Abstract
In this study, the probiotic potential of five bacteriocin-producing lactic acid bacteria (LAB) strains, isolated from meat products, was investigated. They were presumptively identified as Lactococcus lactis subsp. cremoris CTC 204 and CTC 483, L. lactis subsp. hordinae CTC 484, and Lactobacillus plantarum CTC 368 and CTC 469 according to morphological, biochemical, and physiological characteristics. Analysis of genetic variability (random amplified polymorphic (RAPD)-PCR) and whole-cell proteins (SDS-PAGE) revealed similarity between Lactobacillus strains and variability among Lactococcus strains. The evaluation of the probiotic potential showed that the five LAB strains were tolerant to pH 2.0, and only strain CTC 469 was tolerant to the lowest concentration of the bile salts evaluated (0.1%). All strains showed survival or growth ability at 4, 25, and 37 °C, and tolerance at − 20 °C. Although strain CTC 204 in TSB Broth supplemented with MgSO4 showed the highest intensity of biofilm production, this compound was produced by all of them. The safety assessment showed that no thermonuclease, hemolytic, or gelatinase activities were detected. All strains were resistant to erythromycin and sensitive to amoxicillin and phenoxymethylpenicillin; furthermore, CTC 204 was resistant to chloramphenicol, CTC 368 and CTC 469 to chloramphenicol and vancomycin, CTC 483 to tetracycline and vancomycin, and CTC 484 to clindamycin and chloramphenicol. The evaluated strains showed biogenic amine production; the lowest levels were produced by CTC 204 and CTC 368 strains. It was concluded that CTC 204 and CTC 368 strains have the greatest potential for becoming probiotics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the last two decades, interest in functional foods has increased exponentially, especially for those with probiotic properties. Probiotics are defined as live microorganisms, which, when administered in adequate amounts, confer a health benefit on the host [1, 2]. Probiotic research suggests a range of potential health benefits to the host organism, including evidence that supports potential clinical applications of probiotics for prevention and treatment of gastrointestinal (GI) and urogenital tracts as well as respiratory diseases, by stimulation of immune response or anti-mutagenic and anti-carcinogenic activities [3].
Some essential attributes for the use of probiotics in foods are that they should be safe and must contain the appropriate probiotic microorganisms in sufficient numbers at the time of consumption. Hence, the probiotic strains selected should be proper for application in food processing operations, considering their abilities to survive and keep their functional properties during production and storage under deleterious conditions, such as freezing or drying processes along with surviving in human intestinal tract.
Lactic acid bacteria (LAB) are important for food industry, because this group plays a crucial role in fermented food products, and they are associated with mucosal surfaces, such as with the GI tract as well as the oral and vaginal cavities of humans and other mammals. They have received substantial attention regarding their potential health-promoting properties, so some of them have shared the status as Generally Accepted as Safe (GRAS) or Qualified Presumption Safety (QPS). Lactobacillus, Enterococcus, and Streptococcus, and other genera such as Bifidobacterium, Propionibacterium, Bacillus, Escherichia, and Saccharomyces have been applied as probiotics [4].
The human origin is one of the criteria for selection of probiotics intended for human consumption [5]; however, many probiotic products contain bacteria isolated from food products [6, 7]. Actually, it is a complex task to confirm the origin of LAB due to their wide variety in different environments. They can be isolated from food products or possibly be inhabitants of human intestinal tract. The LAB of the human GI tract consists of both autochthonous (true residents) and allochthonous (transient) species [4]. So, only the autochthonous species are able to occupy and to colonize a niche on the mucosal surface due to the presence of specific adhesion factors. Probiotics are considered part of the transient microbiota, and their presence is for a limited time [4, 8].
Use of probiotic cultures for the production of fermented meat products has attracted attention in recent years. The most promising bacteria for use as starter cultures are those which are isolated from the indigenous microbiota of traditional products. In the case of bacteria isolated from meat products, these microorganisms have evolved to become well adapted to the meat environment and are capable of dominating the microbiota of such products. The strains selected as starter or protective cultures must possess the most important technologically compliant properties and/or bacteriocin production [9].
Fermented meat products have been shown to be excellent vehicles for probiotic delivery, since little or no heat treatment is employed during the manufacture of these products, thus providing suitable conditions required to enable survival of the probiotics in question. The well-adapted microbiota of traditional fermented meat is considered a valuable competitor against indigenous contaminant bacteria [10]. Thus, bacteriocinogenic LAB, applied as bioprotective cultures, should be used as an additional hurdle to Good Manufacturing Practices. The huge amount of information produced by genetic and genomic analyses provides powerful tools in order to select new cultures with specific physiological characteristics.
The main objective of this work was to evaluate the probiotic properties of five LAB strains (CTC 204, CTC 368, CTC 469, CTC 483, and CTC 484). These strains were previously selected from a total of 813 bacteriocin-producing LAB isolated meat products [11]. The criteria subsequently used for the selection of these strains were their growth properties and bacteriocin production with activity against pathogenic and spoilage bacteria of importance in food, bacteriocin stability at a wide pH range, and under low and high temperatures. In this study, the strains were evaluated according to the following characteristics: morphological, biochemical, physiological, and molecular; tolerance to gastric acidity and bile salts; ability to survival or growth at different temperatures; biofilms production; presence of virulence factors; resistance to antibiotics and biogenic amines production.
Materials and Methods
Strains, Media, and Culture Conditions
All five strains of LAB, previously isolated from different samples of meat products [11], and other strains of different genera used in this study are listed in Table 1. Stock cultures were maintained at − 80 °C in de Man Rogosa Sharpe Broth (MRS, Oxoid Ltda., UK) for lactic acid bacteria and Trypticase Soya Broth (TSB, Oxoid) supplemented with 0.6% (w/v) yeast extract (Oxoid) for other microorganisms, both media contained 15% (w/v) glycerol (Sinth, Brazil). Before use, the stock strains were cultured for three consecutive times (1%, v/v) under conditions described in Table 1.
