Abstract
Tulasnella spp. are the main fungal symbionts of Brazilian Epidendrum orchids. The taxonomy of these fungi is largely based on ITS rDNA similarity, but culture dependent techniques are still essential to establish the true biological entity of the mycobiont. The aim of this study was to characterize morphologically and molecularly 16 Tulasnella spp. fungi isolated from three different populations of E. secundum and to test the coincidences between morphological and molecular characterization. Two uninucleate rhizoctonia fungi, obtained from Oncidium barbaceniae, and two phytopathogenic isolates were included as outgroups. Qualitative and quantitative morphological characteristics were analyzed using multivariate statistics and were able to distinguish Ceratobasidium, Tulasnella and Thanatephorus genera and separate the isolates of Tulasnella spp. into two groups. Analysis of RAPD (Random Amplified Polymorphic DNA) and ITS rDNA sequences validated the morphological data. Symbionts of O. barbaceniae presented identity to ITS sequences of Ceratobasidium genus, while E. secundum isolates presented identity to two species of Tulasnella. We observed homogeneity among Tulasnella spp. obtained from a single population and from neighboring populations, but there was higher variability among isolates obtained from populations of regions that were farther apart. Morphological data associated with multivariate statistics proved to be a useful tool in the multi-level taxonomy of these orchid-associated fungi and in estimating the diversity of orchid mycorrhizal fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The Tulasnella genus belongs to the rhizoctonia alliance (Roberts 1999; Dearnaley et al. 2012), a polyphyletic grouping of filamentous fungi that share common features in their anamorphic states (García et al. 2006). Tulasnella comprises cosmopolitan fungi, also often described as saprophytic and orchid mycorrhizal fungi (Roberts 1999). In Brazil, many orchid mycorrhizal fungi are described as Epulorhiza (Nogueira et al. 2005; Pereira et al. 2005a, 2009; Pereira and Valadares 2012), the anamorphic stage of Tulasnella. Some Tulasnella isolates have the ability to promote germination of orchid seeds and some of the isolates have been used in the symbiotic propagation of orchids (Zettler 1997; Pereira et al. 2005b, 2011). Moreover, the dependency of lithophytic and epiphytic orchids on mycorrhizal fungi throughout their life cycle is not surprising as the mycobionts may improve access to water and minerals, which can be extremely limiting in these niches, by expanding their surface of nutrient exchange (Osorio-Gil et al. 2008; Dearnaley et al. 2012; Nurfadilah et al. 2013).
The mycorrhizal fungi isolated from Epidendrum secundum Jacq., a very common species in different habitats of the Serra do Brigadeiro State Park (Araponga/MG, Brazil), have also been characterized morphologically and identified as Tulasnella spp. (Pereira et al. 2009). In the previous work, morphological features were assessed from in vitro cultures and successfully used to separate the isolates into three groups. The authors observed low quantitative variability among fungal isolates from the same location but high variability among isolates from different sites.
Isolates discerned by ITS sequences can be different in functional traits. Pereira et al. (2011) selected isolates representing three groups observed by Pereira et al. (2009) and evaluated the ability of different Tulasnella spp. to promote the germination of E. secundum seeds. All isolates of Tulasnella promoted seed germination, but displayed different percentages of seed germination and protocorm development. One isolate was more efficient, even considering others that were grouped together in the morphological analysis (25 % of seed germination and 11 % of seedlings with first leave and root after 44 days of inoculation). It is thus necessary to validate the extent to which different morphological groups truly represent different species and to determine the taxonomic relationship between groups based on fungal multi-level taxonomy. Thus, we need to correlate seed germination data, morphological traits and molecular features to discuss ecological interactions between the orchid and Tulasnella.
Molecular techniques based on DNA sequences are often used to evaluate the variability among isolates of rhizoctonias, both pathogenic and mycorrhizal (Rasmussen 2002; Sharon et al. 2006; Dearnaley 2007; Oliveira et al. 2014). The RAPD technique (Random Amplified Polymorphic DNA) enables revealing a genome-scale polymorphism between isolates, without prior knowledge of the genome sequence (Shan et al. 2002). This technique makes it possible to determine the variability at the species level and at the population level (Horton and Bruns 2001; Shan et al. 2002). Sequence analysis of the ITS region, in turn, has been used for the identification and diversity discovery of rhizoctonia fungi in several studies (Chutima et al. 2011; Cruz et al. 2011; Chen et al. 2012; Martos et al. 2012; Okayama et al. 2012; Valadares et al. 2012; Oliveira et al. 2014). Finally; integration of morphological characterization and ITS analysis has been used to present more accurate identification of fungi (Cruz et al. 2011; Valadares et al. 2012).
