Abstract
We evaluated the probiotic properties of lactic acid bacteria using resistance, safety, and functional assays. A preliminary subtractive screening of nineteen strains was performed based on their survival in simulated gastric and intestinal juice, and cell surface characteristics (hydrophobicity and auto-aggregation). Five strains were selected for further characterization, which included the assessment of their co-aggregation to pathogens, phenol tolerance, antimicrobial activity, and safety. Moreover, their adhesion to Caco-2 and HT-29 cells and the ability to inhibit pathogenic bacteria adhesion were evaluated. All strains had high (≥ 80.0%) survival rates in gastric and intestinal juices. Among them, Lactobacillus brevis CCMA 1284, L. plantarum CCMA 0743, and L. plantarum CCMA 0359 exhibited higher hydrophobicity (95.33, 96.06, and 80.02%, respectively), while L. paracasei CCMA 0504 and L. paracasei CCMA 0505 had the highest auto-aggregation values (45.36 and 52.66%, respectively). However, these last two strains were positive for the DNAse test, which is a safety concern. The CCMA 0359 and CCMA 1284 strains did not show antimicrobial activity, while the CCMA 0505 strain had a higher percentage of adhesion (4.75%) to Caco-2 cells. In the simulated competition and exclusion assays, the CCMA 0743 strain was able to reduce Salmonella adhesion to both cells (Caco-2 and HT-29), but only the CCMA 0743 and CCMA 0505 strains inhibited Escherichia coli adhesion to HT-29 cells in the competition assay. According to the results of these evaluated attributes, this strain showed to be an excellent candidate for probiotic use.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Probiotic microorganisms are recognized for their many health benefits. Among the known probiotic microorganisms, various species of lactic acid bacteria (LAB), especially those of the Lactobacillus genus, are widely used as probiotic cultures, as well as for the development of probiotic fermented products [1, 2]. Although there are several strains with proven probiotic properties on the market, the search for novel strains with functional and technological characteristics remains an attractive goal to satisfy increasingly demanding consumers [1, 2], contributing to improved health and reducing the risk of disease [3].
LAB can be found in a variety of food matrices such as dairy products [4], meats [5], indigenous fermented beverages [6], and cocoa [7]. Thus, these food sources are potential reservoirs of novel probiotic strains. The study and selection of new probiotic strains require a systematic approach consisting of sequential evaluations to reduce the number of candidate strains. Probiotic characteristics are reported as strain-specific [8]; therefore, the evaluation of both wild and novel strains is essential, since isolates belonging to the same species may display different properties and probiotic mechanisms.
The methods and criteria used to characterize probiotic strains include assessing their ability to tolerate stress conditions exerted by the human body, ability to interact with host epithelial cells, safety attributes (such as β-hemolysis, gelatinase, and DNAse enzyme activities), and sensitivity to antibiotics [9], antimicrobial activity, and competition with pathogens [10]. Moreover, cell surface properties (hydrophobicity, auto-aggregation, and co-aggregation with pathogens) and the interaction of candidate strains with human epithelial cell lines and pathogenic bacteria represent different mechanisms that can be considered in the evaluation of probiotic efficacy [11, 12]. In vitro models employing the HT-29 and Caco-2 cell lines isolated from colonic adenocarcinomas have been widely used to investigate probiotic adhesion capacity [13].
Enteropathogenic Escherichia coli (EPEC) and Salmonella enterica are important human pathogens whose virulence traits depend on their ability to adhere to epithelial intestinal cells [14, 15]. The indiscriminate use of antibiotics to combat these microorganisms has contributed to the development of resistance mechanisms [16]. In this sense, probiotics have emerged as an alternative in the treatment of bacterial infections, mainly due to the protection conferred to the host cells [17]. Some studies have indicated that the LAB may prevent or reduce the attachment of pathogens to host cells [17, 18]. Here, we evaluated the in vitro probiotic properties of wild LAB strains isolated from different fermented food products. The antagonistic effects of selected LAB strain on the adhesion of pathogens to Caco-2 and HT-29 cells were evaluated by exclusion and competition assays.
Materials and Methods
Screening of LAB Strains
Nineteen LAB strains belonging to the Culture Collection of Agricultural Microbiology (CCMA) of the Federal University of Lavras and isolated from different substrates were initially employed in this study (Table 1). Sequentially, the pre-selection of LAB cultures was based on their ability to survive simulated gastric and intestinal juices and cell surface characteristics (hydrophobicity and auto-aggregation). Five strains (three with higher hydrophobicity and two with higher auto-aggregation) were selected for further characterization.
Survival to Simulated Gastric and Intestinal Juices
The survival of LAB in simulated gastric and intestinal juices was assessed as previously described [26] with modifications. Cultures were grown for 16 h at 37 °C in sterile MRS both, then 1 mL of each culture was mixed in 9 mL sterile saline solution (NaCl 0.85% w/v) with pH adjusted to 2.0 using 1 M hydrochloric acid (HCl) containing 0.5% pepsin (Fisher Scientific, UK) (w/v). After mixing, the initial bacteria counts were determined by plating. Subsequently, samples were incubated for 90 min at 37 °C and cell viability determined by plating. Then, the simulated intestinal juice was prepared by adding oxgall (Himedia, Mumbai, India) and pancreatin (Dinâmica, Brazil) solutions to obtain final concentrations of 0.3% and 0.1% (w/v), respectively, and the pH was then adjusted to 7.0 by adding 1 M sodium hydroxide (NaOH). After mixing, samples were incubated at 37 °C for 150 min, and then viable cell counts were determined. All sample counts were determined by plating on MRS (Man Rogosa and Shape, Kasvi, Italy) agar. The experiments were repeated three times and performed in triplicate. Results were expressed as mean log colony-forming units per mL (CFU/mL). The survival rate was calculated as follows:
Determination of Cell Surface Characteristics
The hydrophobicity, auto-aggregation, and co-aggregation assays were performed according to Kaktcham et al. [27] with slight modifications.