Morphological, Biochemical, and Physiological Characteristics
Cell morphology, Gram staining, and catalase production were verified by using an optical microscope (Zeiss, Germany). Fermentative metabolism of glucose was evaluated according to Leisner et al. [12]. The characteristics of reduction of nitrate, tolerance to NaCl, and ability to grow in extreme temperatures and conditions of acidic and alkaline pH were analyzed as described by Harrigan [13]. Carbohydrate fermentation and complementary tests were determined using BD BBL Crystal™ Identification Systems, Gram-Positive ID Kit (Becton & Dickinson Microbiology Systems, USA) and API® 50 CHL (Biomérieux, France), according to the manufacturers’ instructions. Previous tests were already performed with strains CTC 204 [14] and CTC 484 [15].
RAPD-PCR Profile Analysis
Differentiation of the LAB strains was performed by random amplified polymorphic DNA (RAPD)-PCR analysis. The genomic DNA of each strain was extracted and purified using GFX Kit (Genomic Blood DNA Purification Kit GE, USA)—Gram-positive bacteria—and used according to the manufacturers’ instructions. The pellets were collected by centrifugation (Sorvall RC14, USA) from 4 mL overnight culture samples in MRS Broth at 37 °C (Table 1). The cells were washed in 1 mL Tris-HCl buffer 10 mM, EDTA 1 mM, NaCl 100 mM, pH 8.0. The concentration and purity of DNA were assessed by determining the optical densities at 260 and 280 nm (Micronal B582, USA), the concentration of each DNA sample was adjusted to approximately 25 ng μL−1. Fifty microliters of the supernatants were used as PCR template. RAPD-PCR analysis was performed with random primers REP 2-I: 5′-ACG ACT TAT CAG GCC TAC-3′, REP: 5′-AAA ACG ACG ACA TCA GGC-3′, ERIC 2: 5′-AAG TAA GTG ACT GGG GTG AGC G-3′, and ERIC 1R: 5′-ATG TAA GCT CCT GGG GAT TCA C-3′ (Invitrogen, Brazil) according to Vila et al. [16], Appuhamy et al. [17], and Herman et al. [18]. Each 25 μL PCR reaction contained 200 μM of each deoxyribonucleotide triphosphates (dNTPs, GE), 1 μM of each primer, 2 U of Taq DNA polymerase (GE), and 25 ng of DNA Taq polymerase buffer (GE). The amplification consisted of 29 cycles at 95 °C for 1 min, 47 °C for 40 s, and 72 °C for 2 min (Mastercicle Gradient, Eppendorf, USA). These cycles were preceded by denaturation at 95 °C for 5 min, and followed by extension at 72 °C for 7 min. The amplicons were separated by electrophoresis at 120 V for 4 h on 1.0% (w/v) agarose gel (GE). The DNA was detected by UV light in Image Master VDS (Pharmacia Biotech, USA) after staining with 5 μg mL−1 of ethidium bromide (GE). A 100-pb PCR-DNA Ladder (GE) was used as marker.
Protein Profile Analysis in SDS-PAGE
The differentiation of LAB strains was carried out by protein profile in SDS-PAGE using a methodology adapted from Valence and Lortal [19]. The pellets were collected by centrifugation (6000×g, 4 °C, 30 min) from 10 mL of the activated cells in MRS Broth, washed twice in 0.9% (w/v) NaCl solution, and resuspended in 1 mL of the same solution. Into this suspension, 50 mg of glass beads and 0.1 g of alumina (Sigma-Aldrich, USA) were added and vigorously shaken for 4 min, alternating cycles of 30 s of agitation and 30 s of ice-bath treatment. Lysozyme (Sigma-Aldrich) at final concentration of 2 mg mL−1 was added to the preparations and incubated at 37 °C for 2.5 h. The extracts obtained by centrifugation (13,000×g, 4 °C, 10 min) were diluted in the TDL buffer 50% (w/v), boiled in water bath for 2 min, and centrifuged (13,000×g, 4 °C, 10 min). Aliquots of 20 μL of these extracts were loaded on the gel PAGE 14% (w/v, pH 8.8) in vertical unit (Hoefer Mini VE, Amersham Biosciences, USA) at 180 V for approximately 1 h. The gels were stained in a solution of Coomassie blue R 250 (Sigma-Aldrich) for 1 h under agitation, and after distaining, they were visualized in Image Master VDS. The protein molecular mass marker LMW with molecular weight ranging from 97,000 to 14,400 KDa was used (Amersham Pharmacia Biotech, USA).
Probiotics Properties
Tolerance to Low pH and Presence of Bile Salts
The tolerance of the LAB strains to acid conditions and in presence of bile salts was tested by determination of viability after their exposition in solutions with low pH values: 1.0, 2.0, 3.0, 4.0, and 5.0; and in final concentrations from 0.1 to 2.0% (w/v) Oxgall (Difco™, USA) solutions. Both methodologies were carried out according to Mishra and Prasad [20].
Survival or Growth at Different Temperatures
The ability of the LAB strains to survive or grow at storage temperatures of probiotic foods and human digestive tract was evaluated. Stationary phase cells were inoculated at 1% (v/v, 105−106 cfu mL−1) into buffered MRS Broth and incubated at the following temperatures: − 20, 4, 25, and 37 °C. At appropriate intervals, mass cell determination by absorbance at OD600 nm in UV-visible spectrophotometer (Varian, Cary I E, USA) and pH (Toledo Mettler, MP125, Switzerland) were performed.
