Abstract
Brain tumor is the most severe nervous system disorder and causes significant damage to health and leads to death. Glioma was a primary intracranial tumor with the most elevated disease and death rate. One of the most widely used medical imaging techniques for brain tumors is magnetic resonance imaging (MRI), which has turned out the principle diagnosis system for the treatment and analysis of glioma. The brain tumor segmentation and classification process was a complicated task to perform. Several problems could be more effectively and efficiently solved by the swarm intelligence technique. In this paper, the fuzzy brain-storm optimization algorithm for medical image segmentation and classification was proposed, a combination of fuzzy and brain-storm optimization techniques. Brain-storm optimization concentrates on the cluster centers and provides them the highest priority; it might fall in local optima like any other swarm algorithm. The fuzzy perform several iterations to present an optimal network structure, and the brain-storm optimization seems promising and outperforms the other techniques with better results in this analysis. The BRATs 2018 dataset was used, and the proposed FBSO was efficient, robust and mainly reduced the segmentation duration of the optimization algorithm with the accuracy of 93.85%, precision of 94.77%, the sensitivity of 95.77%, and F1 score of 95.42%.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Early brain tumor recognition and detection are essential for patients. Brain tumors mainly classified into two types: benign and malignancy. Benign is a non-cancerous tumor, and malicious are those that have cancer cells (Jin et al. 2014).
The most common primary brain tumors are,
-
Gliomas (50.4%)
-
Meningiomas (20.8%)
-
Pituitary adenomas (15%)
-
Nerve sheath tumors (8%)
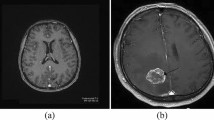
Nowadays, CADe systems were typically utilized for an efficient and explicit recognition of brain anomalies. Computer-aided detection (CADe), also known as computer-aided diagnosis (CADx), is systems that support doctors in the interpretation of the medical images. CADe is an interdisciplinary technology combining elements of AI and computer vision with image processing of radiological and pathology (Manimurugan et al. 2014; Narmatha et al. 2017; Thavasimuthu et al. 2019; Rajendran et al. 2019). The brain tumor is the abnormal development of the central spine or tissue, which could interfere with the best possible brain function. Additionally, the procedures of tumor recognition must be made exceptionally fast and accurate. This is just conceivable by using MRI, and uncertain locales are removed through MRI segmentation from complicated medical images (Prajoona et al. 2020). Figure 1 represents different classifications of MRI.
a T1-weighted MRI; b T2-weighted MRI; c FLAIR; and d FLAIR with contrast enhancement (https://bit.ly/33ab5og)
The detection of brain tumor represents the affected brain area as well as the tumor boundary, size, position, and shape. The tumor structure can be analyzed with a computed tomography (CT) or MRI scan. In any case, the CT scan includes radiation that was impeding to the patient’s body, while MRI provides a precise perception of the anatomy structure of brain tissues. The MRI is a device that directs radio waves and magnetic fields for producing extensive images of the tissues and organs (Muhammad et al. 2018; Manasi and Bhakti 2017; Bjoern et al. 2015).
For brain tumor detection, medical image analysis on the Brain Tumor Segmentation (BRATS) MRI dataset using Fuzzy Brain-Storm Optimization (FBSO) is performed in this work. Medical Image Analysis includes three stages (1) feature extraction and representation, (2) selection of feature that would be used for classification and (3) image classification (Bjoern et al. 2015).
The classification of the medical image has three fundamental steps: preprocessing, feature extraction and classification. After the preprocessing, it needs to extricate features of interest segment from the image for the additional process. The reason behind the classification of pattern framework was to delineate input variables (for example, record information or image information) to turn into the output factors (to portray one particular class (with or without disease class) (Meiyan et al. 2014).
The fundamental target of the classification of medical image is not just to achieve higher accuracy, but to recognize which area of the human body was affected by the disease (Javeria et al. 2018) Image classification is the labelling of a pixel or a group of pixels dependent on its grey value. Classification is one of the regularly used techniques for data extraction. In Classification, several features were used for the pixels set, i.e., several images of the specific object were required. The image classification process that would be used as per the following:
Image acquisition: It is the activity of recovering an Image from the Source.
