Abstract
We examined phylogeographic differentiation of the red-eyed grass snake (Natrix astreptophora) using 1984 bp of mtDNA and 13 microsatellite loci from specimens collected across its distribution range in southwestern Europe and northwestern Africa. Based on phylogenetic analyses of mtDNA, European N. astreptophora constituted the sister clade to a weakly supported North African clade comprised of two deeply divergent and well-supported clades, one corresponding to Moroccan snakes and the other to snakes from Algeria and Tunisia. This tripartite differentiation was confirmed by analyses of microsatellite loci. According to a fossil-calibrated molecular clock, European and North African N. astreptophora diverged 5.44 million years ago (mya), and the two Maghrebian clades split 4.64 mya. These dates suggest that the radiation of the three clades was initiated by the environmental changes related to the Messinian Salinity Crisis and the reflooding of the Mediterranean Basin. The differentiation of N. astreptophora, with distinct clades in the Iberian Peninsula and in the western and eastern Maghreb, corresponds to a general phylogeographic paradigm and resembles the differentiation found in another co-distributed Natrix species, the viperine snake (N. maura). Despite both species being good swimmers, the Strait of Gibraltar constitutes a significant biogeographic barrier for them. The discovery that North Africa harbours two endemic lineages of N. astreptophora necessitates more conservation efforts for these imperilled snakes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For decades, grass snakes (Natrix natrix sensu lato) were thought to be one of the most widely distributed Palearctic snake species, occurring from the North African Maghreb region through the Iberian Peninsula and most of Europe to Lake Baikal in Central Asia (Kabisch 1999). However, two recent investigations have shown that what was previously understood as one polytypic species was composed of three distinct species (Pokrant et al. 2016; Kindler et al. 2017), each with phylogeographic differentiation (Kindler et al. 2013). Ibero-Maghrebian grass snakes are now recognised as the distinct species Natrix astreptophora (Seoane, 1884) (Pokrant et al. 2016). Grass snakes in the remaining western part of the distribution range represent the western species N. helvetica (Lacepède, 1789), which hybridises with the eastern species N. natrix (Linnaeus, 1758) in a narrow belt in the Rhine region (Kindler et al. 2017). Natrix astreptophora differs from the other two grass snake species, N. helvetica and N. natrix, in skull morphology, coloration and pattern, in particular by its unique reddish iris. Moreover, the contact zone of N. astreptophora and N. helvetica is characterised by a parapatric distribution of the two taxa, with rare hybridization (Pokrant et al. 2016).
Two previous studies using mitochondrial DNA (mtDNA) found Iberian and French N. astreptophora deeply divergent from a single sample from Tunisia (Kindler et al. 2013; Pokrant et al. 2016). Natrix astreptophora has a wide relictual distribution in the Maghreb, with few isolated records from Morocco through Algeria to Tunisia (Fig. 1; Bons and Geniez 1996; Schleich et al. 1996). Many amphibian and reptile species, some small mammals and at least one land snail show a pronounced west-east differentiation in the Maghreb (reviews in Husemann et al. 2014; Stuckas et al. 2014; Nicolas et al. 2015), posing the question how Tunisian grass snakes are related to other North African populations. Until now, the rareness of North African grass snakes has prevented examining their differentiation, and populations from Morocco and Algeria remained completely unstudied. For this paper, we expanded upon previous sampling to include novel specimens from most of the North African distribution, including Algeria and Morocco. Using two mitochondrial DNA (mtDNA) fragments and 13 microsatellite loci, we examined the phylogeography of N. astreptophora across its entire range and compared it to other co-distributed taxa.
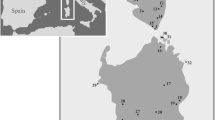
Sampling sites and mitochondrial identity of studied red-eyed grass snakes (n = 56). Olive green areas indicate distribution of Natrix astreptophora in Northern Africa according to Bons and Geniez (1996), Schleich et al. (1996) and Sindaco et al. (2013). Two questionable localities in North Africa are not shown (Atlantic coast of Morocco, Schleich et al. 1996; southern Algeria, Hecht 1930). Colours of sampling sites correspond to Figs. 2, 3, 4. Inset: N. astreptophora from Morocco; photo: Salvador Carranza
Besides the scientific names, we use below the well-established vernacular name ‘barred grass snake’ for N. helvetica and refer to N. natrix as ‘common grass snake’ or ‘eastern grass snake’. For N. astreptophora, we introduce here the name ‘red-eyed grass snake’, acknowledging its unique and highly diagnostic iris coloration.