Hydrophobicity
The cell surface hydrophobicity of each strain was assessed by measuring microbial affinity to xylene. Briefly, cells collected from a 16-h old culture were centrifuged (10,000 rpm for10 min). The resulting pellet was washed twice with sterile phosphate-buffered saline (PBS) (pH = 7.2) and re-suspended in the same buffer. The optical density at the 600 nm wavelength (OD600) of the suspension was measured (A0) using a spectrophotometer (Spectrum, SP-2000UV). Thereafter, 1 mL of xylene was added to 3 mL of cell suspension and mixed by vortexing for 2 min. Then, the water and xylene phases were separated by incubation for 1 h at 37 °C. The aqueous phase was removed and the new OD600 was measured (A1). The percentage of the cell surface hydrophobicity was calculated using the formula:
The strains were then classified into microorganisms of low (0–34%), moderate (35–69%), and high hydrophobicity (70–100%) [28].
Auto-Aggregation
LAB cells were harvested from a 16-h old culture in MRS broth, washed twice with PBS (pH 7.2), re-suspended in the same buffer and diluted to an OD600 of 0.6 ± 0.1 (approximately 7–8 Log CFU/mL). Bacterial cell suspensions were vortexed for 10 s and subsequently incubated at 37 °C for 5 h. The auto-aggregation percentage was determined using the equation:
where At represents the absorbance at time t = 5 h and A0 represents the absorbance at time t = 0 h.
Co-Aggregation
For co-aggregation, the LAB strains were grown in MRS broth for 16 h at 37 °C, while enteropathogenic Escherichia coli (EPEC) CDC 055 and Salmonella enterica serovar Enteritidis ATCC 564 were grown in BHI (Brain-Heart Infusion) broth for 24 h at 37 °C. Bacterial suspensions were prepared as described in the auto-aggregation test above. Equal volumes (2 mL) of LAB and human pathogen suspensions were mixed by vortexing (10 s) and incubated at room temperature without agitation for 4 h. Control tubes contained 2 mL of the suspension of each bacterial strain. The absorbances (OD600) of the mixtures and controls were measured after incubation. The percentage of co-aggregation was calculated using the following formula:
where Alab and Apat refer to the OD600 of the LAB cell suspension and pathogen cell suspension, respectively, in control tubes and Amix represents the absorbance of the mixed bacterial suspension tested after 4 h.
Phenol Tolerance
Phenol tolerance was determined according to the method described by Shehata et al. [29]. Overnight cultures of LAB strains were inoculated (1%) into MRS broth with 0.2 and 0.5% (v/v) of phenol, or without phenol. Bacterial cells in the culture broth were quantified by reading the OD600 after 24 h of incubation at 37 °C. The experiments were performed in duplicate.
Antimicrobial Activity
The antimicrobial activity of LAB cultures was evaluated by the agar spot test according to Arena et al. [30] with modifications. An aliquot of 5 μL of each LAB isolate previously grown in MRS broth was separately spotted on MRS agar and plates were incubated at 37 °C for 48 h to allow the expression and secretion of antimicrobial compounds produced by cultures. The indicator microorganisms were Salmonella enterica serovar Enteritidis ATCC 564, enteropathogenic Escherichia coli (EPEC) CDC 055, Listeria monocytogenes ATCC 19117, Staphylococcus aureus ATCC 5674, and Bacillus cereus ATCC 14579. Overnight cultures of indicator microorganisms were mixed 1:100 with BHI soft agar (0.7% w/v) and overlaid on developed colonies (8 mm diameter) of LAB isolates. After incubation at 37 °C for 24 h, plates were checked for zones of inhibition surrounding the producer colonies. The experiment was repeated three times.
Antibiotic Susceptibility
The antibiotic susceptibility of the strains was determined by the disk diffusion assay. Overnight cultures (100 μL) were spread onto MRS agar media, and antibiotic discs containing ampicillin (10 μg), vancomycin (30 μg), gentamicin (10 μg), streptomycin (10 μg), chloramphenicol (30 μg), erythromycin (15 μg), azithromycin (15 μg), penicillin (10 μg), novobiocin (30 μg), oxacillin (1 μg), and lincomycin (2 μg) were placed on the surface of the inoculated plates using sterile forceps. Inhibition zone diameters were measured after incubation at 37 °C for 24 h. The susceptibility of the isolates was categorized as resistant (R), moderately susceptible (MS), or susceptible (S) according to interpretative values [31]. The experiment was repeated three times.