Image enhancement: Image enhancement supports the qualitative improvement of the image as for a particular application.
Feature extraction: It is the information cleaning stage in which the features pertinent to the classification are extricated from the cleaned images.
Classification: Image classification is the labelling of a pixel or a set of pixels dependent on its gray value (Abdu et al. 2019). The process of image classification is shown in Fig. 2.
The BSO algorithm is basic in concept and simple in implementation. There are three methods in this algorithm: the solution clustering, new individual generation, and selection. In this algorithm, the solutions were divided into many clusters. The population’s best solution will be kept if the newly produced solution at a similar index is not right. A new individual could be generated based on one or two individuals in the clusters. The exploitation capacity is improved when the new individual was close to a better solution so far. While the exploitation capacity was enhanced when the new individual was randomly produced, or produced by individuals in two clusters. This algorithm is a sort of search space reduction; each solution would get into many clusters, respectively. These clusters specify a problem’s local optima. The data of a region includes solutions with better fitness values that were propagated from one cluster to other. This algorithm would explore in decision space initially, and the exploitation and exploration will get into a state of equilibrium after iteration. The fuzzy logic is combined with this proposed algorithm in which the fuzzy performs several iterations to present an optimal network structure.
The rest of the paper includes as Sect. 2 introduces the system’s materials and methods, including implementation using Fuzzy and brain-storm optimization, Sect. 3: Summaries the result analysis. Section 4 concludes the work.
2 Proposed methodology
2.1 Brain-storm optimization
The BSO is another sort of Swarm Intelligence algorithm that relies upon the total human behaviour, which is, the brain-storming method. It acknowledges the Brain-storming technique which people use. People get together contemplating a solution or more to an issue. They develop their ideas, discuss them, access them, and choose the better ideas, then proceed for iterations. The brain-storming procedure proceeds for iterations for accomplishing an efficient solution for the problem close by over participation among people who are solidly related or not to the problem. This technique showed to be compelling about complex practical issues. The process of the BSO algorithm is represented in Fig. 3.
Rabie (2017), developed this method for solving the problems of optimization in different domains of science. The researcher thought about the human as the smartest creature, and adapting their point of view must bring about compelling solutions. This process was transformed into a conventional algorithmic technique to be sensible for optimizations problem comprising clustering processes, generation processes, selector and mutation operator. The actual BSO uses K-Means as the clustering method.
2.2 Process of brain-storming
- Step 1:
-
The solutions are isolated into numerous clusters.
- Step 2:
-
The population’s best solution will be kept if the new produced solution at a similar index is not right.
- Step 3:
-
A new individual could be produced on the basis of one or two cluster individuals.
- Step 4:
-
The ability of exploitation is upgraded when the new individual is close to the better solution identified.
- Step 5:
-
While the ability of exploration is upgraded when the new individual is produced randomly, or produced by two clusters individuals.
The values of the Gaussian random (GR) are consolidated to create new individuals dependent on Eq. (1).
where Xnew is the lately generated individuals and Xold is the chosen individuals to create a new one. n(μ,\( \sigma\)) is the Gaussian function with variance σ and mean μ. ε is a contributing weight of GR value and computed by Eq. (2).
logsig, the function of the logarithmic sigmoid function, miteration is the maximum number of iteration, citeration is the value of present iteration, k is an element for changing the log sigmoid function slope, and rand () is a function of random generator which creates integer around 0 and 1.
2.3 Fuzzy C-mean clustering
FCM is a technique for clustering that partitions one data group into two or more clusters. This technique regularly utilized in pattern detection. This technique performs by allocating membership to every data point comparing to every cluster center (CC) depends on the distance among the cluster and data point. The benefits of the FCM algorithm contain: (1) Giving the best outcome for the overlapped dataset and similarly superior to the k-means algorithm. (2) Contrarily k-means where data point should only have a place with one cluster center, allocating the membership of data point on more than one cluster center. Thus, a data point may have a place with more than one CC. (3) the application of FCM to MR data has demonstrated promising results (Shahariar 2019).