Ability for Biofilm Production
The ability for biofilm production was determined using a protocol based on those described by Lebeer et al. [21], Stepanovic et al. [22], and Freitas et al. [23]. Polystyrene tissue culture plates (NalgeNunc. International, USA) were filled with 180 μL of the following broths: MRS or modified TSB (mTSB) enriched with 20 g L−1 Bacto Proteose Peptone n° 3 (Difco, USA), or mTSB enriched with 0.1 g L−1 MgSO4 and 20 μL of overnight cultures. The inoculated plates were covered with a lid and incubated aerobically at 30 °C for 72 h under static conditions. Volumes of 200 μL of media were discarded every 24 h, and the wells were filled with 200 μL of fresh culture media. Staphylococcus aureus ATCC 12600 was used as positive control in all tests, and the corresponded non-inoculated broths were used as negative controls. After 72 h, the culture media were then discarded, and the wells were gently washed three times with 200 μL of sterile ultra-pure water without disturbing the biofilm at the bottom of the wells. Then, the attached cells were fixed by drying in an oven at 50 °C for 1 h and stained with 2% Hucker’s crystal violet for 15 min. Excess stain was aspirated with a pipette, and the plates were rinsed off under running tap water.
Adherent cells were suspended with 200 μL of ethanol absolute, homogenized, transferred to a new culture plate, and the optical density was measured at 620 nm with a microplate reader (Rayto Life and Analytical Sciences Co., Ltd., RT – 2100 C, China). All the strains were tested in 16 replicates, and the average value for each sample was calculated. The interpretation of the results was performed according to Stepanovic et al. [22].
Safety Assessment
Detection of Virulence Factors
LAB strains were subjected to phenotypic tests to identify its virulence activity. Thermonuclease activity was tested in Brain Heart Infusion Broth (BHI, Oxoid) at 50 °C for 2 h, after which they were observed for pink zones at the edge of the wells indicative of the enzyme activity. S. aureus UNICAMP S6 and S. aureus ATCC 12600 were used as positive control and S. epidermidis ATCC 14990 as negative control [24]. Hemolytic activity was verified in agar containing 0.85% sodium chloride and 5% (w/v) defibrinated sheep blood at 37 °C for 48 h. S. aureus CTC 033 and L. lactis subsp. lactis TL CE 016 were used as positive and negative controls, respectively [13]. Gelatinase activity was assessed in nutrient gelatin broth with the addition of 10–15% gelatin (Difco) final pH 7.2, and incubated at 20–25 °C up to 30 days. Bacillus cereus CTC 011 was used as positive control for gelatin hydrolysis [13].
Resistance to Antibiotics
Resistance of LAB strains to antibiotics was performed using the procedure described by Todorov et al. [25]. Amoxicillin, clindamycin, chloramphenicol, erythromycin, phenoxymethylpenicillin, tetracycline, and vancomycin were dissolved in sterile distilled water and filtered in 0.22 μm membrane (Millipore S.A., France), and the appropriate dilutions were prepared in a twofold series in sterile sodium phosphate buffer (10 mM, pH 7.0) using microplates. Then, a 10-μL aliquot of each antimicrobial dilution was deposited on MRS agar plates containing a 4.5-mL overlay of semi-solid MRS (0.7%) inoculated with 0.5 mL inoculum (final concentration of 105 cfu mL−1). The incubation was performed under aerobic conditions at 37 °C for 48 h. The lowest concentration of the antimicrobial that inhibited bacterial growth was defined as minimal inhibitory concentration (MIC). Resistance of the strains was interpreted by following the breakpoints recommended by the EFSA [26].
Production of Biogenic Amines
The LAB strains were cultured according to the conditions described in Table 1. One milliliter of each culture was spread on MRS agar following incubation at the same conditions. The cell mass formed was suspended in 5 mL of 0.85% saline solution, and the inoculums were adjusted to 1 MacFarland using a Densimat (Biomerieux, France). Aliquots of 100 μL of the suspension of cell mass were inoculated in the meat medium culture. The tubes were stirred in a vortex and incubated at 7 °C for 30 days. The meat culture medium was prepared adding to sterile tubes, 1.25 g of ground beef and 10 mL of 0.85% saline. Putrescine, cadaverine, histamine, spermidine, and spermine were assayed. As a negative control, a tube containing only half the meat sample was used, which remained under the same conditions as inoculated tubes.
According to a modified methodology of Malle et al. [27], 5 g of homogenized beef samples were extracted with 10 mL of 0.2 M perchloric acid, following homogenization and centrifugation. A total of 400 μL of the supernatant was collected, and 800 μL of saturated sodium bicarbonate solution were added. A derivatization step was performed with 0.75% dansylchloride in a water bath at 60 °C for 5 min, followed by the addition of 10% L-proline solution, and then the solution was left to stand in the dark at room temperature for 30 min. After the addition of 2 mL of toluene for phase separation, the organic phase was recovered and evaporated. Acetonitrile was added, and the solution was filtered in PTFE membrane, followed by chromatography using HPLC (Shimadzu, Japan) with C18 reverse phase column (5 μm, 100 Å, 25 cm × 4.6 mm), UV detector at 254 nm, and injection of 20 μL. The chromatograms obtained from the samples were compared with chromatograms of standard solutions, and the analyte peaks were confirmed by retention time. The total peak area of each analyte was interpolated on a standard curve relating to the total area with the analyte concentration.
Statistical Analyses
The collected data from tolerance to pH, bile salts, survival or growth at different temperatures, and biofilm production were transformed into log10 values, and they were analyzed by Microsoft Excel 2010. The mean values and the standard deviation were calculated from the data obtained with duplicate trials.
Results
Characterization of the LAB Strains
In this study, isolates CTC 368, CTC 469, and CTC 483 exhibited the following morphological, biochemical, and physiological characteristics of lactic bacteria: positive reactions in Gram staining; morphologies in rod (CTC 368 and CTC 469) and coccus (CTC 483) forms; non-formation of spore or flagella; and inability to produce catalase and to form carbon dioxide from glucose (homofermentative glucose metabolism).
