Abstract
A method was standardized to isolate quality DNA from cattle and buffalo fat for species identification using QIAamp DNA stool mini kit. The quality of the DNA was sufficient enough to amplify universal primers viz., mt 12S rRNA and mt 16S rRNA, and species specific D loop primers for cattle and buffalo. The sensitivity of the PCR assay in the species specific D loop primer amplification was with a detection level of 0. 47 ng cattle DNA and 0.23 ng buffalo DNA in simplex and, 0. 47 ng cattle DNA and 0.12 ng buffalo DNA in duplex PCR. It is a potentially reliable method for DNA detection to authenticate animal fat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The increased awareness among the consumers on the food safety and food security aspects necessitates the researchers to search for newer techniques to identify adulteration in food items. This has a great impact on the economy since the consumers choice is greatly influenced by the food composition detailed in labelling. In order to assure correct labelling of food products, sensitive techniques are essential to detect various food items so as to discriminate between the different components. Food manufacturers choose to use lard/tallow as a substitute ingredient for oil because it is cheaper and easily available. Beef inclusion in the meat preparations are serious matters in some religions. Further, the Government of India has allowed export of buffalo tallow only and not the beef tallow. Hence, it is an important task for food control laboratories to be able to carry out species differentiation of raw materials to be used for industrial food preparation and the detection of animal species in food products.
Methods developed based on DNA technology will be useful in the case of fat containing materials such as fat tissues and fat itself. In meat specification, either nuclear or mitochondrial DNA (nDNA and mtDNA) genes have been targeted in PCR analysis. Nuclear DNA is larger molecule arranged into chromosomes and contains a greater variation in its type of sequences compared to mtDNA. The mtDNA is commonly used for species identification in food analysis (Meyer et al. 1994; Matsunaga et al. 1999; Girish et al. 2004; Che Man et al. 2007; Zhang et al. 2007; Sahilah et al. 2011; De et al. 2011) especially cytochrome b (cyt b) gene, 12S rRNA and D loop gene. High copy number of mtDNA is found in the cells and, it remains intact during food processing thereby minimizing DNA degradation and does not contain any introns (Unseld et al. 1995). The published methods clearly indicate that PCR offers both the desired sensitivity and the specificity for detection of adulteration of meat and meat products.
Fat is an important ingredient in meat product for the formation of micelles of fat coated with protein dissolved from meat ingredient. In order to reduce the cost of the product, adulteration of low priced animal fat may be mixed in the emulsion making process. However, the identification of species is very difficult after emulsion process. But the identification of different animal fat by its physical characteristics is possible (Singh and Neelam 2011). Further animal fats are also used in the feed industry and their detection becomes crucial wherever it is banned (Fumiere et al. 2009). As per the European Commission requirement, optical microscopy remains the only reference method for the detection of constituents of animal origin. It was reported that no single fatty acid can be used to detect the contamination of fat with tallow in GC and it was demonstrated that PCR technique could detect the contamination level as low as 5 % tallow in a matrix of other fats (Bellorini et al. 2005). Recently, some PCR assays has been developed for identifying feed constituents of animal origin (Dalmasso et al. 2004).
There are very few reports available for the species identification of cattle and buffalo fat. Therefore, the aim of the present study was to find a method to extract DNA from cattle and buffalo fat and evaluate the quality using PCR analysis for species identification.
Materials and methods
Preparation of samples
Animal body fat (cattle and buffalo) were purchased from the municipal slaughterhouse at Hyderabad and preserved at −20 °C until analysis.
DNA extraction
The DNA were extracted from each sample in triplicates by following the Qiagen QIAamp DNA stool mini kit, based on resin tablets that absorb PCR inhibitors and silica-gel columns that allow separation of nucleic acids with some modifications. The amount of fat sample was reduced to 200 mg after several preliminary trials. Briefly, 200 mg tissue was mixed with 1.4 ml ASL buffer and ground using the pestle and mortar followed by sonication for 2 min with 15 s pulse on and 15 s pulse off before proceeding with the instructions given in the kit. DNA concentrations in samples were determined in Nanospectrophotometer (Biospec Nanospectrophotomer, Shimadzu, Japan.