The idea driving FCM was the fuzziness, which impacts each component in the data to different groups. Not at all like the K-Means, where all components have a spot only from a particular set, FCM moderates this difficult constraint to make all components to get a place with a various group with the membership value liable to the addition of every membership values of every point to each cluster should be 1. FCM uses the function of minimization to divide the data sets or the points. Along these, the function of membership U might have components between 0, and 1 with an addition of an information point was proportional to 1 as present in Eq. (3).
The cost/objective function is described as in Eq. 4,
where uij value is in the scope of [0, 1], ci, the focal point of cluster i, dij is the Euclidian distance between the j-data point and ci is the cluster center and m-weight exponents and the values was the scope of (1, ∞). Notwithstanding, the imperatives of ci and uij to achieve the necessary limitation are as per the following Eqs. (5) and (6):
Thus, FCM starts by enabling U as a matrix of membership through random value conditions (3). The fuzzy cluster centers c is hence computed using condition (5). The cost function is thus calculating using condition (4) and relied upon its value the algorithm might end, regardless of whether it was on a specific rate or the enhancement with the last round was lower than a particular threshold. When there is an option for improvement, U is hence assessed using condition (6), and the algorithm is repeated.
2.4 Feature extraction
It is a method toward gathering more high-level information of an image like shape, color, contrast, and texture. Analysis of texture is a significant element of human vision conception and ML frameworks. It was utilized successfully to enhance the precision of the diagnostic framework by choosing significant features. The GLCM and texture feature used for this image analysis process. This method includes two states for the extrication of features out of the images. In the initial state, the GLCM was assessed, and in the following stage, the characteristics of texture dependent on GLCM were computed. Because of the complicated shape of differentiated tissues, for example, Gray Matter (GM), White Matter (WM), and Cerebro-Spinal Fluid (CSF) in the brain MRI, the extrication of the significant feature was a necessary process. Textural results and analysis would enhance the diagnosis, various phases of the tumor (staging of tumor), and therapy reaction evaluation (Nilesh et al. 2017; Sérgio et al. 2016; Aboul 2019).
Contrast: The contrast is an estimation of the intensity of pixel and its neighbor over the image, and it was characterized as in Eq. 7,
Homogeneity or inverse difference moment: IDM was an estimation of the local homogeneity of images. IDM might have a one or an extent of value to decide if the image was non-textured or textured.
Energy: It could be characterized as the computable measure of the range of pixel pair reiterations. It is the metric to quantify the correlation of image. When energy was characterized by GLCM feature, at that point it is additionally meant to as angular second moment, and it was described as in Eq. 9,
Correlation: This feature defines the spatial conditions among the pixels, and it is characterized as in Eq. 10,
where mean is Mx, and SD is σx in the horizontal spatial domain and Mean My, and SD is σy in the vertical domain.
Coarseness: It was an estimation of solidness in the textural review of the images. For a fixed window size, texture with a smaller number of texture elements is said more coarse than one with a larger number. The rough texture refers to high coarseness value. Fine texture obtains lower coarseness value (Andriy 2018). It was characterized as in Eq. 10
FBSO algorithm: The proposed FBSO is discussed in this part. BSO mainly depends on the clustering, so the K-means algorithm is implemented. K-means enable every concept to part of to just a single cluster, and the new Cluster Center (CC) was picked dependent on new cluster individuals/concepts. In a brain-storm, the produced CC idea was considered with a high possibility than various ideas in a cluster. Along these, the global data of the workspace was not completely used. Subsequently, a portion of the best ideas may be lost because of the attention on the CCs.
The predator–prey approach is used when generating new ideas to deviate the search from the local optima. The cluster centers act as predators, having a trend moving towards better and better positions. Whereas the other ideas are preys, escaping from the nearest predator. As a result, the cluster centers can keep the best individuals of the swarm going towards the global best position. At the same time, the prey operation prevents the swarm from trapping in a local optimum.
The predator–prey methodology is used when producing new solutions. Hence the Eq. (1) will be replaced by Eqs. (12) and (13) as given below,
where Xpred, Xprey, and Xcenter are the new positions of a predator, a new location of prey and the closest predator to the present prey, respectively. P is treated as a binary value to determine if the prey can escape or not. The ‘a’ and ‘b’ are two variables that define the complexity of avoiding and calculated as given in Eq. 14.
where processes Xspan is the design variable’s searching range. Pareto dominance is about how good things are from the perspective of two different participants. It is used to choose the best idea generated.