Materials and methods
A total of 56 Natrix astreptophora were studied (Table S1), corresponding to 11 new samples and 45 previously processed ones (Kindler et al. 2013; Pokrant et al. 2016), including GenBank data (Guicking et al. 2006). Our new samples came from Morocco (4), Algeria (2), Tunisia (3) and Spain (2). For these samples, the same two mtDNA fragments were sequenced as in Kindler et al. (2013) and Pokrant et al. (2016), i.e. the partial ND4 gene plus adjacent DNA coding for tRNAs (866 bp) and the cyt b gene (1117 bp). Laboratory procedures were the same as in Kindler et al. (2013). For phylogenetic analyses, the new sequences were concatenated and merged with previously published data for N. astreptophora plus three sequences each for the 14 terminal clades of N. helvetica and N. natrix as identified by Kindler et al. (2013). Sequences of N. maura, N. tessellata and Nerodia sipedon served as outgroups (Table S2). For phylogenetic analyses, the optimal partition scheme and best-fit evolutionary models were assessed using partitionfinder 1.1.1 (Lanfear et al. 2012), with linked branch lengths and the Bayesian Information Criterion (BIC) for model selection. Partitionfinder was run twice, evaluating either evolutionary models implemented in mrbayes 3.2.6 (Ronquist et al. 2012) or implemented in RAxML 8.2.4 (Stamatakis 2014). A user-specific search was run for the following partition schemes: (a) unpartitioned, (b) partitioned by gene with DNA coding for tRNAs merged in one partition, and (c) maximum partitioning, i.e., using each codon position of protein-coding genes and the merged tRNAs as a distinct partition, resulting in scheme (a) and the GTR+G+I model for both tree-building methods. Using mrbayes 3.2.6, two parallel runs were then computed, each with four chains. The chains ran for 10 million generations, with every 100th generation sampled. Convergence of runs was confirmed by the average standard deviation of split frequencies approaching zero. Using the sump command, stationarity was further verified by plotting the generation versus the log probability of the data, with a ‘white noise’ pattern indicating no tendency to increase or decrease over time. For the final 50% majority rule consensus tree, a burn-in of 2.5 million generations (25%) was used. In addition, Maximum Likelihood (ML) analyses were conducted using RAxML 8.2.4. Five independent ML searches were run with different starting conditions and the fast bootstrap algorithm to explore the robustness of the branching patterns by comparing the best trees. Then, 1000 nonparametric thorough bootstrap values were calculated and plotted against the best tree.
In addition, mtDNA sequences were examined using parsimony networks as implemented in tcs 1.21 (Clement et al. 2000), with gaps coded as fifth character state. Because for some samples only one mtDNA fragment was available, each fragment was analysed separately, as in Pokrant et al. (2016) and Kindler et al. (2017), due to the fact that tcs software cannot cope with missing data. Using the default 95% connection limit, unconnected haplotype clusters were obtained for the different genetic lineages, which is why the connection limit was arbitrarily set to 100 steps. Based on haplotypes of each mtDNA fragment (Table S3), uncorrected p distances (average) were computed using mega 7.0.21 (Kumar et al. 2016) and the pairwise deletion option for each of the three grass snake species and clades within each species.
Besides mtDNA, also 13 polymorphic microsatellite loci were studied. For 29 samples of European N. astreptophora, data were available from Pokrant et al. (2016). However, one sample (MTD T 13083) was excluded because it showed genetic impact of N. helvetica. Eleven new samples (Table S1) were genotyped according to the procedures of Pokrant et al. (2016) and data were examined using Principal Component Analyses (PCA) as implemented in the R package adegenet (Jombart 2008). Additional analyses using unsupervised Bayesian cluster analyses are described in the Supporting Information of this article. Pairwise F ST values and Analyses of Molecular Variance (AMOVAs) were calculated for each mtDNA fragment and for microsatellite data using arlequin 3.5.1.3 (Excoffier and Lischer 2010).
