Abstract
Heat-killed lactic acid bacteria perform immunomodulatory functions and are advantageous as probiotics, considering their long product shelf-life, easy storage, and convenient transportation. In this study, we aimed to develop appropriate heat treatments for industrial preparation of probiotics with antioxidant activity. Among 75 heat-killed strains, Lactococcus lactis MG5125 revealed the highest nitric oxide inhibition (86.2%), followed by Lactobacillus acidophilus MG4559 (86.0%), Lactobacillus plantarum MG5270 (85.7%), Lactobacillus fermentum MG4510 (85.3%), L. plantarum MG5239 (83.9%), L. plantarum MG5289 (83.2%), and L. plantarum MG5203 (81.8%). Moreover, the heat-killed selected strains markedly inhibited lipopolysaccharide-induced nitric oxide synthase and cyclooxygenase-2 expression. The use of heat-killed bacteria with intact bio-functionality can elongate the shelf-life and simplify the food processing steps of probiotic foods, given their high stability. The antioxidant and immune-modulatory activities of the heat-killed strains selected in this study indicate a strong potential for their utilization probiotic products manufacturing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Probiotics are defined as “living microorganisms that provide health benefits beyond inherent basic nutrition” when consumed in appropriate quantities [1, 2]. The beneficial effects of probiotics include prevention and treatment of diarrhea, systemic infections, inflammatory bowel disease, immunodeficiency, allergies, cancers, and cholesterolemia [3,4,5]. Functional food products containing probiotics have several therapeutic benefits including anticancer, hypoglycemic, antioxidant, and immunomodulatory effects [6, 7]. Therefore, the identification and isolation of new probiotic strains with health-promoting benefits have garnered immense interest in the medical and industrial sectors [7].
Inflammation is a complex response of vascular tissues to harmful stimuli such as pathogens, damaged cells, and stimulants. It is mediated by various signaling molecules produced by macrophages, monocytes, and mast cells. In chronically inflamed tissue, the stimulus is persistent; therefore, recruitment of monocytes is maintained, and existing macrophages are tethered in place. Macrophages are especially important in innate immunity, as they immediately respond to microbial infections. They can kill pathogens directly by phagocytosis and indirectly via secretion of pro-inflammatory cytokines such as tumor necrosis factor (TNF)-α, interleukin (IL)-1β, and IL-6 [8] as well as excess amounts of mediators such as nitric oxide (NO) and prostanoids in response to lipopolysaccharide (LPS). After stimulation with LPS, pro-inflammatory mediators, NO, and prostaglandin E2 (PGE2) are generated in abundance by inducible NO synthase (iNOS) and cyclooxygenase-2 (COX-2), respectively [9].
The radical produced from L-arginine through the action of NOS [10] is one of the most important inflammatory intermediates and plays a crucial role in biological processes, including neurotransmission, immune defense, and apoptosis. To date, three isoforms of NOS, based on their Ca2+ calmodulin dependence [11] or tissue type, have been identified. Among these isoforms, iNOS produces large amounts of NO when cells are stimulated with LPS and cytokines (TNF-α, IL-1β, IFN-γ), which is further associated with the generation of potent reactive radicals, such as peroxynitrite [12]. Chronic inflammation can contribute to inflammatory pathologies by killing even healthy host cells by NO [13]. COX has two isoforms, COX-1 and COX-2, which convert arachidonic acid to prostaglandins. Similar to iNOS, COX-2 is an inducible form that produces proinflammatory PGs in inflammatory site [14]. As the eicosanoids play a pivotal role in inflammation, pain, and fever, the modulation of iNOS and COX-2 overproduction might represent a therapeutic goal in numerous inflammatory pathologies.
Therefore, in the present study, we aimed to evaluate the antioxidant with inflammation potential of heat-killed lactic acid bacteria (LAB) isolated from human origin and fermented food. The effect of heat-killed selected strains on the expression of proinflammatory mediators and cellular signaling pathways was investigated in LPS-induced murine macrophage, RAW 264.7 cells.