Therefore, the FBSO algorithm could be summarized as the following steps:
-
1.
Specify the count of iterations (Imax), count of the idea (D) to be early produced, count of the cluster (Cmax), Pro1, 2, 3, σ, μ, and m.
-
2.
Attributably produce D ideas.
-
3.
Assess the D generated solution.
-
4.
Use the Pareto dominance on the idea produced
-
5.
Use FCM to cluster, the ideas produced into a count of cluster Cmax.
-
6.
Order the chosen n solution and select the better global solution (Xgbest)
-
7.
Generation of New Individuals:
-
a.
From Pro1, change the chosen clusters with the attributably created idea.
-
b.
From Pro2, attributably select a cluster; else select two clusters.
-
c.
From Pro3, choose the CC(s) and proceed to (d); else choose various ideas from the early picked clusters and proceed to (e); those idea(s) indicate the old ideas.
-
d.
Produce new ideas with the predator process; then proceed to Step e.
-
e.
Produce a new idea with the prey process.
-
f.
Use crossover operation among the newly produced ideas and the old ones; choose the better idea to change the old one, if needed.
-
g.
If n ideas are updated, proceed to step 8, else, to (a).
-
8.
If the present count of iterations < Imax, proceed to Step 3.
-
9.
Assess the present solutions and end.
3 Result analysis
3.1 Description of dataset
Ample multi-institutional routine clinically acquired pre-operative multimodal MRI diagnosis of lower-grade glioma (LGG) and glioblastoma (GBM/HGG) with pathologically affirmed diagnosis and available Overall Survivors, presented as the validation, training, and testing data in this dataset. In this dataset, (a) native (T1) and (b) post-contrast T1-weighted (T1Gd), (c) T2-weighted (T2), and (d) T2 Fluid Attenuated Inversion Recovery (FLAIR) volumes and were acquired with various clinical protocols and different scanners from multiple institutions (SBIA 2020). BRATs 2018 dataset includes multimodal MRI scans from 75 LGG patients, and 210 HGG patients are trained and tested on the presented model. The validation data set consists of 66 patients of unknown class. The dataset gives four MRI T1, T1 + Gd, T2w and FLAIR image volume for every patient, as shown in Fig. 4. Every sort of MRI scan was acquired by utilizing diverse MRI sequences setting, and henceforth tumor areas display contrastingly on every MRI scan (Hu et al. 2019; Anitha and Murugavalli 2015). These images were with the size of 240 × 240 × 155 pixels. The training data set’s ground-truths were given by the organizers of BRATS.
Each color in the ground truth image indicates a tumor class. Where red indicates Necrosis, yellow indicates Enhancing Tumor, blue indicates Non-enhancing Tumor, and green indicates Edema. The labels are classified into five classes, such as (label 0)—healthy tissues, (label 1)—Necrosis, (label 2)—Edema, (label 3)—non-enhancing, and (label 4)—enhancing tumor (Chao et al. 2018; Parasuraman and Vijaykumar 2019).
Based on the protocol in databases of BRATS, the structure of tumors is classified into three sub-regions for every subject.
-
i.
The overall tumor section (four sub-tumoral classes).
-
ii.
The core tumor section (total tumor area but except the “Edema” region).
-
iii.
The enhancing tumor section (just “enhancing” region).
The one-dimensional entropy was used, which represents the complete data of the image to show the measure of data included in the grey value distribution of the image. If the image entropy is higher, it provides a clearer image and better content. The 1D entropy of a grayscale image could be determined as in the Eq. 15,
where pi indicates the probability which a pixel with a grey value of i shows up in the image, the grey value range of image was typically from 0 to 255 in digital image processing. The variance among image entropy and classes could numerically estimate the efficiency of the segmentation of the image. The bigger the value is, the higher the dissimilarity among various classes. Contrasting the variance among classes was to evaluate the condition of the image segmented as per the range of the variation among regions. The variance among classes characterized as in Eq. 16:
where k is the count of CC, A and B indicate the area of the first and second regions, individually, which are commonly the total pixels in nearby regions, V1, V2 is the average grey values of the first and second areas. V is the average grey value of the two regions.