Growth at certain temperatures is mainly used for differentiation of isolated Gram-positive cocci. CTC 483 strains was able to grow at 10 °C, at pH 4.4 and in 6.5% NaCl, but not at 45 °C, 18% NaCl or pH 9.6. The strain CTC 483 also showed growth at pH 4.4. According to the fermentation of carbohydrates and other biochemical reactions obtained on the BBL Crystal and API 50 CHL systems, the strains were presumptively classified as L. lactis subsp. cremoris CTC 483 (85.7%), Lact. plantarum CTC 368 (98.9%), and CTC 469 (98.9%). According to the DNA fingerprints (Fig. 1), CTC 368 and CTC 469 strains presented similarity in the range of 300 up to 1600 pb, and a high degree of differentiation was observed between them and the strains CTC 204, CTC 483, and CTC 484. Similar profiles were observed on the whole-cell proteins in SDS-PAGE (Fig. 2).
RAPD band patterns of the amplicons belonging to LAB strains. a ERIC-RAPD—lane 1, ladder 100 bp; lanes 2 and 8: positive control; lane 3: CTC 204; lane 4: CTC 368; lane 5: CTC 469; lane 6: CTC 483; lane 7: CTC 484; lane 9: negative control. b REP-RAPD—lanes 1 and 2: CTC 204; lanes 3 and 4: CTC 368; lane 5: ladder 100 bp; lanes 6 and 7: CTC 469; lanes 8 and 9: CTC 483; lanes 10 and 11: CTC 484. To analyze the homology between the strains, the range 400–1600 bp was considered
Probiotic and Safety Properties
Tolerance to Low pH and to the Presence of Bile Salts
At pH 2.0, after 3 h of treatment, strains CTC 368 and CTC 469 showed higher survival rates, around 97%, and counts of approximately 8.4 log cfu mL−1 (Table 2). On the other hand, strains CTC 204, CTC 483, and CTC 484 had lower survival rates varying between 65.44 and 74.08% and average counts of 5.4 log cfu mL−1, depending on the strain. The strains exhibited high sensitivity with increasing bile concentration (Table 3). The strain CTC 469 was the only one that survived after 3 h of exposure to 0.1% bile salts. Some strains (CTC 368, CTC 469, and CTC 483) were inhibited by lower concentrations of bile, but were resistant to higher concentrations of 2%.
Survival and Growth Ability at Different Temperatures
All the strains survived at chilling temperature (4 °C) right from the first day of storage (Fig. 3). The freezing temperature caused a slight decreasing in OD of the tested strains. Temperatures of 25 and 37 °C were both adequate conditions for the growth of all strains. A drop in pH values accompanied the growth of the cultures.
Adhesion Ability: Biofilm Production
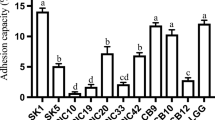
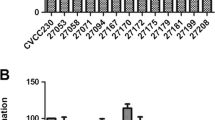
All five strains presented ability for biofilm production in the three media evaluated (Fig. 4); the intensity of biofilms produced varied according to media and strains. The highest capability to produce biofilm was detected with strain CTC 204 in TSB Broth supplemented with MgSO4. This media was also the most effective for strain CTC 484, but did not allow the highest biofilm production among strains CTC 368, CTC 469, and CTC 484.
Detection of Virulence Factors
Among the LAB strains assayed, none showed virulence factors (data not shown); thus, no thermonuclease, hemolytic, or gelatinase activities were detected.
Resistance to Antibiotics
Lactococcus strains did not present the same pattern of sensitivity and resistance against the tested antibiotics, except for the resistance against erythromycin and sensitivity to amoxicillin and phenoxymethylpenicillin; while Lactobacillus strains showed similar pattern of sensitivity (tetracycline, clindamycin, amoxicillin, and phenoxymethylpenicillin) and resistance (chloramphenicol, vancomycin, and erythromycin) (Table 4).
Production of Biogenic Amine
It was verified an increase in the levels of putrescine and cadaverine produced by strains CTC 469, CTC 483, and CTC 484, while the production of these compounds by the others was lower. In relation to spermidine and spermine, a moderate raise of production of all strains was detected after incubation at 7 °C for 30 days (Table 5). Histamine was detected in lower levels. Strains CTC 204 and CTC 368 produced low levels of BA compared to the other LAB evaluated.
Discussion
The LAB strains were presumptively classified by means of their morphological, biochemical, and physiological characteristics: L. lactis subsp. cremoris CTC 204 [14] and CTC 483, L. lactis subsp. hordinae CTC 484 [15], Lact. plantarum CTC 368 and CTC 469. The inability to grow at 45 °C and pH 9.6 were the phenotypic characteristics that allowed the differentiation between the genus Lactococcus from Enterococcus [28].
As the use of enterococci as probiotics is considered controversial, the differentiation between the Lactococcus and Enterococcus strains was focused during the tests. Although the probiotic benefits of some strains of enterococci are established, the recognition of some antibiotic-resistant strains that have emerged and their association with human disease have raised concern regarding their use as probiotics [29]. Growth at pH 9.6 seems to be a feature shared by all species of the genus Enterococcus [30]. In addition, L. lactis spp. CTC 204, CTC 483, and CTC 484 did not present the virulence determinants associated with the pathogenicity of enterococci, such as hemolysin, gelatinase, DNAse, and termonuclease.
The use of the RAPD-PCR in association with SDS-PAGE cell protein analysis was useful to distinguish the diversity of the strains studied in this work, which was also found by Corsetti et al. [31] that classified large adventitious microbial populations using these tools.