In Vitro Assessment of Safety Attributes
The safety of the isolates was investigated by assessing hemolysis, DNAse activity, and gelatin hydrolysis, as described by Singh et al. [32] with modifications. Hemolytic activity was determined by inoculating the strains on blood agar plates containing 5% sheep blood after 48 h incubation at 37 °C. The absence of an effect on blood plaques (γ-hemolysis) was considered non-hemolytic. Green-hued zones around the colonies (α-hemolysis) were considered as partial hemolytic activity, and strains showing clear areas of hydrolysis resulting from blood cell lysis around the colonies were classified as hemolytic strains (β-hemolysis). Gelatinase production by strains was analyzed using tryptone-neopeptone-dextrose (TND) agar (17.0 g tryptone, 3.0 g neopeptone, 2.5 g dextrose, 5.0 g NaCl, 2.5 g K2HPO4, 15 g agar, and 1 L distilled water) containing 0.4% gelatin. The LAB cultures were spot-inoculated onto plates containing the medium and incubated at 37 °C for 48 h. Enzyme production was visualized by the formation of a halo around the colony after addition to a saturated ammonium sulfate solution to confirm gelatin hydrolysis. For the DNAse test, strains were streaked on the DNAse test agar medium (Difco, USA) and the plates were incubated at 37 °C for 48 h. After this time, a 1 M HCl solution was added to the plate. A clear zone around the colonies after incubation was considered positive for DNAse production. For all tests, the Staphylococcus aureus ATCC 25923 strain was used as the positive control. The experiment was repeated three times.
Adhesion of LAB Strains to Caco-2 and HT-29 Cell Lines
Growth and Maintenance of Caco-2 and HT-29 Cells
The Caco-2 and HT-29 cells provided by the Cell Bank of Rio de Janeiro (BCRJ, Rio de Janeiro) were grown in modified Eagle’s minimal essential medium (MEM supplemented with 10% (v/v) heat-inactivated fetal bovine serum), 1× non-essential amino acids, and 0.1 mg/mL gentamicin. All solutions were obtained from Invitrogen, Gibco (Naerum, Denmark). The cells were maintained at 37 °C in a humidified atmosphere of 5% CO2. The culture medium was changed regularly and when the cells reached sub-confluence (80–90%), they were sub-passaged.
Adhesion Assay on Caco-2 and HT-29 Cells
The adhesion capacity test for the five selected strains (based on previous assays) to the human colon adenocarcinoma cell lines (Caco-2 and HT-29) was performed according to Ramos et al. [8] with slight modifications. The Caco-2 and HT-29 cells were sub-cultured (2 × 105 cells/mL) in 24-well tissue culture plates (Sarstedt, Germany) and grown at 37 °C in a humidified atmosphere of 5% CO2 for 21 days to promote differentiation in cell media. The culture medium was changed on alternate days.
For the adhesion assay, bacteria were cultured in MRS broth for 16 h at 37 °C and after washing twice with the phosphate-buffered solution (PBS), the cultures were re-suspended in the media (described above) at a concentration of approximately 108 CFU/mL. One milliliter of each bacteria suspension was added to cells in each well and incubated for 90 min at 37 °C in a 5% CO2 atmosphere. Subsequently, the cells were washed three times with 1 mL of PBS to remove non-adherent bacteria cells and then lysed with 1 mL of Triton-X solution (0.1% v/v in PBS). After 10 min of incubation at 37 °C, the solution with released bacteria cells was serially diluted and plated on MRS agar. The plates were incubated at 37 °C for 48 h. Adhesion ability was expressed as the percentage ratio between the initial counts of bacteria seeded and the counts after the washing steps (CFU/mL). Experiments were performed in duplicate and repeated three times. The probiotic strain L. paracasei LBC-81 (Danisco A/S, Copenhagen, Denmark) was employed as a reference strain.
Inhibition of Pathogenic Bacteria Adhesion to Caco-2 and HT-29 Cells
Cell cultures were maintained as previously described. For the pathogen adhesion inhibition test, two different types of experiments were performed using modifications to previously described procedures [33]. In the competition assay, lactobacilli suspensions (108 CFU/mL) and Salmonella or E. coli (108 CFU/mL) were mixed and co-cultured simultaneously for 90 min with Caco-2 and HT-29 monolayers. In the exclusion assay, Caco-2 and HT-29 cells were first preincubated with lactobacilli suspensions (108 CFU/mL) for 30 min and then a Salmonella or E. coli suspension (108 CFU/mL) was added to each well. Cell cultures in the presence of bacteria were incubated for an additional 90 min at 37 °C in a 5% CO2 atmosphere. Afterward, the cells were washed three times with 1 mL of PBS to remove non-adherent bacteria cells and lysed with 1 mL of Triton-X solution (0.1% v/v in PBS). After 10 min of incubation at 37 °C, the solution of released bacteria cells was spread on BHI agar and the plates were incubated at 37 °C for 16 h. After this time, enumerations of pathogen colonies were performed. Experiments were performed in duplicate and repeated three times.
Statistical Analysis
Data were analyzed by one-way analysis of variance, followed by post hoc Tukey’s and Dunnett’s tests for multiple comparisons. Differences were considered statistically significant when p < 0.05. All statistical analysis was carried out using Statistica software version 10.0 (Statsoft, USA).
Results and Discussion
Survival in Simulated Gastric and Intestinal Juices and Cell Surface Properties
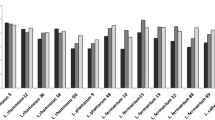
Eighteen isolates had survival rates ≥ 81% after exposure to simulated gastric and intestinal juices, except L. casei CCMA 0411 that showed a survival rate of 79.42% (Table 2). The LAB strains evaluated in the present study exhibited variable hydrophobicity values ranging from 6.67 to 96.06% (Table 2). On the other hand, approximately 68% (13) of the strains had low hydrophobicity (< 35.0%). According to Kaktcham et al. [27], the composition of the bacterial membrane influences the hydrophobicity of the cell surface and, therefore, hydrophobicity evaluation is important in estimating the ability of strains to adhere to host epithelial cells. Regarding auto-aggregation capacity, the strains had values ranging from 16.50 to 52.66% after 5 h of incubation. The highest values were found for L. paracasei strains (CCMA 0504 and CCMA 0505) which exhibited auto-aggregation in the range of 41.00–60.00% (Table 2). This is important because auto-aggregation allows the formation of a barrier that prevents the colonization of pathogens on surfaces of the mucosa [34].
