Abstract
As an important legume plant, Medicago truncatula is a preeminent model for the study of the processes of nitrogen fixation, symbiosis, and legume genomics. The regulatory mechanism of the cytokinin response has been studied in many plants, such as rice, Arabidopsis, tomato, and barley, but information about regulatory pathways and genes involved in the cytokinin response in Medicago truncatula is notably limited. In this study, to better understand the cytokinin response in Medicago truncatula, transcriptome analysis of seedlings grown with 6-benzylaminopurine or lovastatin was performed using RNA-Seq. In this study, 3627 and 3093 transcripts were differentially expressed in cytokinin-induced/control (Cyto/Ctrl) and cytokinin-inhibited/control (Inh/Ctrl) groups, respectively, and differentially expressed genes were tested by quantitative real-time PCR (qRT-PCR). Analysis of the cytokinin response in Medicago truncatula revealed a large number of transcripts involved in signal transduction, metabolic process, secondary metabolite biosynthesis, transport and catabolism, growth and development, defense mechanisms, and transcription. There were 43 transcription factor families, including 1845 transcription factor (TF) genes with 2147 TF transcripts, as detected by RNA-Seq. Additionally, 216 TF genes with 220 transcripts were differentially expressed in Cyto/Ctrl, and 185 TF genes with 189 transcripts were in Inh/Ctrl. A total of 289 and 260 DETs involved in biosynthesis, metabolism, and transduction of plant hormones were identified in the Cyto/Ctrl and Inh/Ctrl groups, respectively. Furthermore, 15 transcripts, including A-ARR, IPT, and CKX, were demonstrated to play roles in cytokinin regulation or signal transduction. These findings were associated with the cytokinin response in other plants. The resulting data provide the first cytokinin transcriptome analysis in Medicago truncatula. Further analysis and identification of cytokinin-regulated transcripts or signal transduction transcripts may help to elucidate the regulatory mechanisms governing the cytokinin response in Medicago truncatula.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Medicago truncatula is a close relative of alfalfa (Medicago sativa) and a small annual legume native to the Mediterranean region. This species has been chosen as a model organism for legume species, because it has a small diploid genome, is self-fertile, has a rapid generation time and prolific seed production, and is amenable to genetic transformation. The genomes of the two cultivars, A17 and R108, have been sequenced and assembled (Branca et al. 2011; Moll et al. 2017; Tang et al. 2014). For relatively higher transformation efficiency and a shorter generation time, R108 has emerged as a heavily researched cultivar in genetics (Tadege et al. 2005). Furthermore, R108 was used to create a large Tnt1-insert population, which has been widely employed around the world for gene functional analysis (Tadege et al. 2008).
Next-generation high-throughput sequencing (NGS) allows us to sequence DNA or RNA conveniently and cost-effectively to analyze differential expression genes or transcripts, and functionally annotate regulatory pathways. With this tool, works on transcriptome analysis have been accomplished in legume plants. As legumes are globally planted forage plants, fall dormancy is a very important characteristic. Dormant and nondormant alfalfa samples at different time points were harvested, and fall dormancy-related DEGs were identified and analyzed by NGS technology (Zhang et al. 2015). To identify single-nucleotide polymorphisms (SNPs) in drought-related genes, sensitive and tolerant cultivars were used for transcriptome sequencing, and more than 6000 SNPs were identified in 2222 drought stress-related genes (Vidal et al. 2012). In our lab, we harvested two developmental stages of red clover leaves, mature and old, and RNA-Seq was used to investigate DEGs and regulatory pathways. A total of 481 genes were identified as DEGs between the two developmental stages (Chao et al. 2018). RNA-Seq was applied to identify pathogenicity-associated DEGs and potential effectors during infection of the Medicago truncatula host by a fungal pathogen (Thatcher et al. 2016).
Cytokinins are a class of molecules that are N6-substituted adenine derivatives, such as isopentenyl adenine and trans- and cis-zeatin, which are common in most plants (Keshishian and Rashotte 2015). Cytokinin is an essential plant hormone that is involved in plant growth and development, such as cell division and differentiation, plant senescence, other hormone biosynthesis and responses, seed development and biomass, and plant tolerances to biotic and abiotic stresses (Jameson and Song 2016; Li et al. 2015; Mok and Mok 2001; Nguyen et al. 2016; Roche et al. 2016; Schaller et al. 2014; Talla et al. 2016; Zurcher and Muller 2016). Isopentenyltransferase (IPT) is a key enzyme in regulating the biosynthesis of cytokinin, and transgenic plants show a delayed senescence phenotype with overexpression of the IPT gene in canola (Kant et al. 2015). In cotton, suppression of the cytokinin metabolism gene GhCKX can significantly improve both seed and fiber yield (Zhao et al. 2015). Cytokinin levels are regulated by a balance between biosynthesis (Kuderova et al. 2008; Kuppu et al. 2013), activation (Kuroha et al. 2009; Tokunaga et al. 2012), inactivation (O-glucosyl transferase) (Polanska et al. 2004), reactivation (β-glucosidase) (Brzobohaty et al. 1993; Kiran et al. 2006), and degradation (cytokinin oxidase/dehydrogenase) (Kollmer et al. 2014; Reid et al. 2016). Recently, some studies have used NGS technology to investigate cytokinin responses, and many key cytokinin-response genes were explored and characterized in plants. In Oryza sativa, the expression of OsNSHB2 was significantly upregulated in the presence of cytokinin (Ross et al. 2004). In Arabidopsis, 4 cytokinin-response factor genes, including CRF1, CRF2, CRF5, and CRF6, are transcriptionally induced by cytokinin (Rashotte et al. 2003). Previous studies, in plants such as Arabidopsis, rice, and tomato, have utilized transcriptomic approaches to investigate cytokinin signaling. (Bhargava et al. 2013; Day et al. 2008; Raines et al. 2016; Shi et al. 2013). Cytokinin transcriptomes in tomato leaves were analyzed by RNA-Seq, and this was the first cytokinin transcriptome analysis in tomato (Shi et al. 2013). RNA-Seq was used to explore the cytokinin-regulated genes in rice, and the results showed that approximately 4700 genes in roots and 2400 genes in shoots were identified as DEGs (Raines et al. 2016). These results may help to identify candidate genes regulated by cytokinin in plants. Previous studies have mainly focused on cytokinin-induced or suppressed genes, but the expression levels of genes with cytokinin inhibitors remain unknown. Little is known about the cytokinin response in legume plants and how it might work in cytokinin responses. As a model plant in legume plants, exploring the cytokinin-response transcriptome of Medicago truncatula remains especially important.