Human origin is one of the criteria for the selection of a probiotic strain [32]. However, genomic rearrangements, which occur during the natural evolution of a microorganism, result in ecological differences between closely related species and among populations within a single species [33,33,34,36]. Due to the difficulty for determining the source of a microorganism, the meat products may not be considered the exclusive ecological niche of the strains of this study.
The main benefit attributed to probiotics is the competitive exclusion of pathogens in the GI tract, which may occur by different mechanisms. The competitive exclusion of pathogenic bacteria is usually performed by the production of antimicrobial substances such as lactic acid and bacteriocins, and by the adhesion to the mucosa and co-aggregation, which can form a barrier that prevents the mucosa colonization by pathogenic microorganisms [37, 38]. In addition to the host health benefits, bacteriocins could be applied as natural ingredients to ensure the quality and safety of manufacture products, extending their shelf life. Bacteriocin-producing bacteria could also be included in fermented food as starter culture, or could be added to fresh products as protective cultures [39, 40]. According to this premise, the antimicrobial activity of the bacteriocins produced by the LAB strains evaluated in this study could act as barrier to inhibit food spoilage and/or pathogenic microorganisms of importance in food [11].
At pH 2.0, the lactobacilli presented higher survival rates than the lactococci. All five strains survived at pH values 3.0, 4.0, and 5.0, with no decrease in viability during 3 h of treatment. According to Martini et al. [41], values of pH 3.0 or higher represent the gastric pH after the food intake, mainly due to the buffering capacity of some food, such as dairy products. Huang and Adams [42] demonstrated that the viability of propionibacteria isolated from milk and cheese was affected at pH 2 and that most of the tested strains did well at pH 3.0 and 4.0. Furthermore, survival of the propionibacteria in simulated gastric juices at pH 2.0 was enhanced by the addition of soymilk and liquid cereal breakfast. The addition of milk proteins singly or in combination with starch enhanced the survival of probiotic lactobacilli strains in simulated gastric juice at pH 2.0, 3.0, and 4.0 [43].
The antimicrobial properties of gastric acid and bile salts secreted by the digestive system are the main defense mechanisms of the body against colonization of GI tract by pathogenic microorganisms [44]. As these mechanisms may also result in adverse effects on beneficial microorganisms, several tools have been evaluated to provide protection or increased survival of probiotics that depend on the viability and physiological activity to perform their beneficial effects [45]. Microencapsulation could use techniques and wall materials compatible and suitable for controlled release of the probiotic at specific target sites in the body. These techniques also have the advantage of protecting the probiotic against adverse conditions during processing and storage of food [46,46,48]. The selection of naturally resistant strains to the physiological conditions of the GI tract, as well as obtaining resistant strains derived by the progressive adaptation procedure, showed favorable results for the use of strains sensitive to bile [49].
Besides the chemical impact of the stomach acidic conditions and the reaction of the bile salts in the duodenum, the thermal variation that a probiotic strain is submitted to during the storage of the food until it achieves the human digestive tract after its consumption could be challeging. A probiotic food is exposed to a wide temperature range from the chilled or freezing storage temperature, the mouth at up to 25 °C, and then the stomach and guts at 37–38 °C, where it will be kept for few hours or days [8]. All the strains tested in this work may survive during the storage of food under low temperatures and are able to maintain themselves during their passage in the human digestive tract.
In the present study, the biofilm production was affected by the strains type, pH, and medium composition. Leeber et al. [21] demonstrated the strong influence of conditions related to the GI tract, including low pH, high osmolarity, presence of bile, mucins, and non-digestible polysaccharides, in the formation of biofilm from Lact. rhamnosus GG.
The presence of intrinsic genes resistant to antibiotics is an acceptable characteristic for a probiotic due to the low potential for horizontal transfer of genes to other organisms. Intrinsic resistance is a specific characteristic for a genus or species of bacteria due to the development of mechanisms of resistance that enable the bacteria to survive in the presence of an antimicrobial compound. On the other hand, acquired resistance generally has a low risk of spreading horizontally when the resistance is a result of a chromosomal mutation, but it is considered as having a high potential to spread horizontally, when the resistance genes are present on mobile genetic elements (transposons and plasmids) [26, 32].
The verification of transferable resistance is recommended prior to considering a probiotic strain safe for human consumption [50]. An important step in the differentiation between the intrinsic and acquired resistance is determination and comparison of the susceptibility patterns of a representative number of different strains of each species [51]. According to Donohue and Gueimond [52], the ability of a probiotic to transfer antibiotic resistance to pathogenic bacteria should always be taken into account when assessing their safety.
Many Lactobacillus species have shown a high level of resistance to vancomycin [38], which is considered an intrinsic property [53]. The vancomycin resistance genes of Lactobacillus appear to be chromosomally located and are not easily transferable to other genera [54]. All the tested strains were considered susceptible to the antibiotics of β-lactams group. The breakpoint for amoxicillin and phenoxymethylpenicillin has not yet been determined [26]. In the present study, the breakpoint of ampicillin (MB ≤ 2) was used as reference for the evaluation, because these antibiotics are derivatives of penicillin. All studied LAB were sensitive to amoxicillin and phenoxymethylpenicillin.
Most of the biogenic amines are produced by decarboxylation of the corresponding amino acids through substrate-specific enzymes derived from microorganisms present in the food. Although the toxicity of biogenic amines is beyond all doubt, determination of the exact toxicity threshold of biogenic amines in a given food product is extremely difficult, because their effect does not depend on their presence alone, but is also influenced by other compounds and by the specific efficiency of the detoxifying mechanisms in different individuals [55]. In addition, bio-active amines acting on the central nervous and the vascular systems like histamine, an hypotensive molecule involved in allergies, and tyramine, a hypertensive metabolite sporadically involved in cerebral hemorrhage, can be produced [56,56,58].