For mitochondrial clades, a dating analysis using a relaxed molecular clock was performed with beast 1.8.4 (Drummond et al. 2012) using the same fossil calibration and settings as in Fritz et al. (2012). A Natrix vertebra from Sardinia (Delfino et al. 2011) of known age (3.6 million years) was applied to constrain the divergence of the Corso-Sardinian clade (clade B in Fig. 2) from its sister clade. The lognormal prior for the TMRCA was set to mean = 0.4, log(stdev) = 1 and offset = 3.6. For finding the optimal partition scheme and model of sequence evolution for the dating analysis, partitionfinder was run again for the models available in beast using the same approach as described above, resulting in the selection of the GTR+G+I model for an unpartitioned data set. As in Fritz et al. (2012), the data set for the molecular clock was based on four mitochondrial genes (cyt b, ND1, ND2 and ND4 without tRNAs) consisting of one sequence of each mitochondrial clade of N. astreptophora, N. helvetica and N. natrix plus two sequences of N. maura and one sequence of N. tessellata (Table S4). For clades not included in the study by Fritz et al. (2012), only cyt b and ND4 data were available (clades A, 2, 5 and 6 of Kindler et al. 2013; western and eastern Maghrebian clades of this study).
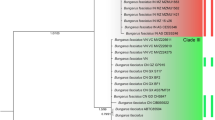
Mitochondrial phylogeny of all three grass snake species inferred from Maximum Likelihood using 1984-bp mtDNA (ND4+tRNAs, cyt b). Terminal clades collapsed to cartoons. Outgroups (Natrix maura, N. tessellata, Nerodia sipedon) removed for clarity. Numbers along nodes indicate branch support under Maximum Likelihood (1000 bootstrap replicates) and Bayesian Inference (posterior probabilities). Asterisks indicate maximum support under both tree-building methods. For Natrix helvetica and N. natrix, clade names correspond to Kindler et al. (2013). Inset: European N. astreptophora (near Nohèdes, south-western France); photo: Philippe Geniez
Results
Under both tree-building methods, sequences from Natrix astreptophora corresponded to three well-supported clades that were placed in a well-supported more inclusive clade (Fig. 2). The three clades matched with sequences from (i) the Iberian Peninsula and France, (ii) the western Maghreb region (Morocco), and (iii) the eastern Maghreb region (Algeria, Tunisia). Clades (ii) and (iii) were, with weak support, together the sister group of (i). The relationships and internal branching patterns of the other two grass snake species (N. helvetica, N. natrix) conformed to expectations from previous studies (Guicking et al. 2006; Fritz et al. 2012; Kindler et al. 2013).
Using the two individual mtDNA fragments of N. astreptophora for parsimony networks, each of the three clades corresponded to a distinct haplotype cluster (Fig. 3). For both mtDNA fragments, the European clade had the highest number of haplotypes, which is likely to be related to the large sample size. For the mtDNA sequences comprised of the partial ND4 gene and adjacent DNA coding for tRNAs, there were 13 haplotypes for the European cluster, three haplotypes for the Moroccan cluster and four haplotypes for the Algerian-Tunisian cluster. The European cluster differed from the Moroccan cluster by a minimum of 26 mutation steps and from the Algerian-Tunisian cluster by 21 steps. The two North African clusters differed by 29 steps from one another. Within the European cluster, a maximum of six steps occurred and within the Moroccan cluster a maximum of five steps. Sequences from the eastern Maghreb differed by a maximum of four steps. For the cyt b gene, the European cluster was comprised of 20 haplotypes, the Moroccan cluster contained three haplotypes, and the cluster with sequences from Algeria and Tunisia had five haplotypes. The European and Moroccan clusters differed by 44 mutation steps; the European and Algerian-Tunisian clusters, by 49 steps, and the two Maghrebian clusters, by 39 steps. Within the European cluster, up to nine steps occurred; within each Maghrebian cluster, up to five steps.
