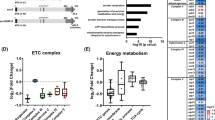
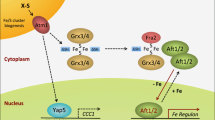
Abstract
Aconitase, a highly conserved protein across all domains of life, functions in converting citrate to isocitrate in the tricarboxylic acid cycle. Cytosolic aconitase is also known to act as an iron regulatory protein in mammals, binding to the RNA hairpin structures known as iron-responsive elements within the untranslated regions of specific RNAs. Aconitase-2 (Aco2) in fission yeast is a fusion protein consisting of an aconitase and a mitochondrial ribosomal protein, bL21, residing not only in mitochondria but also in cytosol and the nucleus. To investigate the role of Aco2 in the nucleus and cytoplasm of fission yeast, we analyzed the transcriptome of aco2ΔN mutant that is deleted of nuclear localization signal (NLS). RNA sequencing revealed that the aco2ΔN mutation caused increase in mRNAs encoding iron uptake transporters, such as Str1, Str3, and Shu1. The half-lives of mRNAs for these genes were found to be significantly longer in the aco2ΔN mutant than the wild-type strain, suggesting the role of Aco2 in mRNA turnover. The three conserved cysteines required for the catalytic activity of aconitase were not necessary for this role. The UV cross-linking RNA immunoprecipitation analysis revealed that Aco2 directly bound to the mRNAs of iron uptake transporters. Aco2-mediated degradation of iron-uptake mRNAs appears to utilize exoribonuclease pathway that involves Rrp6 as evidenced by genetic interactions. These results reveal a novel role of non-mitochondrial aconitase protein in the mRNA turnover in fission yeast to fine-tune iron homeostasis, independent of regulation by transcriptional repressor Fep1.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Alén, C. and Sonenshein, A.L. 1999. Bacillus subtilis aconitase is an RNA-binding protein. Proc. Natl. Acad. Sci. USA 96, 10412–10417.
Beinert, H., Kennedy, M.C., and Stout, C.D. 1996. Aconitase as iron-sulfur protein, enzyme, and iron-regulatory protein. Chem. Rev. 96, 2335–2374.
Benjamin, J.A. and Masse, E. 2014. The iron-sensing aconitase B binds its own mRNA to prevent sRNA-induced mRNA cleavage. Nucleic Acids Res. 42, 10023–10036.
Boukouris, A.E., Zervopoulos, S.D., and Michelakis, E.D. 2016. Metabolic enzymes moonlighting in the nucleus: metabolic regulation of gene transcription. Trends Biochem. Sci. 41, 712–730.
Castello, A., Hentze, M.W., and Preiss, T. 2015. Metabolic enzymes enjoying new partnerships as RNA-binding proteins. Trends Endocrinol. Metab. 26, 746–757.
Chen, H.M., Futcher, B., and Leatherwood, J. 2011. The fission yeast RNA binding protein Mmi1 regulates meiotic genes by controlling intron specific splicing and polyadenylation coupled RNA turnover. PLoS ONE 6, e26804.
Chen, X.J., Wang, X., Kaufman, B.A., and Butow, R.A. 2005. Aconitase couples metabolic regulation to mitochondrial DNA maintenance. Science 307, 714–717.
Dupuy, J., Volbeda, A., Carpentier, P., Darnault, C., Moulis, J.M., and Fontecilla-Camps, J.C. 2006. Crystal structure of human iron regulatory protein 1 as cytosolic aconitase. Structure 14, 129–139.
Egan, E.D., Braun, C.R., Gygi, S.P., and Moazed, D. 2014. Post-transcriptional regulation of meiotic genes by a nuclear RNA silencing complex. RNA 20, 867–881.
Fagerström-Billai, F. and Wright, A.P. 2005. Functional comparison of the Tup11 and Tup12 transcriptional corepressors in fission yeast. Mol. Cell. Biol. 25, 716–727.
Forsburg, S.L. and Rhind, N. 2006. Basic methods for fission yeast. Yeast 23, 173–183.
Gourley, B.L., Parker, S.B., Jones, B.J., Zumbrennen, K.B., and Leibold, E.A. 2003. Cytosolic aconitase and ferritin are regulated by iron in Caenorhabditis elegans. J. Biol. Chem. 278, 3227–3234.