Materials and Methods
Sample Preparation
In this study, 75 LAB were isolated from humans and fermented food [15]. Isolated strains were identified by the 16S rRNA gene sequencing method (SolGent Co., Ltd. Korea). The selected strains were cultivated and maintained in MRS broth (Difco Laboratories, USA) at 37 °C. To evaluate the inflammation potential of these strains, overnight cultivated selected strains were heat-killed at 90 °C for 30 min. Following centrifugation (12,000×g, 5 min), cell pellets were rinsed thrice with phosphate-buffered saline (PBS) and suspended in Dulbecco’s Modified Eagle’s Medium (DMEM, BD Biosciences, Frankin Lakes, NJ, USA) to obtain concentrations of 5 × 108 cells/mL by adjusting the absorbance at 600 nm wavelength.
Cell Culture
The murine macrophage RAW 264.7 cell line was obtained from the Korean Cell Line Bank (KCLB, Korea) and maintained in DMEM (Gibco, NY, USA) supplemented with 10% fetal bovine serum (FBS; Gibco, NY, USA) and 1% penicillin/streptomycin (Gibco, NY, USA) at 37 °C in an atmosphere of 5% CO2. Cells were sub-cultured and plated at 80–90% of confluency.
NO Production and Cell Viability
RAW 264.7 macrophage cells were grown at 37 °C and 5% CO2 in fully humidified air and sub-cultured every 3 days to 95% confluency. For routine subcultures, DMEM was supplemented with 10% FBS, penicillin (100 units/mL), and streptomycin (100 μg/mL). NO formation was detected based on the accumulation of nitrite, an indicator of NO synthesis, in the culture medium via the Griess reaction [16]. RAW 264.7 cells were plated at 2 × 105 cells/well in a 96-well plate and stimulated with 1 μg/mL LPS, followed by the addition of isolated bacterial strains (107 cells/well). After 24 h of incubation, NO concentration was determined by measuring the amount of nitrite in the cell culture supernatant using the Griess reagent. An absorbance measurement at 550 nm wavelength was obtained using the Epoch 2 microplate reader (BioTek, USA). Fresh culture medium was used as the blank control for all experiments.
3-[4,5-Dimethylthiazole-2-yl]-2,5-diphenyltetrazolium bromide (MTT; Sigma, USA) assay was performed to determine the viability of RAW 264.7 cells treated with the strains. RAW 264.7 cells were washed twice with PBS and 100 μL of MTT reagent (0.5 mg/mL) dissolved with PBS was added to each well. After 1 h of incubation, the MTT reagent was discarded and 100 μL of dimethyl sulfoxide (DMSO; Sigma, USA) was added to dissolve the formazan formed as a reactant between the MTT reagent and metabolites of live cells. The absorbance (A) was measured at 570 nm wavelength, and cytotoxicity was calculated in comparison with the result of a negative control group as follows.
Cell viability (%) = (A sample / A negative control) × 100.
In Vitro Antioxidant Properties of the Selected Strains
The 2,2-diphenyl-1-picrylhydrazyl (DPPH; Sigma, USA) radical scavenging assay was performed according to Blois [17], with slight modifications. Briefly, the selected strains adjusted to an OD600 of approximately 1.0 with PBS (pH 7.4) were added to 0.05 mM DPPH solution (1:2 v/v) and mixed well. Thereafter, the mixtures were kept at room temperature for 30 min in the dark. The control reaction was prepared using ethanol added to the DPPH solution. The absorbance of each mixture was quantified at 517 nm wavelength. Each sample assay was performed in triplicate. The results were compared with those of ascorbic acid (10 μg/mL), and the antioxidant activity was calculated using the following formula: Scavenging effect (%) = (Ac-As)/Ac × 100, where As is the absorbance of the test sample and Ac is the absorbance of the control at 517 nm wavelength.