Parsimony networks of mtDNA sequences of Natrix astreptophora. Symbol size corresponds to haplotype frequency; lines connecting haplotypes represent one mutation step, if not otherwise indicated. Small black circles are missing node haplotypes. Haplotype colours correspond to lineages: European lineage in orange, western Maghrebian lineage in green and eastern Maghrebian lineage in brown. Haplotype names in blue. For European Nucleotide Archive (ENA) accession numbers, see Table S3. Differences in mutation counts compared to Pokrant et al. (2016) are due to longer DNA sequences
Uncorrected p distances (averages) between the three grass snake species ranged for the haplotypes corresponding to the partial ND4 gene and adjacent mtDNA from 5.49 to 5.67%; within-species divergences ranged from 1.73 to 3.39% (Table S5). For the cyt b gene, the values between species were 6.32–7.08%, and within species, 1.90–3.59% (Table S6). Regarding the three clades within N. astreptophora, divergences between 2.61 and 3.43% were observed for the ND4 gene plus adjacent mtDNA, and for the cyt b gene, divergences between 3.69 and 4.38% (Tables S7, S8). These values fall into the upper ranges among the clades of the barred grass snake (N. helvetica: 0.63–6.01% for ND4+tRNAs, 0.45–5.22% for cyt b) and the common grass snake (N. natrix: 0.59–5.16% for ND4+tRNAs, 1.37–5.74% for cyt b; Tables S7, S8).
According to an AMOVA for the mtDNA fragment containing ND4+tRNAs, 95.58% of the molecular variance occurred between and 4.42% within the three mitochondrial lineages of N. astreptophora. Pairwise F ST values for the three lineages ranged from 0.93 to 0.96 (Table S9). For cyt b sequences, 94.15% of the molecular variance occurred between and 5.85% within the mitochondrial lineages. Pairwise F ST values varied from 0.93 to 0.94 (Table S10).
The microsatellite loci of N. astreptophora turned out to be highly polymorphic (Table 1). The PCA using these data (Fig. 4) confirmed the distinctness of red-eyed grass snakes representing the three mitochondrial lineages and found three different clusters without any overlap. This was also in line with the unsupervised Bayesian clustering approach (Supporting Information). An AMOVA for microsatellite data revealed that 33.71% of the molecular variance occurred between and 66.29% within the three clusters. Pairwise F ST values varied from 0.25 to 0.45 (Table S11).
Principal Component Analysis (PCA) for microsatellite data. Samples are coloured according to mitochondrial lineages. The oval outlines represent 95% confidential intervals. For axes 1–2 (left), the x axis explains 15.4% and the y axis 11.6% of variance. For axes 1–3 (right), the y axis explains 7.9% of variance. Non-overlapping groups denote significantly different clusters
The dating analysis using mtDNA yielded for N. helvetica similar results as in Fritz et al. (2012). The only exception was clade B, which was inferred to be slightly younger (0.5 million years; Table S12). However, the divergence times of and the branching patterns within N. astreptophora and N. natrix were generally older (0.5–1.89 million years). According to our new estimates, N. astreptophora diverged 10.61 million years ago (mya) and its European clade branched off 5.44 mya, and the western and eastern Maghrebian clades split 4.63 mya (Fig. 5).
Estimated divergence times of grass snakes and their 95% HPD intervals (blue bars). Outgroups removed for clarity. The red arrow highlights the placement of the fossil (3.6 mya; Delfino et al. 2011) used for calibrating the respective node
Discussion
Our new data clearly demonstrate that the red-eyed grass snake (Natrix astreptophora) shows pronounced phylogeographic structuring, with one genetic lineage each in the Iberian Peninsula plus adjacent France as well as in the western Maghreb (Morocco) and the eastern Maghreb (Algeria, Tunisia). The distinctness of these three lineages is concordantly confirmed by mitochondrial and microsatellite data. Moreover, with respect to mitochondrial haplotypes, the two North African lineages are remarkably variable, despite small sample sizes. This is especially obvious for ND4+tRNAs (Fig. 3, top). Among the 13 haplotypes from Europe, a maximum of six mutations occurs. Samples from the western and the eastern Maghreb differ by five and four mutations, respectively, i.e. have a similar degree of variation, although only three haplotypes were found in Morocco and four haplotypes in Algeria and Tunisia. This suggests that the North African populations might be genetically more diverse than the European ones.