In this work, Medicago truncatula under 6-benzylaminopurine-treatment, lovastatin-treatment and no-treatment conditions was investigated, and global changes in the cytokinin response were analyzed by RNA-Seq. We identified DETs, transcriptional factors and plant hormone-related transcripts, which provide abundant gene resources, and the transcriptome data may help to elucidate the regulatory mechanisms governing the cytokinin response in Medicago truncatula.
Materials and Methods
Plant Materials and Growth Conditions
M. truncatula seeds (R108) received as gifts from Samuel Roberts Noble Foundation (Ardmore, Oklahoma, USA) were surface-sterilized and transferred to 1/2 MS medium (pH 5.8) containing 1.5% sucrose in plant tissue culture bottles and then grown in a growth chamber (25/23 °C day/night temperatures, 16-h photoperiod, and 60% relative humidity). For treatment of cytokinin, 0.5 mg/L 6-BA was supplied into 1/2 MS medium, and 1 mg/L lovastatin was added to inhibit cytokinin. Seedlings grown on different MS media for 1 month were harvested for cytokinin analysis, and the cytokinin levels were measured with a plant zeatin (Balb, China) and 6-BA ELISA kit (J&L, China) according to the manual. Seedlings 1 month old on different MS media were harvested for RNA extraction.
Preparation of Libraries and Illumina Sequencing of M. truncatula
Total RNA from whole seedlings with three different growth conditions was extracted and then treated with RNase-free DNase I (Promega, USA) to remove contaminating genomic DNA. Purity, integrity, and quantity of RNA were measured as previously described (Chao et al. 2018). A total of 1 µg of RNA from each sample were used to generate an Illumina library, and index codes (5′ adaptor: 5′-AGATCGGAAGAGCACACGTC-3′ and 3′ adaptor: 5′-AGATCGGAAGAGCGTCGTGT-3′) were added to each sample. Each Illumina library’s quality was measured on the Agilent Bioanalyzer 2100 system, and the libraries were sequenced with HiSeq4000 instruments. After removing reads containing adapters, reads containing poly-N and low-quality reads from raw data, clean data were obtained, and all transcriptome analyses were based on the clean data.
Differentially Expressed Transcripts Analyses in Different M. truncatula Samples
The genome sequences and annotation files of the Medicago truncatula R108 genome (version 1.0) were downloaded from Medicago truncatula HAPMAP Project (Moll et al. 2017). Clean data were aligned to the reference genome using Hisat2 (https://ccb.jhu.edu/software/tophat/index.shtml) with default parameters (Kim et al. 2015). StringTie (https://ccb.jhu.edu/software/stringtie/) was used to assemble mapped data (Pertea et al. 2015). Novel transcripts were identified combined with DIAMOND, NR, and Swiss-Prot. DIAMOND (version 0.8.37.99), HMMER (version 3.1b2), BLAST2GO (version 2.5.0), and KOBAS (version 2.1.1) were used for function annotation (Buchfink et al. 2015; Conesa et al. 2005; Finn et al. 2011; Mao et al. 2005). Then, the FPKM (expected number of Fragments Per Kilobase of transcript sequence per Million base pairs sequenced) of each transcript was calculated using RSEM (version 1.2.31) with default parameters (Li and Dewey 2011). DETs (differentially expressed transcripts) were identified using the DESeq2 (version 1.10.1) between different samples with parameters: adjusted p value < 0.05 and |log2FC|> = 1 (Love et al. 2014).
Quantitative RT-PCR of DETs Identified from RNA-Seq Data of M. truncatula
Nine randomly selected DETs and all DETs involved in cytokinin pathway (Supplementary Table S1) identified from RNA-Seq were confirmed by qRT-PCR, as described previously (Chao et al. 2018). Whole seedlings grown on different 1/2 MS media were harvested, and first-strand cDNAs were synthesized as described above and used for qRT-PCR. The Medicago truncatula GAPDH transcript (MSTRG.23995.2) was used as an internal control. A two-step amplification protocol was performed, and the relative expression level of each transcript was calculated using the 2−∆∆Ct method (Livak and Schmittgen 2001).
Function Annotation of DETs from RNA-Seq Data of M. truncatula
DETs in the Cyto/Ctrl and Inh/Ctrl groups were searched against NR (NCBI non-redundant), NT (NCBI nucleotide sequence), Swiss-Prot, KEGG (Kyoto Encyclopedia of Genes and Genomes) (Kanehisa et al. 2004), and KOG/COG (Cluster of Orthologous Groups) (Tatusov et al. 2000) databases as described previously (Chao et al. 2018). GO (Gene Ontology) annotation was performed using the Blast2GO software (version 2.3.5) under a threshold E value ≤ 10−5 (Ashburner et al. 2000). KEGG pathway annotation was performed with the KEGG Automatic Annotation Server (KAAS1), and HMMER software (Eddy 1998) was used to search the Pfam database.
Assays of TFs and Hormone-Related DETs from RNA-Seq Data of M. truncatula
For transcription factor prediction, the plant transcription factor database PlantTFDB 4.0 (https://planttfdb.cbi.pku.edu.cn/index.php) (Jin et al. 2017) was used. TF prediction was performed with Hmmscan and Blast under a threshold E value ≤ 10−5. The Arabidopsis Hormone Database (AHD) was downloaded from https://ahd.cbi.pku.edu.cn. To predict the plant hormone-related transcripts, DETs from the two groups (Cyto/Ctrl and Inh/Ctrl) were compared with protein sequences from AHD using Blastx (E value ≤ 10−5).