The objective of the present study was to characterize morphologically and molecularly (by combining RAPD and ITS sequence analysis), 16 Tulasnella spp. fungi isolated from different populations of E. secundum, and to test the coincidences between morphological and molecular characterization. Two uninucleate rhizoctonia fungi, obtained from Oncidium barbaceniae Lindl., and two phytopathogenic isolates, Ceratobasidium sp. AGC and Thanatephorus cucumeris (A.B. Frank) Donk AG1IA, were included as outgroups.
2 Materials and methods
2.1 Fungal isolation
Twenty rhizoctonia fungi were used in this work. Sixteen are Tulasnella spp. isolates and two are uninucleate rhizoctonia species, all belonging to the Orchids Mycorrhizal Fungi Collection from the Mycorrhizal Associations Laboratory (Department of Microbiology/BIOAGRO—Institute of Research Applied to Agriculture/Federal University of Viçosa, Brazil). The other two fungi belong to the Phytopathogenic Fungi Collection (Department of Phytopathology, Federal University of Viçosa, Brazil), one being classified as Thanatephorus cucumeris AG1IA (AG1) and the other as Ceratobasidium sp. AGC.
The 16 Tulasnella spp. isolates were isolated from three distinct E. secundum populations, ES1, ES2 and ES3 (Pereira et al. 2009). The two uninucleate rhizoctonia fungi were isolated from O. barbaceniae population OB1. The orchid populations were located at the same high altitude rocky terrain, located in the Serra do Brigadeiro State Park, MG, Brazil (Caiafa and Silva 2005, 2007). The ES1, ES2 and OB1 populations grow in grasslands (Caiafa and Silva 2007) at geographic coordinates S20°42′11.1″/WO42°28′32″, S20°42′10.8″/WO42°28′31.8″ and S20°42′8.38″/WO42°28′30.53″, respectively. The ES3 population grows in grasslands close to shrubs (Caiafa and Silva 2007) at geographic coordinates S20°42′6.5″/WO42°28′30.1″. Isolates were reactivated in Petri dishes containing 25 mL of PDA medium (Potato Dextrose Agar—ACUMEDIA, Neogen Corporation, Lansing, Michigan/MI, USA) from cultures stored at 4 °C in tubes containing 10 mL of PDA medium.
2.2 Morphological characterization
For the morphological characterization, qualitative and quantitative characteristics were evaluated as described by Pereira et al. (2009). Quantitative characteristic data were statistically analyzed using the GENES program version 2007.0.0 (Cruz 2008). The variance was analyzed taking into account the completely randomized design. The averages were compared using the Scott Knott test (P < 0.05). Colony diameter and growth rates were evaluated 48 h after inoculation on both PDA and CMA media (three replicates, obtaining six measurements for each variable). The width (WMC) and length (LMC) of monilioid cells and hyphae diameter (HD) were obtained from 20 replicate cells for each trait. All quantitative measures were arranged in ascending order to select four values: the highest, lowest and two intermediate values to include the entire range of variation. Quantitative and qualitative morphological characteristics were analyzed by biometric techniques to generate distance values and to cluster the isolates using the GENES software (Cruz 2008; Pereira et al. 2009). Qualitative characteristics were analyzed through multi-categorical analyses to calculate the dissimilarity (D) matrix, i.e., 1-similarity, to subsequent clustering of isolates by the UPGMA method (Unweighted Pair Group Method using Arithmetic Averages) (Cruz 2008). Quantitative characteristics were analyzed using canonical variables for graphic dispersion of the isolates based on the first two canonical variables (Cruz 2008). The Mahalanobis distance (D2) was also calculated, generating a distance matrix used for the clustering of isolates by the UPGMA method (Cruz 2008). The quantitative characteristics were subsequently tested using the Singh (1981) criterion, based on D2, to assess the relative contribution of these characteristics in the analysis of diversity and to generate suggestions for disposal (Cruz 2008). The correlation between D and D2 matrices was analyzed using the Mantel test (Mantel 1967).