The ability of the evaluated LAB strains to form biogenic amines under the test conditions does not eliminate by itself the possibility of their use as probiotic cultures. The use of these strains as dietary supplements (tablets, capsules, powders, lozenges, and gums) instead of food (such as a fermented product) that contain beneficial bacteria should be taken into consideration in the selection of probiotic cultures. For instance, LAB strains display a natural ability to tolerate the severe wine environment (pH = 3.0), as well as the very acidic pH of the stomach. This phenotype is based on several strategies to neutralize pH lowering and/or to adapt to acidic environmental and is highly appreciated in starter and probiotic LAB. Unfortunately, some of these metabolic strategies generate undesired molecules such as spoilage amines, like putrescine and cadaverine, which alter the organoleptic properties of food [58, 59]. Hence, excluding a priori a certain LAB strain for such genetic traits is incorrect if the food matrix pH is neutral or alkaline [60].
Conclusions
Morphological, physiological, and biochemical analysis presumptively classified the five LAB strains as L. lactis subsp. cremoris CTC 204 and CTC 483, L. lactis subsp. hordinae CTC 484, and Lact. plantarum CTC 368 and CTC 469. Differentiations in the DNA and whole-cell proteins profiles were observed between Lactococcus and Lactobacillus strains. All strains showed probiotic characteristics: tolerance to pH 2.0, survival or growth at compatible temperatures of the gastrointestinal tract or food storage, and ability for biofilm production. Bile tolerance was observed in strain CTC 469. Moreover, the strains were confirmed safe in relation to the virulence factors. The characteristics of resistance and sensitivity to some antibiotics and the production of biogenic amines under specific conditions are properties that merit additional studies. It is concluded that L. lactis subsp. cremoris CTC 204 and Lact. plantarum CTC 368 are the LAB strains with the greatest potential to be used as a probiotic culture. Further work to evaluate the applicability of these strains in food systems and in vivo is already in progress, since other biological variables may interfere with the results already found.
References
FAO/WHO (2001) Probiotic in food. Health and nutritional properties and guidelines for evaluation. http://www.fao.org/3/a-a0512e.pdf. Accessed 10 March 2016
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, Morelli L, Canani RB, Flint HJ, Salminen S, Calder PC, Sanders ME (2014) Expert consensus document: The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11(8):506–514. https://doi.org/10.1038/nrgastro.2014.66
Tripathi MK, Giri SK (2014) Probiotic functional foods: survival of probiotics during processing and storage. J Funct Foods 9:225–241. https://doi.org/10.1016/j.jff.2014.04.030
Stolaki M, De Vos WM, Kleerebezem M, Zoetendal G (2012) Lactic acid bacteria in the gut. In: Lahtinen S, Ouwehand AC, Salminen S, von Wright A (eds) Lactic acid bacteria. Marcel Dekker, New York, pp 385–401
Singh K, Kallali B, Kumar A, Thaker V (2011) Probiotics: a review. Asian Pac J Trop Biomed 1:S287–S290
Sanders ME, Akkermans LMA, Haller D, Hammerman C, Heimbach J, Hörmannsperger G, Huys G, Levy DD, Lutgendorff F, Mack D, Phothirath P, Solano-Aguilar G, Vaughan E (2010) Safety assessment of probiotics for human use. Gut Microbes 1(3):164–185. https://doi.org/10.4161/gmic.1.3.12127
Zago M, Fornasari ME, Carminati D, Burns P, Suàrez V, Vinderola G, Reinheimer J, Giraffa G (2011) Characterization and probiotic potential of Lactobacillus plantarum strains isolated from cheeses. Food Microbiol 28(5):1033–1040. https://doi.org/10.1016/j.fm.2011.02.009
Antoine JM (2012) Current challenges for probiotics in food: lactic acid bacteria in the gut. In: Lahtinen S, Ouwehand AC, Salminen S, von Wright A (eds) Lactic acid bacteria: microbiology and functional aspects. Marcel Dekker, New York, pp 213–226
Ojha KS, Kerry JP, Duffy G, Beresford T, Tiwari BK (2015) Technological advances for enhancing quality and safety of fermented meat products. Trends Food Sci Technol 44(1):105–116. https://doi.org/10.1016/j.tifs.2015.03.010
Favaro L, Todorov SD (2017) Bacteriocinogenic LAB strains for fermented meat preservation: perspectives, challenges, and limitations. Probiotics Antimicrob Proteins 9(4):444–458. https://doi.org/10.1007/s12602-017-9330-6
Bromberg R, Moreno I, Zagnanini CL, Delboni RR, Oliveira J (2004) Isolation of bacteriocin producing lactic acid bacteria from meat and meat products and its spectrum of inhibitory activity. Braz J Microbiol 35(1-2):137–144. https://doi.org/10.1590/S1517-83822004000100023
Leisner JJ, Vancanneyt M, Rusul G, Pot B, Lefebvre K, Fresi A, Tee LK (2001) Identification of lactic acid bacteria constituting the predominating microflora in acid-fermented condiment (tempoyak) popular in Malaysia. Int J Food Microbiol 63(1-2):149–157. https://doi.org/10.1016/S0168-1605(00)00476-1
Harrigan WF (1998) Laboratory methods in food microbiology. Academic Press, London