PCR assay
For PCR assay, the primers composition given in the Table 1 were used in the present study. Each PCR amplification reaction was set in a volume of 25 μl with 2.5 μl of 10X PCR buffer (100 mM Tris HCl, pH 9.0, 15 mM MgCl2, 500 mM KCl and 0.1 % gelatin), 0.5 μl of 10 mM dNTP mix (Chromous Biotech, Bangalore), 0.5 μl each of (20 pmoles) forward and reverse primers (Bioserve Technologies, Hyderabad), 1 U of Fidelity Taq DNA polymerase (Chromous Biotech, Bangalore), 20 μg of BSA (SRL, Mumbai) and 50 ng of purified DNA. Volume was made up to 25 μl by adding nuclease free water. Conditions on a master cycler gradient thermocycler (Eppendorf, Germany or PEQSTAR, Germany) were as per the published protocol for each primer.
Quantitative simplex and duplex PCR assay
This assay was carried out only for mt D loop primer. The sensitivity of simplex and duplex PCR assay was quantified by means of another set of PCR amplification with binary mixtures of known amount of cattle and buffalo DNA as template. The template DNA was sourced from either cattle or buffalo fat and concentration was measured spectrophotometrically. DNA was diluted accordingly to get 30 ng of diluted DNA stock before starting the PCR reaction. Binary mixture of DNA was prepared by mixing appropriate quantities of cattle and buffalo DNA from their respective stocks. A total of five independent series, each having 11 different proportions of DNA mixture containing 30, 15, 7.5, 3.8, 1.9, 0.94, 0.47, 0.23, 0.12, 0.06 and 0 ng cattle DNA in 0, 0.06, 0.12, 0.23, 0.47, 0.94, 1.9, 3.8, 7.5, 15 and 30 ng buffalo DNA, respectively, were prepared. The PCR amplification was performed in 25 μl reaction volume with a cattle specific primer set (Primer 1 and 3 as given in Table 1). In parallel PCR control assay, buffalo specific primer set (Primer 1 and 2 as given in Table 1) was used to detect different concentration of buffalo DNA in the binary mixture of cattle and buffalo DNA. The PCR cycling conditions were same as mentioned in published article (Table 1). Finally, to test whether the sensitivity shown in simplex PCR is reflected on duplex PCR as well, the amplification conditions of duplex PCR with both cattle and buffalo specific primers were simulated on these different series of diluted DNA. Amplified products were electrophoresed on agarose (2 %) gels and stained with ethidium bromide (0.5 μg/ml) and images were captured through gel documentation system (Alpha Image, USA).
Results
An extraction method for DNA isolation from fat tissue was accomplished using a commercially available kit with some modifications, and the amplification of universal primer as well as mt D loop region with species specific primers has been described for species identification of fat from cattle and buffalo. The quality of DNA obtained from fat tissues was not examined by agarose gel electrophoresis as the concentration was very less i.e. 14.56 ± 5.50 for cattle and 14.38 ± 5.44 ng/μl. The agarose gel electrophoretic method enables to verify the DNA integrity since variations of DNA fragment length, which is a parameter for DNA integrity, are dependent on the material under examination, the degree of processing and the DNA extraction method. The purity of the DNA obtained from fat tissue samples were not high, since the ratio A260/A280 ranged between 1.4 and 3.21. This method under this report was standardized after several trials of DNA extraction using conventional PCI method/commercially available tissue DNA extraction kits which could not provide DNA for PCR assay. Of the several methods tested, the most consistent results were achieved with the Qiagen QIAamp DNA stool extraction kit with modifications; the other kits gave more erratic PCR amplifications (data not shown). Though the DNA extracted from fat tissue was small amount, the concentration of DNA did not appear to be limiting; rather, successful amplification likely depended on the ability of the Qiagen QIAamp DNA stool mini kit to free DNA from inhibitors of PCR present in the samples.
DNA from fat tissue produce specific and concordant amplifications
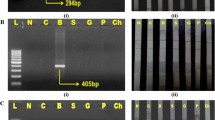
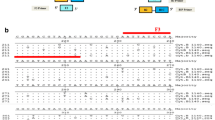
The presence of DNA was initially put in evidence by the universal mt 12S rRNA amplification which we subsequently extended further by other PCR assays using the mt D loop cattle and mt D loop buffalo and mt 16S rRNA ruminant specific primers. In spite of the DNA obtained from fat tissues of cattle and buffalo with less concentration and purity, amplification was obtained correctly (Fig 1). The mt 12S rRNA and mt D loop amplification was observed correctly. While mt 16 s rRNA amplification was observed only in cattle as expected since the primer was designed on the sequences of Bos taurus, Capra hircus and Ovis aries.