2.3 Biological material and DNA extraction
Fungal mycelia were cultivated for DNA extraction. Disks (9 mm in diameter) containing an active growing mycelium of each isolate were inoculated into 50 mL Erlenmeyer flasks containing 10 mL of PDB medium (Potato Dextrose Broth—ACUMEDIA, Neogen Corporation, Lansing, Michigan/MI, USA) and incubated at 28 °C for 5 to 10 days, depending on fungal growth rate.. Obtained mycelia were transferred to 1.5 mL microtubes, frozen at −86 °C and lyophilized prior to DNA isolation, according to Schäfer and Wöstemeyer (1992). DNA samples were suspended in 20 μL of ultra-pure water (MilliQ water) and quantified by electrophoresis on a 0.8 % agarose gel (Promega Corporation, Madison, USA) stained with 0.5 mg mL−1 ethidium bromide. Different concentrations of λ vector were used as a standard. DNA samples were stored at −20 °C.
2.4 Molecular analysis by RAPD
RAPD reactions were carried out as described by Junghans et al. (1998). The amplification products were electrophoresed through a 1.5 % agarose gel containing 0.5 mg mL−1 ethidium bromide. The gels were scanned using an image processing system (Eagle Eye II - Stratagene, La Jolla, USA). Scanned images were analyzed by the Gel-Pro Analyzer (version 3.1.00.00; NIPPON ROPER Co., Ltd., Tokyo, Japan) to construct tables showing the presence or absence of bands. Thereafter, the data were analyzed using GENES software for construction of the Jaccard distance (J) matrix (Cruz 2008) and grouping of the isolates using the UPGMA technique (Cruz 2008). The Mantel test (Mantel 1967) was applied to compare J matrix against the D and D2 matrices.
2.5 Amplification, sequencing and analysis of the ITS region
PCR was carried out using primers ITS1-ITS4 (White et al. 1990) and the reaction conditions described by Gardes and Bruns (1993). PCR products were purified using Exo-Sap (USB Corporation, Cleveland, Ohio), as recommended by the manufacturer. Both strands (forward and reverse) were sequenced by the Macrogen Inc. company (South Korea). Amplified ITS sequences were assembled and edited with the software Sequencher version 4.5 (GeneCodes, Ann Arbor, USA). Subsequently, the sequences were aligned and forwarded to Mega software 4.0 (Tamura et al. 2007). Dendrograms were constructed using the neighbor-joining method with two-parameter Kimura nucleotide analysis and bootstrap with 5,000 repetitions, excluding gaps and missing data (Sharon et al. 2008). Random representative sequences from each clade were selected and searched with the algorithm BLASTn (Altschul et al. 1997) to select sequences from the NCBI database (GenBank, http://www.ncbi.nlm.nih.gov). Tulasnellaceae sequences obtained from the database and the Tulasnella spp. isolate sequences were then included in the alignment. Clustering of these organisms was performed as described above. The same procedure was performed for the T. cucumeris AG1IA, Ceratobasidium sp. AGC and uninucleate rhizoctonia sequences, as well as for their most similar sequences obtained from the Genbank database. Fungal sequences from this work were deposited in the NCBI database with the accession numbers HQ127084, HQ127085, HQ127086 and JX456553 to JX456569.
3 Results
3.1 Morphological characterization
The morphological characteristics analyzed showed variability among the isolates, making it possible to assign them to different genera of rhizoctonia fungi (Table 1). “Aerial mycelium” was the qualitative characteristic that displayed the most variability among the 16 isolates of E. secundum (Table 1), but there was no clear variation between the two uninucleate rhizoctonias obtained from O. barbaceniae with respect to this morphological characteristic. Additionally, pathogenic isolates Ceratobasidium sp. AGC and T. cucumeris AG1IA showed similarities with respect to qualitative features, but Ceratobasidium sp. AGC was more similar to the uninucleate rhizoctonias when comparing quantitative characteristics (Table 1).
Multi-categorical analysis of qualitative characteristics revealed low dissimilarity (D) between mycorrhizal fungi isolated from the same E. secundum population and from O. barbaceniae isolates. After analyzing these distances by the UPGMA method, we observed clustering of isolates belonging to each of the different rhizoctonia fungi genera (Fig. 1a). Symbionts of E. secundum were then distributed into two neighboring groups, with one group represented by isolates from populations very far apart from the other (Fig. 1a).