Scavenging of the 1 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) (ABTS; Sigma, USA) radical was measured according to the method reported by Re et al. [18]. Briefly, the radical cation was prepared by mixing 7 mM of ABTS with 2.45 mM potassium persulfate (1:1 v/v), and the mixture was kept at room temperature in the dark for 24 h. Thereafter, 50 μL of the selected strain samples and 100 μL of ABTS solution were mixed and incubated for 10 min at room temperature. The absorbance of the mixture was measured at 734 nm wavelength. Each sample assay was performed in triplicate, and the scavenging rate was calculated as follows: scavenging rate (%) = (Ac-As)/Ac × 100, where As is the absorbance of the test sample and Ac is the absorbance of the control at 734 nm.
Semi-quantitative Reverse Transcriptase-Polymerase Chain Reaction
Semi-quantitative reverse transcriptase-polymerase chain reaction (RT-PCR) was performed to determine the expression of COX-2 and iNOS mRNA. Total RNA was extracted from RAW 264.7 cells using TRI REGENT™ (Sigma Chemical Co., St. Louis, MO), according to the manufacturer’s recommendation. iNOS and COX-2 primers were designed for RT-PCR. Glyceraldehydes-3-phosphate dehydrogenase (GAPDH; Sigma, USA) was used as a housekeeping gene to normalize all samples. Table 1 lists the sequences of primer pairs used to amplify iNOS, COX-2, and GAPDH. RT-PCR was performed using the ONE-STEP RT-PCR PreMix kit™ (Qiagen Inc. Valencia, CA, USA), according to the manufacturer’s instructions. Each of the primers and 1 μg of the RNA template were mixed with ONE-STEP RT-PCR PreMix™. These samples were processed by one-step RT-PCR, under the following conditions: predenaturation of RNA at 95 °C for 5 min; 40 cycles of 95 °C for 15 s, 61 °C for 30 s, and 61 °C for 30 s; and a final elongation step of 30 s at 61 °C. The extent of iNOS and COX-2 mRNA expression was quantified using a densitometer with Quantity One software (Bio-Rad Lab., Hercules, CA, USA).
Statistical Analysis
Results are expressed as means ± S.D. of three experiments. Difference between groups was evaluated using the Student’s t test, and a P value of < 0.05 was considered as statistically significant.
Results and Discussion
NO Productive Capacity and Cell Viability of Heat-Killed LAB
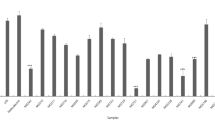
NO is a multi-functional mediator and plays a pivotal role in the immune response to inflammatory activity. The physiological or normal NO production in phagocytes is beneficial for the host defense against microorganisms, parasites, and tumor cells [19]. According to the results of the NO assay, the probiotic strains revealed a wide range of NO production inhibition rates (Table 2). This result indicated that bacterial strains would have different functional properties, even if they belong to the same species. Among the 75 probiotic strains, Lac. lactis MG5125 (86.2%) exhibited highest NO inhibition in LPS-induced cells, followed by L. acidophilus MG4559 (86.0%), L. plantarum MG5270 (85.7%), L. fermentum MG4510 (85.3%), L. plantarum MG5239 (83.9%), L. plantarum MG5289 (83.2%), and L. plantarum MG5203 (81.8%) (Table 2).
The heat-killed selected strains showed low toxicity in RAW 264.7 cells, with cell viability of approximately 82.06–111.66%. The effect of killed selected strains on cell viability increased in a dose-dependent manner (Table 3).