PCR amplification of mt 12S rRNA, mt D loop, mt 16S rRNA primers. DNA ladder (L), BM = fat tissue of cattle male, CM = fat tissue of male buffalo. 12S rRNA = mt 12S rRNA amplification (456b6), Cattle D loop = cattle specific mt D loop amplification (126 bp), Buff D Loop = buffalo specific mt D loop amplification (226 bp), 16S rRNA = mt 16S rRNA amplification (104 bp)
Simplex and duplex PCR assay for species identification
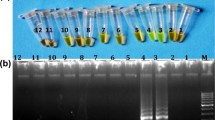
The specificity of primers was confirmed from the preliminary PCR experiments. The primers produced only species specific amplification and no false positive amplification was observed. Desired amplification of 126 bp fragment was obtained only with cattle DNA when cattle specific primers were used (Fig. 2). Similarly, buffalo specific primers amplified 226 bp fragment from buffalo DNA and no amplification was found with cattle DNA (Fig. 3). In simplex PCR, amplification with cattle specific primer set showed a consistent PCR signal corresponding to product size of 126 bp from mixture of DNA. The quantity of PCR products was directly proportional to the concentration of DNA in the mixture with a detection threshold of 0.47 ng of cattle fat DNA. The other assay with buffalo specific primers also showed specific amplification of a 226 bp PCR product with a detection threshold of 0.23 ng of buffalo fat DNA. In duplex PCR assay by primer combinations used in this study, the amplification suggest that as low as 0. 47 ng cattle DNA and 0.12 ng buffalo DNA could be detected (Fig. 4).
Amplification of D loop cattle and buffalo specific primers in duplex PCR DNA ladder (L), binary mixture containing serially diluted cattle DNA (ng) in decreasing order = 30, 15, 7.5, 3.8, 1.9, 0.94, 0.47, 0.23, 0.12, 0.06 and 0.0 (bottom line with amplicons at 126 bp) and serially diluted buffalo DNA (ng) in increasing order =0, 0.06, 0.12, 0.23, 0.47, 0.94, 1.9, 3.8, 7.5, 15 and 30 ng (top line with amplicons at 226 bp) and negative control [NC]
Discussion
DNA based identification of species in meat and meat (Floren et al. 2015; Kesmen et al. 2009; Girish et al. 2005) has received an important attention in food industry due to high sensitivity, specificity and simplicity of the technique. In the present study, a method has been standardized using Qiagen QIAamp DNA stool kit with slight modifications for the isolation of DNA from the fat tissues of cattle and buffalo followed by PCR assay amplifying universal mitochondrial primer (mt 12S), mitochondrial primer 16S RNA specific for ruminants (cattle, sheep and goat) and mt DNA D loop primer (cattle and buffalo) for the detection of cattle and buffalo fat. The quality of DNA isolated was sufficient enough to amplify and identify the species of the source material (fat).
Numerous factors have known to influence the outcome of successful PCR amplification e.g. quality of DNA, specificity of primers, etc. A variety of components of in the fat tissue including calcium ion (Bickley et al. 1996), fat and proteases (Wilson 1997) can potentially inhibit PCR amplification. Few modifications designed to improve the amount and quality of DNA extracted from fat tissue did not lead to substantial improvements, and the DNA recovered seldom exceeded 20 ng/μL. Of the kits assayed, the most consistent results were achieved with the Qiagen QIAamp DNA stool extraction kit; the other kits gave more erratic PCR amplifications. Aida et al. (2005) have reported the use of DNeasy kit for the isolation of genomic and mitochondrial DNA from beef fat samples and used them in successful PCR assay. Testolin and Lain (2005) have reported the successful use of QIAamp DNA stool kit for DNA isolation from olive oil and its use in the PCR amplification. However, the concentration of DNA did not appear to be limiting; rather, successful amplification likely depended on the ability of the different kits to free DNA from inhibitors of PCR present in the samples.