Cluster analysis of rhizoctonia fungal isolates based on qualitative (a) and quantitative (b) characteristics. a Qualitative data were converted to a matrix of dissimilarity using a multi-categorical technique, and the grouping was generated by the UPGMA method. b Quantitative data were converted to a matrix of Mahalanobis distance and the grouping was generated using the UPGMA method. Codes presented in the dendrogram correspond to the following: AG1, T. cucumeris AG1IA; AGC, Ceratobasidium sp. AGC; the other codes are related to strains of E. secundum as shown in Table 1. D, dissimilarity; D2, Mahalanobis distance; %, percentage relative to the maximum dissimilarity or Mahalanobis distance
Concerning the quantitative parameters, the largest calculated Mahalanobis distances (D2) were observed among isolates of Tulasnella spp. and T. cucumeris AG1IA. Among the isolates of Tulasnella spp. from different populations, the D2 values ranged from 1.01 to 353.69, whereas the D2 values were less than 64.00 among the isolates from the same population. An exception was observed for isolate ES2.2B, which presented D2 values greater than 149.00 when compared to isolates from the same population and was closer to isolates from the population ES3. The Mantel test shows that the correlation between D and D2 matrices was significant at 1 % of probability, with value of 0.81.
The dendrogram constructed using the UPGMA method based on Mahalanobis distances (Fig. 1b) and graphic dispersion of canonical variables separated the isolates into four groups. Isolates of Tulasnella spp. formed a group clearly distinct from the others and two sub-groups could also be distinguished. We also observed great similarity between the two rhizoctonia uninucleate fungi and Ceratobasidium sp. (Fig. 1b).
Using the Singh criterion (1981) based on the Mahalanobis distances, we observed that the quantitative characteristic RCMA (growth rate determined on CMA medium) contributed more to the estimation of diversity among the isolates, followed by the RPDA (growth rate determined on PDA medium) feature (Table 2). HD was the trait with the smallest contribution to the determination of this diversity. HD disposal was suggested by the Singh criterion (1981) (Table 2).
3.2 Molecular analysis by RAPD
From the comparison of rhizoctonia fungi using the RAPD technique, which was performed with nine random primers and 140 loci, the shortest distance in the dissimilarity matrix was observed between isolates of Tulasnella sp. The longest distance was between Tulasnella sp. ES1.2B and the uninucleate rhizoctonia OB1.3H isolate. Isolates from the same population of orchid had distances lower than 0.387, except for isolate ES2.2B, which showed a distance greater than 0.723 when compared to isolates from the ES2 population. By grouping the isolates using the dissimilarity matrix and the UPGMA method, seven groups were formed with a cut-off at 50 % dissimilarity (Fig. 2). Fungal isolates from the same orchid were placed in the same group, except for the ES2.2B isolate, which was not grouped with isolates from population 2 (Fig. 2). The ES2.2B, T. cucumeris AG1IA (AG1) and Ceratobasidium sp. AGC isolates showed no similarity to any other isolate, so each one of these isolates formed a distinct group (Fig. 2).
Cluster analysis of rhizoctonia fungal isolates based on RAPD results. The pattern of RAPD bands was converted to a dissimilarity matrix based on Jaccard distance to group isolates using the UPGMA method. Codes presented in the dendrogram correspond to the following: AG1, T. cucumeris AG1IA; AGC, Ceratobasidium sp. AGC; the other codes are related to strains of E. secundum as shown in Table 1. D, dissimilarity; %, percentage relative to the maximum dissimilarity
We observed significance, at 1 % of probability, after comparison between D against J (D x J) and D2 against J (D2 x J) matrices. When we compare the matrices generated with data obtained from the 20 isolates, the correlation of D x J was 0.60 and the correlation D2 x J was 0.37. However, the correlations between the matrices comprising data only from Tulasnella isolates were 0.81 to D x J and 0.72 to D2 x J.
3.3 ITS analysis
The ITS fragments of the 20 rhizoctonia fungi studied were approximately 600 bp long. The analysis of the Kimura 2-parameter based matrix led to the identification of two groups among isolates of Tulasnella spp.: the first group included isolates from the ES1 and ES2 populations, except for isolate ES2.2B, and the second group included ES2.2B with the isolates from the ES3 population (Fig. 3).