In Vitro Antioxidant Properties of the Selected Strains
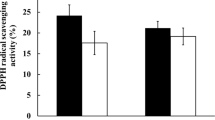
The antioxidant properties of the four selected strains, which revealed high inhibition of NO production, were evaluated by DPPH and ABTS radical scavenging activities. The DPPH free radical scavenging activities of the probiotic strains ranged from 14.7 to 20.9% (Fig. 1a). L. acidophilus MG4559 exhibited the highest radical scavenging activity (20.9%) and similar antioxidant activity when compared with that of ascorbic acid (10 μg/mL) control (17.9%), followed by L. plantarum MG5239 (20.1%). Regarding ABTS radical scavenging activities, results of the strains ranged from 43.0 to 53.3% (Fig. 1b). L. acidophilus MG4559 exhibited the highest radical scavenging activity (53.3%) and similar antioxidant activity when compared with that of ascorbic acid (10 μg/mL) control (56.7%), followed by Lac. lactis MG5125 (47.8%).
All seven strains presented high antioxidant activities, indicating that the selected probiotic strains possess the ability to reduce ROS. The trends of antioxidant activity and NO production inhibition rate were similar. Our results were in accordance with those of other studies regarding antioxidant activities of Lactobacilli. Li et al. [21] reported the antioxidant activities of L. plantarum strains derived from food, and Afify et al. [22] reported the ABTS radical scavenging effects of L. reuteri. Lin and Yen [23] evaluated the inhibitory effect of Bifidobacterium longum, and Kim et al. [24] isolated antioxidative Bifidobacterium species from infant fecal samples. Notably, probiotics produce bioactive compounds with beneficial properties, including antioxidant activity, and may act via specific molecular mechanisms responsible for defense against oxidative stress based on the strain specificity [20, 25].
Immunomodulatory Activity of Heat-Killed Selected Strains on Murine Macrophage RAW 264.7 Cells Via RT-PCR
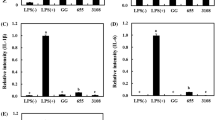
To evaluate the immunomodulatory ability, RT-PCR assays were performed. Cells were treated with heat-killed bacteria as described earlier, and the results are illustrated in Fig. 2. The iNOS and COX-2 gene expression markedly increased following LPS stimulation; however, heat-killed selected strains remarkably inhibited LPS-induced iNOS (Fig. 2a) and COX-2 (Fig. 2b) expression. The expression of the housekeeping gene GAPDH was not affected by the heat-killed selected strains. Raw 264.7 macrophages are representative antigen-presenting cells (APCs); they participate in the first stage of innate immunity by swallowing pathogens and inducing the release of intercellular signaling molecules, such as NO and COX-2. When the body’s immune system confronts immune-challenging elements, these cells rapidly initiate colonization, secrete cytokines, and activate natural killer cells and dendritic cells [26]. NO, induced by iNOS and cytokines, acts as a protective agent against pathogens and decreases their leukocyte adherent-activity [27]. In previous studies, the immuno-modulatory abilities of polysaccharides in cell walls have been reported by several researchers. Immunomodulatory polysaccharides activate innate and adaptive immune responses via direct and indirect interactions. Since probiotics are equipped with varying compositions of intrinsic enzymes, their immunomodulatory properties could be strain-specific [28].
RT-PCR analysis of mRNA expression of iNOS (a) and COX-2 (b). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as a housekeeping gene to normalize all samples. Data are representative of three experiments, respectively. Values represented indicate the means ± SD of three independent experiments. Different letters (a–d) indicate significant difference at p < 0.05
Conclusion
We aimed to select superior probiotic strains with desired antioxidant activity from 75 strains of probiotic candidates obtained from human origin and fermented foods by evaluating their inhibitory activity on NO production. In this study, molecular mechanisms were not elucidated and the heat-killed strains metabolites related to the inhibition of NO production should be investigated. However, we selected seven probiotic strains that exhibited high antioxidant activities. The use of heat-killed cells, which still maintain their bio-functionality, can elongate the shelf-life and simplify the food-processing steps of probiotic foods, given their high stability. Recently, LAB have been studied to prevent and treat inflammatory conditions in vivo and in vitro. These positive effects might be related to direct and indirect molecular mechanisms. Finally, the antioxidant and immunomodulatory activities of the heat-killed strains selected in this study indicate strong potential for their utilization in probiotic product manufacturing.