Based on the results obtained for survival in simulated gastric and intestinal juices as well as assessments of hydrophobicity and auto-aggregation, five LAB strains were selected (Tables 3 and 4). The selected strains showed high percentages (90.06–96.50%) of survival in simulated gastric and intestinal juices, indicating that they were able to tolerate stressful conditions imposed by GIT (Table 3).
Researchers have suggested a correlation between hydrophobicity and aggregation capacity [35, 36]. All of the five selected strains, which exhibited hydrophobicity, also displayed auto-aggregation capacities after 5 h of incubation (Table 4). All selected strains were able to co-aggregate with EPEC and S. enteritidis, except L. paracasei CCMA 0504, which did not co-aggregate with S. enteritidis (Table 4). The co-aggregation abilities of Lactobacillus strains can prevent intestinal colonization by pathogenic bacteria and represent an important host defense mechanism [37, 38].
Phenol Tolerance
The effect of the two different phenol concentrations (0.2% and 0.5%) [39, 40] evaluated on the growth of the five selected LAB is shown in Fig. 1. There were differences in the sensitivities of the strains for the different evaluated phenol concentrations. As expected, the strains were more tolerant to 0.2% phenol than 0.5%. In 0.2% phenol, L. paracasei CCMA 0504 was the most tolerant (58.93%), followed by L. plantarum CCMA 0743 (53.11%). In 0.5% phenol, all the evaluated strains had less than 5% relative growth. Divisekera et al. [41] evaluated three Lactobacillus spp. and reported that they were not able to tolerate 0.5% phenol.
Phenols are compounds formed after bacterial degradation of aromatic amino acids and have been shown to exert toxic effects. Their presence can be affected by many factors such as diet, endogenous proteins, and gut microbiota composition [42, 43]. Phenols inhibit various species of bacteria and may, therefore, affect the diversity and metabolic activity of the intestinal microbiota, mainly by the formation of more potent inhibitory compounds, such as phenolic acids, resulting from microbial transformations of flavonols, flavan-3-ols, flavones, and anthocyanins obtained from the diet [44]. Furthermore, most polyphenols follow through the colon, where they maybe converted by many intestinal bacteria in short-chain fatty acids that can modulate intestinal microbiota composition, increasing Lactobacillus genus and other beneficial bacteria populations [45]. Taken together, these results suggest that phenolic compounds resulting from deamination of aromatic amino acids by the intestinal microbiota have bacteriostatic effects against some probiotic strains [9]. Thus, phenol tolerance is an interesting issue for probiotic strains characterization.
Antimicrobial Activity
Regarding antimicrobial activity, L. plantarum CCMA 0743 and L. paracasei (CCMA 0504 and CCMA 0505) demonstrated inhibitory activity against all pathogens evaluated (Table 5). The two L. paracasei strains and L. plantarum CCMA 0743 showed the highest activity against S. aureus and B. cereus, respectively. Conversely, L. brevis CCMA 1284 and L. plantarum CCMA 0359 showed no inhibitory activity for the evaluated pathogenic microorganisms. Lactobacillus spp. have been identified with different antibacterial activities against a range of human pathogens [46].
Antibiotic Susceptibility
The evaluated strains were sensitive to at least one antibiotic from the cell wall synthesis inhibitor and protein synthesis inhibitor classes. All strains were sensitive to ampicillin and chloramphenicol, resistant to vancomycin, streptomycin, and gentamicin and showed some sensitivity to erythromycin (Table 6). Only the two L. paracasei strains were moderately susceptible or susceptible to lincomycin, azithromycin, and penicillin, while the L. brevis CCMA 1284 strain was resistant to these three antibiotics. Several Lactobacillus spp. have been reported to be vancomycin-resistant [26, 31], which was also observed in the present study. Antibiotic resistance may become a risk if associated with gene transfer [47]. However, in most cases, it is not a cause for concern as it maybe not of the transmissible type, nor is it a specific characteristic of the microbial genus or species [48]. Therefore, these resistance mechanisms may be intrinsic to the strain as demonstrated by Handwerger et al. [49], who reported on vancomycin-resistant Lactobacillus spp. These authors suggested that the antibiotic-resistant strains, which are not associated with gene transfer, are interesting candidates for concomitant therapy or after antibiotic use, thereby decreasing the adverse effects of these drugs. However, this parameter was not evaluated in the present study and should be considered for further characterization of selected candidates. Finally, all strains showed some degree of susceptibility to novobiocin (a gyrase inhibitor), while only the two L. paracasei strains were susceptible to 1 μg oxacillin (an inhibitor of cell wall synthesis); both antibiotics had no proposed scores.
In Vitro Assessment of Safety Attributes
The determination of safety characteristics is one of the criteria for selecting novel probiotic strains [50]. All five isolates did not show hemolytic and gelatinase activities when compared to the positive control strain, S. aureus ATCC 25923. Regarding DNAse activity, L. paracasei CCMA 0504 and CCMA 0505 strains were positive. Previous studies have reported the presence of extracellular DNAse in L. plantarum [51] and that the secretion of this enzyme may be found in milk-related Lactobacillus [52]. In the present study, the positive DNAse strains were isolated from kefir. Although the production of these enzymes has been considered as a virulence factor [53], nucleases secreted by Lactobacillus spp. have demonstrated activity against Gram-negative bacteria and bacteriophages and may be associated with nutritional functions [52].