Results
Exogenous of 6-BA or Lovastatin Both Decreased the Zeatin Levels in M. truncatula for a Long Term
Grown in different 1/2 MS media for 1 month, whole seedlings were harvested for further experiments (Fig. 1a). The height of plants under cytokinin or lovastatin conditions was considerably smaller than those under normal conditions (Fig. 1a). Plants with cytokinin produced more branches with more leaflets, which were considerably smaller in size than those of plants with no treatment (Fig. 1b). Seedlings with cytokinin inhibitor exhibited a slow growth phenotype, and some compound leaves appeared in deformed states (Fig. 1b). We measured the contents of plant hormone zeatin, and the results showed that the cytokinin levels in plants with 6-BA or lovastatin treatments were lower than those in plants without treatment (Fig. 1c). Meanwhile, we also measured the 6-BA contents in plants, and the results showed that the 6-BA contents in cytokinin-induced plants were improved largely compared with those without cytokinin treatment (Fig. S1).
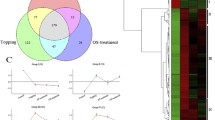
Phenotypes and transcripts detected by NGS from three samples. a Phenotypes of Medicago truncatula of 1 month old at different growing conditions. Ctrl, Medicago truncatula with un-treatment; Cyto, Medicago truncatula with 6-BA treatment; Inh, Medicago truncatula with lovastatin treatment. b Leaflets from Medicago truncatula of 1 month old at different growing conditions. Bar = 1 mm. c The levels of plant zeatin measured in different samples. Error bars represent the SD from three independent experiments. *Indicates significant differences of the means at p < 0.01 between samples for plant zeatin measurements (n = 3). d Vern diagram of transcripts detected in the three samples. e Length distributions of all transcripts and novel transcripts detected in RNA-Seq
Sequencing and Statistical Analysis of RNA-Seq Data from all M. truncatula Samples
Each library was sequenced with an Illumina platform. Approximately 0.45 billion raw reads were generated (SRA accession number: SRP159219), and the GC content ranged from 42.70 to 43.65% from the nine Illumina libraries (Table 1). After filtering out low-quality reads, the number of clean reads from each library was 41–65 millon (Table 1). By mapping to the reference genome, more than 97% of clean reads were mapped to the Medicago truncatula R108 genome (Table 2). A total of 33,782 genes with 55,558 transcripts were detected in the three samples with a min length of 132 bp and a max length of 16,535 bp. The mean length of transcripts was 1801 bp with an N50 of 2.32 kb. A total of 27,810 transcripts were detected in all three samples (Fig. 1d). In total, 49,238 transcripts (88.62%) were longer than 500 bp, and 39,602 transcripts (71.28%) were longer than 1 kb (Fig. 1e). By mapping to the reference genome, 22,407 transcripts and 1079 genes were identified as novel. The mean length of novel transcripts was 2,285 bp with an N50 of 2.73 kb. By analysis, 21,716 novel transcripts (96.92%) were longer than 500 bp, and 19,234 novel transcripts (85.84%) were longer than 1 kb (Fig. 1e).
Function Annotation of Novel Transcripts Identified from RNA-Seq
Novel transcripts were functionally annotated by searching multiple databases, and 21,939 novel transcripts were annotated (Fig. 2a; Supplementary Table S2). By searching the NR database, we found the largest ten transcripts distributed in Medicago truncatula (20,766), Cicer arietinum (627), Glycine max (115), Glycine soja (53), Phaseolus vulgaris (47), Medicago sativa (20), Citrus sinensis (17), Vitis vinifera (16), Arabidopsis thaliana (14), and Lotus japonicas (12) (Fig. 2b). GO analysis demonstrated that 15,242 transcripts were divided into biological processes, molecular functions, and cellular components (Fig. 2c). The KEGG analysis showed that 4941 novel transcripts were mapped to 116 KEGG pathways (Fig. 2d).
Function annotation of novel transcripts. a Function annotation of transcripts in all databases. Nr Non-Redundant Protein Database, GO Gene Ontology, COG/KOG Cluster of Orthologous Groups of proteins, KEGG Kyoto Encyclopedia of Genes and Genomes. b Nr Homologous species distribution diagram of transcripts. c Percentages and numbers of GO terms for novel transcripts in cellular component, molecular function, and biological process. d KEGG pathways enriched of transcripts with number more than 50
DET Analysis Between 6-BA and Lovastatin-Treated Samples with Control Samples
By DET analysis, there were 3627 and 3093 DETs in the Cyto/Ctrl and Inh/Ctrl groups, respectively. In the Cyto/Ctrl group, 2498 transcripts were upregulated and 1129 transcripts were downregulated (Fig. 3a; Supplementary Table S3). In the Inh/Ctrl group, 2069 transcripts were upregulated, and 1024 transcripts were downregulated (Fig. 3b; Supplementary Table S4). We randomly selected 9 transcripts for qRT-PCR analysis to verify the DET results, and the qRT-PCR results displayed expression patterns that were in keeping with those of the RNA-Seq analysis (Fig. 3c) in Medicago truncatula.
DETs’ identification and RT-PCR validation. a Volcano plot of DETs in Cyto/Ctrl. b Volcano plot of DETs in Inh/Ctrl. Each plot represents one transcript with three colors, including red (up regulated), green (downregulated), and gray (not changed). The X-axis represents the value of log2 (Fold Change) between the two samples, and the Y-axis indicates the negative value of log10(P-adjust). c The qRT-PCR validation of nine randomly selected DETs. Columns represent the value of log2 (Fold Change) (Y-axis). Error bars represent the SD from three independent experiments. *Indicates significant differences of the means at p < 0.01 between samples for lovastatin treatment with control (n = 3). **Indicates significant differences of the means at p < 0.01 between samples for 6-BA treatment with control (n = 3)
Function Annotation of DETs Between 6-BA and Lovastatin-Treated Samples with Control Samples
All DETs from the two groups were functionally annotated, and a total of 3597 (99.17%) and 3066 (99.13%) transcripts were annotated for the Cyto/Ctrl and Inh/Ctrl groups, respectively (Table 3). When searching sequences in the NR database, a total of 3596 and 3066 transcripts were annotated in the Cyto/Ctrl and Inh/Ctrl groups, respectively (Table 3). GO analysis showed that the enrichment of 2550 and 2177 transcripts could be divided into three groups (Fig. 4a-b). A total of 687 transcripts were mapped to 103 KEGG pathways in Cyto/Ctrl, and 548 transcripts were mapped to 94 KEGG pathways in Inh/Ctrl (Fig. 4c-d).