Gray, N.K., Pantopoulos, K., Dandekar, T., Ackrell, B.A., and Hentze, M.W. 1996. Translational regulation of mammalian and Drosophila citric acid cycle enzymes via iron-responsive elements. Proc. Natl. Acad. Sci. USA 93, 4925–4930.
Gupta, M. and Outten, C.E. 2020. Iron-sulfur cluster signaling: the common thread in fungal iron regulation. Curr. Opin. Chem. Biol. 55, 189–201.
Harigaya, Y., Tanaka, H., Yamanaka, S., Tanaka, K., Watanabe, Y., Tsutsumi, C., Chikashige, Y., Hiraoka, Y., Yamashita, A., and Yamamoto, M. 2006. Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442, 45–50.
Hocquet, C., Robellet, X., Modolo, L., Sun, X.M., Burny, C., Cuylen-Haering, S., Toselli, E., Clauder-Münster, S., Steinmetz, L., Haering, C.H., et al. 2018. Condensin controls cellular RNA levels through the accurate segregation of chromosomes instead of directly regulating transcription. eLife 7, e38517.
Jacques, J.F., Mercier, A., Brault, A., Mourer, T., and Labbé, S. 2014. Fra2 is a co-regulator of Fep1 inhibition in response to iron starvation. PLoS ONE 9, e98959.
Jeffery, C.J. 2015. Why study moonlighting proteins? Front. Genet. 6, 211.
Jung, S.J., Choi, Y., Lee, D., and Roe, J.H. 2019. Nuclear aconitase antagonizes heterochromatic silencing by interfering with Chp1 binding to DNA. Biochem. Biophys. Res. Commun. 516, 806–811.
Jung, S.J., Seo, Y., Lee, K.C., Lee, D., and Roe, J.H. 2015. Essential function of Aco2, a fusion protein of aconitase and mitochondrial ribosomal protein bL21, in mitochondrial translation in fission yeast. FEBS Lett. 589, 822–828.
Kilchert, C., Wittmann, S., and Vasiljeva, L. 2016. The regulation and functions of the nuclear RNA exosome complex. Nat. Rev. Mol. Cell Biol. 17, 227–239.
Kim, E.J., Cho, Y.J., Chung, W.H., and Roe, J.H. 2020. The role of Rsv1 in the transcriptional regulation of genes involved in sugar metabolism for long-term survival. FEBS J. 287, 878–896.
Kim, K.D., Kim, H.J., Lee, K.C., and Roe, J.H. 2011. Multi-domain CGFS-type glutaredoxin Grx4 regulates iron homeostasis via direct interaction with a repressor Fep1 in fission yeast. Biochem. Biophys. Res. Commun. 408, 609–614.
Kim, H.J., Lee, K.L., Kim, K.D., and Roe, J.H. 2016. The iron uptake repressor Fep1 in the fission yeast binds Fe-S cluster through conserved cysteines. Biochem. Biophys. Res. Commun. 478, 187–192.
Li, X., Egervari, G., Wang, Y., Berger, S.L., and Lu, Z. 2018. Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat. Rev. Mol. Cell Biol. 19, 563–578.
Lind, M.I., Missirlis, F., Melefors, Ö., Uhrigshardt, H., Kirby, K., Phillips, J.P., Söderhäll, K., and Rouault, T.A. 2006. Of two cytosolic aconitases expressed in Drosophila, only one functions as an iron-regulatory protein. J. Biol. Chem. 281, 18707–18714.
Mercier, A., Watt, S., Bähler, J., and Labbé, S. 2008. Key function for the CCAAT-binding factor Php4 to regulate gene expression in response to iron deficiency in fission yeast. Eukaryot. Cell 7, 493–508.
Mourer, T., Jacques, J.F., Brault, A., Bisaillon, M., and Labbé, S. 2015. Shu1 is a cell-surface protein involved in iron acquisition from heme in Schizosaccharomyces pombe. J. Biol. Chem. 290, 10176–10190.
Mukherjee, K., Gardin, J., Futcher, B., and Leatherwood, J. 2016. Relative contributions of the structural and catalytic roles of Rrp6 in exosomal degradation of individual mRNAs. RNA 22, 1311–1319.
