Abstract
For the disulfide-bonded β-sheet-forming peptides which include protegrins, the presence of positively charged amino acid residues allows their strong interaction with the lipid matrix of the plasma membrane, as opposed to a protein target on the surface of the cell. We used high-resolution NMR spectroscopy for 3D structure determination of several protegrins in a solution with perdeuterated dodecylphosphocholine (DPC) micelles which is a commonly used zwitterionic detergent for the solubilization of membrane peptides and proteins. Structural studies by NMR spectroscopy of protegrins PG-1 (PDB ID: 1PG1), PG-2 (PDB ID: 2MUH), and PG-3 (PDB ID: 2MZ6) shown that the sidechains of Leu5, Phe12, Val14, and Val16 form a relatively well-ordered apolar cluster. Due to the fact that a membrane surface with positive curvature allows the hydrophobic cluster to be buried while the charged residues remain solvated in water, here we can conclude that this area could bind to the bacterial cell walls via hydrophobic and positively charged amphipathic surfaces.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Antimicrobial peptides (AMPs) are small peptides with a strong antibiotic activity which play an important role in the immune system of many different animals. Naturally occurring AMPs probably represent one of the very first evolved forms of chemical defense of living eukaryotic cells against invasion by bacteria, protozoa, fungi, and virus. AMPs are less susceptible to the development of bacterial resistance because they disrupt the membrane of bacteria through non-specific peptide–lipid interactions. Natural AMPs have such properties as the broad-spectrum antibacterial activity, high selectivity, and the disruption of bacterial cell membranes, which allow to suggest that these molecules are potentially useful as antibiotics; so, it is important to investigate their structure and function at high resolution in order to increase their potency and selectivity. Protegrins are one of the best characterized β-sheet cytolytic peptides isolated from porcine leukocytes that contain 16–18 residues whose structure has been shown to be a β-hairpin stabilized by two disulfide bonds [1–4]. There are five known protegrins: PG-1 (RGGRL5CYCRR10RFCVC15VGR18), PG-2 (RGGRL5CYCRR10RFCIC15V16), PG-3 (RGGGL5CYCRR10RFCVC15VGR18), PG-4 (RGGRL5CYCRG10WICFC15VGR18), and PG-5 (RGGRL5CYCRP10RFCVC15VGR18) with a high content of cysteine (Cys) and several positively charged arginine (Arg) residues. For the disulfide-bonded β-sheet-forming peptides, which include protegrins, the presence of positively charged amino acid residues allows their strong interaction with the lipid matrix of the plasma membrane, as opposed to a protein target on the surface of the cell.
2 Material and Methods
The protegrin peptides were synthesized by solid-phase peptide synthesis by Dr. Andrey Filippov in the Chemistry of Interfaces Laboratory at the Luleå University of Technology.
The NMR investigation was done on a Bruker Avance 700 spectrometer equipped with a cryoprobe. The peptide (4 mg) was solubilized in an aqueous solution (H2O or 2H2O, 500 μl) containing 20 mg perdeuterated dodecylphosphocholine (DPC) (molar ratio ∼1:12). 3-(Trimethylsilyl)-propionic-2,2,3,3–2H4 acid (TMSP-2,2,3,3-2H4) (98 % atom 2H, Aldrich) was added as an internal chemical shift standard for 1H NMR spectroscopy. Perdeuterated d38 DPC (98 % 2H) and TSP-d4 were purchased from Aldrich. Chemical shift assignments of PG-2 and PG-3 in DPC micelles were obtained using standard methods of protein NMR spectroscopy by 2D NMR 1H–1H TOCSY and NOESY (Nuclear Overhauser Effect SpectroscopY) experiments. Interproton NOE distance constraints determined from NOESY experiments were used for the structural calculations by molecular dynamic method calculations of the Xplor-NIH program [5].
3 Results and Discussion
Protegrins are antibacterial peptides of 16–18 residues whose structure has been shown to be a β-hairpin stabilized by two disulfide bonds [3]. The most studied of these is the protegrin-1 (PG-1) which is composed of 18-amino acid residues (PDB IDs: 1PG1 and 1ZY6) and supposed that PG-1 forms ion channels in membranes and kills bacteria involved in oligomeric peptide toroidal pores in anionic lipid bilayer membranes, which mimic the inner membrane of Gram-negative bacteria. However, even for PG-1, there are a lot of experimental and theoretical studies; for other protegrins (PG-2, PG-3, PG-4, PG-5), structural studies are almost have not been investigated and biological functions among the different protegrins are also still limited.