Function annotation of DETs. a Percentages and numbers of GO terms for DETs in cellular component, molecular function, and biological process in Cyto/Ctrl. b Percentages and numbers of GO terms for DETs in cellular component, molecular function, and biological process in Inh/Ctrl. c KEGG pathways enriched of DETs with number more than 10 in Cyto/Ctrl. d KEGG pathways enriched of DETs with number more than 10 in Inh/Ctrl
Plant Hormone DETs Between 6-BA and Lovastatin-Treated Samples with Control Samples
The Arabidopsis Hormone Database was used to predict plant hormone DETs. The prediction results indicated that 289 DETs in Cyto/Ctrl and 260 DETs in Inh/Ctrl were involved in plant hormones, including ABA, auxin, brassinosteroid, cytokinin, ethylene, gibberellin, jasmonic acid, and salicylic acid (Table 4; Supplementary Table S5–S6). In this work, for cytokinin in Cyto/Ctrl, five transcripts (236:maker-Contig51-snap-gene-11.43-mRNA-1, 276:maker-Contig20 -augustus-gene-5.34-mRNA-1, 276:maker-Contig20-snap-gene-28.48-mRNA-1, 507:augustus_masked-Contig195 -processed-gene-13.13-mRNA-1, and 510:maker-Contig21-augustus- gene-44.43-mRNA-1) were upregulated, and two transcripts (576:augustus_masked-Contig77-processed-gene-98.13-mRNA-1 and 71:snap_masked-Contig26-processed-gene-95.34-mRNA-1) were downregulated. To confirm the expression patterns of those cytokinin-response transcripts, qRT-PCR was conducted. The results showed that the expression trends of those transcripts were consistent with the analysis results from RNA-Seq (Supplementary Fig. S2). In the Inh/Ctrl group, five transcripts (782:augustus_masked-Contig247-processed-gene-1.11-mRNA-1, 782:augustus_masked-Contig247- processed-gene-11.5-mRNA-1, 782:augustus_masked-Contig247-processed-gene-90.12-mRNA-1, 782:maker- Contig247-augustus-gene-4.33-mRNA-1 and 782:maker-Contig247-snap-gene-31.59-mRNA-1) were upregulated, and three transcripts (75:maker-Contig8-augustus-gene-125.25-mRNA-1, 782:maker-Contig247-augustus-gene-49.32-mRNA-1, and 858:augustus_masked-Contig100-processed-gene-113.5-mRNA-1) were downregulated. For example, two transcripts, 236:maker-Contig51-snap-gene-11.43-mRNA-1 (homolog of AT2G41510.1 in Arabidopsis) and 276:maker-Contig20-snap-gene-28.48-mRNA-1 (homolog of AT1G75450.1 in Arabidopsis), which were identified as cytokininoxidase/dehydrogenase (CKX) homologs, showed upregulated expression levels in Cyto/Ctrl. The expression levels of the transcript 75:maker-Contig8-augustus-gene-125.25-mRNA-1, homolog of EXPANSIN 1 (AT1G69530) in Arabidopsis involved in cytokinin signal response was drastically decreased in Inh/Ctrl.
A total of 22 and 29 DETs from Cyto/Ctrl and Inh/Ctrl, respectively, were enriched in the plant hormone signal pathway by KEGG analysis (Fig. 5). In the cytokinin signal pathway, cytokinin receptor (CRE1) can respond to cytokinins (Franco-Zorrilla et al. 2005). Only one DET was detected in the Inh/Ctrl group, and the expression was decreased by inhibition of cytokinin. Histidine-containing phosphotransfer proteins (AHP) function as positive regulators of cytokinin responses (D’Agostino and Kieber 1999). The AHP transcript 877:maker-Contig37-augustus-gene-283.41-mRNA-1 was upregulated by cytokinin treatment, and another transcript, 660:maker-Contig15-augustus-gene-25.68-mRNA-1, was downregulated to 38.9% of the control by lovastatin treatment. Type-B Arabidopsis thaliana response regulators (ARRs) act as positive regulators of cytokinin signal transduction (Mason et al. 2005), and the homologs in Medicago truncatula showed decreased levels in both treatments. Type-A Arabidopsis thaliana response regulators (A-ARR) act as negative regulators of cytokinin responses, and MSTRG.18714.2 was decreased by cytokinin treatment (Hwang and Sheen 2001). Six A-ARR homolog transcripts were identified as DETs in Inh/Ctrl, of which three were upregulated and three were downregulated. For the auxin signaling pathway, 10 transcripts were differentially expressed in Cyto/Ctrl, and 6 were identified as DETs by cytokinin disruption. GA-insensitive dwarf 2 (GID2) regulates the expression of SLEEPY1 (SLR1) gene, which plays a role in the GA signal pathway in plants, and one homolog transcript (MSTRG.18582.1) was downregulated by lovastatin treatment in the GA signal pathway (Gomi et al. 2004). Two and five DETs were detected in the two treatments for the ABA signal pathway. For the JA signaling pathway, DET was not detected in Cyto/Ctrl, while one jasmonate resistant 1 (JAR1) homolog transcript, MSTRG.6326.4, and one jasmonate ZIM-domain (JAZ) homolog transcript, 169:maker-Contig176-augustus-gene-35.42-mRNA-1, displayed decreased expression levels in addition to cytokinin inhibitor. For the SA signaling pathway, two transcripts showed enhanced expression levels, and two showed decreased expression levels in Cyto/Ctrl. Three transcripts were identified as DETs, which were all upregulated in Inh/Ctrl. Pathogenesis-related 1 (PR-1) gene expression is induced in response to a variety of pathogens and SA, representing a useful molecular marker for the SA response (Rogers and Ausubel 1997). In our work, 138:augustus_masked-Contig16-processed-gene-8.3-mRNA-1 in Cyto/Ctrl and 169:augustus_masked-Contig176- processed-gene-52.13-mRNA-1 in Inh/Ctrl were both upregulated.