In this analysis, T1, T1 + Gd, T2 and FLAIR images are selected as input images, and the results are obtained the algorithm performed on chosen ten images. As shown in Fig. 5, the selected image is used for the different classification of the process. The evaluation of image entropy, variance and time computations are calculated and computed, and the calculated values are correlated with the existing methods like particle swarm optimization (PSO), glow swarm optimization (GSO), and whale swarm optimization (WSO).
The FBSO algorithm has high image entropy over the other algorithms and could select its parameters suitably, which results in producing the best segmentation outcomes than other algorithms. The variance values are additionally higher than the other algorithms, which represents that the FBSO algorithm preserves more data while segmenting grayscale images. The results of the segmented images are very similar to the original images. The comparison results among the other classes are better. The performance evaluation of precision, sensitivity, accuracy, and F1-score has been evaluated and compared with other techniques, as shown below (Shanmuganathan et al. 2020).
As shown in the above Eqs. 17–20, true positive (TP) indicates the total cases detected adequately as affected, FP indicates the total cases improperly detected as affected, TN indicates total cases correctly identified as non-affected and FN indicates total cases incorrectly detected as non-affected.
Sensitivity: From Table 1, it is the proportion of adequately anticipated positive perceptions of the total predicted positive perceptions. The computation of sensitivity analysis of the proposed technique FBSO algorithm accomplished 94.39%, whereas the PSO, WSO, GSO, and CBSO are lower than the proposed algorithm, as shown in Fig. 6.
Specificity: In terms of specificity from Table 2, it is the proportion of adequately anticipated negative perceptions of each observation in the real class. The proposed technique has obtained a high specificity rate compared to the other methods, as shown in Fig. 7.
Accuracy: From Table 3, it is significant natural performance estimation, and it is just a proportion of correctly anticipated perceptions of the total perceptions. Accuracy is termed as one of the main parameters in result analysis. The accuracy of the proposed technique has achieved better results compared to the other methods, as represented in Fig. 8.
F1-Score: From Table 4, it was the weighted average of Recall and Precision. Hence, this rate acquires both FP and FN into account. It is difficult to comprehend as accuracy, but F1 is very effective than accuracy, particularly if we have an irregular class distribution. As shown in the Fig. 9, the proposed technique has obtained a better F1-score result with 95.42%.
It can be seen from all the review of the result parameters which the fuzzy BSO algorithm has obtained the best results in terms of all the proposed parameters. Compared to (Muhammad et al. 2018) PCA-RELM technique, the proposed Fuzzy Brain-Storm Optimization technique has achieved 2.34% more accuracy, 8.85% more accuracy than the SOM-kNN technique (Aboul 2019), and 2.68% more accuracy than Ensemble classifier by (Hu et al. 2019). The fuzzy BSO can be utilized for any class of segmentation and classification analysis.
4 Conclusion
The fuzzy brain-storm optimization for medical image segmentation and classification was proposed. Analyzing brain MRI image is considered as the most significant concept for research and analysis. The brain tumor segmentation and classification process is a complex task to perform, and the proposed FBSO technique has achieved better results in terms of all parameters. For the brain MRI images, BRATs 2018 data set was used. The proposed FBSO was efficient, robust, and mainly reduced the segmentation time of the optimization algorithm with the accuracy of 93.85%, sensitivity of 94.39%, the specificity of 88.96%, and F1 score of 95.42%. In the future, the algorithm with BSO and fuzzy not only be used for segmenting and classifying medical images but also, can be used for different processes like classifications and detection.