Cluster analysis of rhizoctonia fungal isolates based on ITS sequence. The sequence alignment was analyzed using the Kimura 2-parameter distance, the neighbor-joining method and bootstraps with 5,000 repetitions. Codes presented in the dendrogram correspond to the following: AG1, T. cucumeris AG1IA; AGC, Ceratobasidium sp. AGC; the other codes are related to strains of E. secundum as shown in Table 1
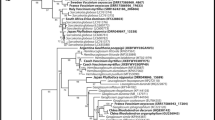
A comparison of the ITS sequences of our Tulasnella spp. with those in the NCBI database of Tulasnella showed that the isolates from the ES1 and ES2 populations, except for isolate ES2.2B, grouped with Tulasnella calospora Aut (Fig. 4). In contrast, the sequences of isolates from the ES3 population and ES2.2B were closer to Tulasnella albida Bourdot & Galzin, Tulasnella pruinosa Bourdot & Galzin and Tulasnella violea (Quél.) Bourdot & Galzin (Fig. 4). Sequences of uninucleate isolates and T. cucumeris AG1IA were grouped into a larger group (Fig. 5) that comprises sequences from R. solani (AF472512) and Thanatephorus cucumeris (Frank) Donk (EU244843). Uninucleate isolates showed identity to the sequences of the Ceratobasidium genus.
Cluster analysis of Tulasnella spp. isolates based on ITS sequence. The sequence alignment was analyzed using the Kimura 2-parameter distance, the neighbor-joining method and bootstraps with 5,000 repetitions. Codes in bold are related to strains of E. secundum as shown in Table 1
4 Discussion
The Tulasnella strains studied in this work had already been characterized morphologically by Pereira et al. (2009), but the authors did not conclude whether the observed variability was due to variation at population level or if each group of isolates represented different species. Based on morphological and molecular characterization, it was possible to confirm that there is variability among the fungi obtained from different populations of E. secundum and that the fungal isolates belong to two species of the genus Tulasnella. The morphological characteristics allowed the distinction of Tulasnella, Ceratobasidium and Thanatephorus (Table 1, Fig. 1) and molecular techniques further corroborated the morphological differences among the genera (Figs. 2 and 3).
The quantitative traits evaluated in this study had different relative contributions to our estimation of the diversity between rhizoctonia fungi (Table 2). Growth rates (RPDA and RCMA) followed by colony diameter (DPDA and DCMA) were the characteristics that contributed most to the observed variability. HD contributed less than 1 % to the variability observed and the Singh criterion suggested that HD be discarded. We can therefore suggest that HD is not useful in the estimation of variability in studies that aim to analyze diversity. However, HD and the monilioid cell dimensions (WMC and LMC) are important features in the screening between rhizoctonia genera and these features should be considered during the description of fungal species (Currah and Zelmer 1992; Pereira et al. 2003).
Morphological characteristics, such as the vegetative and reproductive structures, are widely used in the identification of fungal isolates (Currah and Zelmer 1992; Cruz et al. 2011). Given the existence of variation between the characteristics of cultures of the same species and the great difficulty of inducing the formation of reproductive structures of rhizoctonia fungi, the application of molecular techniques to aid in identification has been seen as essential (Horton and Bruns 2001; Dearnaley et al. 2012).
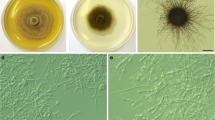
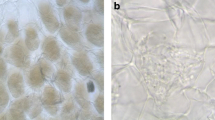
The 16 E. secundum symbionts showed the typical morphology described for the Tulasnella genus (Table 1) (Currah and Zelmer 1992; Pereira et al. 2003, 2005a, 2009; García et al. 2006). Morphological features and ITS analysis allowed for the separation of Tulasnella isolates into two sub-groups (Figs. 1, 3, 4 and 5) and these results showed that these isolates represent two species: Tulasnella sp.1 and Tulasnella sp.2.
The Tulasnella sp.1 comprises ES1 and ES2 population isolates, except for ES2.2B. The Tulasnella sp.1 ITS sequences showed identity to T. calospora (Fig. 4), which is a teleomorph without a corresponding anamorph. This species was proposed by Hadley (1970) to be a universal orchid symbiont due to its ability to establish in vitro symbiotic associations with a wide diversity of orchid species. The Tulasnella sp.2 are represented by the ES2.2B isolate and isolates of the ES3 population. The ITS sequence of Tulasnella sp.2 isolates did not show any identity to sequences of Tulasnella species in the NCBI database (Fig. 4).