Adhesion of LAB Strains to Caco-2 and HT-29 Cell Lines
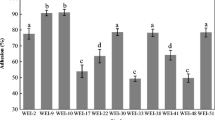
The adhesion capacities of beneficial bacteria and pathogens may be affected by the in vitro cell line used for evaluation as well as the mechanisms of the strain interacting with superficial components of intestinal cells; this is mainly related to the production (or not) of mucus [13]. The present study evaluated LAB adhesion to two different cell lines, Caco-2 and HT-29. The percentage adhesion of the strains did not differ (p > 0.05) from the positive control strain L. paracasei LBC-81 on HT-29 cells. On the other hand, L. paracasei CCMA 0505 showed higher (p < 0.001) adhesion capacity (4.75%) than the positive control strain (0.85%) on Caco-2 cells (Fig. 2). Adhesion capacity is influenced by cell surface components and by specific adhesive proteins expressed on this surface that can confer varying degrees of adhesive properties [54].
Adhesion capacity of LAB strains to Caco-2 and HT-29 cells. The adhesion capacity is calculated using the ratio of the number of bacterial cells that remained attached to the total number of bacterial cells initially added to each well. Asterisks indicate statistically significant differences: *p < 0.05; **p < 0.01; ***p < 0.001 compared to the control, using Dunnett’s test. The results are expressed as the mean ± SEM of three independent assays
Inhibition of Pathogenic Bacteria Adhesion to Caco-2 and HT-29 Cells
The inhibition of Escherichia coli (EPEC) CDC 055 and Salmonella enterica serovar Enteritidis ATCC 564 adhesion by the five selected LAB strains was evaluated by competition and exclusion assays. A significant reduction (0.7 to 1.7 Log CFU/mL) in E. coli adhesion to Caco-2 cells by the competition test was observed in the presence of all evaluated strains except CCMA 1284, while in the exclusion test, CCMA 1284 and CCMA 0743 were not able to reduce pathogen counts (p ≥ 0.05) (Fig. 3a). On the other hand, for the assays using HT-29 cells, only CCMA 0743 and CCMA 0505 were able to significantly (p < 0.01) reduce E. coli adhesion (Fig. 3b).
Effect of Lactobacillus strains on the adhesion of enteropathogens to intestinal cell lines. Caco-2 (a, c) and HT-29 (b, d) cells were incubated with E. coli and S. enteritidis alone, or in the presence of Lactobacillus strains. Asterisks indicate significant differences: *p < 0.05; **p < 0.01; ***p < 0.001, compared to the control, according to Dunnett’s test. The results are expressed as the mean ± SEM of three independent assays
The adhesion of S. enteritidis to Caco-2 cells was significantly (p < 0.05) reduced by competition with CCMA 1284, CCMA 0743, and CCMA 0359 strains, but in the exclusion assay, only the CCMA 0743 strain reduced (p < 0.001) pathogen counts (approximately 2.5 Log CFU/mL) (Fig. 3c). All the five evaluated LAB strains were able to inhibit (p < 0.05) the adhesion of S. enteritidis to HT-29 cells in both exclusion and competition assays (Fig. 3d).
Studies have reported that Lactobacillus strains may inhibit pathogen adhesion by preventing their colonization through competitive exclusion, a highly specific mechanism that is strain-dependent for both probiotics and pathogens [18]. However, adhesion inhibition may be related to different mechanisms such as antimicrobial substances produced by LAB, competition for eukaryotic cell receptors and substrates, the intestinal-mucosal barrier, immunomodulation, and co-aggregation [33]. In the present study, there was a reduction of approximately 1 Log CFU/mL of S. enteritidis to HT-29 cells in the presence of L. plantarum CCMA 0743 by the exclusion assay.
According to Gagnon et al. [13], Salmonella has higher adhesion and invasion capacities in mucus-producing intestinal cell models (HT-29-MTX) than in non-mucus-producing cells (Caco-2). The adhesion of S. enteritidis to the Caco-2 and HT-29 cells observed in this study corroborate the results of these authors, as we observed adherent bacteria counts of 5.4 Log CFU/mL and 7.3 Log CFU/mL in non-mucus-producing (Caco-2) and low mucus-producing (HT-29) models, respectively (Fig. 3c and d). In contrast, EPEC adhered more strongly to Caco-2 cells (6.8 Log CFU/mL) than HT-29 cells (Fig. 3a). There is evidence that the ability of pathogenic bacteria to colonize and invade cells of different mucosal surfaces is directly related to the expression of specific proteins, pili, fimbriae, and flagella [55].
Conclusion
Of the 19 strains that were able to tolerate and survive in simulated gastric and intestinal juices, only five expressed remarkable cell surface characteristics (hydrophobicity and auto-aggregation). These strains were able to reduce the colonization and invasion of EPEC and S. enteritidis in human epithelial cells (Caco-2 and HT-29), with L. plantarum CCMA 0743, L. paracasei CCMA 0504, and L. paracasei CCMA 0505 also exhibiting antimicrobial activity towards pathogenic bacteria. However, special attention should be given to L. paracasei strains due to their ability to secrete DNAse, whose properties and mechanisms of action must be elucidated.
Among these strains, L. plantarum CCMA 0743, which is isolated from cauim, an indigenous fermented beverage for infants, exhibited interesting probiotic properties, making it the most promising candidate. However, in vivo evaluation of these probiotic effects is required to confirm these findings. Moreover, the analysis of the potential of this probiotic strain for biotechnological development is still needed before its therapeutic applications can be defined.