Numerous taxa in North Africa display a phylogeographic pattern resembling that of N. astreptophora, with different lineages in the western and eastern Maghreb, in many cases with the Moroccan Moulouya River as a divide. Moroccan populations west of the Moulouya River are often closely related to or indistinguishable from Iberian conspecifics. However, in some instances, Morocco can also harbour one or more clades being distinct from those in the Iberian Peninsula and the eastern Maghreb (Husemann et al. 2014; Stuckas et al. 2014; Martínez-Freiría et al. 2017). Such endemic lineages may occur in Morocco alone or in addition to the ‘Iberian’ ones. Examples for the general west-east paradigm in the Maghreb are the following snake species: Coronella girondica (Santos et al. 2012), Macroprotodon spp. (Carranza et al. 2004), Malpolon spp. (Carranza et al. 2006), Natrix maura (Barata et al. 2008; Guicking et al. 2008), the Vipera latastei complex (Velo-Antón et al. 2012) and Daboia mauritanica (Martínez-Freiría et al. 2017). Shared genetic lineages between the Iberian Peninsula and Morocco occur in Macroprotodon brevis (Carranza et al. 2004) and Malpolon monspessulanus (Carranza et al. 2006). Other taxa are reviewed in Barata et al. (2008), Husemann et al. (2014) and Stuckas et al. (2014). The lack of genetic differentiation between Moroccan and Iberian conspecifics is explained either by natural transoceanic dispersal across the Strait of Gibraltar or translocation by humans (Veith et al. 2004; Fritz et al. 2006; Recuero et al. 2007; Velo-Antón et al. 2015; Veríssimo et al. 2016). Since grass snakes are excellent swimmers (Kabisch 1999), the differentiation of Iberian and Moroccan N. astreptophora was not necessarily expected. Today, the Strait of Gibraltar is only 15 km wide and during glacial low sea level stands it was further reduced to 5 km (Zazo 1999); distances that should be easily bridged by semiaquatic species. Accordingly, both freshwater turtle species with similar distribution patterns as N. astreptophora have mitochondrial clades and haplotypes occurring on both sides of the Strait of Gibraltar (Emys orbicularis: Stuckas et al. 2014; Velo-Antón et al. 2015; Mauremys leprosa: Fritz et al. 2006; Veríssimo et al. 2016). However, as in the red-eyed grass snake, the Strait of Gibraltar constitutes a significant phylogeographic barrier for the semiaquatic viperine snake N. maura (Barata et al. 2008; Guicking et al. 2008).
Moroccan N. astreptophora are neither closely related to nor undifferentiated from Iberian conspecifics, but represent a genetic lineage being deeply divergent from both European and eastern Maghrebian N. astreptophora. Their genetic divergences fall into the upper ranges observed between the genetic lineages of N. helvetica and N. natrix. Our divergence time estimates suggest that the clades from the western and eastern Maghreb split 4.63 mya and that the European clade is slightly older, with 5.44 mya (Fig. 5). These dates resemble the inferred age of the Messinian Salinity Crisis (MSC), which commenced ca. 7.25 mya with the gradual restriction of the Atlantic-Mediterranean connection, leading to the complete isolation of the Mediterranean Basin and a huge sea level fall or even complete drying-out of the Mediterranean Basin. It was reflooded in a catastrophic event at end of the Messinian, 5.33 mya (Roveri et al. 2014), and it seems likely that, as a corollary of the MSC, massive environmental changes triggered the radiation of the extant lineages of N. astreptophora. Yet, the reestablishment of a sea strait between the Iberian Peninsula and North Africa was not necessarily the only responsible vicariance event. During the Late Miocene and Early Pliocene, severe climatic fluctuations caused large-scale environmental shifts (Roveri et al. 2014) that probably contributed to repeated range contractions and expansions. It seems likely that these events played a major role in shaping the current genetic structure of many Mediterranean animal and plant species. Using mitochondrial and nuclear markers, Guicking et al. (2008) found in the viperine snake (N. maura) a similar phylogeographic pattern as in N. astreptophora. These authors, however, concluded that the reflooding of the Mediterranean at the end of the MSC caused the genetic divergence of European and North African viperine snakes.