Pelletier, B., Beaudoin, J., Mukai, Y., and Labbé, S. 2002. Fep1, an iron sensor regulating iron transporter gene expression in Schizosaccharomyces pombe. J. Biol. Chem. 277, 22950–22958.
Pelletier, B., Beaudoin, J., Philpott, C.C., and Labbé, S. 2003. Fep1 represses expression of the fission yeast Schizosaccharomyces pombe siderophore-iron transport system. Nucleic Acids Res. 31, 4332–4344.
Pelletier, B., Trott, A., Morano, K.A., and Labbé, S. 2005. Functional characterization of the iron-regulatory transcription factor Fep1 from Schizosaccharomyces pombe. J. Biol. Chem. 280, 25146–25161.
Piccinelli, P. and Samuelsson, T. 2007. Evolution of the iron-responsive element. RNA 13, 952–966.
Pouliot, B., Jbel, M., Mercier, A., and Labbé, S. 2010. abc3+ encodes an iron-regulated vacuolar ABC-type transporter in Schizosaccharomyces pombe. Eukaryot. Cell 9, 59–73.
Satoh, R., Morita, T., Takada, H., Kita, A., Ishiwata, S., Doi, A., Hagihara, K., Taga, A., Matsumura, Y., Tohda, H., et al. 2009. Role of the RNA-binding protein Nrd1 and Pmk1 mitogen-activated protein kinase in the regulation of myosin mRNA stability in fission yeast. Mol. Biol. Cell 20, 2473–2485.
Shichino, Y., Otsubo, Y., Yamamoto, M., and Yamashita, A. 2020. Meiotic gene silencing complex MTREC/NURS recruits the nuclear exosome to YTH-RNA-binding protein Mmi1. PLoS Genet 16, e1008598.
Telekawa, C., Boisvert, F.M., and Bachand, F. 2018. Proteomic profiling and functional characterization of post-translational modifications of the fission yeast RNA exosome. Nucleic Acids Res. 46, 11169–11183.
Touat-Todeschini, L., Shichino, Y., Dangin, M., Thierry-Mieg, N., Gilquin, B., Hiriart, E., Sachidanandam, R., Lambert, E., Brettschneider, J., Reuter, M., et al. 2017. Selective termination of lncRNA transcription promotes heterochromatin silencing and cell differentiation. EMBO J. 36, 2626–2641.
Volz, K. 2008. The functional duality of iron regulatory protein 1. Curr. Opin. Struct. Biol. 18, 106–111.
Yamamoto, M. 2010. The selective elimination of messenger RNA underlies the mitosis-meiosis switch in fission yeast. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 86, 788–797.
Yamanaka, S., Yamashita, A., Harigaya, Y., Iwata, R., and Yamamoto, M. 2010. Importance of polyadenylation in the selective elimination of meiotic mRNAs in growing S. pombe cells. EMBO J. 29, 2173–2181.
Yamashita, A., Shichino, Y., Tanaka, H., Hiriart, E., Touat-Todeschini, L., Vavasseur, A., Ding, D.Q., Hiraoka, Y., Verdel, A., and Yamamoto, M. 2012. Hexanucleotide motifs mediate recruitment of the RNA elimination machinery to silent meiotic genes. Open Biol. 2, 120014.
Znaidi, S., Pelletier, B., Mukai, Y., and Labbé, S. 2004. The Schizosaccharomyces pombe corepressor Tup11 interacts with the iron-responsive transcription factor Fep1. J. Biol. Chem. 279, 9462–9474.
Acknowledgements
This work was supported by grants from the National Research Foundation of Korea (NRF) to K.K. (2019R1A4A-1024764). SY Cho was supported by the BK21-Plus fellowship at SNU and the BK21 FOUR fellowship at CAU.
Author information
Authors and Affiliations
Contributions
Conception and experimental design: S. C., K. K., and J. R.; methodology and data acquisition: S. C. and S. J.; paper writing: S. C., K. K., and J. R.
Corresponding authors
Ethics declarations
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Cho, SY., Jung, SJ., Kim, KD. et al. Non-mitochondrial aconitase regulates the expression of iron-uptake genes by controlling the RNA turnover process in fission yeast. J Microbiol. 59, 1075–1082 (2021). https://doi.org/10.1007/s12275-021-1438-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-021-1438-4