DETs in plant hormone signal transduction of KEGG pathways. a Heat map of the expression levels of DETs involved in plant hormones in the three samples. b Six plant hormone signal transduction pathways. Numbers in boxes represent DET numbers involved in plant hormone signal transduction pathways identified in Cyto/Ctrl (left) or Inh/Ctrl (right). p represents phosphate. All signal transduction pathways were simplified from the pathways in Medicago truncatula
TF Identification of Transcripts Identified from RNA-Seq Data
TFs are key regulatory proteins involved in regulating plant growth and development. By sequence analysis, a total of 2147 TF transcripts and 1845 TF genes were identified from 43 TF families from RNA-Seq, and the top 10 were the B3 superfamily, AP2/ERF, MYB, bHLH, MYB-related, FAR1, MADS, WRKY, C2C2, and NAC (Fig. 6a; Supplementary Table S7). For cytokinin treatment, the expression levels of 117 TF transcripts were clearly changed, including 86 upregulated and 29 downregulated. A total of 102 TF transcripts were differentially expressed in Inh/Ctrl, with 76 upregulated and 26 downregulated (Fig. 6b). A C2H2 TF transcript 169:augustus_masked-Contig176-processed-gene-42.2-mRNA-1 showed an increased expression level. The transcript 738:augustus_masked-Contig125-processed-gene-24.12-mRNA-1, encoding an AP/ERF TF, showed a 4.7-fold increase in expression, while another AP/ERF transcript 877:maker-Contig37-snap-gene-240.37-mRNA-1 showed a decreased expression level under 6-BA treatment. For cytokinin inhibitor treatment, 98 DETs were identified as TF transcripts, including 71 upregulated and 26 downregulated. A bHLH transcript, 138:maker-Contig16-augustus-gene-28.26-mRNA-1, was upregulated to more than four times the expression level, and the WRKY transcript 276:maker-Contig20-augustus-gene-123.43-mRNA-1 showed a similar expression trend under lovastatin treatment. An HD-ZIP transcript, 169:maker-Contig176-augustus-gene-56.30-mRNA-1, was downregulated to ~ 50% of the expression level in Inh/Ctrl.
Differentially expressed transcription factor transcripts in cytokinin response. a A total of 34 DET TF families identified by NGS. The X-axis represents transcription factor family, and the Y-axis represents the number of transcripts. b Heat map of the expression levels of DETs involved in TF families in the three samples
Discussion
In this work, new insights into the Medicago truncatula transcriptome were provided, and this transcriptome analysis is the first work to study the cytokinin response of a legume species with NGS technology. In this work, we applied lovastatin to inhibit the regulatory effect of cytokinin. We determined that the zeatin contents and the results showed that the levels of cytokinin in plants supplemented with 6-BA or cytokinin inhibitor were both drastically decreased (Fig. 1c). The findings were consistent with the previous reports. Low concentrations of lovastatin have been shown to inhibit normal growth of plant seedlings and cell (Hashizume et al. 1983; Hata et al. 1987). The inhibition of growth by lovastatin can be reversed with the presence of cytokinin, and these data suggest that lovastatin specifically inhibits cytokinin biosynthesis (Crowell and Salaz 1992). When we tested the zeatin and 6-BA contents, we found that the 6-BA contents in cytokinin-treated plants were much higher compared with those without cytokinin treatment, while the zeatin contents were decreased (Fig. 1c; Fig. S1). The 6-BA is a zeatin analogue that plays a similar role in regulating plant growth and development. Grown with 6-BA, plants accumulate the zeatin analogue, and a high concentration of cytokinin prevents normal growth and development of plants. To weaken the negative effect of cytokinin, plants have to downregulate the biosynthesis of endogenous zeatin levels. In the previous work, another zeatin analogue, N- (2-chloro-4-pyridyl) -N′-phenylurea (CPPU), was used to regulate endogenous plant hormones. The results showed that the zeatin contents were decreased in Momordica charantia with CPPU treatment (Xia et al. 2009).
We used high-throughput Illumina sequencing technology to explore the DETs and analyze the regulatory mechanisms of cytokinin response in Medicago truncatula. Under different growth conditions, multiple transcripts are up- or downregulated, and these transcripts are involved in different signaling pathways of the cytokinin response. In this work, 3627 and 3093 DETs in the Cyto/Ctrl and Inh/Ctrl groups were identified by transcriptome analysis. In previous work on the cytokinin-response transcriptome, genes regulated by cytokinin were identified, but information on genes involved in cytokinin inhibitor was limited. The cytokinin-regulated transcripts in tomato leaves were investigated using RNA-Seq, and over 1000 were identified as responses to cytokinin (Shi et al. 2013). To explore cytokinin-regulated gene expression in Arabidopsis, RNA-Seq was used to characterize the response of the transcriptome to cytokinin, and the results revealed that 573 genes were differentially regulated by cytokinin with 423 upregulated and 150 downregulated (Bhargava et al. 2013).
Two CKX transcripts, MSTRG.27704.1 and 236:maker-Contig51-snap-gene-11.43-mRNA-1, showed considerably increased expression in Inh/Ctrl. The CKX transcript encoding a cytokinin oxidase/dehydrogenase plays a negative role in the biosynthesis of cytokinin (Schmulling et al. 2003). We cloned the CKX transcript, 236:maker-Contig51-snap-gene-11.43-mRNA-1 and overexpression of the transcript in Medicago truncatula displayed typical phenotypes of accumulation of cytokinin oxidase/dehydrogenase (Supplementary Fig. S3a). Those are maize CKX gene ZmCKO1, encoding the enzyme, catalyzes one molecule of cytokinin, yielding one molecule of H2O2 (Kopecny et al. 2005). In barley, downregulation of a CKX gene decreased cytokinin oxidase/dehydrogenase activities, resulting in higher plant productivity (Zalewski et al. 2010). These data reveal that lovastatin decreases cytokinin accumulation by improving the activities of CKX enzymes. In the Cyto/Ctrl group, four CKX transcripts showed upregulated expression levels, and we noticed that the transcript 236:maker-Contig51-snap-gene-11.43-mRNA-1 was both increased under cytokinin or lovastatin conditions. These results indicated that the transcript can be induced under different conditions via different regulatory mechanisms. Under cytokinin conditions, the plants accumulate CKX enzymes to maintain the balance of cytokinin levels. These findings were consistent with the expression pattern of three genes, BnCKX5-1, BnCKX5-2, and BnCKX6-1, induced by 6-BA in Brassica napus (Liu et al. 2018). The transcript 71:snap_masked-Contig26-processed-gene-95.34-mRNA-1, which encodes an isopentenyltransferase (IPT) and plays a role in cytokinin biosynthesis, was downregulated under cytokinin inhibitor treatments. We also cloned the IPT transcript, and transgenic plants displayed delayed leaf senescence phenotype (Supplementary Fig. S3). There are two classes of IPTs in plants. ATP/ADP ITPs are required for tZ-type cytokinin synthesis, and tRNA IPTs are required for cZ-type cytokinin (Miyawaki et al. 2006). The downregulated expression levels of the cytokinin biosynthesis gene under cytokinin conditions also confirmed the hypothesis about balanced regulation of cytokinin in Medicago truncatula.