Bromberg R, Moreno I, Delboni RR, Cintra HC, Oliveira PTV (2005) Characteristics of the bacteriocin produced by Lactococcus lactis subsp. cremoris CTC 204 and the effect of this compound on the mesophilic bacteria associated with raw beef. World J Microbiol Biotechnol 21(3):351–358. https://doi.org/10.1007/s11274-004-2610-9
Bromberg R, Moreno I, Delboni RR, Cintra HC, Oliveira PTV (2006) Características da bacteriocina produzida por Lactococcus lactis ssp. hordinae CTC 484 e seu efeito sobre Listeria monocytogenes em carne bovina. Ciênc Tecnol Aliment 26(1):135–144. https://doi.org/10.1590/S0101-20612006000100023
Vila J, Marcos MA, Jimenes de Anta MTA (1996) Comparative study of different PCR-based DNA fingerprinting techniques for typing of the Acinetobacter calcoaceticus – A. baumannii complex. J Med Microbiol 44(6):482–489. https://doi.org/10.1099/00222615-44-6-482
Appuhamy S, Parton R, Coote JG, Gibbs HA (1997) Genomic fingerprinting of Haemophilus somnus by a combination of PCR methods. J Clin Microbiol 35(1):288–291
Herman L, Heyndrickx M, Waes G (1998) Typing of Bacillus sporothermodurans and others Bacillus species isolated from milk by repetitive element sequence based PCR. Lett Appl Microbiol 26:183–188
Valence F, Lortal S (1995) Zymogram and preliminary characterization of Lactobacillus helveticus autolysins. Appl Environ Microbiol 61(9):3391–3399
Mishra V, Prasad DN (2005) Application of in vitro methods for selection of Lactobacillus case in strains as potential probiotics. Int J Food Microbiol 103(1):109–115. https://doi.org/10.1016/j.ijfoodmicro.2004.10.047
Lebeer S, Verhoeven TLA, Vélez MP, Vanderleyden J, Keersmaecker SCJ (2007) Impact of environmental and genetic factors on biofilm formation by the probiotic strain Lactobacillus rhamnosus GG. Appl Environ Microbiol 73(21):6768–6775. https://doi.org/10.1128/AEM.01393-07
Stepanovic S, Vukovic D, Hola V, Boaventura GD, Djukic S, Cirkovic I, Ruzicka F (2007) Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. Acta Pathol Microbiol Immunol Scand 115(8):891–899. https://doi.org/10.1111/j.1600-0463.2007.apm_630.x
Freitas VR, van der Sand ST, Simonetti AB (2010) Formação in vitro de biofilme por Pseudomonas aeruginosa e Staphylococcus aureus na superfície de canetas de alta rotação. Rev Odontol UNESP 39:193–200
Henning DR, Flowers R, Reiser R, Ryser ET (2004) Pathogens in milk and milk products. In: Wehr HM, Frank JF (eds) Standard methods for the examination of dairy products. APHA, Seventeenth, Washington, pp 103–125. https://doi.org/10.2105/9780875530024ch05
Todorov SD, Botes M, Guigas C, Schillinger U, Wiid I, Wachsman MB, Holzappfel WH (2008) Boza, a natural source of probiotic lactic acid bacteria. J Appl Microbiol 104:465–477
EFSA European Food Safety Authority (2012) Guidance on the assessment of bacterial susceptibility to antimicrobial of human or veterinary importance. EFSA J 10:2740
Malle P, Valle M, Bouquelet S (1996) Assay for biogenic amines involved in fish decomposition. J AOAC Int 79:43–49
Axelsson L (2004) Lactic acid bacteria: classification and physiology. In: Salminem S, von Wright A, Ouwehand A (eds) Lactic acid bacteria: microbiology and functional aspects. Marcel Dekker, New York, pp 01–66
Franz CMAP, Holzapfel WH (2004) The genus Enterococcus: biotechnological and safety issues. In: Salminen A, von Wright A, Ouwehand A (eds) Lactic acid bacteria: microbiological and functional aspects. Marcel Dekker, Inc, New York, pp 199–248. https://doi.org/10.1201/9780824752033.ch6
Franzetti L, Pompei M, Scarpellini M, Galli A (2004) Phenotypic and genotypic characterization of Enterococcus spp. of different origins. FEMS Microbiol Lett 236(2):257–260
Corsetti A, De Angelis M, Dellaglio F, Paparella A, Fox PF, Settanni L, Gobetti M (2003) Characterization of sourdough lactic acid bacteria based on genotypic and cell-wall protein analysis. J Appl Microbiol 94:641–654
von Wright A, Axelsson L (2012) Lactic acid bacteria: an introduction In: Lactic acid bacteria: microbiological and functional aspects. In: Lahtinen S, Ouwehand AC, Salminen S, von Wright A (eds) Lactic acid bacteria, Forth edn. CRC Press, New York, p 779
Sharma P, Tomar SK, Goswami P, Sangwan V, Singh R (2014) Antibiotic resistance among commercially available probiotics. Food Res Int 57:176–195. https://doi.org/10.1016/j.foodres.2014.01.025
Bačun-Družina V, Mrvčić J, Butorac A, Stehlik-Tomas V, Gjuračić K (2009) The influence of gene transfer on lactic acid bacteria evolution. Mljekarstvo 59:181–192
Wiedenbecke J, Cohen FM (2011) Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol Rev 35(5):957–976. https://doi.org/10.1111/j.1574-6976.2011.00292.x
Patel AR, Shah NP, Prajapati JB (2012) Antibiotic resistance profile of lactic acid bacteria and their implications in food chain. World J Dairy Food Sci 7:202–211
Redondo-Lopez V, Cook RL, Sobel JD (1990) Emerging role of lactobacilli in the control and maintenance of the vaginal bacterial microflora. Rev Infect Dis 12(5):856–872. https://doi.org/10.1093/clinids/12.5.856
Luchese RH (2012) Microbial interactions in the gut: the role of bioactive components milk and honey. In: Rigobelo EC (ed) Probiotics. http://www.intechopen.com/books/probiotics/microbial-interactions-in-the-gut-the-role-of-bioactive-components-in-milk-and-honey. Accessed 15 Feb 2016