Data Availability
The authors declare that all data and materials support published claims and comply with field standards.
References
Hawley HB, Shepherd PA, Wheater DM (1960) Factors affecting the implantation of lactobacilli in the intestine. J Appl Bacteriol 22:360–367
Klein G, Pack A, Bonaparte C, Reuter G (1998) Taxonomy and physiology of probiotic lactic acid bacteria. Int J Food Microbiol 41:103–125. https://doi.org/10.1016/S0168-1605(98)00049-X
Gill HS, Guarner F (2004) Probiotics and human health: a clinical perspective. Postgrad Med J 80:16–26. https://doi.org/10.1136/pgmj.2003.008664
Spahaak S, Havenaar R, Schaafsma G (1998) The effect of consumption of milk fermented by Latobacillus casei strain Shirota on the intestinal microflora and immune parameters in humans. Eur J Clin Nutr 52:899–9070. https://doi.org/10.1038/sj.ejcn.1600663
Stiles ME, Holzapfel WH (1997) Lactic acid bacteria of foods and their current taxonomy. Int J Food Microbiol 36(1):1–29. https://doi.org/10.1016/S0168-1605(96)01233-0
Clare DA, Swaisgood HE (2000) Bioactive milk peptides: a prospectus. J Dairy Sci 83(6):1187–1195. https://doi.org/10.3168/jds.S0022-0302(00)74983-6
Khan SU (2014) Probiotics in dairy foods: a review. Nutr Food Sci 44(1):71–88. https://doi.org/10.1108/NFS-04-2013-0051
Stuehr DJ, Cho HJ, Kwon NS, Weise M, Nathan CF (1991) Purification and characterization of the cytokine-induced macrophage nitric oxide synthase: an FAD- and FMN –containing flavoprotein. Proc Natl Acad Sci USA 88(17):7773–7777. https://doi.org/10.1073/pnas.88.17.7773
Lee HJ, Kim NY, Jang MK, Son HJ, Kim KM, Sohn DH, Lee SH, Ryu JH (1999) A sesquiterpene, dehydrocostus lactone, inhibits the expression of inducible nitric oxide synthase and TNF-alpha in LPS-activated macrophage. Planta Med 65(2):104–108. https://doi.org/10.1055/s-1999-13968
Moncada S, Higgs A (1993) The L-arginine-nitric oxide pathway. N Engl J Med 329(27):2002–2012. https://doi.org/10.1056/NEJM199312303292706
Bredt DS, Snyder SH (1990) Isolation of nitric oxide synthetase, a calmodulin-requiring enzyme. Proc Natl Acad Sci USA 87(2):682–685. https://doi.org/10.1073/pnas.87.2.682
Boota A, Zar H, Kim YM, Johnson B, Pitt B, Davies P (1996) IL-1 beta stimulates superoxide and delayed peroxynitrite production by pulmonary vascular smooth muscle cells. Am J Physiol 271:L932-938. https://doi.org/10.1152/ajplung.1996.271.6.L932
Brown GC (2003) NO says yes to mitochondria. Science 299:838–839. https://doi.org/10.1126/science.1082028
Weisz A, Cicatiello L, Esumi H (1996) Regulation of the mouse inducible-type nitric oxide synthase gene promoter by interferon-gamma, bacterial lipopolysaccharide and NG-monomethyl-L-arginine. Biol Chem J 316:209–215. https://doi.org/10.1042/bj3160209
Kim HM, Kim JS, Kim YG, Jeong YA, Kim JE, Paek NS, Kang CH (2020) Antioxidant and probiotic properties of lactobacilli and bifidobacteria of human origins. Biotechnol Bioprocess Eng 25:421–430. https://doi.org/10.1007/s12257-020-0147-x