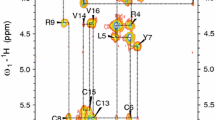
We used high-resolution NMR spectroscopy for 3D structure determination of protegrins PG-2 and PG-3 in a solution with perdeuterated dodecylphosphocholine (DPC) micelles which is a commonly used zwitterionic detergent for the solubilization of membrane peptides and proteins, and found a tendency for dimerization of PG-3 the same as for PG-1 [6–8]. Protegrins PG-1, PG-2, and PG-3 form a well-defined structure composed of a two-stranded antiparallel β-sheet in the presence of DPC micelles [3, 4, 7, 8]. In present paper, we focused on the protegrin peptide–micelle binding via hydrophobic and positively charged amphipathic surfaces. Structural studies by NMR spectroscopy of protegrins PG-1 (PDB ID: 1PG1), PG-2 (PDB ID: 2MUH), and PG-3 (PDB ID: 2MZ6) shown that the sidechains of Leu5, Phe12, Val14, and Val16 form a relatively well-ordered apolar cluster (Fig. 1). Due to the fact that a membrane surface with positive curvature allows the hydrophobic cluster to be buried while the charged residues remain solvated in water, here we can conclude that this area could bind to the bacterial cell walls via hydrophobic and positively charged amphipathic surfaces. These observations also support the hypothesis of the requirement of hydrophobic/hydrophilic cluster proximity to induce a cytolytic activity of protegrins [1]. Here, we should mentioned that even though solution NMR experiments on DPC micelles provide high-resolution structures, the structural results obtained from a detergent micelle need to be substantiated by studies in lipid bilayers, which do not have a curved surface and are a better approximation to cell membranes. We plan to investigate these problems in our further researches of AMPs.
Kyte–Doolittle hydrophobicity surface structural model for PG-1 1 (PDB ID: 1PG1), PG-2 (PDB ID: 2MUH), and PG-3 (PDB ID: 2MZ6): from blue for the most hydrophilic to white and to red for the most hydrophobic (50). An apolar cluster (Leu5, Phe12, Val14, and Val16) is marked by a dotted line. The structures were drawn using chimera (https://www.cgl.ucsf.edu/chimera/)
4 Conclusions
Structural studies by NMR spectroscopy of protegrins PG-1 (PDB ID: 1PG1), PG-2 (PDB ID: 2MUH), and PG-3 (PDB ID: 2MZ6) shown that the sidechains of Leu5, Phe12, Val14, and Val16 form a relatively well-ordered apolar cluster. Due to the fact that a membrane surface with positive curvature allows the hydrophobic cluster to be buried while the charged residues remain solvated in water, here we can conclude that this area could bind to the bacterial cell walls via hydrophobic and positively charged amphipathic surfaces.
References
Aumelas, A., Mangoni, M., Roumestand, C., Chiche, L., Despaux, E., Grassy, G., et al. (1996). Synthesis and solution structure of the antimicrobial peptide protegrin-1. European Journal of Biochemistry, 237, 575–583.
Cho, Y., Turner, J. S., Dinh, N. N., Lehrer, R. I. (1998). Activity of protegrins against yeast-phase Candida albicans. Infection and Immunity, 66, 2486–2493.
Fahrner, R. L., Dieckmann, T., Harwig, S. S. L., Lehrer, R. I., Eisenberg, D., Feigon, J. (1996). Solution structure of protegrin-1, a broad-spectrum antimicrobial peptide from porcine leukocytes. Chemical Biology, 3, 543–550.
Kokryakov, V. N., Harwig, S. S. L., Panyutich, E. A., Shevchenko, A. A., Aleshina, G. M., Shamova, O. V., et al. (1993). Protegrins—leukocyte antimicrobial peptides that combine features of corticostatic defensins and tachyplesins. FEBS Letters, 327, 231–236.
Schwieters, C. D., Kuszewski, J. J., Tjandra, N., Clore, G. M. (2003). The Xplor-NIH NMR molecular structure determination package. Journal of Magnetic Resonance, 160, 65–73.
Usachev, K. S., Filippov, A. V., Khairutdinov, B. I., Antzutkin, O. N., Klochkov, V. V. (2014). NMR structure of the arctic mutation of the Alzheimer’s Aβ(1-40) peptide docked to SDS micelles. Journal of Molecular Structure, 1076, 518–523.
Usachev, K. S., Efimov, S. V., Kolosova, O. A., Filippov, A. V., Klochkov, V. V. (2015). High-resolution NMR structure of the antimicrobial peptide protegrin-2 in the presence of DPC micelles. Journal of Biomolecular NMR, 61, 227–234.
Usachev, K. S., Efimov, S. V., Kolosova, O. A., Klochkova, E. A., Aganov, A. V., Klochkov, V. V. (2015). Antimicrobial peptide protegrin-3 adopt an antiparallel dimer in the presence of DPC micelles: a high-resolution NMR study. Journal of Biomolecular NMR, 62, 71–79.
Acknowledgments
The work is performed in accordance with the Russian Government Program of Competitive Growth of Kazan Federal University, received a subsidy allocated to Kazan Federal University for the project part of the state assignment in the sphere of scientific activities, and funded by RFBR, according to the research project No. 16-34-60001 mol_a_dk.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kolosova, O.A., Usachev, K.S., Aganov, A.V. et al. Antimicrobial Peptide Protegrins Interact with DPC Micelles by Apolar Hydrophobic Cluster: Structural Studies by High-Resolution NMR Spectroscopy. BioNanoSci. 6, 317–319 (2016). https://doi.org/10.1007/s12668-016-0218-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12668-016-0218-9