De Vuyst L, Leroy F (2007) Bacteriocin from lactic acid bacteria: production, purification and food application. J Mol Microbiol Biotechnol 13(4):194–199. https://doi.org/10.1159/000104752
Butel MJ (2014) Probiotics, gut microbiota and health. Med Mal Infect 44(1):1–8. https://doi.org/10.1016/j.medmal.2013.10.002
Martini MC, Bollweg GL, Levitt MD, Savaiano DA (1987) Lactose digestion by yogurt β-galactosidase: influence of pH and microbial cell integrity. Am J Clin Nutr 45(2):432–436. https://doi.org/10.1093/ajcn/45.2.432
Huang Y, Adams MC (2004) In vitro assessment of the upper gastrointestinal tolerance of potential probiotic dairy propionibacteria. Int J Food Microbiol 91(3):253–260. https://doi.org/10.1016/j.ijfoodmicro.2003.07.001
El-Shafei K, Tawfic NF, Dabiza NMA, Sharaf OM, Effat BA (2010) In vitro assessment of gastrointestinal viability of potentially probiotic Lactobacilli. J Am Sci 6:357–367
Marteau P (2004) Facteurs de contrôle de la flore. Définitions et mode d’actions des probiotiques et prébiotiques. In: Rambaud J-C, Buts J-P, Corthier G, Flourié B (eds) Flore microbienne intestinale: physiologie et pathologie digestives. John Libbey Eurotext, Montrouge, pp 37–58
Jungersen M, Wind A, Johansen E, Christensen JE, Stuer-Lauridsen B, Eskesen D (2014) The science behind the probiotic strain Bifidobacterium animalis subsp. lactis BB-12®. Microorganisms 2(2):92–110. https://doi.org/10.3390/microorganisms2020092
Park JK, Chang HN (2000) Microencapsulation of microbial cells. Biotechnol Adv 18(4):303–319. https://doi.org/10.1016/S0734-9750(00)00040-9
Cook MT, Tzortz G, Charalampopoulos D, Khutorryanskiy V (2012) Microencapsulation of probiotics for gastrointestinal delivery. J Control Release 162(1):56–67. https://doi.org/10.1016/j.jconrel.2012.06.003
Gbassi GK, Vandamme T (2012) Probiotic encapsulation technology: from microencapsulation to release into the gut. Pharmaceutics 4:149–163
Burns P, Vinderola G, Binetti A, Quiberoni A, de los Gavilán-Reyes CG, Reinheimer J (2008) Bile-resistant derivatives obtained from non-intestinal dairy lactobacilli. Int Dairy J 18:377–385
Caggia C, De Angelis M, Pitino I, Pino A, Randazzo CL (2015) Probiotic features of Lactobacillus strains isolated from Ragusano and Pecorino Siciliano cheeses. Food Microbiol 50:109–117
Gueimond M, Sánchez B, Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:1–6
Donohue DC, Gueimond M (2012) Some considerations for the safety of novel probiotic bacteria. In: Lahtinen S, Ouweland AC, Salminen S, von Wright A (eds) Lactic acid bacteria: microbiology and functional aspects. Marcel Dekker, New York, pp 423–437
Aymerich T, Martín B, Garriga M, Vidal-Carou MC, Bover-Cid S, Hugas M (2006) Safety properties and molecular strain typing of lactic acid bacteria from slightly fermented sausages. J Appl Microbiol 100(1):40–49. https://doi.org/10.1111/j.1365-2672.2005.02772.x
Borriello SP, Hammes WP, Holzapfel W, Marteau P, Schrezenmeir J, Vaara M, Valtonen V (2003) Safety of probiotics that contain lactobacilli or bifidobacteria. Clin Infect Dis 36(6):775–780. https://doi.org/10.1086/368080
Stadnik J, Dolatowski ZJ (2010) Biogenic amines in meat and fermented meat products. Acta Sci Pol Techno Aliment 9:251–263
Moreno-Arribas MV, Polo MC, Jorganes F, Muñoz R (2003) Screening of biogenic amine production by lactic acid bacteria isolated from grape must and wine. Int J Food Microbiol 84(1):117–123. https://doi.org/10.1016/S0168-1605(02)00391-4
Konings WN (2006) Microbial transport: adaptation to natural environments. Antonie Van Leeuwenhoek 90(4):325–342. https://doi.org/10.1007/s10482-006-9089-3
Mangiapane E, Mazzoli R, Pessione A, Svensson B, Riedel K, Pessione E (2015) Ten years of subproteome investigations in lactic acid bacteria: a key for food starter and probiotic typing. J Proteome 127(Pt B):332–339. https://doi.org/10.1016/j.jprot.2015.04.028
Costantini A, Pietroniro R, Doria F, Pessione E, Garcia-Moruno E (2013) Putrescine production from different amino acid precursors by lactic acid bactéria from wine and cider. Int J Food Microbiol 165:11–17
Pessione A, Lamberti C, Pessione E (2010) Proteomics as a tool to study biogenic amine production. Mol Biosyst 6(8):1419–1430. https://doi.org/10.1039/c001948h
Acknowledgements
This work was supported by the São Paulo Research Foundation (FAPESP), SP, Brazil and National Council for Scientific and Technological Development (CNPq), DF, Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This research does not contain any studies with human or animal subjects.
Rights and permissions
About this article
Cite this article
Moreno, I., Marasca, E.T.G., de Sá, P.B.Z.R. et al. Evaluation of Probiotic Potential of Bacteriocinogenic Lactic Acid Bacteria Strains Isolated from Meat Products. Probiotics & Antimicro. Prot. 10, 762–774 (2018). https://doi.org/10.1007/s12602-018-9388-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-018-9388-9






 Abs;
Abs;  pH); 25 °C (
pH); 25 °C ( Abs;
Abs;  pH); 4 °C (
pH); 4 °C ( Abs;
Abs;  pH) and −20 °C (
pH) and −20 °C ( Abs;
Abs;  pH).
pH).