Pereira et al. (2011) observed that different isolates of Tulasnella sp.1 and sp.2 are able to promote the germination of E. secundum seeds. However, the protocorms germinated in the presence of Tulasnella sp.1 developed much faster than in the presence of Tulasnella sp.2. In addition, Tulasnella sp.1 grows three times faster than that Tulasnella sp.2 in culture medium (Table 1). These two factors may be related to higher root colonisation of ES1 and ES2 populations by Tulasnella sp.1. Furthermore, factors linked to the interaction between fungus and adult plants in situ, which were not assessed in this study, may also contribute to the observed pattern. Indeed, in nature, orchid seed germination and root colonization are affected by factors related to orchid habitat and abiotic conditions, which influence the variation of mycorrhizal diversity, specificity and functioning (Martos et al. 2012).
Isolates with similar ITS sequences display different potential to promote seed germination. Eight isolates of Tulasnella sp.1 and five isolates of Tulasnella sp.2 promoted seed germination of E. secundum (Pereira et al. 2011). However, isolates from the same “species” were not similar in the percentage of total germination, development stage of protocorm and growth index. ITS similarity does not reflect the ability to promote seed germination; i.e., it is not obvious that isolates from the same ITS clade display the same germination potential. If morphological features reflect intraspecific variability, it would be possible to select different isolates from the same fungal species to test in orchid germination experiments based on its morphological variability.
We observed significant correlation between morphological and RAPD data. Nevertheless, the RAPD clusters were slightly different from morphological grouping. For example, the ES2.2B isolate showed great dissimilarity to the others Tulasnella isolates in RAPD grouping (Fig. 2) but grouped with isolates from ES3 population when considering morphological parameters. This discrepancy can be explained by the sensitivity of RAPD technique to variations between populations and between species (Shan et al. 2002; Sharon et al. 2006). Thus, it is likely that the ES2.2B isolate was not grouped together with the isolates from its population because it belongs to a different species. Additionally, ES2.2B and ES3 isolates, despite belonging to the same species, do not group together, most likely because they were obtained from different populations.
Uninucleate isolates of rhizoctonia fungi have been obtained from orchids in Brazil and Puerto Rico (Otero et al. 2002; Valadares et al. 2012). The isolates obtained from O. barbaceniae showed similar qualitative morphological features to the Ceratobasidium sp. obtained from Brazilian orchids (Table 1; Nogueira et al. 2005; Pereira et al. 2005a, c). These features suggest that these isolates belong to the genus Ceratobasidium. Uninucleate rhizoctonia isolates were also seen as aggressive pathogens in roots of Pinus sylvestris L. and showed growth characteristics and appearance similar to Ceratobasidium genera (Lilja et al. 1992; García et al. 2006). As Ceratobasidium fungi are more commonly binucleate (Currah and Zelmer 1992), ITS analysis was essential to confirm the identification of uninucleate isolates as Ceratobasidium sp. (Fig. 5). Nonetheless, the uninucleate fungi isolated from O. barbaceniae are mycorrhizal fungi, because they were able to induce germination of seeds from their host (data not published).
Analysis of the conserved regions of DNA by nucleotide sequencing agrees and adds significant information to the morphological identification (Pereira et al. 2005a; Chen et al. 2012; Valadares et al. 2012). Other studies have eliminated morphological characterization, and even isolation, using ITS sequence analysis to identify orchid mycorrhizal fungi (Chutima et al. 2011; Cruz et al. 2011; Okayama et al. 2012). As Tulasnella taxonomy is unclear, a multi-method approach should be applied to increase identification reliability. Species identification of isolates using only molecular techniques can be limited by the absence of identified reference cultures. Thus, in the possession of pure cultures of Tulasnella sp.1 and Tulasnella sp.2, the isolates can be used for species description and the culture can be stored for further applications.