In North Africa, and in Morocco in particular, N. astreptophora is extremely rare and confined to relictual, humid and relatively cool habitats (Bons and Geniez 1996), which are threatened by destruction. With the discovery of two endemic genetic lineages in North Africa, it is clear that their conservation needs more efforts than before. Furthermore, the presented genetic data indicate that a full taxonomic revision of N. astreptophora is needed, which should include a morphological comparison of all three genetic lineages.
References
Barata, M., Harris, D. J., & Castilho, R. (2008). Comparative phylogeography of northwest African Natrix maura (Serpentes: Colubridae) inferred from mtDNA sequences. African Zoology, 43, 1–7.
Blouin-Demers, G., & Gibbs, H. L. (2003). Isolation and characterization of microsatellite loci in the black rat snake (Elaphe obsoleta). Molecular Ecology Notes, 3, 98–99.
Bons, J., & Geniez, P. (1996). Amphibiens et reptiles du Maroc (Sahara Occidental compris). Atlas biogéographique. Barcelona: AHE.
Burns, E. L., & Houlden, B. A. (1999). Isolation and characterization of microsatellite markers in the broad-headed snake Hoplocephalus bungaroides. Molecular Ecology, 8, 520–521.
Carranza, S., Arnold, E. N., Wade, E., & Fahd, S. (2004). Phylogeography of the false smooth snakes, Macroprotodon (Serpentes, Colubridae): mitochondrial DNA sequences show European populations arrived recently from Northwest Africa. Molecular Phylogenetics and Evolution, 33, 523–532.
Carranza, S., Arnold, E. N., & Pleguezuelos, J. M. (2006). Phylogeny, biogeography, and evolution of two Mediterranean snakes, Malpolon monspessulanus and Hemorrhois hippocrepis (Squamata, Colubridae), using mtDNA sequences. Molecular Phylogenetics and Evolution, 40, 532–546.
Clement, M., Posada, D., & Crandall, K. A. (2000). tcs: a computer program to estimate gene genealogies. Molecular Ecology, 9, 1657–1660.
Delfino, M., Bailon, S., & Pitruzella, G. (2011). The Late Pliocene amphibians and reptiles from “Capo Mannu D1 Local Fauna” (Mandriola, Sardinia, Italy). Geodiversitas, 33, 357–382.
Drummond, A. J., Suchard, M. A., Xie, D., & Rambaut, A. (2012). Bayesian phylogenetics with beauti and the beast 1.7. Molecular Biology and Evolution, 29, 1969–1973.
Excoffier, L., & Lischer, H. E. L. (2010). arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10, 564–567.
Fritz, U., Barata, M., Busack, S. D., Fritzsch, G., & Castilho, R. (2006). Impact of mountain chains, sea straits and peripheral populations on genetic and taxonomic structure of a freshwater turtle, Mauremys leprosa. Zoologica Scripta, 35, 97–108.
Fritz, U., Corti, C., & Päckert, M. (2012). Mitochondrial DNA sequences suggest unexpected phylogenetic position of Corso-Sardinian grass snakes (Natrix cetti) and do not support their species status, with notes on phylogeography and subspecies delineation of grass snakes. Organisms, Diversity & Evolution, 12, 71–80.
Garner, T. W. J., Gregory, P. T., McCracken, G. F., Burghardt, G. M., Koop, B. F., McLain, S. E., & Nelson, R. J. (2002). Geographic variation of multiple paternity in the common garter snake (Thamnophis sirtalis). Copeia, 2002, 15–23.
Gautschi, B., Widmer, A., & Koella, J. (2000). Isolation and characterization of microsatellite loci in the dice snake (Natrix tessellata). Molecular Ecology, 9, 2192–2193.
Guicking, D., Lawson, R., Joger, U., & Wink, M. (2006). Evolution and phylogeny of the genus Natrix (Serpentes: Colubridae). Biological Journal of the Linnean Society, 87, 127–143.
Guicking, D., Joger, U., & Wink, M. (2008). Molecular phylogeography of the viperine snake Natrix maura (Serpentes: Colubridae): evidence for strong intraspecific differentiation. Organisms, Diversity & Evolution, 8, 130–145.
Hecht, G. (1930). Systematik, Ausbreitungsgeschichte und Ökologie der europäischen Arten der Gattung Tropidonotus (Kuhl) H. Boie. Mitteilungen aus dem Zoologischen Museum Berlin, 16, 244–393.