References
Argyri AA, Zoumpopoulou G, Karatzas KAG, Tsakalidou E, Nychas GJE, Panagou EZ, Tassou CC (2013) Selection of potential probiotic lactic acid bacteria from fermented olives by in vitro tests. Food Microbiol 33:282–291. https://doi.org/10.1016/j.fm.2012.10.005
Ayeni FA, Sánchez B, Adeniyi BA, de los Reyes-Gavilán CG, Margolles A, Ruas-Madiedo P (2011) Evaluation of the functional potential of Weissella and Lactobacillus isolates obtained from Nigerian traditional fermented foods and cow’s intestine. Int J Food Microbiol 147:97–104. https://doi.org/10.1016/j.ijfoodmicro.2011.03.014
Sanders ME, Merenstein DJ, Reid G, Gibson GR, Rastall RA (2019) Probiotics and prebiotics in intestinal health and disease: from biology to the clinic. Nat Rev Gastroenterol Hepatol 16:605–616. https://doi.org/10.1038/s41575-019-0173-3
de Almeida Júnior WLG, da Silva FÍ, de Souza JV et al (2015) Characterization and evaluation of lactic acid bacteria isolated from goat milk. Food Control 53:96–103. https://doi.org/10.1016/j.foodcont.2015.01.013
Dias FS, Santos MRRM, Schwan RF (2015) Enumeration, identification and safety proprieties of lactic acid bacteria isolated from pork sausage. Arq Bras Med Vet e Zootec 67:918–926. https://doi.org/10.1590/1678-4162-8119
Almeida EG, Rachid CCTC, Schwan RF (2007) Microbial population present in fermented beverage “cauim” produced by Brazilian Amerindians. Int J Food Microbiol 120:146–151. https://doi.org/10.1016/j.ijfoodmicro.2007.06.020
de Melo PGV, da Cruz Pedrozo Miguel MG, Ramos CL, Schwan RF (2012) Microbiological and physicochemical characterization of small-scale cocoa fermentations and screening of yeast and bacterial strains to develop a defined starter culture. Appl Environ Microbiol 78:5395–5405. https://doi.org/10.1128/AEM.01144-12
Ramos CL, Thorsen L, Schwan RF, Jespersen L (2013) Strain-specific probiotics properties of Lactobacillus fermentum, Lactobacillus plantarum and Lactobacillus brevis isolates from Brazilian food products. Food Microbiol 36:22–29. https://doi.org/10.1016/j.fm.2013.03.010
Choudhary J, Dubey RC, Sengar G, Dheeman S (2019) Evaluation of probiotic potential and safety assessment of Lactobacillus pentosus MMP4 isolated from mare’s lactation. Probiotics Antimicrob Proteins 11:403–412. https://doi.org/10.1007/s12602-018-9431-x
Gharbi Y, Fhoula I, Ruas-Madiedo P, Afef N, Boudabous A, Gueimonde M, Ouzari HI (2019) In-vitro characterization of potentially probiotic Lactobacillus strains isolated from human microbiota: interaction with pathogenic bacteria and the enteric cell line HT29. Ann Microbiol 69:61–72. https://doi.org/10.1007/s13213-018-1396-1
Wang J, Wang J, Yang K, Liu M, Zhang J, Wei X, Fan M (2018) Screening for potential probiotic from spontaneously fermented non-dairy foods based on in vitro probiotic and safety properties. Ann Microbiol 68:803–813. https://doi.org/10.1007/s13213-018-1386-3
Falah F, Vasiee A, Behbahani BA, Yazdi FT, Moradi S, Mortazavi SA, Roshanak S (2019) Evaluation of adherence and anti-infective properties of probiotic Lactobacillus fermentum strain 4-17 against Escherichia coli causing urinary tract infection in humans. Microb Pathog 131:246–253. https://doi.org/10.1016/j.micpath.2019.04.006
Gagnon M, Zihler Berner A, Chervet N, Chassard C, Lacroix C (2013) Comparison of the Caco-2, HT-29 and the mucus-secreting HT29-MTX intestinal cell models to investigate Salmonella adhesion and invasion. J Microbiol Methods 94:274–279. https://doi.org/10.1016/j.mimet.2013.06.027
Bardiau M, Szalo M, Mainil JG (2010) Initial adherence of EPEC, EHEC and VTEC to host cells. Vet Res 41:57. https://doi.org/10.1051/vetres/2010029
Rehman T, Yin L, Latif MB, Chen J, Wang K, Geng Y, Huang X, Abaidullah M, Guo H, Ouyang P (2019) Adhesive mechanism of different Salmonella fimbrial adhesins. Microb Pathog 137:103748. https://doi.org/10.1016/j.micpath.2019.103748
Kadwalia A, Thakur P, Vivekanandhan R et al (2019) Antimicrobial resistant : a glance on emergence , spread and combat. J Pharmacogn Phytochem 8:995–998
Khan I, Kang SC (2016) Probiotic potential of nutritionally improved Lactobacillus plantarum DGK-17 isolated from Kimchi - a traditional Korean fermented food. Food Control 60:88–94. https://doi.org/10.1016/j.foodcont.2015.07.010
Xu H, Jeong HS, Lee HY, Ahn J (2009) Assessment of cell surface properties and adhesion potential of selected probiotic strains. Lett Appl Microbiol 49:434–442. https://doi.org/10.1111/j.1472-765X.2009.02684.x
Viana RO, Magalhães-Guedes KT, Braga RA et al (2017) Fermentation process for production of apple-based kefir vinegar: microbiological, chemical and sensory analysis. Braz J Microbiol 48:592–601. https://doi.org/10.1016/j.bjm.2016.11.006