Transcription factors are proteins that regulate the biosynthesis rate of messenger RNA from DNA by recognizing and binding to a specific DNA motif (Latchman 1997). We identified 2,147 TF transcripts from 43 TF families from RNA-Seq (Supplementary Table S7). For cytokinin treatment, the expression levels of 108 TF transcripts were clearly changed, including 82 upregulated and 26 downregulated. A total of 97 TF transcripts were differentially expressed in Inh/Ctrl with 71 upregulated and 26 downregulated (Fig. 6b). TFs play important roles in regulating the biosynthesis, metabolism, and signal induction of cytokinin. Overexpression of a cytokinin-induced transcription factor gene, ASL9, causes an altered sensitivity to cytokinin (Naito et al. 2007). In tomato, the CLAU gene encodes an MYB transcription factor that promotes leaf differentiation by negatively regulating cytokinin signaling (Bar et al. 2016). In our work, under 6-BA treatment, 4 MYB TF transcripts were upregulated, and 3 MYB transcripts were downregulated. Under lovastatin treatment, 7 and 2 MYB transcripts were upregulated and downregulated, respectively. In Arabidopsis, as a basic Helix–Loop–Helix (bHLH) transcription factor, SPATULA was shown to be involved in regulating cytokinin signaling (Reyes-Olalde et al. 2017). Induced by cytokinin, a total of 12 bHLH transcripts were notably changed, including 10 upregulated and 2 downregulated, in Medicago truncatula. Under lovastatin treatment, 7 bHLH transcripts were upregulated, and 2 were downregulated. The type-A ARRs are cytokinin response genes that function as repressors of cytokinin signaling (Hwang and Sheen 2001). The type-A ARR7 and ARR15 genes play negative roles in regulating cytokinin signaling in Arabidopsis thaliana (Muller and Sheen 2008). The AP2/ERF transcript factor DRNL controls gynoecium development by affecting its response to cytokinin (Duran-Medina et al. 2017). In Medicago truncatula, the transcription factor MtSERF1 of the AP2/ERF family is shown to induce expression levels by auxin plus cytokinin (Mantiri et al. 2008). Eight AP2/ERF transcripts were upregulated, and two were downregulated in the Cyto/Ctrl group.
By DET analysis, we noticed that there were 2 DETs showing upregulated expression levels in Cyto/Ctrl and downregulated expression levels in Inh/Ctrl, and there were 9 DETs showing upregulated expression in Inh/Ctrl and downregulated expression in Cyto/Ctrl. The transcript 660:maker-Contig15-augustus-gene-166.50-mRNA-1, encoding a cysteine proteinase, was expressed more than twofold under cytokinin treatment, while the transcript showed drastically decreased expression levels under lovastatin treatment. MSTRG.16266.1 encoding an ABA 8′-hydroxylase was expressed in a similar manner with more than threefold increased expression level under cytokinin treatment and 23.5% of expression under lovastatin treatment. An uncharacterized transcript, 738:augustus_masked-Contig125-processed-gene-128.9-mRNA-1, was identified as downregulated DET in Cyto/Ctrl and upregulated DET in Inh/Ctrl. We believe that these genes may play key roles in the cytokinin response and signal transduction, and the functions of these genes will be the focus of future research.
Conclusions
In this study, we identified DETs, transcriptional factors, and plant hormone-related transcripts. This study provides rich gene resources and the transcriptome data will give access to exploring regulatory mechanisms of cytokinin response in Medicago truncatula, which will help to understand this response and further explore of legume plants.
References
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Harris MA (2000) Gene ontology: tool for the unification of biology. Nat Genet 25:25–29
Bar M, Israeli A, Levy M, Gera H, Jiménez-Gómez J, Kouril S (2016) CLAUSA Is a MYB transcription factor that promotes leaf differentiation by attenuating cytokinin signaling. Plant Cell 28(7):1602–1615
Bhargava A, Clabaugh I, To JP, Maxwell BB, Chiang YH, Schaller GE, Kieber JJ (2013) Identification of cytokinin-responsive genes using microarray meta-analysis and RNA-Seq in arabidopsis. Plant Physiol 162:272–294
Branca A, Paape TD, Zhou P, Briskine R, Farmer AD, Mudge J, Ben C (2011) Whole-genome nucleotide diversity, recombination, and linkage disequilibrium in the model legume Medicago truncatula. P Natl Acad Sci USA 108:E864–E870
Brzobohaty B, Moore I, Kristoffersen P, Bako L, Campos N, Schell J, Palme K (1993) Release of active cytokinin by a beta-glucosidase localized to the maize root meristem. Science 262:1051–1054
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60
Chao Y, Xie L, Yuan J, Guo T, Li Y, Liu F, Han L (2018) Transcriptome analysis of leaf senescence in red clover (Trifolium pratense L.). Physiol Mol Biol Pla 24:753–765
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Crowell DN, Salaz MS (1992) Inhibition of growth of cultured tobacco cells at low concentrations of lovastatin is reversed by cytokinin. Plant Physiol 100:2090–2095
D’Agostino IB, Kieber JJ (1999) Molecular mechanisms of cytokinin action. Curr Opin Plant Bio 2(5):359–364
Day RC, Herridge RP, Ambrose BA, Macknight RC (2008) Transcriptome analysis of proliferating Arabidopsis endosperm reveals biological implications for the control of syncytial division, cytokinin signaling, and gene expression regulation. Plant Physiol 148:1964–1984
Durán-Medina Y, Serwatowska J, Reyes-Olalde JI, De Folter S, Marsch-Martínez N (2017) The AP2/ERF transcription factor DRNL modulates gynoecium development and affects its response to cytokinin. Front Plant Sci 8:1841