Husemann, M., Schmitt, T., Zachos, F. E., Ulrich, W., & Habel, J. C. (2014). Palaearctic biogeography revisited: evidence for the existence of a north African refugium for Western Palaearctic biota. Journal of Biogeography, 41, 81–94.
Jombart, T. (2008). adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics, 24, 1403–1405.
Kabisch, K. (1999). Natrix natrix (Linnaeus, 1758) – Ringelnatter. In W. Böhme (Ed.), Handbuch der Reptilien und Amphibien Europas. Band 3/IIA: Schlangen II (pp. 513–580). Aula: Wiebelsheim.
Kindler, C., Böhme, W., Corti, C., Gvoždík, V., Jablonski, D., Jandzik, D., Metallinou, M., Široký, P., & Fritz, U. (2013). Mitochondrial phylogeography, contact zones and taxonomy of grass snakes (Natrix natrix, N. megalocephala). Zoologica Scripta, 42, 458–472.
Kindler, C., Chèvre, M., Ursenbacher, S., Böhme, W., Hille, A., Jablonski, D., Vamberger, M., & Fritz, U. (2017). Hybridization patterns in two contact zones of grass snakes reveal a new Central European snake species. Scientific Reports, 7, 7378.
Kumar, S., Stecher, G., & Tamura, K. (2016). mega7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33, 1870–1874.
Lacepède, B.-G.-É. (1789). Histoire naturelle des quadrupèdes ovipares et des serpens, vol. 2 (Histoire naturelle des serpens). Paris: Hôtel de Thou.
Lanfear, R., Calcott, B., Ho, S. Y. W., & Guindon, S. (2012). partitionfinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molecular Biology and Evolution, 29, 1695–1701.
Linnaeus, C. (1758). Systema Naturae per regna Tria Naturae, Secundum Classes, Ordines, Genera, Species, cum Characteribus, Differentiis, Synonymis, Locis. Tomus I. Regnum Animale. Editio Decima. Stockholm: Laurentius Salvius.
Martínez-Freiría, F., Crochet, P.-A., Fahd, S., Geniez, P., Brito, J. C., & Velo-Antón, G. (2017). Integrative phylogeographic and ecological analyses reveal multiple Pleistocene refugia for Mediterranean Daboia vipers in north-west Africa. Biological Journal of the Linnean Society, 122, 366–384.
Meister, B., Armbruster, F. J., Frauenfelder, N., & Bauer, B. (2009). Novel microsatellite loci in the grass snake (Natrix natrix) and cross-amplification in the dice snake (Natrix tessellata). Molecular Ecology Resources, 9, 604–606.
Nicolas, V., Mataame, A., Crochet, P.-A., Geniez, P., & Ohler, A. (2015). Phylogeographic patterns in North African water frog Pelophylax saharicus (Anura: Ranidae). Journal of Zoological Systematics and Evolutionary Research, 53, 239–248.
Pokrant, F., Kindler, C., Ivanov, M., Cheylan, M., Geniez, P., Böhme, W., & Fritz, U. (2016). Integrative taxonomy provides evidence for the species status of the Ibero-Maghrebian grass snake Natrix astreptophora. Biological Journal of the Linnean Society, 118, 873–888.
Prosser, M. R., Gibbs, H. L., & Weatherhead, P. J. (1999). Microgeographic population genetic structure in the northern water snake, Nerodia sipedon sipedon detected using microsatellite DNA loci. Molecular Ecology, 8, 329–333.
Recuero, E., Iraola, A., Rubio, X., Machordom, A., & García-Paris, M. (2007). Mitochondrial differentiation and biogeography of Hyla meridionalis (Anura: Hylidae): an unusual phylogeographical pattern. Journal of Biogeography, 34, 1207–1219.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M. A., & Huelsenbeck, J. P. (2012). mrbayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61, 539–542.
Roveri, M., Flecker, R., Krijgsman, W., Lofi, J., Lugli, S., Manzi, V., Sierro, F. J., Bertini, A., Camerlenghi, A., De Lange, G., & Govers, R. (2014). The Messinian Salinity Crisis: past and future of a great challenge for marine sciences. Marine Geology, 352, 25–58.