da Silva FI, de Souza JV, Ramos CL et al (2016) Selection of autochthonous lactic acid bacteria from goat dairies and their addition to evaluate the inhibition of Salmonella typhi in artisanal cheese. Food Microbiol 60:29–38. https://doi.org/10.1016/j.fm.2016.06.014
Ramos CL, de Almeida EG, de Melo PGV et al (2010) Determination of dynamic characteristics of microbiota in a fermented beverage produced by Brazilian Amerindians using culture-dependent and culture-independent methods. Int J Food Microbiol 140:225–231. https://doi.org/10.1016/j.ijfoodmicro.2010.03.029
Ramos CL, de Almeida EG, Freire AL, Freitas Schwan R (2011) Diversity of bacteria and yeast in the naturally fermented cotton seed and rice beverage produced by Brazilian Amerindians. Food Microbiol 28:1380–1386. https://doi.org/10.1016/j.fm.2011.06.012
Puerari C, Magalhães-Guedes KT, Schwan RF (2015) Physicochemical and microbiological characterization of chicha, a rice-based fermented beverage produced by Umutina Brazilian Amerindians. Food Microbiol 46:210–217. https://doi.org/10.1016/j.fm.2014.08.009
de Oliveira dos Santos A, da Silva ÁCL, Soares C et al (2019) Lactic acid bacteria diversity in corn silage produced in Minas Gerais (Brazil). Ann Microbiol 69:1445–1459. https://doi.org/10.1007/s13213-019-01528-w
Freire AL, Ramos CL, de Almeida EG, Duarte WF, Schwan RF (2014) Study of the physicochemical parameters and spontaneous fermentation during the traditional production of yakupa, an indigenous beverage produced by Brazilian Amerindians. World J Microbiol Biotechnol 30:567–577. https://doi.org/10.1007/s11274-013-1476-0
Saini K, Tomar SK (2017) In vitro evaluation of probiotic potential of Lactobacillus cultures of human origin capable of selenium bioaccumulation. LWT - Food Sci Technol 84:497–504. https://doi.org/10.1016/j.lwt.2017.05.034
Kaktcham PM, Temgoua JB, Zambou FN, Diaz-Ruiz G, Wacher C, Pérez-Chabela ML (2018) In vitro evaluation of the probiotic and safety properties of bacteriocinogenic and non-bacteriocinogenic lactic acid bacteria from the intestines of Nile tilapia and common carp for their use as probiotics in aquaculture. Probiotics Antimicrob Proteins 10:98–109. https://doi.org/10.1007/s12602-017-9312-8
Colloca M, Ahumada M, López M, Nader-Macías M (2000) Surface properties of lactobacilli isolated from healthy subjects. Oral Dis 6:227–233. https://doi.org/10.1076/noph.22.3.195.3726
Shehata MG, El Sohaimy SA, El-Sahn MA, Youssef MM (2016) Screening of isolated potential probiotic lactic acid bacteria for cholesterol lowering property and bile salt hydrolase activity. Ann Agric Sci 61:65–75. https://doi.org/10.1016/j.aoas.2016.03.001
Arena MP, Silvain A, Normanno G, Grieco F, Drider D, Spano G, Fiocco D (2016) Use of Lactobacillus plantarum strains as a bio-control strategy against food-borne pathogenic microorganisms. Front Microbiol 7:1–10. https://doi.org/10.3389/fmicb.2016.00464
Charteris WP, Kelly PM, Morelli L, Collins JK (1998) Antibiotic susceptibility of potentially probiotic Lactobacillus species. J Food Prot 61:1636–1643
Singh TP, Kaur G, Malik RK, Schillinger U, Guigas C, Kapila S (2012) Characterization of intestinal Lactobacillus reuteri strains as potential probiotics. Probiotics Antimicrob Proteins 4:47–58. https://doi.org/10.1007/s12602-012-9090-2
Yu Q, Wang Z, Yang Q (2011) Ability of Lactobacillus to inhibit enteric pathogenic bacteria adhesion on Caco-2 cells. World J Microbiol Biotechnol 27:881–886. https://doi.org/10.1007/s11274-010-0530-4
Pithva S, Shekh S, Dave J, Vyas BRM (2014) Probiotic attributes of autochthonous Lactobacillus rhamnosus strains of human origin. Appl Biochem Biotechnol 173:259–277. https://doi.org/10.1007/s12010-014-0839-9
Maldonado NC, Ficoseco CA, Mansilla FI, Melián C, Hébert EM, Vignolo GM, Nader-Macías MEF (2018) Identification, characterization and selection of autochthonous lactic acid bacteria as probiotic for feedlot cattle. Livest Sci 212:99–110. https://doi.org/10.1016/j.livsci.2018.04.003
Hector Momo Kenfack C, Zambou Ngoufack F, Marie Kaktcham P et al (2018) Safety and antioxidant properties of five probiotic Lactobacillus plantarum strains isolated from the digestive tract of honey bees. Am J Microbiol Res 6:1–8. https://doi.org/10.12691/ajmr-6-1-1
Ekmekci H, Aslim B, Ozturk S (2009) Characterization of vaginal lactobacilli coaggregation ability with Escherichia coli. Microbiol Immunol 53:59–65. https://doi.org/10.1111/j.1348-0421.2009.00115.x
Silva Dias F, Ferreira Duarte W, Freitas Schwan R (2013) Evaluation of adhesive properties of presumptive probiotic Lactobacillus plantarum strains. Biosci J 29:1678–1686