Qualitative and quantitative morphological data made it possible to distinguish between the three genera of rhizoctonia fungi studied. The biometric analysis of quantitative features detected variability among isolates with similar ITS sequences and corroborated RAPD data. Thus, the integration of morphological traits with ITS sequences in a multilevel taxonomy can improve the understanding of the diversity and ecology of cultivable orchid mycorrhizal fungi and is useful for fungal selection for germination tests. This understanding allows a better comprehension of the specificity in orchid mycorrhizal association, which is essential to establish conservation strategies based on the symbiotic propagation of orchids (Zettler 1997; Dearnaley et al. 2012).
References
Altschul SF, Madden TL, Schaffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Caiafa AN, Silva AF (2005) Composição florística e espectro biológico de um campo de altitude no Parque Estadual da Serra do Brigadeiro, Minas Gerais—Brasil. Rodriguésia 56:163–173
Caiafa AN, Silva AF (2007) Structural analysis of the vegetation on a highland granitic rock outcrop in Southeast Brazil. Rev Brasil Bot 30:657–664
Chen J, Wang H, Guo SX (2012) Isolation and identification of endophytic and mycorrhizal fungi from seeds and roots of Dendrobium (Orchidaceae). Mycorrhiza 22:297–307
Chutima R, Dell B, Lumyong S (2011) Effects of mycorrhizal fungi on symbiotic seed germination of Pecteilis susannae (L.) Rafin (Orchidaceae), a terrestrial orchid in Thailand. Symbiosis 53:149–156
Cruz CD (2008) Programa genes: diversidade genética. Editora UFV, Viçosa
Cruz D, Suárez JP, Kottke I, Piepenbring M, Oberwinkler F (2011) Defining species in Tulasnella by correlating morphology and nrDNA ITS-5.8S sequence data of basidiomata from a tropical Andean Forest. Mycol Prog 10:229–238
Currah RS, Zelmer CD (1992) A key and notes for the genera of fungi with orchids and a new species in the genus Epulorhiza. Rep Tottori Mycol Inst 30:43–59
Dearnaley JDW (2007) Further advances in orchid mycorrhizal research. Mycorrhiza 17:475–486
Dearnaley JDW, Martos F, Selosse MA (2012) Orchid mycorrhizas: molecular ecology, physiology, evolution and conservation aspects. In: Hock B (ed) The Mycota IX: fungal associations, 2nd edn. Springer, Berlin
García VG, Onco MAP, Susan VR (2006) Review. Biology and Systematics of the form genus Rhizoctonia. Span J Agric Res 4:55–79
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycets—application to the identification of mycorrhizae and rust. Mol Ecol 2:113–118
Hadley G (1970) Non-specificity of symbiotic infection in orchid mycorrhiza. New Phytologist 69:1015–1023
Horton TR, Bruns TD (2001) The molecular revolution in ectomycorrhizal ecology: peeking into the black-box. Mol Ecol 10:1855–1871
Junghans DT, Gomes EA, Barros EG, Guimarães WV, Araújo EF (1998) Genetic diversity of the ectomycorrhizal fungus Pisolithus tinctorius based on RAPD-PCR analysis. Mycorrhiza 7:243–248
Lilja A, Lilja S, Poteri M, Ziren L (1992) Conifer seedling root fungi and root dieback in Finnish nurseries. Scand J For Res 7:547–556
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Martos F, Munoz F, Pailler T, Kottke I, Gonneau C, Selosse MA (2012) The role of epiphytism in architecture and evolutionary constraint within mycorrhizal networks of tropical orchids. Mol Ecol 21:5098–5109
Nogueira RE, Pereira OL, Kasuya MCM, Lanna MC, Mendonça M (2005) Fungos micorrízicos associados e orquídeas em campos rupestres na Região do Quadrilátero Ferrífero, Minas Gerais, Brasil. Acta Bot Bras 3:417–424
Nurfadilah S, Swarts ND, Dixon KW, Lambers H, Merritt DJ (2013) Variation in nutrient-acquisition patterns by mycorrhizal fungi of rare and common orchids explains diversification in a global biodiversity hotspot. Ann Bot 111:1233–1241
Okayama M, Yamato M, Yagame T, Iwase K (2012) Mycorrhizal diversity and specificity in Lecanorchis (Orchidaceae). Mycorrhiza 22(7):545–553