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37
Franco-Zorrilla JM, Martín AC, Leyva A, Paz-Ares J (2005) Interaction between phosphate-starvation, sugar, and cytokinin signaling in Arabidopsis and the roles of cytokinin receptors CRE1/AHK4 and AHK3. Plant Physiol 138(2):847–857
Gomi K, Sasaki A, Itoh H, Ueguchi-Tanaka M, Ashikari M, Kitano H, Matsuoka M (2004) GID2, an F-box subunit of the SCF E3 complex, specifically interacts with phosphorylated SLR1 protein and regulates the gibberellin-dependent degradation of SLR1 in rice. Plant J 37(4):626–634
Hashizume T, Matsubara S, Endo A (1983) Compactin (ML-236B) as a new growth inhibitor of plant callus. Agric Biol Chem 47:1401–1403
Hata S, Takagishi H, Kouchi H (1987) Variation in the content and composition of sterols in alfalfa seedlings treated with compactin (ML-236B) and mevalonic acid. Plant Cell Physiol 28:709–714
Hwang I, Sheen J (2001) Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413:383–389
Jameson PE, Song J (2016) Cytokinin: a key driver of seed yield. J Exp Bot 67:593–606
Jin JP, Tian F, Yang DC, Meng YQ, Kong L, Luo JC, Gao G (2017) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45:D1040–D1045
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Kant S, Burch D, Badenhorst P, Palanisamy R, Mason J, Spangenberg G (2015) Regulated expression of a cytokinin biosynthesis gene IPT delays leaf senescence and improves yield under rainfed and irrigated conditions in canola (Brassica napus L.). PLoS ONE 10:e0116349
Keshishian EA, Rashotte AM (2015) Plant cytokinin signalling. Essays Biochem 58:13–27
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Kiran NS, Polanská L, Fohlerová R, Mazura P, Válková M, Šmeral M, Brzobohatý B (2006) Ectopic over-expression of the maize beta-glucosidase Zm-p60.1 perturbs cytokinin homeostasis in transgenic tobacco. J Exp Bot 57:985–996
Kollmer I, Novak O, Strnad M, Schmulling T, Werner T (2014) Overexpression of the cytosolic cytokinin oxidase/dehydrogenase (CKX7) from Arabidopsis causes specific changes in root growth and xylem differentiation. Plant J 78:359–371
Kopecny D, Pethe C, Sebela M, Houba-Herin N, Madzak C, Majira A, Laloue M (2005) High-level expression and characterization of Zea mays cytokinin oxidase/dehydrogenase in Yarrowia lipolytica. Biochimie 87:1011–1022
Kuderova A, Urbankova I, Valkova M, Malbeck J, Brzobohaty B, Nemethova D, Hejatko J (2008) Effects of conditional IPT-dependent cytokinin overproduction on root architecture of Arabidopsis seedlings. Plant Cell Physiol 49:570–582
Kuppu S, Mishra N, Hu R, Sun L, Zhu X, Shen G, Zhang H (2013) Water-deficit inducible expression of a cytokinin biosynthetic gene IPT improves drought tolerance in cotton. PLoS ONE 8:e64190
Kuroha T, Tokunaga H, Kojima M, Ueda N, Ishida T, Nagawa S, Sakakibara H (2009) Functional analyses of LONELY GUY cytokinin-activating enzymes reveal the importance of the direct activation pathway in Arabidopsis. Plant Cell 21:3152–3169
Latchman DS (1997) Transcription factors: an overview. NT J Biochem Cell B 29:1305–1312
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12(1):323
Li YJ, Wang B, Dong RR, Hou BK (2015) AtUGT76C2, an Arabidopsis cytokinin glycosyltransferase is involved in drought stress adaptation. Plant Sci 236:157–167
Liu P, Zhang C, Ma JQ, Zhang LY, Yang B, Tang XY, Li JN (2018) Genome-wide identification and expression profiling of cytokinin oxidase/dehydrogenase (CKX) genes reveal likely roles in pod development and stress responses in oilseed rape (Brassica napus L.). Genes 9(3):168
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative pcr and the 2−ΔΔCT method. Methods 25:402–408
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550
Mantiri FR, Kurdyukov S, Lohar DP, Sharopova N, Saeed NA, Wang XD, Rose RJ (2008) The transcription factor MtSERF1 of the ERF subfamily identified by transcriptional profiling is required for somatic embryogenesis induced by auxin plus cytokinin in Medicago truncatula. Plant Physiol 146:1622–1636
Mao XZ, Cai T, Olyarchuk JG, Wei LP (2005) Automated genome annotation and pathway identification using the KEGG orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793
Mason MG, Mathews DE, Argyros DA, Maxwell BB, Kieber JJ, Alonso JM, Schaller GE (2005) Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 17(11):3007–3018
Miyawaki K, Tarkowski P, Matsumoto-Kitano M, Kato T, Sato S, Tarkowska D, Kakimoto T (2006) Roles of arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. P Natl Acad Sci USA 103:16598–16603
Mok DW, Mok MC (2001) Cytokinin metabolism and action. Annu Rev Plant Physiol Plant Mol Biol 52:89–118
Moll KM, Zhou P, Ramaraj T, Fajardo D, Devitt NP, Sadowsky MJ, Silverstein KA (2017) Strategies for optimizing BioNano and dovetail explored through a second reference quality assembly for the legume model Medicago truncatula. BMC Genomics 18(1):578
Muller B, Sheen J (2008) Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453:1094–1097
Naito T, Yamashino T, Kiba T, Koizumi N, Kojima M, Sakakibara H, Mizuno T (2007) A link between cytokinin and ASL9 (ASYMMETRIC LEAVES 2 LIKE 9) that belongs to the AS2/LOB (LATERAL ORGAN BOUNDARIES) family genes in Arabidopsis thaliana. Biosci Biotech Bioch 71:1269–1278
Nguyen KH, HaC V, Nishiyama R, Watanabe Y, Leyva-González MA, Fujita Y, Schaller GE (2016) Arabidopsis type B cytokinin response regulators ARR1, ARR10, and ARR12 negatively regulate plant responses to drought. P Natl Acad Sci USA 113(11):3090–3095
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295