Mohd Yusof H, Mohamad R, Zaidan UH, Rahman NA (2020) Sustainable microbial cell nanofactory for zinc oxide nanoparticles production by zinc-tolerant probiotic Lactobacillus plantarum strain TA4. Microb Cell Factories 19:1–17. https://doi.org/10.1186/s12934-020-1279-6
Padmavathi T, Bhargavi R, Priyanka PR, Niranjan NR, Pavitra PV (2018) Screening of potential probiotic lactic acid bacteria and production of amylase and its partial purification. J Genet Eng Biotechnol 16:357–362. https://doi.org/10.1016/j.jgeb.2018.03.005
Divisekera DMWD, Samarasekera JKRR, Hettiarachchi C et al (2019) Lactic acid bacteria isolated from fermented flour of finger millet, its probiotic attributes and bioactive properties. Ann Microbiol 69:79–92. https://doi.org/10.1007/s13213-018-1399-y
Smith EA, Macfarlane GT (1997) Formation of phenolic and indolic compounds by anaerobic bacteria in the human large intestine. Microb Ecol 33:180–188. https://doi.org/10.1007/s002489900020
Hughes R, Magee EAM, Bingham S (2000) Protein degradation in large intestine. Curr Issues Intest Microbiol 1:51–58
Cueva C, Moreno-Arribas MV, Martín-Álvarez PJ, Bills G, Vicente MF, Basilio A, Rivas CL, Requena T, Rodríguez JM, Bartolomé B (2010) Antimicrobial activity of phenolic acids against commensal, probiotic and pathogenic bacteria. Res Microbiol 161:372–382. https://doi.org/10.1016/j.resmic.2010.04.006
De Oliveira VPS, Dolinsky M, Barroso SG et al (2017) Dark polyphenols-rich chocolate and gut microbiota: a literature review. DEMETRA Aliment Nutr Saúde 12:399–410. https://doi.org/10.12957/demetra.2017.25475
Halder D, Mandal M, Chatterjee S, Pal N, Mandal S (2017) Indigenous probiotic Lactobacillus isolates presenting antibiotic like activity against human pathogenic bacteria. Biomedicines 5:31. https://doi.org/10.3390/biomedicines5020031
Gueimonde M, Sánchez B, de los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:1–6. https://doi.org/10.3389/fmicb.2013.00202
Salminen S, von Wright A, Morelli L, Marteau P, Brassart D, de Vos WM, Fondén R, Saxelin M, Collins K, Mogensen G, Birkeland SE, Mattila-Sandholm T (1998) Demonstration of safety of probiotics — a review. Int J Food Microbiol 44:93–106
Handwerger S, Pucci MJ, Volk KJ, Liu J, Lee MS (1994) Vancomycin-resistant Leuconostoc mesenteroides and Lactobacillus. J Bacteriol 176:260–264
FAO, WHO (2002) Guidelines for the evaluation of probiotics in food - report of a joint FAO/WHO working group on drafting guidelines for the evaluation of probiotics in food. London Ontario, Canada
Caso JL, Suárez JE (1997) Detection and analysis of extra- and intracellular deoxyribonuclease activities in Lactobacillus plantarum. Lett Appl Microbiol 25:148–152. https://doi.org/10.1046/j.1472-765X.1997.00192.x
Péant B, LaPointe G (2004) Identification and characterization of a conserved nuclease secreted by strains of the Lactobacillus casei group. J Appl Microbiol 96:367–374. https://doi.org/10.1046/j.1365-2672.2004.02160.x
Hasegawa T, Minami M, Okamoto A, Tatsuno I, Isaka M, Ohta M (2010) Characterization of a virulence-associated and cell-wall-located DNase of Streptococcus pyogenes. Microbiology 156:184–190. https://doi.org/10.1099/mic.0.031955-0
Wang G, Zhang M, Zhao J, Xia Y, Lai PFH, Ai L (2018) A surface protein from Lactobacillus plantarum increases the adhesion of Lactobacillus strains to human epithelial cells. Front Microbiol 9:1–9. https://doi.org/10.3389/fmicb.2018.02858
Ye J, Song L, Liu Y, Pan Q, Zhong X, Li S, Shang Y, Tian Y, He Y, Chen L, Chen W, Peng Z, Wang R (2015) Core 2 Mucin-type O-glycan is related to EPEC and EHEC O157:H7 adherence to human colon carcinoma HT-29 epithelial cells. Dig Dis Sci 60:1977–1990. https://doi.org/10.1007/s10620-015-3548-5
Acknowledgments
The authors thank the CCMA for providing the strains used in this work.
Funding
This work was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico do Brasil (CNPq), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG), and CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior).
Author information
Authors and Affiliations
Contributions
H.C.F. and D.S.M. performed the experiments; H.C.F. wrote the original draft preparation; D.S.M., C.L.R., D.R.D., R.F.S. supervised the study; C.L.R., D.R.D., R.F.S.; revised and edited the final manuscript; R.F.S. managed the project and acquired financing.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Research Involving Human Participants and/or Animals
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fonseca, H.C., de Sousa Melo, D., Ramos, C.L. et al. Probiotic Properties of Lactobacilli and Their Ability to Inhibit the Adhesion of Enteropathogenic Bacteria to Caco-2 and HT-29 Cells. Probiotics & Antimicro. Prot. 13, 102–112 (2021). https://doi.org/10.1007/s12602-020-09659-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-020-09659-2