Oliveira SF, Bocayuva MF, Veloso TGR, Bazzolli DMS, Silva CC, Pereira OL, Kasuya MCM (2014) Endophytic and mycorrhizal fungi associated with roots of endangered native orchids from the Atlantic Forest, Brazil. Mycorrhiza 24(1):55–64
Otero JT, Ackerman JD, Bayman P (2002) Diversity and host specificity of endophytic Rhizoctonia-like fungi from tropical orchids. Am J Bot 89:1852–1858
Pereira MC, Valadares RBS (2012) Diversidade e aplicação dos fungos micorrízicos de orquídeas brasileiras. In: Pazza R, Souza EA, Pereira JD, Kavalco KF, Lisboa LCO, Reis LB, Pereira MC (eds) Biodiversidade em foco. Araucária Comunicação Integrada, Rio Paranaíba
Pereira OL, Rollemberg CL, Borges AC, Matsuoka K, Kasuya MCM (2003) Epulorhiza epiphytica sp. nov. isolated from mycorrhizal roots of epiphytic orchids in Brazil. Mycoscience 44:153–155
Pereira OL, Kasuya MCM, Borges AC, Araújo EF (2005a) Morphological and molecular characterization of mycorrhizal fungi isolated from neotropical orchids in Brazil. Can J Bot 83:54–65
Pereira OL, Kasuya MCM, Rollemberg CL, Borges AC (2005b) Indução in vitro da germinação de sementes de Oncidium flexuosum (Orchidaceae) por fungos micorrízicos rizoctonióides. Rev Bras Ciências Solo 29:199–206
Pereira OL, Kasuya MCM, Rollemberg CL, Borges GM (2005c) Isolamento e identificação de fungos micorrízicos rizoctonióides associados a três espécies de orquídeas epífitas neotropicais no Brasil. Rev Bras Ciências Solo 29:191–197
Pereira MC, Kasuya MCM, Pereira OL, Costa MD, Rocha RB (2009) Diversidade de fungos micorrízicos Epulorhiza spp. isolados de Epidendrum secundum (Orchidaceae). Rev Bras Ciências Solo 33:1187–1197
Pereira MC, Torres DP, Guimarães FAR, Pereira OL, Kasuya MCM (2011) Germinação de sementes e desenvolvimento de protocormos de Epidendrum secundum Jacq. (Orchidaceae) em associação com fungos micorrízicos do gênero Epulorhiza. Acta Bot Bras 25(3):534–541
Rasmussen HN (2002) Recent developments in the study of orchid mycorrhiza. Plant Soil 244:149–163
Roberts P (1999) Rhizoctonia-forming fungi: a taxonomic guide. Royal Botanic Gardens, Kew
Schäfer C, Wöstemeyer J (1992) Random primer dependent PCR differentiates aggressive from non-aggressive isolates of the oilseed rape pathogen Phoma lingam (Leptosphaeria maculans). J Phytopathol 136:124–136
Shan XC, Liew EC, Weatherhead MA, Hodkiss LJ (2002) Characterization and taxonomy placement of Rhizoctonia-like endophytes from orchid roots. Mycologia 94(2):230–239
Sharon M, Kuninaga S, Hyakumachi M, Sneh B (2006) The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience 47:299–316
Sharon M, Kuninaga S, Hyakumachi M, Naito S, Sneh B (2008) Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience 49(2):93–114
Singh D (1981) The relative importance of characters affecting genetic divergence. Indian J Genet Plant Breed 41:237–245
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Valadares RBS, Pereira MC, Otero JT, Cardoso EJ (2012) Narrow fungal mycorrhizal diversity in a population of the orchid Coppensia doniana. Biotropica 44(1):114–122
White TJ, Bruns TD, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic, San Diego
Zettler LW (1997) Terrestrial orchid conservation by symbiotic seed germination: techniques and perspectives. Selbyana 18:188–194
Acknowledgments
The authors thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) for financial support of project and for the PhD scholarship of the first author; the Instituto Estadual de Florestas (IEF) for granting us permission to collect material in the conservation area Serra do Brigadeiro, Minas Gerais; Gilmar Valente for his help during collections; two anonymous referees and Manuscript Editor of Symbiosis for improvements and advice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pereira, M.C., da Silva Coelho, I., da Silva Valadares, R.B. et al. Morphological and molecular characterization of Tulasnella spp. fungi isolated from the roots of Epidendrum secundum, a widespread Brazilian orchid. Symbiosis 62, 111–121 (2014). https://doi.org/10.1007/s13199-014-0276-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-014-0276-0