The variable regions of the mitochondrial gene are present in thousands of copies per cell (Greenwood and Paboo 1999), which increases the probability of achieving a positive result even in severe DNA fragmentation due to intense processing conditions (Bellagamba et al. 2001) as well as in very small amount (present study), thus making it ideal for identification of species origin of processed meat and meat products. In the present study, all 3 primers tested gave DNA amplicons of the correct size (Fig. 1). The results showed the applicability of the primers proposed by De et al. (2011) for the identification of cattle and buffalo species and by Dalmasso et al. (2004) for the identification of ruminants (in the present study, cattle species).
The specificity and sensitivity of primers was further confirmed with simplex and duplex PCR assay using cattle and buffalo specific D loop primers. The sensitivity of the simplex as well as duplex PCR assay was determined for species specific identification of fat tissue. The sensitivity of a PCR depends on successful amplification from a minimal amount of template DNA present in a sample. As DNA from particular proportion of fat tissue (e.g. 0.1 %) in a meat product mixture may vary, therefore, the sensitivity of a PCR assay can be better estimated by quantifying the minimal amount of template DNA from which successful amplification is obtained. In the present assay, sensitivity of PCR assay was validated by determining the minimal concentration of DNA in cattle and buffalo DNA mixture, for which the primers produced a successful amplification. This was carried out for cattle and buffalo specific simplex PCR as well as duplex PCR. Cycling conditions and parameters for these simplex and duplex PCR amplifications were kept similar except that known amount of DNA was used as template. The result of these PCR amplification suggest that as low as 0. 47 ng cattle DNA and 0.23 ng buffalo DNA could be detected in simplex (Figs 2 and 3) and, 0. 47 ng cattle DNA and 0.12 ng buffalo DNA in duplex PCR by primer combinations used in this study (Fig 4). Amplification from mixture of template DNA was species specific and without any false positive amplification. De et al. (2011) has reported a similar duplex PCR assay for identification of adulteration of 0.1 % cattle and buffalo milk. Soares et al. (2010) has reported a similar duplex PCR assay for identification of pork in chicken meat mixture with a sensitivity of 0.1 % pork in chicken meat. The assays used in this research have a high potential as a molecular tool that can be used in quality control laboratories for the verification and control of adulteration in animal fat tissue to verify it origin. This study urges the need for fast, highly sensitive, specific and obviously cost-effective identification techniques to examine adulterated animal fat tissue and evaluation of their label authenticity. The developed assay technique is a novel technique to isolate quality DNA from animal fat tissue and has the ability to perform simultaneous amplification in the duplex quantitative PCR assay without a need for real time PCR.
Conclusion
Species identification of food ingredient in a complex food matrix is a challenge for the food control laboratories. Thus, there is a need for reliable, fast and sensitive methodologies that ensure an effective detection of frauds in the meat sector by the identification of species present in the meat products. In the present work, a method was standardized and reported which is useful to isolate quality DNA from animal fat tissue and it was sufficient enough to deploy in the PCR assay to detect species of source materials. The fat tissue DNA isolation method and the PCR assay for species identification may be used by food inspection departments as well as manufacturers who have much to rely on supplier’s claim on source of the fat tissue.