Santos, X., Rato, C., Carranza, S., Carretero, M. A., & Pleguezuelos, J. M. (2012). Complex phylogeography in the southern smooth snake (Coronella girondica) supported by mtDNA sequences. Journal of Zoological Systematics and Evolutionary Research, 50, 210–219.
Schleich, H. H., Kästle, W., & Kabisch, K. (1996). Amphibians and reptiles of North Africa. Königstein: Koeltz Scientific Publishers.
Seoane, V. L. (1884). Identidad de Lacerta schreiberi (Bedriaga) y Lacerta viridis var. gadovi (Boulenger) e investigaciones herpetológicas de Galicia. A Coruña: Vicente Abad.
Sindaco, R., Venchi, A., & Grieco, C. (2013). The reptiles of the Western Palearctic, volume 2: Annotated checklist and distributional atlas of the snakes of Europe, North Africa, Middle East and Central Asia, with an update to volume 1. Latina: Edizioni Belvedere.
Sloss, B. L., Schuurman, G. W., Paloski, R. A., Boyle, O. D., & Kapfer, J. M. (2012). Novel microsatellite loci for studies of Thamnophis gartersnake genetic identity and hybridization. Conservation Genetics Resources, 4, 383–386.
Stamatakis, A. (2014). RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30, 1312–1313.
Stuckas, H., Velo-Antón, G., Fahd, S., Kalboussi, M., Rouag, R., Arculeo, M., Marrone, F., Sacco, F., Vamberger, M., & Fritz, U. (2014). Where are you from, stranger? The enigmatic biogeography of North African pond turtles (Emys orbicularis). Organisms, Diversity & Evolution, 14, 295–306.
Veith, M., Mayer, C., Samraoui, B., Donaire Barroso, D., & Bogaerts, S. (2004). From Europe to Africa and vice versa: evidence for multiple intercontinental dispersal in ribbed salamanders (genus Pleurodeles). Journal of Biogeography, 31, 159–171.
Velo-Antón, G., Godinho, R., Harris, D. J., Santos, F., Martínez-Freiría, F., Fahd, S., Larbes, S., Pleguezuelos, J. M., & Brito, J. C. (2012). Deep evolutionary lineages in a Western Mediterranean snake (Vipera latastei/monticola group) and high genetic structuring in Southern Iberian populations. Molecular Phylogenetics and Evolution, 65, 965–973.
Velo-Antón, G., Pereira, P., Fahd, S., Teixeira, J., & Fritz, U. (2015). Out of Africa: did Emys orbicularis occidentalis cross the Strait of Gibraltar twice? Amphibia-Reptilia, 36, 133–140.
Veríssimo, J., Znari, M., Stuckas, H., Fritz, U., Pereira, P., Teixeira, J., Arculeo, M., Marrone, F., Sacco, F., Naimi, M., Kehlmaier, C., & Velo-Antón, G. (2016). Pleistocene diversification in Morocco and recent demographic expansion in the Mediterranean pond turtle Mauremys leprosa. Biological Journal of the Linnean Society, 119, 943–959.
Zazo, C. (1999). Interglacial sea levels. Quaternary International, 55, 101–113.
Acknowledgements
Markus Auer, José Carlos Brito, Pierre-André Crochet, Boualem Dellaoui, Marco Favelli, Daniel Jablonski, Jérôme Maran, Olivier Peyre, and Ulrich Scheidt donated samples. Philippe Evrard provided a photo of a red-eyed grass snake from Spain. Cäcilia Spitzweg helped with laboratory work.
Funding
This study was funded by the German Research Foundation (DFG; FR 1435/11-2). Menad Beddek’s PhD research was supported by a grant from ANRT/Naturalia Environnement CIFRE (no. 2013/0145); field work in Algeria was funded by Naturalia Environnement. Salvador Carranza’s work was funded by CGL2015-70390-P from the Ministerio de Economía y Competitividad, Spain (cofunded by FEDER).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(PDF 1.24 mb).
Rights and permissions
About this article
Cite this article
Kindler, C., de Pous, P., Carranza, S. et al. Phylogeography of the Ibero-Maghrebian red-eyed grass snake (Natrix astreptophora). Org Divers Evol 18, 143–150 (2018). https://doi.org/10.1007/s13127-017-0354-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-017-0354-2