Microbiol 327:108654. https://doi.org/10.1016/j.ijfoodmicro.2020.108654
Takenaka S, Ogawa C, Uemura M, Umeki T, Kimura Y, Yokota S, Doi M (2021) Identification and characterization of extracellular enzymes secreted by Aspergillus spp. involved in lipolysis and lipid-antioxidation during katsuobushi fermentation and ripening. Int J Food Microbiol 353:109299. https://doi.org/10.1016/j.ijfoodmicro.2021.109299
Verma VV, Gupta R, Goel M (2015) Phylogenetic and evolutionary analysis of functional divergence among gamma glutamyl transpeptidase (GGT) subfamilies. Biol Direct 10:49. https://doi.org/10.1186/s13062-015-0080-7
Virgili R, Saccani G, Gabba L, Tanzi E, Bordini CS (2007) Changes of free amino acids and biogenic amines during extended ageing of Italian dry-cured ham. LWT-Food Sci Technol 40(5):871–878. https://doi.org/10.1016/j.lwt.2006.03.024
Wada K, Irie M, Suzuki H, Fukuyama K (2010) Crystal structure of the salt-tolerant γ-glutamyltranspeptidase from Bacillus subtilis in complex with glutamate reveals a unique architecture of the solvent-exposed catalytic pocket. FEBS J 277(4):1000–1009. https://doi.org/10.1111/j.1742-4658.2009.07543.x
West MB, Wickham S, Quinalty LM, Pavlovicz RE, Li C, Hanigan MH (2011) Protein structure and folding autocatalytic cleavage of human γ-glutamyl transpeptidase is highly dependent on N-glycosylation at Asparagine 95. J Biol Chem 286(33):28876–28888. https://doi.org/10.1074/jbc.M111.248823
West MB, Chen Y, Wickham S, Heroux A, Cahill K, Hanigan MH, Mooers BHM (2013) Novel insights into eukaryotic γ-glutamyltranspeptidase 1 from the crystal structure of the glutamate-bound human enzyme. J Biol Chem 288(44):31902–31913. https://doi.org/10.1074/jbc.M113.498139
Williams JP, Hallsworth JE (2009) Limits of life in hostile environments: no barriers to biosphere function? Environ Microbiol 11(12):3292–3308. https://doi.org/10.1111/j.1462-2920.2009.02079.x
Wu G (2020) Important roles of dietary taurine, creatine, carnosine, anserine and 4-hydroxyproline in human nutrition and health. Amino Acids 52:329–360. https://doi.org/10.1007/s00726-020-02823-6
Yang JC, Liang WC, Chen YY, Chi MC, Lo HF, Chen HL, Lin LL (2011) Biophysical characterization of Bacillus licheniformis and Escherichia coli γ-glutamyltranspeptidases: a comparative analysis. Int J Biol Macromol 48(3):414–422. https://doi.org/10.1016/j.ijbiomac.2011.01.006
Yang J, Bai W, Zeng X, Cui C (2019) Gamma glutamyl peptides: The food source, enzymatic synthesis, kokumi-active and the potential functional properties – A review. Trends Food Sci Technol 91:339–346. https://doi.org/10.1016/j.tifs.2019.07.022
Zacharius RM, Zell TE, Morrison JH, Woodlock JJ (1969) Glycoprotein staining following electrophoresis on acrylamide gels. Anal Biochem 30(1):148–152. https://doi.org/10.1016/0003-2697(69)90383-2
Acknowledgements
We thank Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Funding
This work was supported by the Japan Society for the Promotion of Science KAKENHI [Grant No. 20K05808].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
No human or animal experiments were concluded in this study.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nishikawa, A., Senba, H., Kimura, Y. et al. Isolation and characterization of a salt-tolerant γ-glutamyl transpeptidase from xerophilic Aspergillus sydowii. 3 Biotech 12, 253 (2022). https://doi.org/10.1007/s13205-022-03259-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03259-3