Lyons CR, Orloff GJ, Cunningham JM (1992) Molecular cloning and functional expression of an inducible nitric oxide synthase from a murine macrophage cell line. J Biol Chem 267(9):6370–6374. https://doi.org/10.1016/S0021-9258(18)42704-4
Blois MS (1958) Antioxidant determinations by the use of a stable free radical. Nature 181:1199–1200. https://doi.org/10.1038/1811199a0
Re R, Pellegrini N, Proteggente A, Pannala A, Yang M, Rice-Evans C (1999) Antioxidant activity applying an improved ABTS radical cation decolorization assay. Free Radic Biol Med 26:1231–1237. https://doi.org/10.1016/S0891-5849(98)00315-3
Lin YL, Lin JK (1997) (−)-Epigallocatechin-3-gallate blocks the induction of nitric oxide synthase by down-regulating lipopolysaccharide-induced activity of transcription factor nuclear factor-kB. Mol Pharmacol 52(3):465–472. https://doi.org/10.1124/mol.52.3.465
Amaretti A, di Nunzio M, Pompei A, Raimondi S, Rossi M, Bordoni A (2013) Antioxidant properties of potentially probiotic bacteria: in vitro and in vivo activities. Appl Microbiol Biotechnol 97:809–817. https://doi.org/10.1007/S00253-012-4241-7
Li S, Zhao Y, Zhang L, Zhang X, Huang L, Li D, Niu C, Yang Z, Wang Q (2012) Antioxidant activity of Lactobacillus plantarum strains isolated from traditional Chinese fermented foods. Food Chem 135(3):1914–1919. https://doi.org/10.1016/j.foodchem.2012.06.048
Afify AEMMR, Romeilah RM, Sultan SIM, Hussein MM (2012) Antioxidant activity and biological evaluations of probiotic bacteria strains. Int J Acad Res 4(6):131–139. https://doi.org/10.7813/2075-4124.2012/4-6/A.18
Lin MY, Yen CL (1999) Inhibition of lipid peroxidation by Lactobacillus acidophilus and Bifidobacterium longum. J Agric Food Chem 47(9):3661–3664. https://doi.org/10.1021/jf981235l
Kim JY, Choi SI, Heo TR (2003) Screening of antioxidative activity of Bifidobacterium species isolated from Korean infant feces and their identification. Biotechnol Bioprocess Eng 8:199–204. https://doi.org/10.1007/BF02935897
Di Cerbo A, Palmieri B, Aponte M, Morales-Medina JC, Iannitti T (2016) Mechanisms and therapeutic effectiveness of lactobacilli. J Clin Pathol 69(3):187–203. https://doi.org/10.1136/jclinpath-2015-202976
Chalamaiah M, Yu W, Wu J (2018) Immunomodulatory and anticancer protein hydrolysates (peptides) from food proteins: a review. Food Chem 245:205–222. https://doi.org/10.1016/j.foodchem.2017.10.087
Soufli I, Toumi R, Rafa H, Touil-Boukoffa C (2016) Overview of cytokines and nitric oxide involvement in immuno-pathogenesis of inflammatory bowel diseases. World J Gastrointest Pharmacol Ther 7(3):353–360. https://doi.org/10.4292/wjgpt.v7.i3.353
Yin HM, Wang SN, Nie SP, Xie MY (2018) Coix polysaccharides: gut microbiota regulation and immunomodulatory. Bioact Carbohydr and Diet Fibre 16:53–61. https://doi.org/10.1016/j.bcdf.2018.04.002
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kang, CH., Kim, JS., Kim, H. et al. Heat-Killed Lactic Acid Bacteria Inhibit Nitric Oxide Production via Inducible Nitric Oxide Synthase and Cyclooxygenase-2 in RAW 264.7 Cells. Probiotics & Antimicro. Prot. 13, 1530–1538 (2021). https://doi.org/10.1007/s12602-021-09781-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-021-09781-9