References
Abdu G, Mohammad MH, Md Rafiul H, Abdulhameed A, Fortino G (2019) A hybrid feature extraction method with regularized extreme learning machine for brain tumor classification. IEEE Access 7:36266–36273
Aboul EH (2019) Machine learning paradigms: theory and application. Springer Nature, Geneva
Andriy M (2018) 3D MRI brain tumor segmentation using autoencoder regularization. Comput Vision Pattern Recog. arXiv:1810.11654
Anitha V, Murugavalli S (2015) Brain tumour classification using two-tier classifier with adaptive segmentation technique. IET Comput Vis 10:1–9
Bjoern HM et al (2015) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE T Med Imaging 34(10):1993–2024
Chao M, Gongning L, Kuanquan W (2018) Concatenated and connected random forests with multiscale patch driven active contour model for automated brain tumor segmentation of MR images. IEEE T Med Imaging 37(8):1943–1953
Hu K et al (2019) Brain tumor segmentation using multi-cascaded convolutional neural networks and conditional random field. IEEE Access 7:92615–92629
Javeria A, Muhammad S, Mudassar R, Mussarat Y (2018) Detection of brain tumor based on features fusion and machine learning. J Ambient Intell Human Comput. https://doi.org/10.1007/s12652-018-1092-9
Jin L, Min L, Jianxin W, Fangxiang W, Tianming L, Yi P (2014) A survey of MRI-based brain tumor segmentation methods. Tsinghua Sci Technol 19(6):578–595
Manasi K, Bhakti S (2017) A survey on medical image classification techniques. Int J Innov Res Comput Commun Eng 5(7):13510–13516
Manimurugan S, Porkumaran K, Narmatha C (2014) The new block pixel sort algorithm for TVC-encrypted medical image. Imaging Sci J 62(8):403–414
Meiyan H, Wei Y, Yao W, Jun J, Wufan C, Qianjin F (2014) Brain tumor segmentation based on local independent projection-based classification. IEEE T Bio-Med Eng 61(10):2633–2645
Muhammad S, Uroosha T, Ehsan UM, Muhammad AK, Mussarat Y (2018) Brain tumor segmentation and classification by improved binomial thresholding and multi-features selection. J Ambient Intell Human Comput. https://doi.org/10.1007/s12652-018-1075-x
Narmatha C, Manimegalai P, Manimurugan S (2017) A lossless compression scheme for grayscale medical images using a P2-bit short technique. J Med Imaging Health Inf 7(6):1196–1204
Nilesh BB, Arun KR, Har PT (2017) Image analysis for MRI based brain tumor detection and feature extraction using biologically inspired BWT and SVM. Int J Bio-Med Imaging 2017:9749108. https://doi.org/10.1155/2017/9749108
Parasuraman K, Vijaykumar B (2019) Brain tumor MRI segmentation and classification using ensemble classifier. Int J Recent Tech Eng 8(1S4):244–252
Prajoona V, Sriramakrishnan P, Sridhar S, Charlyn Pushpa Latha G, Priya A, Ramkumar S, Robert Singh A, Rajendran T (2020) Knowledge based fuzzy c-means method for rapid brain tissues segmentation of magnetic resonance imaging scans with CUDA enabled GPU machine. J Ambient Intell Human Comput. https://doi.org/10.1007/s12652-020-02132-6
Rabie AR (2017) Fuzzy brain storming optimization (FBSO) algorithm. Int J Intell Eng Inform 10(150):1
Rajendran T, Sridhar KP, Manimurugan S, Deepa S (2019) Advanced algorithms for medical image processing. Open Biomed Eng J 13:102
Section for Biomedical Image Analysis (SBIA) (2020). https://www.med.upenn.edu/sbia/brats2018/data.html. Accessed 25 Jun 2020
Sérgio P, Adriano P, Victor A, Carlos AS (2016) Brain tumor segmentation using convolutional neural networks in MRI images. IEEE T Med Imaging 35(5):1240–1251
Shahariar A et al (2019) Automatic human brain tumor detection in MRI image using template-based K means and improved fuzzy C means clustering algorithm. Big Data Cognit Comput 3(27):1–18
Shanmuganathan M, Saad A, Majed MA, Subramaniam G, Varatharajan R (2020) A review on advanced computational approaches on multiple sclerosis segmentation and classification. IET Signal Proc https://doi.org/10.1049/iet-spr.2019.0543
Thavasimuthu R, Sridhar KP, Manimurugan S, Deepa S (2019) Recent innovations in soft computing applications. Curr Signal Transduction Ther 14(2):129–130
Acknowledgements
We would like to thank the University of Tabuk for the immense support and encouragement for our research work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Narmatha, C., Eljack, S.M., Tuka, A.A.R.M. et al. A hybrid fuzzy brain-storm optimization algorithm for the classification of brain tumor MRI images. J Ambient Intell Human Comput (2020). https://doi.org/10.1007/s12652-020-02470-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12652-020-02470-5