References
Aida AA, Che Man YB, Wong CMVL, Raha AR, Son R (2005) Analysis of raw meats and fats of pigs using polymerase chain reaction for halal authentication. Meat Sci 69:47–52
Bellagamba F, Moretti VM, Comincini S, Valfre F (2001) Identification of species in animal feedstuffs by polymerase chain reaction–restriction fragment length polymorphism analysis of mitochondrial DNA. J Agric Food Chem 49:3775–3781
Bellorini S, Strathmann S, von Holst C (2005) Discriminating animal fats and their origins: assessing the potentials of Fourier transform infrared spectroscopy, gas chromatography, immunoassay and polymerase chain reaction techniques. Anal Bioanal Chem 383:1073–1083
Bickley J, Short JK, McDowell DG, Parkes HC (1996) Polymerase chain reaction (PCR) detection of listeria monocytogenes in diluted milk and reversal of PCR inhibition caused by calcium ions. Lett Appl Microbiol 22:153–158
Che Man Y, Aida A, Raha A, Son R (2007) Identification of pork derivatives in food products by species-specific polymerase chain reaction (PCR) for halal verification. Food Control 18:885–889
Dalmasso A, Fontanella E, Piatti P, Civer T, Rosati S, Bottero MT (2004) A multiplex PCR assay for the identification of animal species in feedstuffs. Mol Cell Probes 18:81–87
De S, Brahma B, Polley S, Mukherjee A, Banerjee D, Gohaina M, Singh KP, Singh R, Datta TK, Goswami SL (2011) Simplex and duplex PCR assays for species specific identification of cattle and buffalo milk and cheese. Food Control 22:690–696
Floren C, Wiedemann I, Brenig B, Schutz E, Beck J (2015) Species identification and quantification in meat and meat products using droplet digital PCR (ddPCR). Food Chem 173:1054–1058
Fumiere O, Veys P, Boix A, von Holst C, Baeten V, Berben G (2009) Methods of detection, species identification and quantification of processed animal proteins in feeding stuffs. Biotechnol Agron Soc Enrviron 13:59–70
Girish PS, Anjaneyulu ASR, Viswas KN, Anand M, Rajkumar N, Shivakumar BM, Bhaskar S (2004) Sequence analysis of mitochondrial 12S rRNA gene can identify meat species. Meat Sci 66:551–556
Girish PS, Anjaneyulu ASR, Viswas KN, Shivakumar BM, Anand M, Patel M, Sharma B (2005) Meat species identification by polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) of mitochondrial 12S rRNA gene. Meat Sci 70:107–112
Greenwood A, Paboo S (1999) Nuclear insertion sequences of mitochondrial DNA predominate in hair but not in blood of elephants. Mol Ecol 8:133–137
Kesmen Z, Gulluce A, Sahin F, Yetim H (2009) Identification of meat species by TaqMan-based real-time PCR assay. Meat Sci 82:444–449
Kocher TD, Thomas WK, Meyer A, Edwards SV, Paabo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA sequence evolution in animals. Proc Natl Acad Sci U S A 86:6196–6200
Matsunaga T, Chikuni K, Tanabe R, Muoya S, Shibata K, Yamada J, Shinmura Y (1999) A quick and simple method for the identification of meat species and meat products by PCR assay. Meat Sci 51:143–148
Meyer R, Candrian U, Lüthy J (1994) Detection of pork in heated meat products by the polymerase chain reaction. J AOAC Int 77:617–622
Sahilah AM, Norhayati Y, Wan Aida WM, Aminah A (2011) Halal authentication of raw meats using PCR amplification of mitochondrial DNA. Inter Food Res J 18:1489–1491
Singh VP, Neelam S (2011) Meat species specifications to ensure the quality of meat-A review. Inter J Meat Sci 1:15–26
Soares S, Amaral JS, Mafra I, Beatriz M, Oliveira PP (2010) Quantitative detection of poultry meat adulteration with pork by a duplex PCR assay. Meat Sci 85:531–536
Testolin R, Lain O (2005) DNA Extraction from Olive Oil and PCR Amplification of microsatellite markers. J Food Sci 70:1–5
Unseld M, Beyermann B, Brandt P, Hiesel R (1995) Identification of the species of origin of highly processed meat products by mitochondrial DNA sequences. PCR Meth Appl 4:241–243
Wilson IG (1997) Inhibition and facilitation of nucleic acid amplification. Appl Environ Microbiol 63:3741–3751
Zhang C, Fowler MR, Scott NW, Lawson G, Slater A (2007) A TaqMan real-time PCR system for the identification and quantification of bovine DNA in meats, milks and cheeses. Food Control 18:1149–1158
Acknowledgments
The authors thank the National Research on Meat for providing the necessary facilities to carry out this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Highlights
• A method standardized to isolate DNA from cattle and buffalo fat for PCR assay.
• The quality of DNA was sufficient enough to amplify universal primers and species specific primers, and to quantitatively identify species of fat
• The method and assay may be used by food inspection departments as well as manufacturers
Rights and permissions
About this article
Cite this article
Vaithiyanathan, S., Kulkarni, V.V. Species identification of cattle and buffalo fat through PCR assay. J Food Sci Technol 53, 2077–2082 (2016). https://doi.org/10.1007/s13197-016-2198-8
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13197-016-2198-8