Polanska L, Malbeck J, Travnickova A, Vankova R, Machackova I (2004) Cytokinin occurrence in chloroplasts of tobacco plants harboring zeatin O-glucosyl transferase gene (Zog1). Acta Physiol Plant 26:39–39
Raines T, Blakley IC, Tsai YC, Worthen JM, Franco-Zorrilla JM, Solano R, Kieber JJ (2016) Characterization of the cytokinin-responsive transcriptome in rice. BMC Plant Biol 16:260
Rashotte AM, Carson SD, To JP, Kieber JJ (2003) Expression profiling of cytokinin action in Arabidopsis. Plant Physiol 132:1998–2011
Reid DE, Heckmann AB, Novak O, Kelly S, Stougaard J (2016) CYTOKININ OXIDASE/DEHYDROGENASE3 maintains cytokinin homeostasis during root and nodule development in lotus japonicus. Plant Physiol 170:1060–1074
Reyes-Olalde JI, Zúñiga-Mayo VM, Serwatowska J, Montes RAC, Lozano-Sotomayor P, Herrera-Ubaldo H, Paolo D (2017) The bHLH transcription factor SPATULA enables cytokinin signaling, and both activate auxin biosynthesis and transport genes at the medial domain of the gynoecium. PLoS Genet 13:e1006726
Roche J, Love J, Guo Q, Song J, Cao M, Fraser K, Jameson PE (2016) Metabolic changes and associated cytokinin signals in response to nitrate assimilation in roots and shoots of Lolium perenne. Physiol Plant 156:497–511
Rogers EE, Ausubel FM (1997) Arabidopsis enhanced disease susceptibility mutants exhibit enhanced susceptibility to several bacterial pathogens and alterations in PR-1 gene expression. Plant Cell 9(3):305–316
Ross EJ, Stone JM, Elowsky CG, Arredondo-Peter R, Klucas RV, Sarath G (2004) Activation of the Oryza sativa non-symbiotic haemoglobin-2 promoter by the cytokinin-regulated transcription factor, ARR1. J Exp Bot 55:1721–1731
Schaller GE, Street IH, Kieber JJ (2014) Cytokinin and the cell cycle. Curr Opin Plant Biol 21:7–15
Schmulling T, Werner T, Riefler M, Krupkova E, Bartrina y Manns I (2003) Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J Plant Res 116:241–252
Shi X, Gupta S, Lindquist IE, Cameron CT, Mudge J, Rashotte AM (2013) Transcriptome analysis of cytokinin response in tomato leaves. PLoS ONE 8:e55090
Tadege M, Ratet P, Mysore KS (2005) Insertional mutagenesis: a Swiss army knife for functional genomics of Medicago truncatula. Trends Plant Sci 10:229–235
Tadege M, Wen J, He J, Tu H, Kwak Y, Eschstruth A, Ratet P (2008) Large-scale insertional mutagenesis using the Tnt1 retrotransposon in the model legume Medicago truncatula. Plant J 54:335–347
Talla SK, Panigrahy M, Kappara S, Nirosha P, Neelamraju S, Ramanan R (2016) Cytokinin delays dark-induced senescence in rice by maintaining the chlorophyll cycle and photosynthetic complexes. J Exp Bot 67:1839–1851
Tang H, Krishnakumar V, Bidwell S, Rosen B, Chan A, Zhou S, Mayer KF (2014) An improved genome release (version Mt4.0) for the model legume Medicago truncatula. BMC Genomics 15(1):312
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33–36
Thatcher LF, Williams AH, Garg G, Buck SG, Singh KB (2016) Transcriptome analysis of the fungal pathogen Fusarium oxysporum f. sp. medicaginis during colonisation of resistant and susceptible Medicago truncatula hosts identifies differential pathogenicity profiles and novel candidate effectors. BMC Genomics 17:860
Tokunaga H, Kojima M, Kuroha T, Ishida T, Sugimoto K, Kiba T, Sakakibara H (2012) Arabidopsis lonely guy (LOG) multiple mutants reveal a central role of the LOG-dependent pathway in cytokinin activation. Plant J 69:355–365
Vidal RO, do Nascimento LC, Mondego JM, Pereira GA, Carazzolle MF (2012) Identification of SNPs in RNA-seq data of two cultivars of glycine max (soybean) differing in drought resistance. Genet Mol Biol 35:331–334
Xia Z, Zeng X, Ma G (2009) Regulation of CPPU on dynamic changes of endogenous phytohormone in Momordica charantia L. Southwest China J Agricu Sci 22:5
Zalewski W, Galuszka P, Gasparis S, Orczyk W, Nadolska-Orczyk A (2010) Silencing of the HvCKX1 gene decreases the cytokinin oxidase/dehydrogenase level in barley and leads to higher plant productivity. J Exp Bot 61:1839–1851
Zhang S, Shi Y, Cheng N, Du H, Fan W, Wang C (2015) De novo characterization of fall dormant and nondormant alfalfa (Medicago sativa L.) leaf transcriptome and identification of candidate genes related to fall dormancy. PLoS ONE 10:0122170
Zhao J, Ba W, Zeng Q, Song S, Zhang M, Li X, Luo X (2015) Moderately enhancing cytokinin level by down-regulation of expression in cotton concurrently increases fiber and seed yield. Mol Beeding 35:60
Zurcher E, Muller B (2016) Cytokinin synthesis, signaling, and function-advances and new insights. Int Rev Cel Mol Bio 324:1–38
Acknowledgements
The work is supported by the Fundamental Research Funds for the Central Universities (2019ZY11) and Tibet Science and Technology Major Projects of Pratacultural Industry. The data were analyzed on the free online platform of Majorbio I-Sanger Cloud Platform (www.i-sanger.com).
Author information
Authors and Affiliations
Contributions
YHC and LBH planned and designed the experiments, and wrote the main manuscript text; ZXZ and HCL performed majorly the experiments and data acquisition; CNM participated partially the experiments and data analysis; YHC and LBH participated the figures preparation and MS English editing.
Corresponding author
Ethics declarations
Conflict of Interest
All the authors declare that they do not have conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, Z., Liu, H., Ma, C. et al. Transcriptome Analysis of the Cytokinin Response in Medicago truncatula. J. Plant Biol. 63, 189–202 (2020). https://doi.org/10.1007/s12374-020-09244-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-020-09244-8